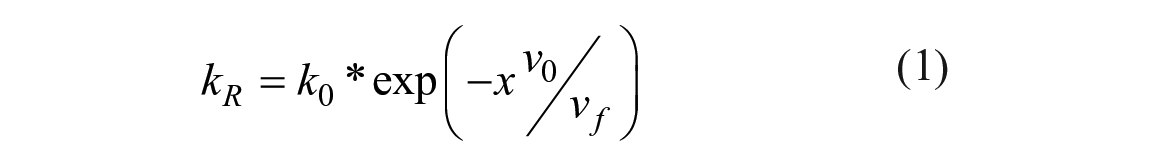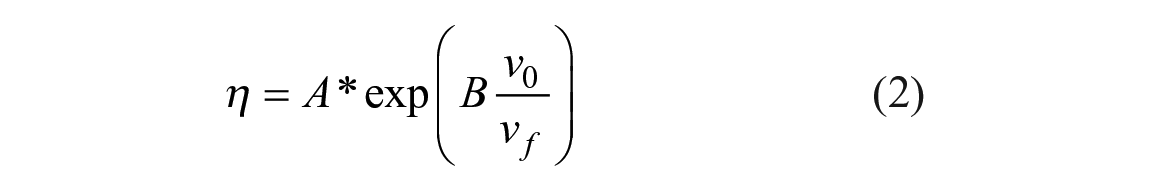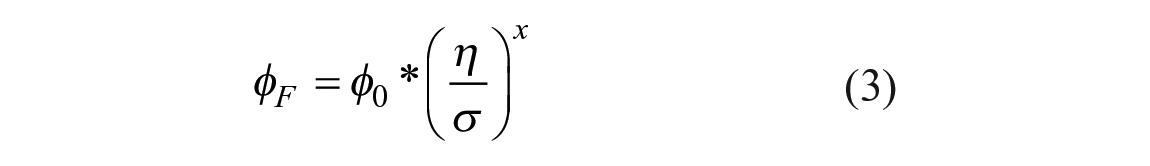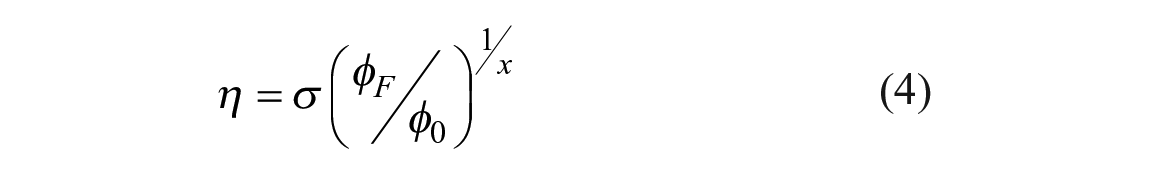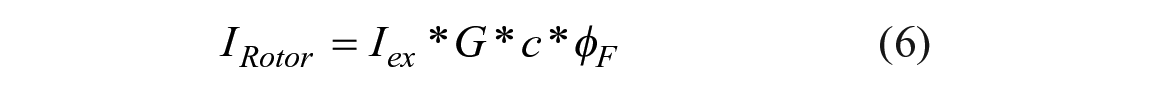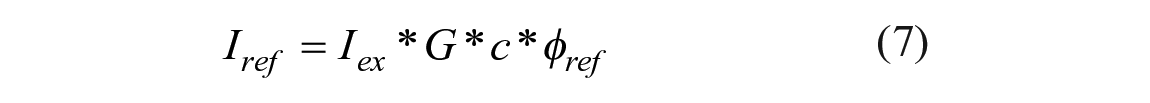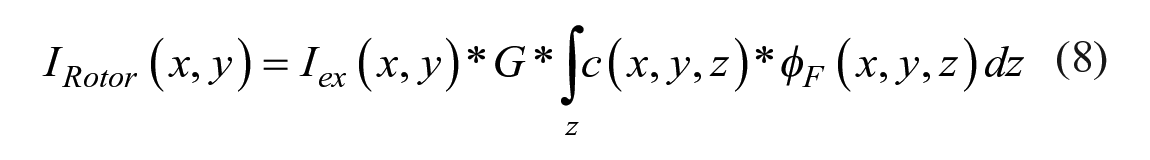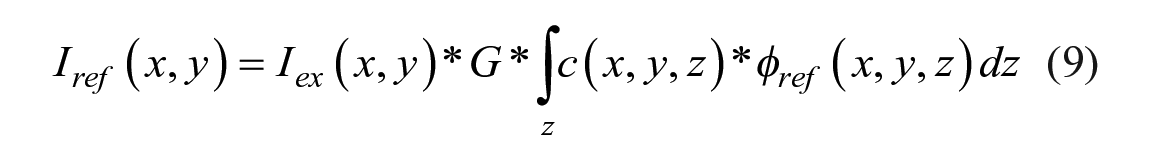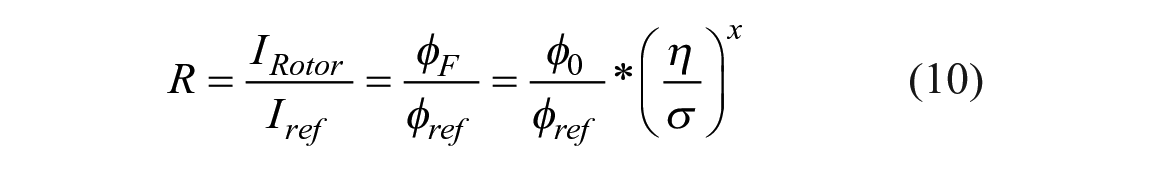Abstract
New insights into the biomechanical properties of cells are revealing the importance of these properties and how they relate to underlying molecular, architectural, and behavioral changes associated with cell state and disease processes. However, the current understanding of how these in vitro biomechanical properties are associated with in vivo processes has been developed based on the traditional monolayer (two-dimensional [2D]) cell culture, which traditionally has not translated well to the three-dimensional (3D) cell culture and in vivo function. Many gold standard methods and tools used to observe the biomechanical properties of 2D cell cultures cannot be used with 3D cell cultures. Fluorescent molecules can respond to external factors almost instantaneously and require relatively low-cost instrumentation. In this review, we provide the background on fluorescent molecular rotors, which are attractive tools due to the relationship of their emission quantum yield with environmental microviscosity. We make the case for their use in both 2D and 3D cell cultures and speculate on their fundamental and practical applications in cell biology.
Introduction
In 1971, Förster and Hoffmann proposed a theory based on the Debye–Stokes–Einstein (DSE) microfriction concept that quantitatively explained the relationship between solvent microviscosity and the emission quantum yield of dyes with the ability to perform intramolecular rotation. 1 At the same time, Rotkiewicz and Grabowsky proposed the hypothesis that 1,4-dimethylamino benzonitrile (DMABN) fluorescence is due to the existence of twisted intramolecular charge transfer (TICT) states, 2 which provides a general model for a family of fluorescent molecules referred to as molecular rotors. Recently, these molecules have gained enormous popularity because of their ability to monitor polymerization processes, 3 image flow patterns in microfluidic systems, 4 and study cell physiology parameters.5,6 Implementation of molecular rotors into the plasma membrane of endothelial cells made it possible to demonstrate that increased fluid shear stress (FSS) increases membrane fluidity in the endothelial cell membrane. 7 Based on these observations, it was hypothesized that changes in membrane fluidity are the primary mechanism of mechanotransduction that converts the mechanical stimulus of FSS into the biochemical signals that have been associated with maintaining a specific level of flow or associated shear stress in blood vessels. 7
Recently, three-dimensional (3D) cell cultures have gained significant attention, and it is now widely accepted that 3D in vitro platforms support more physiologically relevant cell cultures when compared with cells grown as a monolayer (two-dimensional [2D]) on a more traditional substrate. One relevant example involves SH-SY5Y neuroblastoma cells grown on a 3D platform, where voltage-gated calcium channel (VGCC) activation via high potassium depolarization yields a smaller intracellular calcium transient signal in comparison with 2D cultures of the same line.8,9 Kisaalita and colleagues hypothesized that the cell membrane plays a role in those observations as the membrane is thought to be stretched more tautly in a monolayer grown over a rigid 2D platform, opening invaginations in the membrane termed caveolae and exposing more VGCCs to the extracellular environment. At the time, the tools to corroborate this hypothesis were not being explored for this application. Many methods exist that directly or indirectly report membrane physical properties, but these methods are time-consuming and of limited spatial resolution. Molecular rotors lend themselves to the notion of viscosity imaging because they require relatively simple steady-state fluorescence instrumentation, they report changes in real time, and their spatial resolution is only limited by the optical system.
We set the stage by first presenting molecular rotors and their potential for viscosity sensing at the single cell level. This is important given the current understanding of cell–cell physical microenvironment interaction. We review the state-of-the-art physical property measurement techniques and how these methods fall short in 3D culture settings—a gap molecular rotors are hypothesized to close. Finally, we suggest other potential practical applications of molecular rotors beyond viscosity imaging.
The Physical Principle of Molecular Rotors
Single-Emission Molecular Rotors
A typical molecular rotor consists of three subunits: an electron donor unit, an electron-rich spacer unit that is composed of a network of alternating single and double bonds, and an electron acceptor unit (
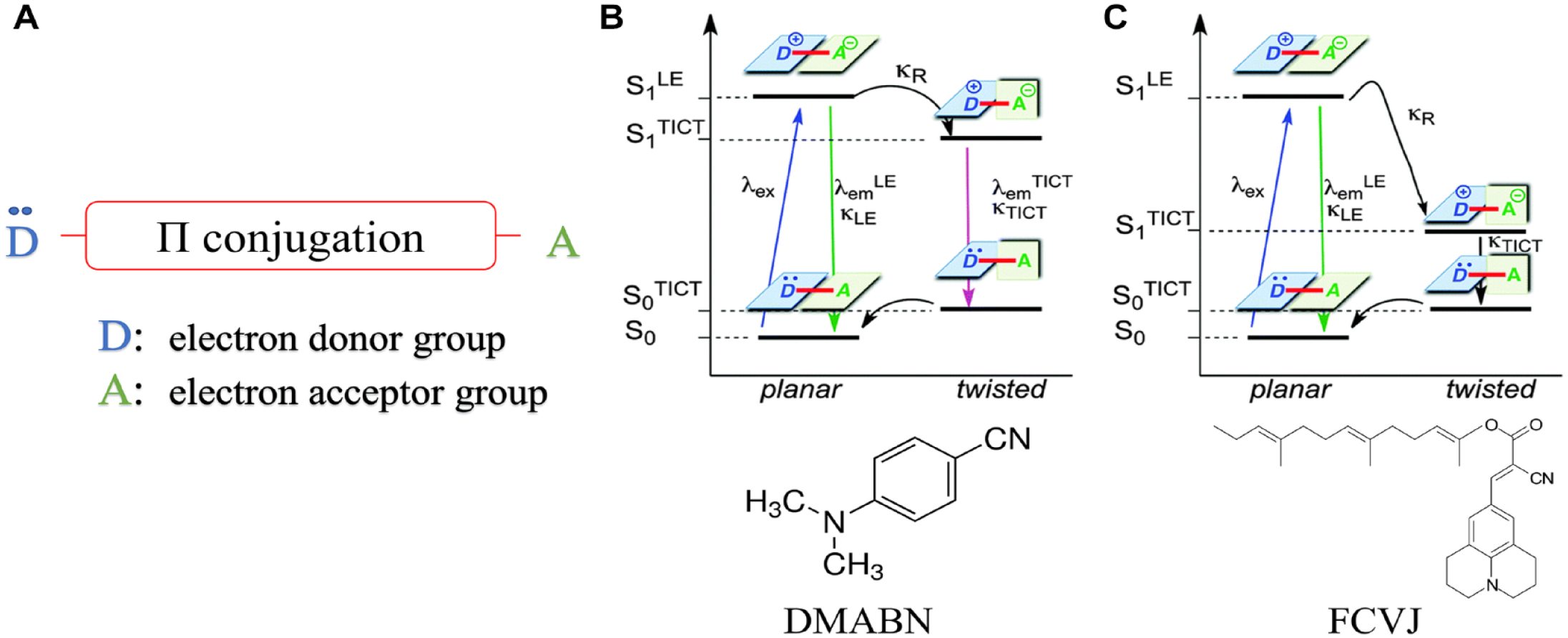
Structure and TICT states and kinetics for molecular rotors. (
For some fluorescent molecular rotors, including the most popular and well-explored molecular rotor, DMABN, relaxation from the twisted conformation to the planar ground state can occur through two channels, resulting in dual emission (
While the reorientation relaxation of molecules in low-viscosity solution follows the DSE hydrodynamic model with a stick boundary condition, Loutfy and Arnold observed deviations from the model in medium- and high-viscosity solvents. 13 They accurately described the nonradiative decay rate (i.e., intramolecular rotation) kR of molecular rotors and its relation to the molecular free volume of these solvents through a free-volume concept:
where k0 is the free-rotor reorientation rate, v0 is the limiting specific volume of the solvent, vf is the free volume, and x is a constant for the particular probe. In aggregates of macromolecules, which include the phospholipid bilayer, molecular free volume and viscosity are directly related. This relationship was experimentally provided by Doolittle 14 and given as
where A and B are solvent-dependent constants (and B ≈ 1).
In solutions with low viscosity, intramolecular rotation is the predominant deexcitation pathway, while intramolecular rotation is impeded in solutions with high viscosity.13,15 Furthermore, the fluorescence quantum yield from the planar excited state increases with increasing solvent viscosity. 11 By recognizing that intramolecular rotation and fluorescence emission are competing deactivation pathways (and thus an increase of kR is associated with a lower quantum yield), substitution of the term v0/vf with eq 2 leads to a form very similar to the Förster–Hoffmann equation:
where φ0 is the dye’s intrinsic quantum yield, σ is a dye-specific constant, and x depends on both the dye and solvent. This relationship holds over several magnitudes of viscosity.
Stretching of the cellular membrane, by FSS, 7 for example, leads to a higher free volume and thus to a lower apparent viscosity. In fact, fluidity (the inverse of viscosity) has been used in the fluid mosaic model of the cell membrane 16 to explain many of the physiological responses of the cell membrane, notably the behavior of membrane-bound proteins. Although amended (e.g., Vereb et al. 17 and Engelman 18 ), the fluid mosaic model is still widely accepted. In this respect, we believe that apparent viscosity (and, related, free volume) serves as a suitable proxy for the freedom of motion of individual phospholipids, membrane-bound proteins, and whole-membrane partitions, such as lipid rafts. The same freedom of motion clearly applies to membrane-bound molecular rotors, which justifies their use as membrane viscosity sensors.
Molecular Rotors with Engineered Dual Emission
It is possible to invert eq 3 and solve for the viscosity:
but the quantum yield is difficult to obtain. One can use time-resolved spectroscopy where fluorescence quantum yield φF and measured lifetime τ are directly related through
where τN is the fluorophore’s natural lifetime (i.e., the lifetime where intramolecular rotation is inhibited and nonradiative deexcitation is absent). Lifetime measurements allow for the sequential calculation of a fluorophore’s quantum yield and the microviscosity of the environment.19,20 However, several molecular rotors have a fluorescence lifetime in the range of 100 ps or less and show a low quantum yield in most solvents. Time-resolved spectroscopy and even fluorescent lifetime imaging (FLIM) depend on expensive equipment, and because the lifetime of many molecular rotors is below the nanosecond range, experiments using lifetime instrumentation can become complex. Steady-state instruments, like microscopes for steady-state fluorescence, are an attractive alternative because they are readily available in most research environments and are simple to use. This was the rationale for the development of self-calibrating dyes that provide an artificial reference,21–23 termed ratiometric molecular rotors. Although many TICT molecules exhibit dual emission (e.g., DMABN), the general design of a ratiometric molecular rotor comprises covalent linking of a reference fluorophore and a molecular rotor (
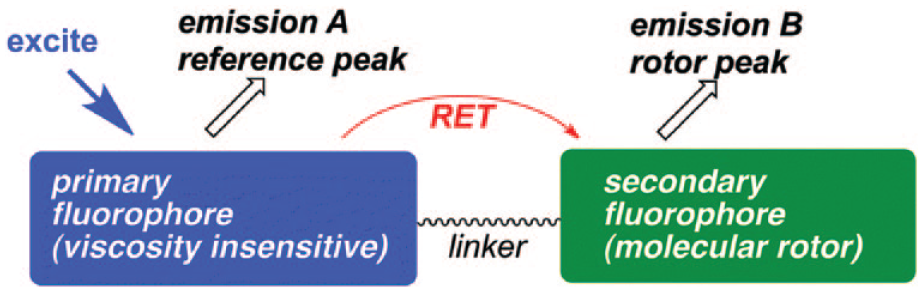
Generic motif of ratiometric TICT nanoviscometers based on the RET concept. The viscosity insensitive reference fluorophore is excited and emits at reference peak A. The reference emission peak A has a significant overlap with the covalently linked molecular rotor excitation spectrum, allowing the viscosity-sensitive molecular rotor to be excited via RET. The molecular rotor emits at rotor peak B, with the emission quantum yield depending on environmental microviscosity. Reprinted from Haidekker and Theodorakis 21 with permission from the Royal Society of Chemistry.
The peak emission intensity IRotor of a molecular rotor with a low dye concentration c can be approximated as follows:
where Iex is the intensity of the excitation light and G is an instrument gain factor that reflects sensitivity, collection efficiency, and signal processing. The same equation is valid for the emission of the reference dye,
where φref is the quantum yield of the reference fluorophore, where usually φref ≈ 1. For fluorescence microscopy of thin layers, eqs 6 and 7 become spatially resolved:
where inhomogeneous excitation signal Iex(x, y) and inhomogeneous dye distribution c(x, y, z) are assumed, and the fluorescence intensity is acquired by a projection along the excitation signal path z. Since the molecular rotor and the reference dye are covalently linked, the local concentration c(x, y, z) should be the same. Additionally, the instrument-dependent factors Iex(x, y) and G, as well as the concentration, cancel out, leaving a ratio R of steady-state rotor to reference emission:
Equation 10 has two implications. First, assuming a known and constant reference dye quantum yield φref, the ratiometric intensity gives access to the molecular rotor’s quantum yield, which is otherwise only obtainable with expensive lifetime equipment. Second, eq 10 can be solved for viscosity η, 23 and the ratiometric intensity R allows direct computation of viscosity where x, σ, and φ0 can be considered calibration constants and be obtained from model solvents. To keep the emission spectra of these dual-emission molecular rotors as simple as possible, a molecular rotor with a single-emission band is preferred. This allows the ratio R to produce a concentration-independent self-calibrating measurement of the environmental microviscosity.
Molecular Rotors as Viscosity Sensors
Molecular rotors have been used to quantitatively determine phospholipid microviscosity 12 and observe temperature-dependent viscosity changes of phospholipid bilayers and their sol-gel transition.24,25 In addition, molecular rotors have already been used in cell cultures to elucidate the influence of FSS on endothelial cell membranes. 7 In the context of Kisaalita and colleagues’ main hypothesis that SH-SY5Y neuroblastoma cells exhibit physiologically abnormal membrane stretch when cultured on glass (2D),8,9 membrane stretching and the increase in free volume are quite intuitive, which links stretching back to the free volume discussion around eq 3. The recent development of ratiometric nanoviscometers also presents new opportunities for studying the effects of a cell’s microenvironment on membrane stretch. Using membrane viscosity as a proxy for membrane stretch, with the proposed ratiometric molecular rotors, it will not only be possible to explain the calcium signal difference between 2D and 3D neuroblastoma cultures, but also for the first time enable the examination of the membrane viscosity of a cell embedded in a 3D microtissue.
Cell–Cell Physical Microenvironment Interaction
It is well established that cells continuously “sense” the mechanical properties of their environment. Normal physiological functions, including morphogenesis, 26 tissue remodeling,27,28 and differentiation,29,30 are driven in part by the biophysical microenvironment. Cells use their adhesive and contractile molecular machinery to transmit forces to their surroundings and then transduce those forces to biochemical signals. Fibroblasts have been shown to adapt their internal stiffness to match that of their substrate for a stiffness range spanning that of soft tissues (between 800 Pa and 4 kPa) before reaching a saturation limit (>10 kPa), with an increasing cell stiffness strongly correlating to increased cell area. 31 This suggests that within the stiffness range in which native fibroblasts reside, a cell can quantitatively adapt and match its mechanical environment. Similarly, epithelial cells grown on compliant substrates were less spread and contained irregular ruffles on the ventricle surface. 32 These observations point to two different states of cell adhesion and spreading: stiff substrates result in remodeling of the actin cytoskeleton into stress fibers and increased cell area, and soft substrates on which the cell does not spread out and stress fibers do not exist. Alternatively, motor neurons derived from mouse spinal cords adopt extensive neurite branches on soft but not stiff substrates, 33 while some cell types cannot grow on soft (<50 Pa) substrates.32,34–36 It is clear that not all adherent cell types respond to biophysical cues from the cellular microenvironment in the same manner.
Although the regulation of focal adhesions and the cytoskeleton has garnered a lot of attention, the plasma membrane is far from a passive participant in mechanotransduction. In addition to maintaining a specific level of blood flow and associated shear stress,
7
changes in membrane viscosity and activation of flow-sensitive ion channels facilitate vascular endothelial wound closure under FSS.
37
Membrane viscosity (or membrane fluidity, which is its reciprocal) describes the ease of movement within the phospholipid bilayer, and directly influences physiological properties such as membrane-bound enzyme activity, carrier-mediated transport, and membrane-bound receptor binding.
38
The diffusion rate of molecules and therefore the reaction rates of diffusion-controlled processes are also affected by membrane viscosity. The rate of mass transport of reagents within and across the domains in heterogenous systems like the lipid bilayer are often determined by diffusion-controlled processes, and many complex intracellular processes rely on diffusion as a rate-limiting step. In this sense, the dynamic state of the lipid bilayer is akin to other characteristic properties, such as gene and protein expression, which are used to phenotype cell populations. An open question arising from this collection of observations is how substrate stiffness affects membrane viscosity. Although flattening of membrane ruffles in response to the application of linear strain has been demonstrated,39–41 a link between membrane viscosity and the observable changes in cell area because of tactile sensing of substrate stiffness has not been conclusively established. Intuition tells us that when cell area increases in response to a stiff substrate, the plasma membrane adopts a physiologically abnormal stretched or “taut” condition (
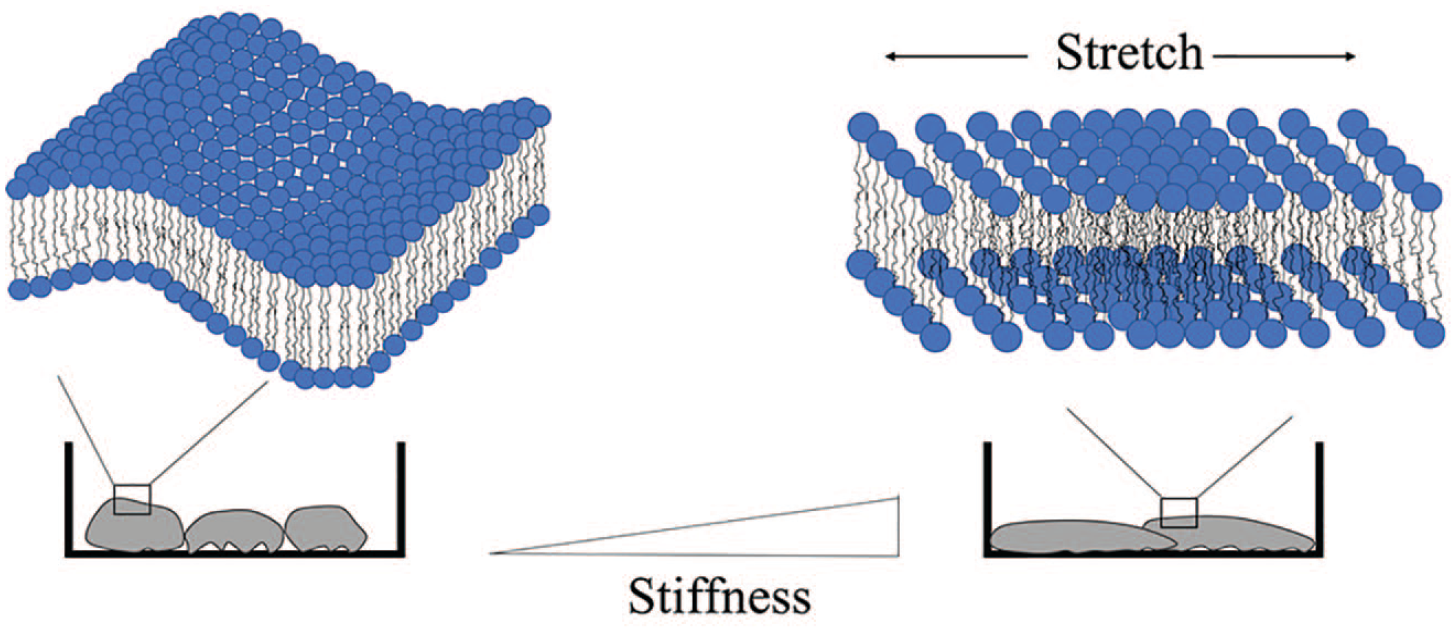
Differences in substrate stiffness have been linked to changes in cell shape, 82 and the bilayer membrane must accommodate these changes. Passive remodeling of the membrane includes flattening of membrane ruffles in response to cell stretch, 39 which we hypothesize is associated with an increase in macromolecular free volume.
Measuring Membrane Physical Properties
Atomic force microscopy (AFM) is currently the gold standard to measure the mechanical properties of cells with a resolution down to nanometer scales. However, differences in assumptions and constitutive models as well as experimental setups, such as cantilever characteristics and probe settings, 45 make a direct quantitative comparison across studies using AFM infeasible. Probing individual cells is time-consuming, and measuring live cells can be problematic if the ability to control environmental parameters (i.e., temperature and percent CO2) is not available. AFM, along with optical tweezers and micropipette aspiration, is an invasive technique limited to surface topologies and is not a direct method for probing membrane viscosity; rather, it measures the “deformability” by applying nanoindentation, shear stresses, and pressure gradients that may elicit an unintended cellular response. The result is the sum of the force needed to deform the membrane (in-plane membrane tension) and the force needed to deform membrane-to-cytoskeleton attachments and membrane proteins. While AFM is typically used to test cells adhered to a surface, micropipette aspiration usually measures cells in suspension. Therefore, cytoskeletal differences associated with adherence or suspension significantly influence the measured properties; adhered cells measured by AFM appear to be less viscous than those in suspension measured by micropipette aspiration. 46
Noninvasive elastography techniques like fluorescence recovery after photobleaching (FRAP) overcome this issue by indirectly reporting membrane viscosity through diffusivity of a fluorophore; however, these techniques are time-consuming and of limited spatial resolution. Depending on instrumentation, a FRAP image may take up to 30 min, including a 6–9 min photobleaching process 47 during which dye will begin migrating into the irradiated spot, creating a zone of reduced dye concentration around the bleached zone that reduces dye recovery. While molecular rotors report free volume in the immediate surroundings, FRAP reports diffusivity over a relatively large area (e.g., 30 µm in the case of Majd and Mayer 47 ).
The notion of viscosity imaging lends itself as a possible alternative method. Molecular rotors require relatively simple steady-state fluorescence instrumentation and report changes in real time, and their spatial resolution is only limited by the optical system. Similar to noninvasive elastography techniques, molecular rotors do not deform the plasma membrane. The result is a direct measurement of membrane microviscosity, which can be reported in viscosity units (mPa·s). 23 The changes of membrane microviscosity can then be interpreted with respect to the testing parameters, such as substrate stiffness and FSS. If membrane microviscosity can be used as a proxy for membrane stretch, it should be possible to establish not only a relationship between membrane microviscosity and the cell stiffness as measured by AFM, but also a relationship between membrane microviscosity and substrate stiffness.
Adding a Third Dimension
It has become widely accepted that 3D platforms support more physiologically relevant cell cultures when compared with cells grown on a simple 2D substrate, because 3D platforms better resemble the natural microenvironment of the cells.48,49 As one example, human hepatocarcinoma (HepG2) cells cultured in alginate hydrogels better match in vivo tissue samples with respect to morphology, gene expression, and metabolic activity. 50 Hepatocytes are widely used in high-throughput drug screening (HTS) platforms for drug discovery, and evidence points to a more physiologically relevant cell culture providing more physiologically relevant drug responses. “Physiologically more relevant” claims are often made if the phenotypic characteristics attributed to the culture platform are different between 2D cell cultures and cells grown on a platform that provides a loosely defined 3D architecture. Apart from the 3D matrix adhesion 51 as a possible “biomarker” for the three-dimensionality of a cell culture, research in cell biology has not provided a foundation on which these physiologically more relevant claims could be validated. Lai and colleagues make the case for cytokines to be the chemical cues monitored to determine the physiological relevance of a platform, 52 but the variability and role of cytokine concentrations in 3D cultures are not well understood. In the absence of a biomarker of three-dimensionality (i.e., a quantifiable entity), the optimum composition of the 3D microenvironment will remain elusive. As such, a consensus for the three-dimensionality of a cell culture should be established.
The microenvironmental factors (MEFs) that support the formation of physiologically relevant 3D cultures (microtissues) have been defined as the following: (1) chemical or biochemical composition, (2) spatial cues (geometric 3D), (3) temporal cues, and (4) substrate biophysical properties.53,54 Many different 3D cell culture platforms provide different combinations of MEFs but elicit similar functional and/or structural responses from various cell lines. In fact, many of these cell culture platforms are commercially available and are advertised as 3D. For instance, HepG2 cells cultivated in Matrigel (Corning Matrigel Growth Factor Reduced Basement Membrane Matrix, Corning, NY), collagen sandwich (Corning Collagen I), and Alvetex (porous polystyrene scaffolds) formed structural bile canaliculi networks, a characteristic associated with hepatocytes in vivo. 55 Interestingly, these platforms have been described in literature as 3D but are very different in nature. The collagen sandwich consists mostly of one protein element, while Matrigel is a mixture of laminin, collagen, fibronectin, and numerous other components. Unlike hydrogels, synthetic polymers like polystyrene have Young’s moduli above the physiological range of native liver tissue and provide a rigid pore structure, preventing tissue growth past the size of the pores. To complicate things further, Luckert et al. 55 suggest that the comparable metabolic activity of 2D and 3D HepG2 cells might be a consequence of the duration of cultivation, not the result of 3D culture as previously reported. 56 Based on the evidence from literature, we should ask the following questions: What is the minimum level of exogenous MEFs that engender an in vitro tissue model that best emulates the desired in vivo structural and/or function features, and what is the optimum composition of the culture platform to promote this? A precise emulation of in vivo conditions might be necessary for many regenerative medicine applications. In turn, evidence suggests it might not be necessary in the field of drug discovery.55,56 Conflicting reports on the importance of providing a 3D architecture for hepatocyte cultures used as a liver model for toxicity studies, and their associated metabolic activity, bring to light the potential redundancy in MEFs for certain applications. The identification and elimination of redundant MEFs may make cell culture platforms more predictive, compatible, and affordable for research as well as HTS instrumentation.
The widely accepted consensus is that in order to achieve in vivo-like microtissues in vitro, the stiffness of a 3D culture platform should be within the physiological range of the native tissue of interest. This consensus is based on cellular responses to 2D substrates, as described in the “Cell–Cell Physical Microenvironment Interaction” section, and studying single cells embedded in a 3D matrix. However, the behavior of isolated cells grown on a 2D substrate or within a 3D matrix might differ from the behavior of an individual cell within a tissue. Maintaining cell–cell adhesion is crucial for tissue morphogenesis and homeostasis, while perturbation of the keratin–desmosome complex, to name just one example, severely compromises cell and tissue integrity.57,58 Furthermore, Doyle et al. 59 suggest that the average stiffness of a 3D collagen gel obtained using bulk rheological measurements does not indicate the local differences in fiber stiffness a cell may “sense” at the fiber/cell adhesion scale. A better understanding of how the stiffness of various 3D culture platforms affects microtissues growing within the platform could provide valuable insight into potential redundancy of substrate biophysical MEFs.
While single endothelial cells “sense” the pliability of their substrate and respond with cytoskeleton remodeling and morphology changes, an endothelial cell monolayer grown to confluency has indistinguishable morphologies on both pliable and rigid surfaces. 60 A possible explanation for the lack of morphological changes mentioned above comes from the “cell-on-cell” hypothesis, 29 where cell-to-cell contacts appear to influence the cellular response to the mechanical properties of the environment. For example, when cultured on stiff substrates, myotubes form stress fibers and strong focal adhesions, suggesting a state of isometric contraction; on soft substrates, actomyosin appears scattered. Both cellular responses are consistent with other 2D mechanotransduction observations discussed in the “Cell–Cell Physical Microenvironment Interaction” section. Moreover, when a gel matching relaxed muscle bundles (E ≈ 10 kPa) is used as a substrate, definitive actomyosin striations are observed. The cell-on-cell hypothesis comes into the picture when myotubes are cultured on top of an initial myotube monolayer. While the lower myotube monolayer attaches strongly to glass and forms numerous stress fibers, the upper myotubes differentiate to the more physiological state, with prominent actomyosin striations and cell elasticity in the range of normal muscle tissue. 29 Similarly, a basal layer of astrocytes cultured on glass seems to impart a pliable brainlike environment for the branching of neurons to grow. 61 These observations suggest that these cell-to-cell contacts provide a soft “barrier” between the cell being studied and the substrate, which lends credibility to the concept that physiologically relevant microtissues can be cultured on substrates that have a physiologically abnormal stiffness.
The hypothesis provided by Kisaalita and colleagues to explain differences in high potassium depolarization-induced calcium entry for SH-SY5Y neuroblastoma cells grown on 2D substrates or as 3D microtissues is interesting in the context of the cell-on-cell hypothesis. In these experiments, the same substrate materials (poly (
Concluding Remarks
The focus of this review has been on the application of fluorescent molecular rotors to basic 2D and more complex 3D cell cultures for studying the effects of biophysical signals on cell state. In addition to the fundamental interest, there is a practical need to quantitatively measure changes in cell mechanics. Cell culture applications of molecular rotors are not just limited to the study of physical properties but also have the potential to be used as a tool for studying stem and cancer cells. Evidence from literature shows that stem cells of various lineages and cancer cells with different levels of invasiveness have significant differences in cell physical properties, and there remains a need to identify cells within heterogenous populations.
It has been suggested that cancer cell stiffness plays a role in the invasion of surrounding tissues. Although cancer tissues have shown to be stiffer than normal tissues, recent reports suggest that metastatic cancer cells themselves are more compliant than normal cells.62,63 Swaminathan et al. demonstrate that the stiffness of patient tumor cells and cancer cell lines inversely correlates with migration and invasion through 3D basement membranes. 64 While a better understanding of the onset and progression of cancer can lead to improved detection and more efficacious anticancer drug testing, the potential for single-cell mechanical properties to be used as a biomarker for disease is promising. If a change in mechanical phenotype is an indicator of metastasis, then molecular rotors could be used to study the utility of various cancer cell lines as in vitro cancer models. Molecular rotors are an attractive tool for mechanical property-based screening because, unlike benchtop methods like AFM and micropipette aspiration, they have the potential to be translated to an off-the-shelf assay that can be applied to fluorescence-activated cell sorting (FACS) instrumentation. Other potential applications include combination with microfluidic devices to study single-cell mechanics during migration and intravascular adhesion of metastatic cell lines.
Studies of single-cell mechanical properties for adipose-derived stem cell (ASC) enrichment has shown the potential for distinguishing differences in cell subpopulations, disease state, and tissue source based on mechanical properties.
65
As one example, the cellular biomechanics are remodeled during osteogenic differentiation, which may precede upregulation of the osteogenic markers.
66
Membrane viscosity of undifferentiated stem cells could be correlated with lineage-specific metabolite production, potentially providing a more complete understanding of differentiation. While fundamental stem cell research aims to understand differentiation patterns, clinical applications of stem cells have shown great potential as a novel therapeutic alternative for musculoskeletal, hematopoietic, neurological, and cardiovascular disorders.67,68 Autologous ASCs are just one attractive cell source for cell-based regenerative therapies due to their immunogenicity and multipotent characteristics and are available in large quantities compared with mesenchymal stem cells (MSCs) derived from other tissues. However, adipose tissue contains multiple cell types and only a portion of MSCs satisfy the stem cell criteria, while others are more committed to a specific lineage.69,70 Furthermore, the long-term and large-scale in vitro adherent culture expansion of MSCs necessary to obtain a clinically relevant number of cells for therapeutic use results in reduced multilineage potential.71,72 Although studies suggest the potential for cellular mechanical properties as a novel “biomarker” of cell phenotype, quantitative comparisons across different studies have proven difficult. Mechanical properties within a specific cell lineage have varied by up to an order of magnitude between different research groups, as shown in
Elastic Moduli of Several Stem Cell Lineages Measured by Common Techniques.

TCP, tissue culture plastic.
In vitro cell culture has evolved into complex systems, which require new tools to study cell phenomena. Cell physical properties have become a popular “marker” for cell physiological studies, but limitations in instrumentation make it difficult to study this in complex systems. Molecular rotors lend themselves to the notion of viscosity imaging, which could prove to be a useful alternative. This tool provides the opportunity to study complex 3D cell systems and has the potential to become a useful approach to sorting heterogenous cell samples.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.