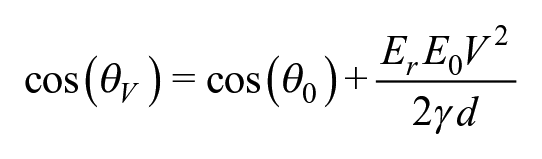Abstract
Digital microfluidics (DMF) is a liquid handling technique that has been demonstrated to automate biological experimentation in a low-cost, rapid, and programmable manner. This review discusses the role of DMF as a “digital bioconverter”—a tool to connect the digital aspects of the design–build–learn cycle with the physical execution of experiments. Several applications are reviewed to demonstrate the utility of DMF as a digital bioconverter, namely, genetic engineering, sample preparation for sequencing and mass spectrometry, and enzyme-, immuno-, and cell-based screening assays. These applications show that DMF has great potential in the role of a centralized execution platform in a fully integrated pipeline for the production of novel organisms and biomolecules. In this paper, we discuss how the function of a DMF device within such a pipeline is highly dependent on integration with different sensing techniques and methodologies from machine learning and big data. In addition to that, we examine how the capacity of DMF can in some cases be limited by known technical and operational challenges and how consolidated efforts in overcoming these challenges will be key to the development of DMF as a major enabling technology in the computer-aided biology framework.
Introduction
With increasing use of engineering and data-led approaches in biological experimentation, several attempts have been made at consolidating these approaches within a framework that gives structure to the pursuit of a rational understanding of biological systems. Some notable concepts are “digital biology,”
1
“biomanufacturing,”
2
and, more recently, “computer-aided biology” (CAB).
3
CAB talks about enhancing human abilities with an ecosystem of digital and automation tools (
Following this, the paper describes the CAB framework and the design–build–test–learn (DBTL) cycle that it drives. This approach has fostered the development of biofoundries, which are currently the model biomanufacturing platform for the production of rationally designed bio-based innovations for global challenges and bringing in the next-generation bioeconomy (
The CAB framework is the driving principle of biofoundries, which are a source of rationally designed bio-based innovations for global challenges.
As a digital bioconverter, DMF has found many applications in popular workflows. This paper covers workflows that could fit in a production pipeline for novel organisms and biomolecules. These include gene assembly, editing and cloning, sample and library preparation for sequencing, sample preparation for mass spectrometry (MS), and enzymatic, immuno-, and cell-based screening assays. A study of these applications of DMF has revealed that there are no monolithic platforms that can fulfill every requirement, insofar as these requirements differ between applications. Therefore, the ideal DMF-based platform is application specific.
In this regard, the role of DMF in driving a DBTL cycle can be shaped by the tools with which it can be integrated. Alongside hardware integrations highlighted in the previously mentioned workflows, there is a tremendous opportunity to incorporate machine learning methods as a means to tap into an increase in data generated when executing workflows.
The question of whether DMF is the appropriate automated execution platform in a DBTL-driven CAB framework was distilled as a matter of the following questions:
Can a DMF platform rapidly respond to changes made in the design and build stage? If analytics of biological experimentation were to inform experimental design, it should be expected that such a tool should have the dynamic flexibility to modify execution parameters at a speed matching the rate of incoming data. To illustrate, imagine the speed at which stock price projection algorithms respond to breaking global news.
Can DMF technology reliably perform multiplexing?
Can DMF enable the integration of tools throughout the entire pipeline?
Limitations in integration, reliable electrowetting performance, matching sample throughput to a standard 96-well plate format for analysis, and the general complexity of using a multidisciplinary technology are all known barriers against the implementation of DMF as a digital bioconverter in the CAB framework. However, these barriers are not insurmountable, and with continued and consolidated efforts to build a CAB framework, the performance and functionality of DMF could be further developed such that it can be an important tool in enabling laboratory automation and making biology programmable.
Digital Microfluidics
DMF is a liquid handling technique for processing droplets ranging from a few picoliters to milliliters.7,8 Different droplet operations can be implemented using forces like light (i.e., optoelectrowetting
9
) over photoconductor arrays or by modulating electric voltage (i.e., EWOD) to modify a substrate’s wettability and actuate discrete liquid volumes. In EWOD-type systems, droplets are usually manipulated over an array of electrodes. When these electrodes are enclosed within a dielectric and an electric potential is applied on them (see
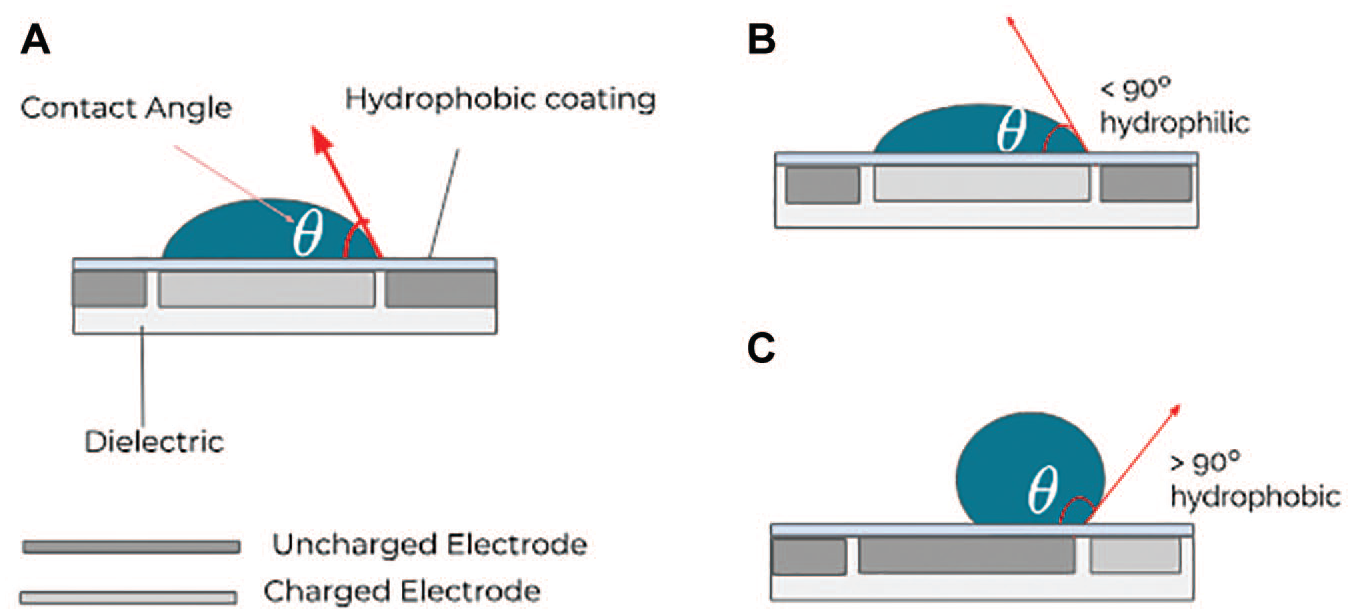
Schematic representation of an EWOD-type digital microfluidics device, specifically showing the contact angle of the droplet.
Manipulating the electric field by varying the voltage applied on the electrodes modifies the wettability
10
of the hydrophobic surface upon which the droplet moves (i.e., contact angle) to allow an implementation of different droplet operations like dispensing, splitting, merging, and mixing (see
where E0 is the permittivity of vacuum, Er is the relative permittivity of the dielectric, γ is the interfacial liquid–gas surface tension, and d is the thickness of the dielectric.
These EWOD-type DMF devices can have either an open or a closed configuration. This is defined by the absence and presence (respectively) of a ground electrode on the top (as shown in
In a closed configuration, the ratio of the area of the bottom electrode to the gap between the top and bottom layers defines the volume and shape of the actuated droplet. As described by Yafia et al., 14 the gap between the two layers plays an important role in how well different droplet operations are executed on the chip. Unlike an open configuration, a closed EWOD configuration can be used with an air-filled or oil-filled operation where the space between the top continuous ground electrode and the bottom array of conducting electrodes is filled with either air or an immiscible oil (e.g., silicone oil). Depending on the composition of the droplet, employing an oil environment can reduce sample evaporation, ensure ideal temperature conditions, and, in some cases, improve droplet actuation and minimize surface adsorption. 15 At the same time, an oil-filled environment could hinder the use of DMF devices for growing cells, 16 implementing liquid–liquid extraction 17 and surface-based assays. 18 Meanwhile, air-filled operation is usually preferred when working with droplets containing hydrophobic components like proteins and lipids, to avoid dispersion within the filler oil and ensure separation of the components from the surrounding oil.
This technique allows users to precisely address individual droplets by sending a sequence of activation and inactivation signals to apply electric voltages on the array of conducting electrodes under the droplets. This droplet-level addressability that the technique provides allows each of the manipulated droplets to be treated as a separate microreactor upon which different droplet operations can be applied. The droplet can contain different components like DNA, RNA, cells, proteins, or chemical reagents. By specifying activation and inactivation sequences on electrodes, a number of complex workflows can be automated, be it liquid handling in molecular biology labs or in chemical synthesis laboratories. With a number of workflows in labs being repetitive and manual, automation is increasingly seen as a solution to some of the current challenges we face with reproducibility in biological workflows. 19
Over the years, there have been sustained efforts to expand the repertoire of workflows that DMF-based tools support, especially for supporting applications in life science laboratories. Workflows in these labs usually involve samples that are expensive. Tools that therefore minimize the volume of samples required are advantageous, especially when the execution of workflows is manual and has a number of repetitive steps where the outcome of the experiment is driven by the experience and skillset of the operator. The flexibility of droplet-level control offerred by DMF is different from what other flow-based microfluidics setups offer. It is ideal for workflows where low-volume sample processing is required along with a precise control over what happens to samples (or individual droplets) of interest with a small footprint and simpler build. The dead volumes in traditional flow-based microfluidics or conventional liquid handlers are much higher than with DMF. In comparison to traditional flow-based microfluidics, DMF offers capabilities for manipulating and sorting individual droplets of interest using programmable biochips that can support multiple workflows. DMF also does not require external pumping and pressure control systems, which are costly and difficult to maintain. This not only increases the ease of operation but also decreases capital and operation expenses and is more efficient, as it reduces the overall footprint of an automated workflow compared with a traditional flow-based setup. An additional commonly cited benefit of using DMF is the absence of blockages, a consistent challenge found in flow-based microfluidics devices.20,21 In addition to this, integration with different detector modules,22,23 and support for purification, 24 DMF can allow a convergence of all steps in an experimental workflow on a single device. Workflows like running heterogeneous immunoassays (enzyme-linked immunosorbent assay [ELISA])24,25 and enzymatic assays 26 for diagnostics, sample preparation for proteomics, 27 gene editing, 28 and drug screening via MS 29 all give an indication of the breadth of workflows that can be automated using DMF.
CAB and Biofoundries
CAB refers to a collective approach to biological research through digital integration and augmentation, and the application of the DBTL approach (
CAB is the guiding principle behind biofoundries.44–46 Biofoundries are platforms that integrate tools to automate steps in the DBTL cycle. 47 In their review, Chao et al.47 provide an overview of the components of a typical biofoundry, which correlate with different CAD, CAM, and data management and analysis tools. Some examples of well-known biofoundries include iBioFab, 46 Edinburgh Genome Foundry, 45 and the start-ups Ginkgo Bioworks and Zymergen. However, the first attempt at such a style of research and development was demonstrated by Sparkes et al. in 2010 in their prototype systems “Adam” for the study of functional genomics in yeast and “Eve” for closed-loop learning for drug screening and design. 43 Another notable and more recent example of a (proof-of-concept) full implementation of the DBTL cycle is the BioAutomata, used to optimize the production pathway for lycopene. 46 Nevertheless, biofoundries are a promising platform for next-generation biological engineering. For example, the Ginkgo Bioworks biofoundry can accomplish the rapid prototyping of cell factories for the production of various biomolecules at a rate 25 times faster than manual methods. Additionally, the output from their foundry has doubled every 6 months since 2015. 48
Beyond conceptual frameworks, the CAB approach introduces opportunities for the use of data-driven technologies that are increasingly established in other industries. Especially in the “learn” phase, the combination of IoT, computer vision, machine learning, semantic technology, sensor network models, and so forth, can greatly enhance decision-making processes at the “design” and “build” stages. This type of approach, which is focused on big data analysis of disparate and multisource databases to generate decision-making input, has found applications in innovation management,49–51 smart farming, 52 and smart cities, 53 among others. Indeed, restructuring current approaches in biological experimentation to fit the DBTL approach would make clearer any avenues for improvement based on emerging digital and automation technologies in other industries.
DMF as an Enabling Technology in the DBTL Cycle
In their paper, Minie and Samudrala
1
described the digital biology loop, a concept that is centered around a “digital bioconverter,” an instrument that is capable of translating digitized genes into cells, viruses, and biological molecules (
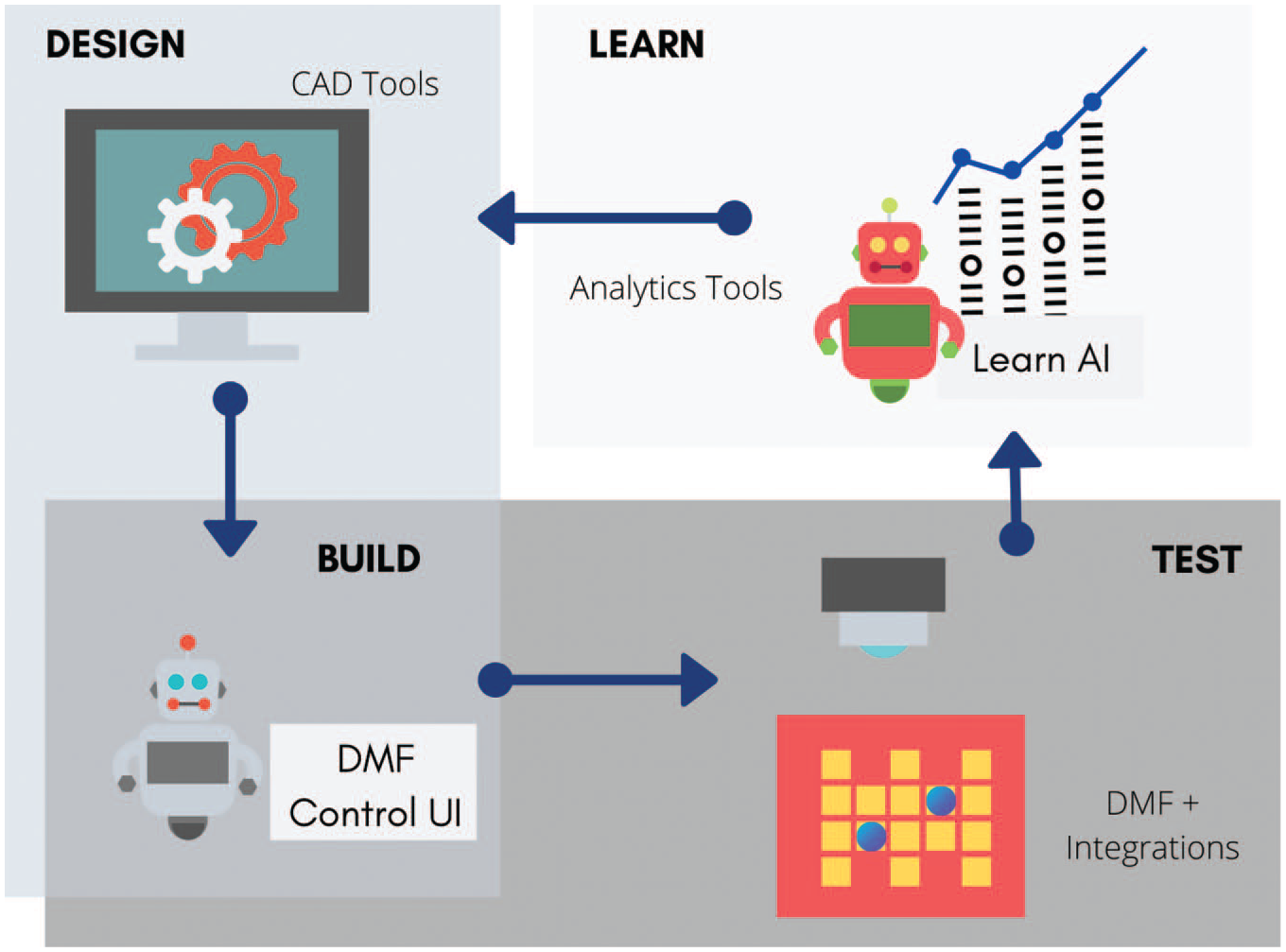
Using DMF as an execution platform, CAD and analytics tools can be integrated upstream and downstream to complete the DBTL cycle.
The potential utility of DMF as an enabling technology can be demonstrated in the workflows that are currently automated using DMF, covered in the following section.
Workflows Currently Automated Using DMF
Most common workflows in a modern biology laboratory involve working with oligonucleotides, proteins, and cells. They usually have multiple steps that are often implemented manually and across several devices. As a DMF device functions as a “build” and “test” tool, the following workflows focus on protocols used in construct assembly and validation, as well as protocols used in the detection and quantification of interesting biomolecules. The protocols covered in this section can be arranged in different ways in a single pipeline for the production of novel organisms and biomolecules of interest (
While several papers have discussed the utility of DMF systems for cell culture,54–56 it is notably missing in our discussion as it is still unclear if cell growth in this format can generate results that are statistically comparable to phenotypic observations resulting from plate-based growth.
Gene Assembly, Editing, and Cloning
Traditionally, assembly or editing of long oligonucleotides has been achieved by the fusion of shorter DNA fragments using ligation-based 57 or polymerase-based 58 reactions. Several challenges are commonly observed with these approaches. First, these protocols come with a certain degree of loss due to limitations in sequence fidelity, purification, transfection, and transformation efficiencies. 59 This is typically compensated by starting with larger than required amounts of DNA, vectors, reagents, and enzymes, which increases the total cost of the workflow. An additional challenge in the genetic engineering pipeline is supporting the construction of rationally designed synthetic circuits, which are important tools in systematically understanding and designing cellular pathways.60–62 Rational design approaches often require the assembly of a very large number of diverse fragments using expensive and precious genetic materials and enzymes. DMF has been examined as a potential solution to the aforementioned challenges, 63 , 64 as it has been suggested to reduce DNA loss and support complex protocols using low sample volumes.
For example, Yehezkel et al.
65
developed ad hoc DMF methods for de novo synthesis, combinatorial assembly, and cell-free cloning. Using their programmable order polymerization (POP) method, the authors show that ad hoc, on-chip de novo synthesis of eight 2.5 kb fragments can be achieved with error rates conforming to those in literature reports on other de novo synthesis methods within 1.5 h. Additionally, the POP method showed a higher yield of correct assemblies and lower mis-hybridizations, compared with fragments generated through a one-pot assembly reaction (Gibson). The authors also demonstrated an ad hoc combinatorial assembly of rationally designed variants. This microfluidics combinatorial assembly of DNA (M-CAD) method showed 50-fold and 10-fold reduction in reagent volume and turnaround time, respectively, along with an error rate that is comparable to that found in the literature. Yehezkel et al. further prove the utility of DMF by demonstrating a high-fidelity cloning method (microfluidics in vitro cloning [MIC]) based on a combination of an on-cartridge serial dilution PCR and DNA barcoding. Another approach to improving the reliability of current assembly methods using DMF is the addition of error correction. Khilko et al.
66
show this where they implemented a one-pot Gibson assembly coupled with an enzyme-mediated error correction step to obtain 339 bp fragments within 15 min in comparison with off-chip benchtop methods, which typically take 15–60 min. In comparison with the ad hoc assembly method demonstrated by Yehezkel et al., sequence fidelity is also comparable (2.2 errors kb–1 in Yehezkel et al. and 1.8 errors kb–1 in Khilko et al.). While these results were promising, Khilko et al.’s method did require the use of additional amounts of MgCl2, polymerase, and PEG8000 to achieve amplification. Using a hybrid flow–DMF platform, Shih et al.
64
showed the execution of DNA assembly (using three different assembly methods: Golden Gate, Gibson, and TAR cloning), incubation, and electrotransformation on a single chip using low total volumes (<1 µL). These studies are summarized in
Summary of Implementation of DMF in Gene Assembly, Editing, and Cloning.
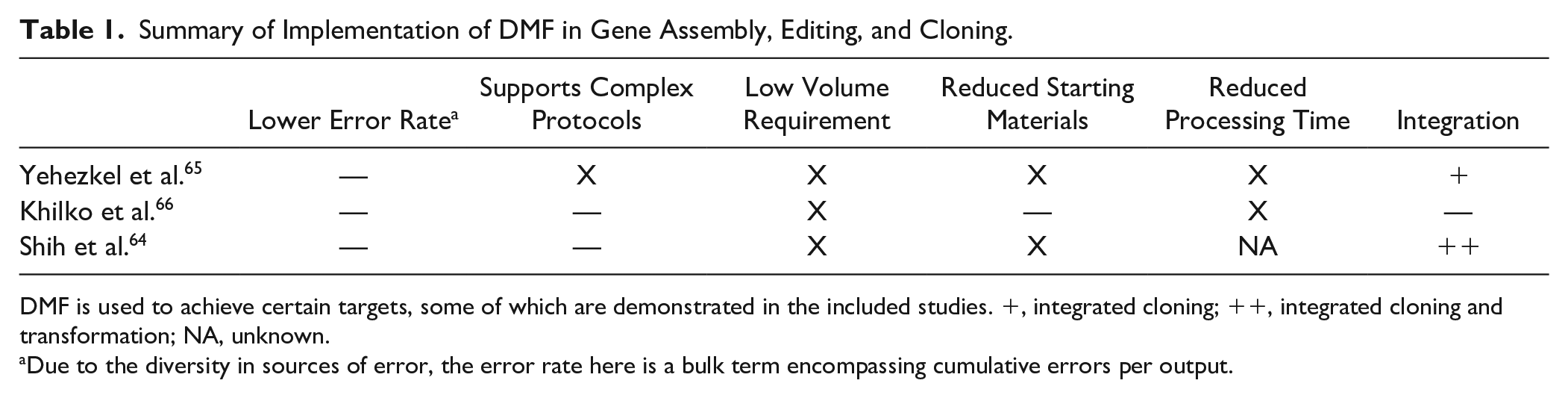

DMF is used to achieve certain targets, some of which are demonstrated in the included studies. +, integrated cloning; ++, integrated cloning and transformation; NA, unknown.
Due to the diversity in sources of error, the error rate here is a bulk term encompassing cumulative errors per output.
Summary of Implementation of DMF in Sample/Library Prep for Sequencing.
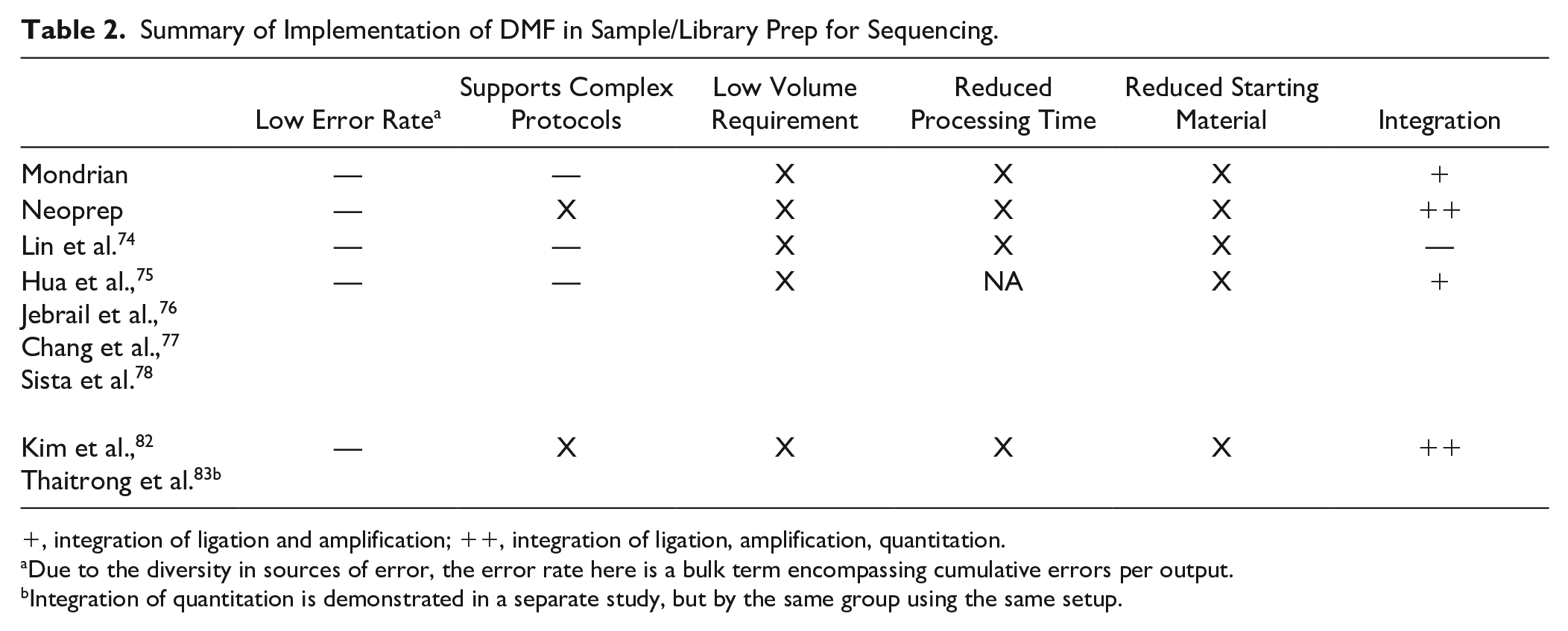

, integration of ligation and amplification; ++, integration of ligation, amplification, quantitation.
Due to the diversity in sources of error, the error rate here is a bulk term encompassing cumulative errors per output.
Integration of quantitation is demonstrated in a separate study, but by the same group using the same setup.
Summary of Implementation of DMF for Assays.
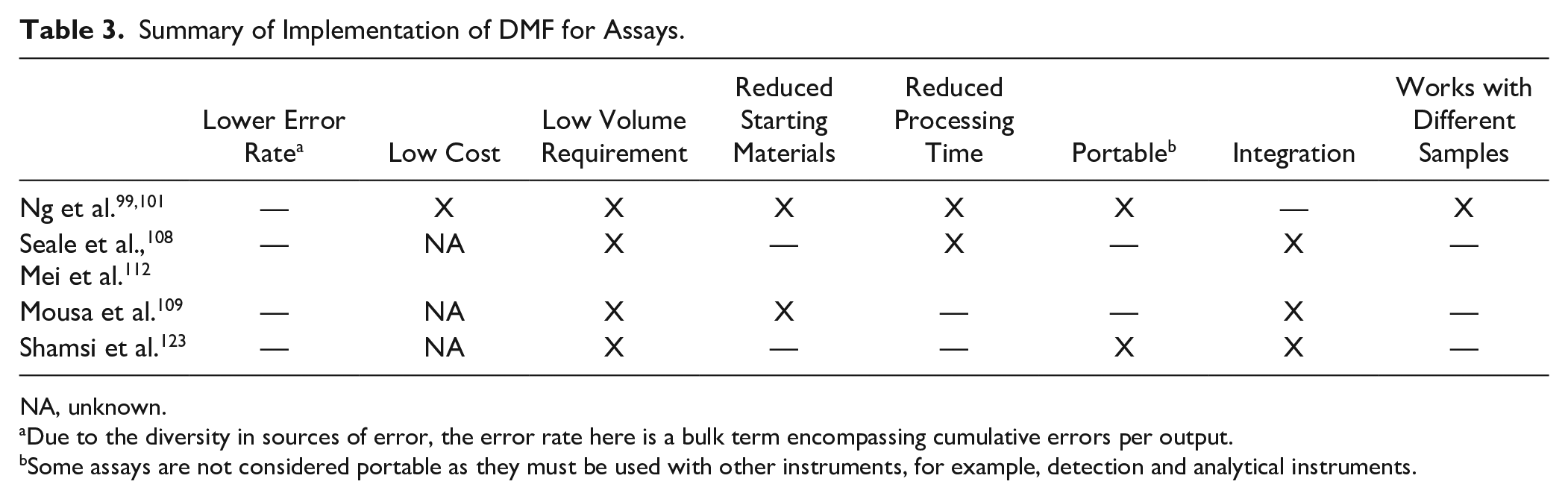
NA, unknown.
Due to the diversity in sources of error, the error rate here is a bulk term encompassing cumulative errors per output.
Some assays are not considered portable as they must be used with other instruments, for example, detection and analytical instruments.
CRISPR-Cas9 systems could be an interesting pairing with DMF (Suppl. Table S2) as its functionality in multiplexed mutagenesis can greatly scale with the parallelization capabilities of DMF. 67 Furthermore, the ad hoc nature of DMF protocols can be used to reduce the incidence of off-site mutagenesis associated with CRISPR-Cas9, for example, by optimizing the concentration of Cas9 per reaction against the number of targeted edits. 68 However, very few examples exist to demonstrate the use of CRISPR-Cas9 systems in a DMF environment. Iwai et al. 69 show single deletions in Saccharomyces cerevisiae using a hybrid flow–DMF system. Sinha et al. 28 cultured and edited H1299 lung cancer cells on a single chip to develop an automated genome editing pipeline using CRISPR-Cas9. The transfection efficiency was observed to be approximately 35% on the DMF platform, 28 which is comparable to well plate experiments. Additionally, knockout efficiencies were comparable across loci. The authors attributed the observed knockout efficiencies to the theory that culturing on a DMF device is more homogenous and reproducible compared with well plate methods. 70
Sample and Library Preparation for Sequencing
Sample and library preparation protocols are a crucial precursor to high-quality DNA sequencing data. Many different library preparation protocols have been developed to address the variety of genetic data that need to be sequenced, and the potential for biases and errors associated with specific samples or libraries. Most of these protocols require multiple tools and often involve several stages of manual processing. From this aspect, realizing high efficiencies can be challenging because errors accumulate at each step.71–73 This becomes especially problematic for preparing libraries for next-generation sequencing, which can comprise 25–100 steps. Furthermore, the quality of sequencing data is also affected by low-volume samples. This represents a considerable limitation as many applications involve limited and/or expensive samples. DMF has been investigated widely in order to reduce the accumulation of errors throughout a library prep protocol and to support low-volume sample preparation.
The execution of library prep protocols on DMF systems has focused on consolidating as many parts of the protocol as possible on a single platform. Steps in a sample/library preparation protocol are shown in
Commercial solutions also exist in this space, though no literature can be found that fairly demonstrates performance. The Mondrian SP+ Workstation takes 3.5 h to process 100 ng of input and produces a library yield of 2.25 µg of DNA. The instrument automates the a-tailing and adaptor ligation stages. The NeoPrep Library Prep Systems covers size selection to quantitation and processes 25–100 ng samples in 7 h.
Sample Preparation for MS
Next to sequencing, MS is another valuable validation technique used in synthetic biology pipelines. Several reviews84–87 discuss the application of DMF in sample preparation for off-line and in-line mass spectrometric analysis. Applications are typically targeted at creating a portable sample processing and analysis platform for rapid diagnostics. DMF-MS integrations are also focused on reducing errors that arise when there is manual sample processing, minimizing sample loss between chip-to-MS transfers, and lowering the sample volume requirements for analysis. Many studies have shown DMF coupled with a variety of MS methods. Either samples can be processed on chip and then removed from the chip before analysis (indirect off-line method) or a sample can be transported directly from a chip to an instrument for analysis without further processing (direct off-line method). A third mode of integration is the in-line analysis where quantification is made using an integrated analytical instrument.
Indirect off-line methods are fairly straightforward and mainly focus on sample preparation prior to MS. An example is seen in Abdulwahab et al., 88 where the authors used a DMF system for on-chip tissue extraction and magnetic particle-based immunoassay to detect estradiol from mammalian tissue biopsies using high-performance liquid chromatography (HPLC)–MS/MS. The method can be executed from extraction to analysis within 40 min. This short processing time and the portable format of this system make it suitable for use in the clinician’s office.
Direct off-line methods have been well characterized in the early works of the Wheeler Lab, where samples are dried, diluted, resuspended in a matrix, and dried again prior to analysis by matrix-assisted laser desorption/ionization (MALDI)–MS via a hole drilled through the chip.89,90 Next to that, Lapierre et al. 91 developed a method involving a patterned superhydrophobic-superhydrophilic silicon nanowire interface for off-line matrix-free laser desorption/ionization (LDI)–MS analysis of low-molecular-weight compounds (700 m/z) within the pmol L–1 range (down to 10 fmol L–1).
With in-line analysis, a common method to interface DMF with MS is through electrospray ionization, where enabling differences in pressure and voltage necessitates additional instrumentation. For example, Shih et al. 90 and Kirby et al. 29 employ sandwiched capillary emitters, while Baker et al. 92 and Hu et al. 93 employ Venturi pump-based capillaries connected to external emitters. A different method for in-line analysis was also shown by Liu et al., 94 where the authors designed a chip that can be used directly with a standard autosampler for delivery to a HPLC-MS instrument. Their technique allows for the analysis of 1–5 µL aqueous and methanolic samples, a 10× reduction compared with a similar method described previously. 95 This method was used in a later study by Choi et al. 96 that incorporates DMF, solid-phase microextraction (SPME), and HPLC-MS for the quantification of free steroid hormones in urine within the picogram per milliliter range. Their technique supported a 25-fold concentration of sample extracts, allowing for lower detection limits, which are necessary for the detection of free hormones in urine.
Some studies have also focused on directly integrating MS with the DMF platform without the need for intermediate equipment. For example, Sathyanarayanan et al.
97
show in their work a DMF device integrated MS via desorption atmospheric pressure photoionization (DAPPI-MS). The device was developed for the implementation and in situ quantification of an on-chip drug distribution and drug metabolism assay while minimizing sample loss. This method allowed for detection within the range of 0.25–1.0 µg mL–1, which makes it suitable for use in the detection of in vivo administered drugs in bodily fluids. In another study, Heinemann et al.
98
used nanostructure-initiator MS (NIMS) integrated with a hybrid flow and a DMF platform to quantify 20 enzyme assays in parallel using 150 nL droplets. For more examples of papers discussing DMF for sample prep in MS, refer to
Enzymatic Assay, Immunoassays, and Cell-Based Screening Assays
DMF is also an attractive platform for implementing a number of different types of assays like enzymatic assays, 26 heterogeneous immunoassays99–101 or oligonucleotide binding assays. 81 , 102 , 103 These assays can be employed in a number of different application areas, for example, protein design, metabolic engineering, diagnostics, and drug discovery. One of the most popular applications of these protocols is in the development of point-of-care (POC) diagnostics tools. The adaptability of DMF technology to low-cost and portable instrument design 13 , 104 makes DMF systems ideal for diagnostic assay development 78 , 105 and serological surveys. 106 Additionally, there is evidence showing that the use of DMF in developing laboratory-quality assays can deliver accurate results 107 that are comparable to those of benchtop methods.
In the case of heterogeneous immunoassays, the use of DMF has demonstrated various advantages over typical methods. 99 Ng et al. 99 demonstrated a 100-fold reduction in reagent use and 10-fold reduction in assay time while maintaining clinically relevant accuracy for the detection of 17b estradiol and thyroid stimulating hormone (TSH). Ng et al. 101 developed a seven-step ELISA-based method to detect rubella virus (RV) antibodies. This uses 1.8 µL samples to generate four assay replicates within 25 min for RV IgG assays and 35 min for RV IgM assays, with a sensitivity and specificity comparable to those for standard laboratory-based methods. The method works with human serum, plasma, and whole-blood samples.
Most notably, assays on a DMF platform can be integrated with various sample preparation protocols, such as immunoprecipitation, 108 solid-phase and liquid–liquid extraction 109 , 110 transfer of pathogen-containing aerosol into droplets, 111 and removal of abundant protein from human plasma. 112 The integration of sample preparation protocols on the device allows a DMF platform to work with a variety of samples, like blood serum, 108 , 109 plasma, 101 , 112 whole-blood samples, 76 , 113 solids like tissues, 109 , 114 dried blood, 90 , 115 hydrogels, 116 , 117 and monoliths. 118 , 119
Examples in the literature show the use of optical detection of analytes through absorbance, 120 chemiluminescence, 121 or fluorescence, 122 and, in some cases, electrochemical detection such as amperometry. 123 Shamsi et al. demonstrated the use of amperometry with DMF 123 to detect diimines generated from the reaction between horseradish peroxidase (HRP) and 3,3′,5,5′-tetramethylbenzidine (TMB), with a detection limit of 2.4 µIU mL–1. This value is lower than the clinically required cutoff value, but slightly higher than values reported for chemiluminescence detection. There is also the potential for use of electrochemical detection, but in some cases, depending on the application and requirements on strength of the output signal, integration of sensing instrumentation can be complex and expensive.122,104 While DMF-based immunoassays offer the possibility of a generalized low-cost and portable application for the diagnosis of diseases, there exist challenges when examining ways in which detection methods can be integrated with sample handling. The challenges can be due to issues like limits of detection, where insufficient signal molecules are present in the droplet. Difficulties with integrating sensors on a DMF chip are an additional challenge, as some might require transparent surfaces that will affect the substrate used for building the chip. In general, the presence of additional electronics on the device will require further optimization of design, software development, and, in some cases, a reevaluation of the coatings used to ensure droplet processing.
The Ideal DMF Platform
From the examples of automated workflows presented in the previous section, it can be seen that DMF technology is mainly employed to address the following requirements:
Low cost
Fewer errors
Low volume
Reduced starting materials
Reduced processing time
Complex protocols supported
Processing support for different types of samples (blood, serum, human plasma)
Portability
End to end workflow integration
There are only a few overlaps in requirements between the different protocols ( Table 4 ). The ideal DMF platform would tick all the above boxes. Realistically, designing the ideal DMF platform would require concessions and trade-offs based on a balance of priorities. For example, to support a diagnostics assay for rubella testing in a rural setting, a DMF device needs to be low cost, portable, fast, and able to handle different types of samples. Such a device would not necessarily need to support complex ad hoc protocols as the assays are already defined. For compound screening, a DMF device should reduce errors from transfer loss and support low volumes to preserve precious samples. It would not be so important to maintain costs as low as expected for a diagnostics assay. A compound screening device would benefit from the addition of processing modules, such as for purification and electrophoresis, which would make this device not portable. The ideal DMF platform therefore looks different in different applications.
Requirements of Different Workflows That We Discussed in Khilko et al. 66
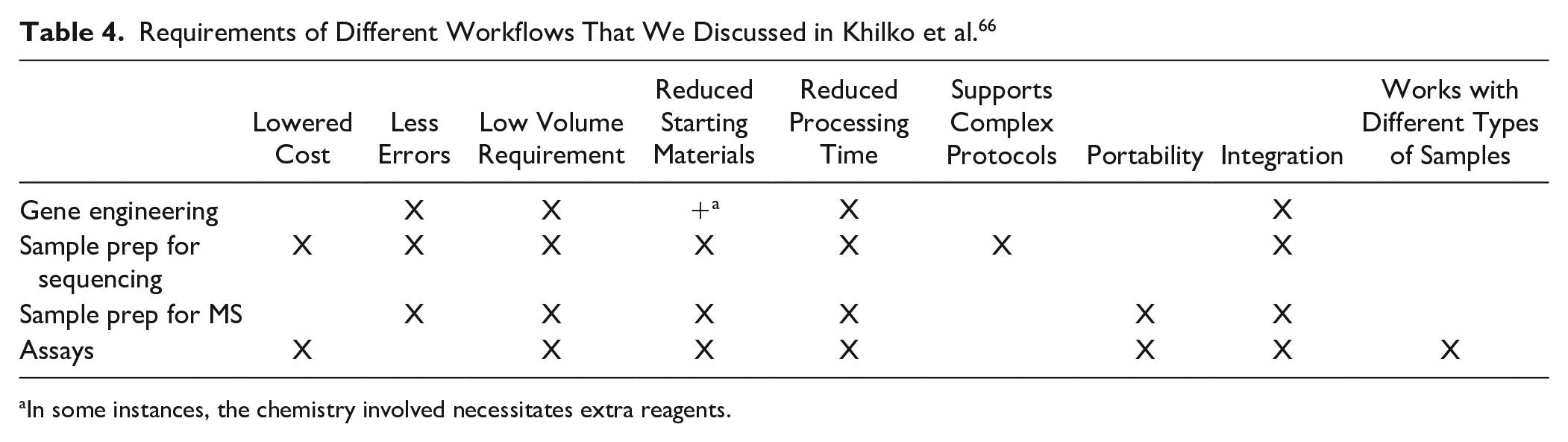

In some instances, the chemistry involved necessitates extra reagents.
DMF-Driven CAB
When employing DMF as a digital bioconverter, ensuring that the DMF device works as part of a DBTL cycle is highly dependent on the target application. For example, for rapid diagnostics of hormonal markers in athletes, the requirements at the “design” and “build” stages are not very high: the tests are already defined, and designing and building the microfluidics protocols would not differ between runs. However, there are many opportunities at the “learn” stage in this context. Given the known differences in upper and lower limits of hormonal levels between sex, race, age, medical history, and so forth, integrating additional “learn” tools could result in more precise diagnoses at the “test” stage, by careful adjustments at the “design” and “build” stages where necessary. An example of such a “learn” tool would be the use of available databases, statistical analysis, and machine learning to determine a preliminary diagnosis, as demonstrated by IBM’s Watson. 124 Meanwhile, heavy requirements for “design” and “build” tools are necessary in discovering novel therapeutics. Here, “design” and “build” tools can automatically generate a construct variant based on screening input from the “learn” stage. There are currently many options available, for example LabGenius’s EVA, 125 which uses machine learning to generate promising new protein therapeutics. Given the vast potential for data led analytics and tools to significantly innovate biological discovery, a digital bioconverter must have the flexibility to rapidly respond to changes at the “learn,” “design,” and “build” stages.
The most important theme in a CAB framework is integration between the DBTL stages. Wang et al. 126 pinpoints in their analysis of the metabolic engineering pipeline that the “test” phase, that is, screening or phenotype evaluation, is considered the major bottleneck in the DBTL cycle. According to Rogers and Church, 2 the number of strains that can be tested is orders of magnitude lower than the strains that can be designed, with design variants produced in the range of billions per day, and test variants in the range of thousands per day. In their paper, the authors suggest that the most impressive gains can be made through multiplexing. Meanwhile, Burgard et al. 127 and Kim et al. 128 both posit that the solution to this bottleneck is the use of an integrated technology platform that connects the entire biomanufacturing pipeline from strain development to fermentation and process engineering. Both approaches are adopted by biofoundries using robotic automation.
Theoretically speaking, DMF technology is entirely programmable and naturally suited to full pipeline integration.
Challenges for DMF in a DBTL Cycle
DMF is known to be associated with certain challenges that impede its widespread use as an automation solution. Although, theoretically speaking, DMF can fit the mold of a digital bioconverter capacity for multiplexing, rapidly respond to digital instructions, and support upstream and downstream integrations, there are some limitations in current versions of DMF, along with potential solutions to improve the applicability of this technology within a DBTL cycle.
Integration Challenges
A major challenge for any automation tool is in integrating with other tools within an automation pipeline. It should go without saying that there is no one-size-fits-all solution in designing a DBTL cycle, and a full platform integration would entail the use of other automation solutions (
Software Integration
A general issue with software integration in a DBTL cycle is in connecting “learn” and “design” tools to automation platforms that will execute the “build” and “test” stages. Connecting “test” and “learn” can be easily done as there are plenty of analysis tools for different applications that are currently available, such as ImageJ 130 and analysis of variance (ANOVA). 131 However, connecting the “learn” and “design” components and the “design” and “Build” components is not common. This is likely due to a lack of standard conversions between these stages. The challenge is in converting insights gained from the “learn” stage into instructions to be executed by the “build” and “test” tool. A solution can be potentially found in the use of machine learning algorithms in other industries, for example:
Generation of recommended treatments from analysis of medical symptoms (MedTech)
Generation of recommended sentencing/settlement based on analysis of legal precedents (LegalTech)
One recent example of connecting the “learn” and “design” stages is Transcriptic’s REST API, 132 which is a software tool for algorithmically driven experimentation and data analysis. A common solution to integration issues between different software tools is the use of open APIs or web hooks. These particular tools do not have widespread use in the life science industry due to proprietary ownership of most available software. However, based on the dearth of open-source design and analytics tools, it can be assumed that if a DBTL approach gains more momentum, this constraint would be lessened.
An advantage in using DMF in the context of software integration is that there is a 1:1 connection between a digital instruction and physical execution. This is in contrast to other automation tools that depend on transferring digital instructions across tools, each with their own conversions. It is hoped that with an increasing focus on the use of a CAB/DBTL approach, this gap can be filled.
Physical Integration
A key challenge with adopting DMF-based tools currently is their nonstandard format, which differs from the most commonly used plate format, like the 96-well plate. This limits integration of DMF tools within existing workflows, which are mostly dependent on a 96-well plate output. Work by Moazami et al. 133 shows the integration of a world-to-chip interface for implementing bacterial transformation and enzymatic assays. Products like Nicoya’s DMF-based tool Alto 134 promise an ability to integrate with existing standard well plate format. SEEKER by Baebies 135 also uses a similar interface to consolidate their screening assay on one device. The interface design depends on the workflow being supported by the platform and the returns that an increase in efficiency can bring to the workflow (reduction in errors, increased accuracy of outcomes, reduction in costs, and reagent usage).
An additional challenge is in ensuring an adequate supply of reagents, a solution for which is shown by Jebrail et al. 136 An excellent review by Hitzbleck and Delamarche 137 shows that reagents can be supplied into and taken out of a microfluidics device in active, passive, and hybrid ways, using dry or liquid forms.
Technical Challenges
Another major concern with using DMF is preventing dielectric breakdown and biofouling. It is difficult to determine a constant solution for both issues, as both are highly dependent on how a DMF device is used (e.g., operating voltage and frequency, type of reagents used). Additionally, there is a large difference between the design of DMF chips between studies. An added dimension to the aforementioned constraint is the question of manufacturability. Given that the phenomena leading to dielectric breakdown are a matter of balancing dielectric thickness, applied voltage, and surface resistance, 138 it is imperative that DMF chips can be manufactured reliably within a set tolerance. Different attempts at quantifying and predicting performance to reduce the probability of dielectric breakdown have been made in previous studies.139–140
Throughput Challenges
Beyond addressing technical challenges of implementing consolidated workflows and configuring protocols for new workflows, a DMF-based automation tool also has to match what users want from their automation tools. The current lab automation market is filled with high-throughput tools from companies like Tecan, LabCyte, Analytik Jena, and High Res Bio, often tailored toward blue chip biopharma research. These tools are geared toward supporting high-volume workflows aimed at processing thousands of samples a day, and the niche nature of this market means that they are significantly more expensive. In addition to that, these are large automated pipelines and their minimum throughput is significantly higher than required by most experiments conducted in the average laboratories. An exception to this is Opentrons, an open-source personal lab robot. Most academic laboratories and small to medium enterprise (SME) research experimental workflows still operate in the low- to medium-throughput regimes, have tougher cost restrictions, and require a higher degree of flexibility (i.e., support for multiple protocols). This makes them ideal users for DMF platforms.
Operational Complexity Challenges
When there is not a DMF product optimized to support a specific workflow, a new user interested in adopting DMF is often faced with the need to have specialist knowledge of several disciplines like electronics, surface chemistry, and software. This is required as there are several parameters of a protocol, reagents employed, coatings used, and electronics that have to be optimized when implementing a new workflow. For example, choosing if a DMF chip with a closed configuration is to be operated with an air-filled or oil-filled configuration, choosing the right filler oils, understanding if there is a requirement for including additional salts and surfactants with the droplet to improve actuation, and managing contact angle saturation 141 determine how well droplets can be transported on the chip. In the event that some droplets contain proteins, workflows could fail due to nonspecific adsorption of the proteins, which makes it very difficult to move droplets and causes biofouling. 13 ,82,142 Additives like Pluronic F127144 and other Pluronic additives with >50% poly (ethylene oxide) (PEO)143–144 can be used to reduce biofouling by affecting the contact angle of the droplet and improving its on-chip actuation. There are also options for including intermediate washing steps using different buffers to reduce biofouling when running a protocol. While it is common to troubleshoot research instruments, an automation tool should require minimal intervention. However, similar to the technical challenges mentioned, operational complexity issues arise due to the large variety of ways a DMF platform is used and the large variations in the specific characteristics of DMF platforms that exist. In this regard, there needs to be a consolidated database that correlates chip characteristics, chip performance, and experiment conditions.
Building mature DMF-based products benefiting from high-quality manufacturing facilities from the electronics industry can be the alternative to specialized, inflexible, high-throughput-only technologies as a flexible means of automation across low to medium throughput at a lower cost and physical footprint. By easing user adoption of automation through well-designed plug-n-play products where users can optimize protocols, have more control over experimental design, design better experiments, and implement workflows that were previously not possible due to constraints of manual experimentation, DMF can further support the increasing standardization in biological experimentation leading to improved outcomes. With this, DMF-based automation tools can drive a wider adoption of automation and become digital bioconverters that make biology more programmable.
Conclusions
Digital microfluidics is a promising technology for the rapid, automated processing of low-volume samples and has a potential to function well in workflows that require the integration of operationally distinct protocols, such as amplification, fluorescence detection, and MS. In this regard, DMF is more than just a liquid handling automation platform.
This potential of DMF as a digital bioconverter within a DBTL cycle merits further consolidated efforts in overcoming known limitations with this technology. Droplet-level addressability, along with the possibility of collecting more information about an experimental execution than possible with other techniques, makes DMF amenable to integration with big data and machine learning-based methods of analysis, which has the potential to vastly improve experimentation and output quality. With increased focus on incorporating the CAB framework in everyday labs, the development of DMF as a technology and as a tool remains a promising trajectory of research.
Supplemental Material
Kothamachu_et_al_SI – Supplemental material for Role of Digital Microfluidics in Enabling Access to Laboratory Automation and Making Biology Programmable
Supplemental material, Kothamachu_et_al_SI for Role of Digital Microfluidics in Enabling Access to Laboratory Automation and Making Biology Programmable by Varun B. Kothamachu, Sabrina Zaini and Federico Muffatto in SLAS Technology
Footnotes
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: All authors of this paper are currently working at Digi.Bio, a company that aims to build digital microfluidics-based platforms to bring laboratory automation to a wider audience, and their research and authorship of this article was completed within the scope of their employment with Digi.Bio.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.