Abstract
Cell lines are essential models for biomedical research. However, they have a common and important problem that needs to be addressed. Cell lines can be misidentified, meaning that they no longer correspond to the donor from whom the cells were first obtained. This problem may arise due to cross-contamination: the accidental introduction of cells from another culture. The contaminant, which is often a rapidly dividing cell line, will overgrow and replace the original culture. The end result is a false cell line, also known as a misidentified or imposter cell line. False cell lines may come from an entirely different species, tissue, or cell type than the original donor. If undetected, false cell lines produce unreliable and irreproducible results that pollute the biomedical literature and threaten the development of reliable drug discovery and meaningful patient treatments.
The goal of this study was to ascertain how widespread this problem is and how it affects the literature, as well as to estimate how much funding has been used to produce pools of scientific literature of questionable value. We focus on HEp-2 [HeLa] and Intestine 407 [HeLa], two false cell lines that are widely used in the scientific literature but were shown to be cross-contaminated in 1967. These two cell lines have been used in 8497 and 1397 published articles and extensively described as laryngeal cancer and normal intestine, respectively, rather than their true identity: the cervical cancer cell line HeLa. Discussed are tools, approaches, and resources that can address this issue—both retrospectively and prospectively.
Introduction
Consequences of Using False Cell Lines: Pollution of the Scientific Literature
The ideal cell line is a pure culture of genetically, epigenetically, and phenotypically stable cells that are used to mimic an organism, a tissue, or a biological process. Cell lines are an immensely valuable tool for studying biological processes. Laboratories use cell lines for basic studies of enzymology and molecular biology, to understand interactions between microbes and animal cells—including the infection of cells by pathogenic organisms—and to understand the mechanisms of cancer. Some of the findings presented herein were described earlier. 1
Past History: Many Cell Lines Were Cross-Contaminated
In 1951, George Gey established the first human cell line, HeLa. 2 The cells were quickly put to use for testing of the Salk polio vaccine, 3 but the process used to establish the cell line was equally important, teaching the research community how to establish and culture human cell lines. Unfortunately, many of the early cell lines were plagued by cross-contamination with microbes and both interspecies and intraspecies contaminants. Such contaminations undermined the utility of a cell line as a model of specific biological processes and the reproducibility of published findings and conclusions. These problems have contributed to a “reproducibility crisis”: an inability to replicate many of the reports within the scientific literature, which hampers progress in our understanding of different biological processes.4–6
After Gey’s initial success, there was a steady rise in the establishment of new cell lines. Unfortunately, many of these “new” cell lines turned out to be derivatives of HeLa cells,7–12 a fact unrecognized by many researchers, even to this day. These cells are hardy, grow vigorously, and can outcompete many other cell lines in culture. They can even withstand being dispersed in aerosols and by other mechanical means. 13 The difficulties encountered in handling HeLa by safe and sterile means, and the risk of infection from microbial contaminants, led to the development of biological safety hoods with laminar flow ventilation.14,15
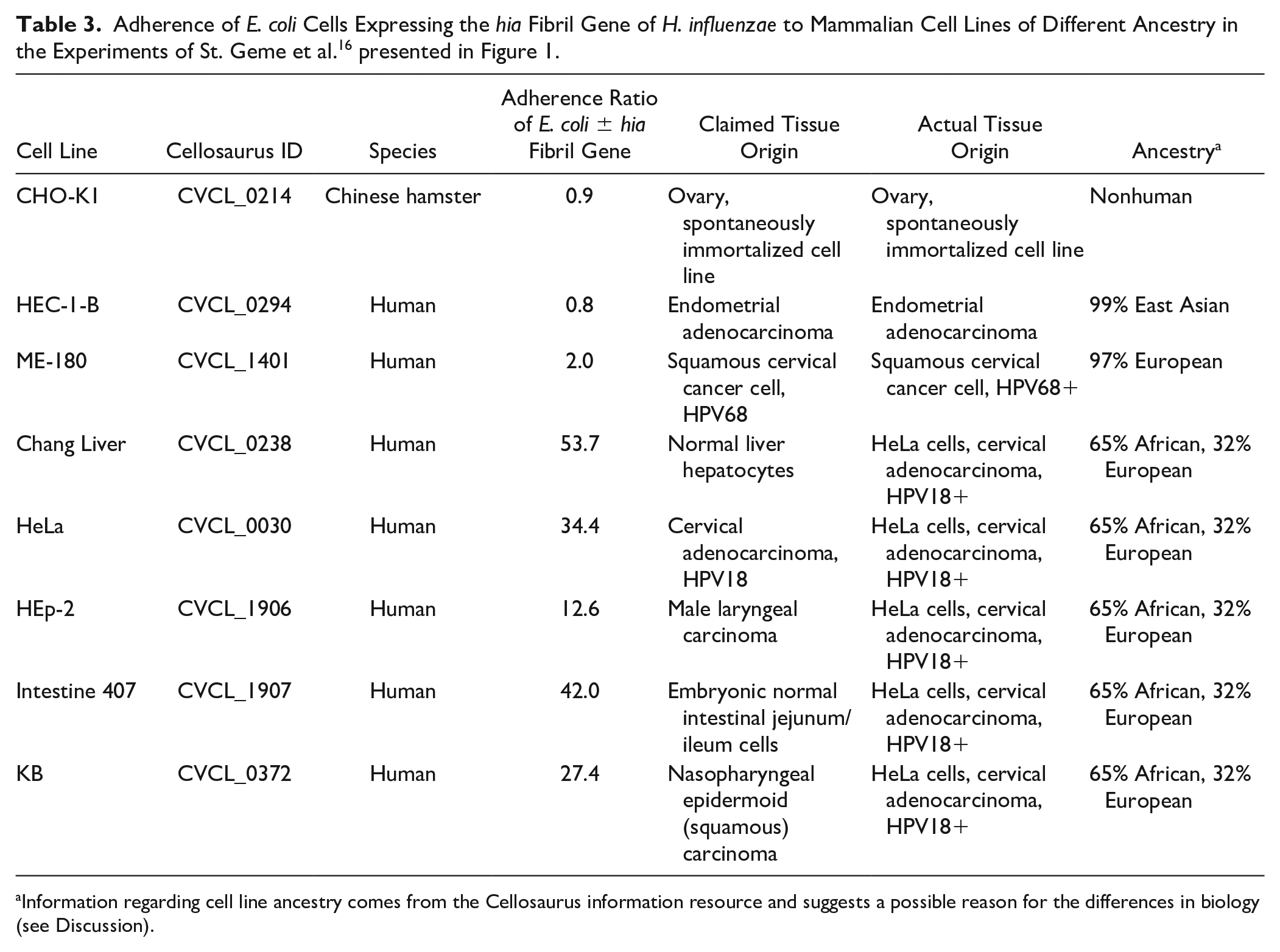
Researchers commonly use panels of multiple cell lines to demonstrate that a biological response either is shared by cell lines from different tissues or is unique to a specific tissue. This approach can be used to show the impact of using false cell lines in publications. For example, a 1996 publication studied how Haemophilus influenzae uses fibrils to attach to human cell lines. 16 The authors examined how Escherichia coli transformed with the H. influenza hia gene could attach to multiple cell lines, as illustrated in Figure 1 . The five cell lines to which the transformed E. coli bound were HeLa (RRID:CVCL_0030), Chang Liver (RRID:CVCL_0238), HEp-2 (RRID:CVCL_1906), Intestine 407 (RRID:CVCL_1907), and KB (RRID:CVCL_0372). All were originally reported to come from different tissues, as listed in Table 3. However, all five of these cell lines were actually derived from HeLa, as reported by Stanley Gartler in 1967.7,8 A better interpretation of these experiments would be that the hia-encoded fibril protein stimulates binding to the cervical cancer cell line HeLa and its derivatives, and not necessarily to the tissues that the different cell lines purportedly represent. It should be noted that the transformed E. coli did not bind to the two other human cell lines used in these experiments, HEC-1-B (endometrial carcinoma; RRID:CVCL_0294) and ME-180 (squamous cell cervical carcinoma; RRID:CVCL_1401).
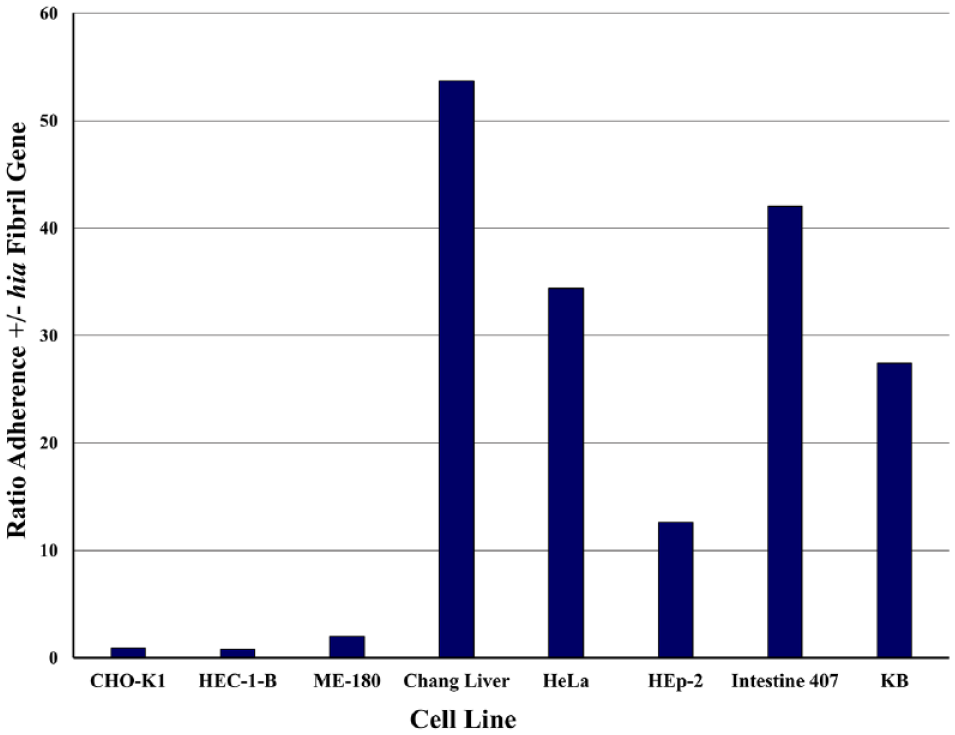
Ratio of adherence of E. coli that expressed the hia fibril gene to human cell lines. Based on the data of St. Geme et al., 16 the change in the number of adhering E. coli bacteria carrying a plasmid vector with the hia fibril gene relative to bacterial cells carrying only the empty vector is shown. The ratios are highest in HeLa and its derivatives, demonstrating how using multiple misidentified cell lines derived from the same cell line does not convey the generality of a phenomenon, that is, bacterial adhesion to human cells of different tissue types.
Present Day: Cross-Contamination Persists and the Problem Is Worsening
The problem of cell line cross-contamination persists to this day and is growing in frequency and seriousness, despite long-standing efforts to raise awareness and propose solutions.7–12,17–23 The proportion of false cell lines reported in the literature ranges between 10% and 100%,24,25 even when collections are studied where those cell lines were deposited by their originators. The International Cell Line Authentication Committee (ICLAC) maintains a Register of Misidentified Cell Lines (https://iclac.org/databases/cross-contaminations/).
21
The latest version of the ICLAC Register (released in June 2021) lists 531 cell lines that are known to be misidentified, with no authentic stocks. However, the ICLAC Register only lists cell lines where there are sufficient data to draw a firm conclusion. Many other cell lines are candidates to be added to the ICLAC Register but have insufficient data or are awaiting review. A 2018 analysis by Korch and Varella-Garcia (
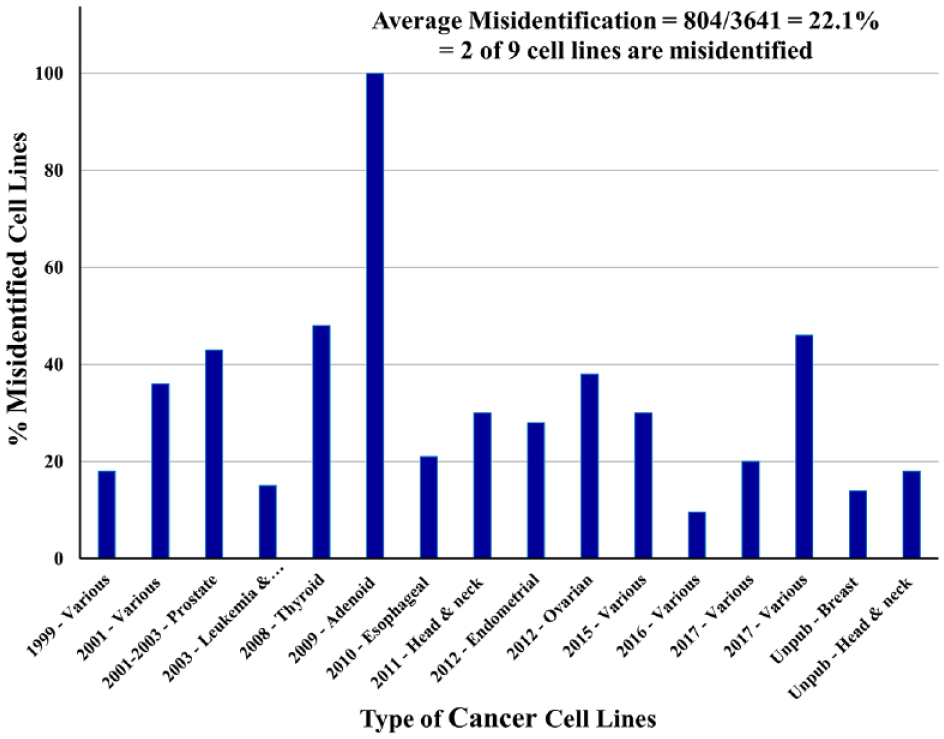
Incidence of misidentified cell lines from 16 published and unpublished studies. These data were reported earlier in a tabular format. 24 Of 3641 cell lines that were characterized primarily by STR genotyping, 804 were misidentified, giving an average incidence of 22.1%, or 2 of 9 cell lines.
Although cross-contamination was originally considered a “HeLa problem,” it can affect any cell line used in a laboratory and the contaminant can be any rapidly growing cell line. For example, M14 is a hardy, rapidly growing cell line derived from cutaneous melanoma obtained from a male patient, and it has contaminated different cell lines that were used for studying thyroid cancer, 26 female breast cancer,27,28 and uveal melanoma.28,29 This finding questions the validity of multiple publications based on these derivative cell lines. 28
False cell lines are being used with their incorrect identities long after they were shown to be misidentified. Vaughan et al. 30 studied the use of the HeLa-derived cell line KB with different descriptions in publications from 2000 to 2014. They found 631 publications in which the authors clearly ascribed an identity to the KB cell line. Of these, 574 described the cell line identity incorrectly as being from an oral epidermoid carcinoma, and only 57 described KB correctly as being a derivative of HeLa cells or from a cervical carcinoma. The continued use of the original description may be due to differing interpretations of the terms “contamination” and “cross-contamination.” For example, HeLa-derived cell lines have been described in the American Type Culture Collection (ATCC) catalog as “contaminated,” which has been interpreted as being a mixture of two different cell lines.31–33 However, these terms were meant to convey that the original culture was “contaminated” during its initial culturing with another cell that took over the culture and became the only component in the culture. Other terms, such as “false,” “misidentified,” or “imposter” cell lines, are less likely to cause confusion.
The continuous usage of false cell lines is infuriating given the current availability of resources that can be used to address the problem. Such resources include the ICLAC Register of Misidentified Cell Lines 21 and the genotype-based techniques that are now available to authenticate human cell lines. Researchers are advised to verify the authenticity of their cell lines by checking them against the ICLAC Register and confirming their identity and purity using short tandem repeat (STR) genotyping. STR genotyping before and after completion of the study must be a prerequisite for publishing the research results in the journals. This would dramatically reduce the number of publications with false cell lines.
To understand the current usage of false cell lines, we selected two false cell lines, Intestine 407 [HeLa] and HEp-2 [HeLa], for analysis. These were both established in the 1950s and were extensively used as models for normal intestine and laryngeal cancer,34–39 respectively, before found to be misidentified in 1967. Both cell lines were included as derivatives of HeLa in the ICLAC Register in 2010 based on STR genotyping. We evaluated usage of these two false cell lines using four databases of publications and supplementing these findings with searches in the websites of Google Scholar, Web of Science, and PLoS.
Materials and Methods
Search Methodology to Explore the Extent of the Cell Line Contamination Problem
We assembled a data set of publications using Intestine 407 [HeLa] (RRID:CVCL_1907) and HEp-2 [HeLa] (RRID:CVCL_1906) from various databases. Henceforth, the cell line names will not include the RRID identifiers for simplicity. PubMed was the primary source of our findings but was not as searchable as some other databases—neither when looking for specific cell lines nor for their use in older references. Recently (April 2020), it was greatly improved by showing the textual context of how the search terms appear in the search results within the information that is a component of each entry, that is, title, authors, abstract, and some limited amount of information in the medical search heading (MeSH) terms. The latter includes only a limited number of cell lines. This search engine does not examine the text of the whole article, where in many cases the cell line name is found in the Materials and Methods section. PubMed Central can search in the full text of those articles to which it has access, but it does not report the textual context where the search term is located. In many cases, the search terms were only listed in the title of a reference in the article and one had to search the whole article before learning this.
The HighWire publication database at Stanford University (which could include PubMed data) was useful but did not include several major journals (e.g., Cell, Science, Nature) until the last year or two of operation. Unfortunately, HighWire ceased to operate at the end of 2014. It did provide the textual context of the search terms in the whole body of the article. The HighWire database had an error rate of approximately 1% in its entries (e.g., two publications accidently merged into one). Its search tool found articles where the search terms were included in the references, which was not useful, but from the surrounding text in the output one could discern whether this was the case.
The American Society for Microbiology (ASM) database of its publications and the Elsevier database Embase were also useful sources of citations using cell lines. These two databases included searchable copies of many earlier publications, making it easier to identify the articles using specific cell lines prior to 1985. Embase provides extensive lists of index terms (i.e., keywords), which usually include the cell lines used in a report. Using Embase, we were able to find very recent (2021) papers that describe HEp-2 as both a head and neck squamous cell carcinoma and a larynx squamous carcinoma; these incorrect descriptions were both included in Embase’s index terms.
Results
On reviewing the extensive literature in which Intestine 407 [HeLa] and HEp-2 [HeLa] have been used, we discovered various ways in which these false cell lines are used in publications. Some articles cite the purported origin of a cell line and use the line to model a specific type of tissue but do not acknowledge that the cell line is actually derived from HeLa cells or a cervical carcinoma, even though this fact has been known since 1967.7,8 Other articles do not cite the tissue of origin (at least as discernible by various complex Boolean search algorithms [see Supplemental Information for examples used to search PubMed, Embase, and other databases]), nor do they cite that the cells are derivatives of HeLa cells. A third type of article gives the cell line a nonstandard name, which makes it difficult to identify its provenance. Only a handful of studies documented the identity and purity of the cell lines they have used.
Publications Using the HeLa Derivative HEp-2 [HeLa]
The cell line HEp-2 [HeLa], originally called H.Ep. #2, was first described by Toolan 39 and colleagues 40 as being derived from human epidermoid cancer cells (now called squamous cell carcinoma) from a laryngeal carcinoma, taken from a Caucasian male. This cell line has appeared in the literature as HEp-2, HEp 2, Hep2, Hep-2c, Hep 2c, Hep2c, H.Ep.-2, H. Ep.-2, H.Ep. #2, H. Ep.-2, or H. Ep. #2 and had various descriptions as described in the Supplemental Information (page 4). In accordance with a proposal from 2000 41 and an early deposit in the ATCC catalog, its currently accepted name should be HEp-2 [HeLa].
In 1967, these cells were shown by Stanley Gartler using isozyme genetics to be HeLa cells,7,8 from an aggressive adenocarcinoma of the cervix. 42 The true identity of the HEp-2 [HeLa] cell line was confirmed by Walter Nelson-Rees in the 1970s and 1980s by various additional genetic tests.9,10,43 In 1978, Lavappa confirmed that ATCC samples of this cell line from 1962, as well as other samples of early cell lines, were actually HeLa cells. 44 In 1990, the DNA profile of the HEp-2 [HeLa] cell line was first published using mini-satellite genotyping by Southern blot analysis, which again confirmed that this cell line was derived from HeLa. 45 It was clearly stated in the first ATCC catalog from 1975, and in subsequent versions, that it is not a true cell line but is a derivative of HeLa that arose by cross-contamination. HEp-2 [HeLa] and other HeLa derivatives were reviewed multiple times by Masters and others,17–21,23,46 and a warning of this cross-contamination was announced in 1993 to the microbiology community by Kenny. 47
Despite these publications, the cell line is persistently described as epidermoid cells from a laryngeal carcinoma. It is used extensively not only to model laryngeal or head and neck cancers but also by microbiologists to study adherence and infection of human cells by various microbes as illustrated by the works of Jones et al. 48 and St. Geme et al. 16 in Figure 1 . Figure 3 compares the frequency of publications using this cell line as a model of laryngeal cancer or simply as an unauthenticated research model. As of June 2021, we found 8497 articles that used HEp-2 [HeLa] inappropriately, published in 2130 journals ( Table 1 ). Within this data set, 3162 (37%) articles described it as a laryngeal or head and neck carcinoma model. The HEp-2 [HeLa] literature is currently growing at about 250 publications annually.
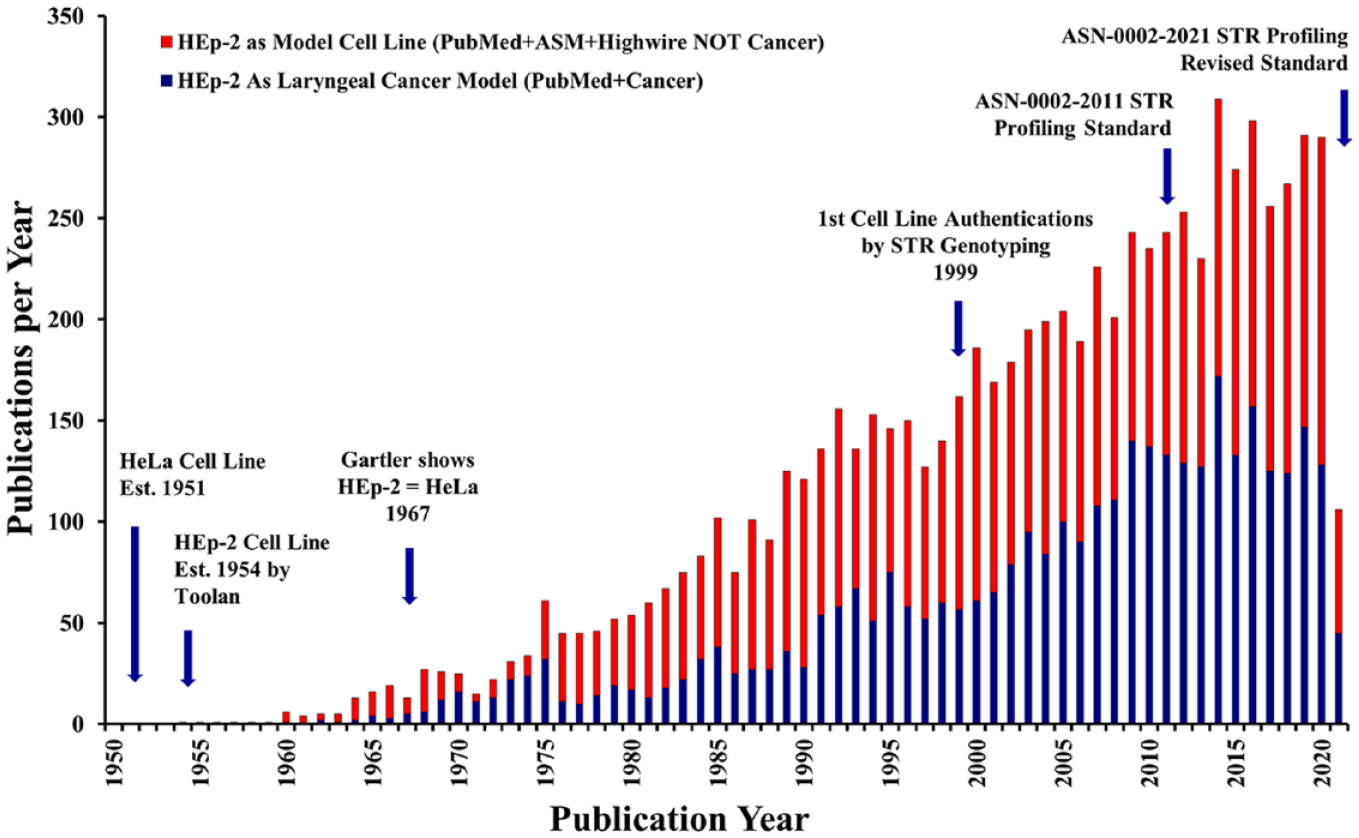
Annual incidence of publications using the false cell line HEp-2 [HeLa], reported as a model of laryngeal cancer (blue bars) versus a cell line without mention of its false tissue of origin (red bars). Key events in the use of this cell line and the authentication of cell lines are noted in the figure. Note that the lower number of articles shown for 2021 (102) comes from only 5 months (January through May), which explains the lower number compared with previous years.
Summary of Articles Using HEp-2 [HeLa] and Intestine 407 [HeLa] Cells Incorrectly and without Authentication.
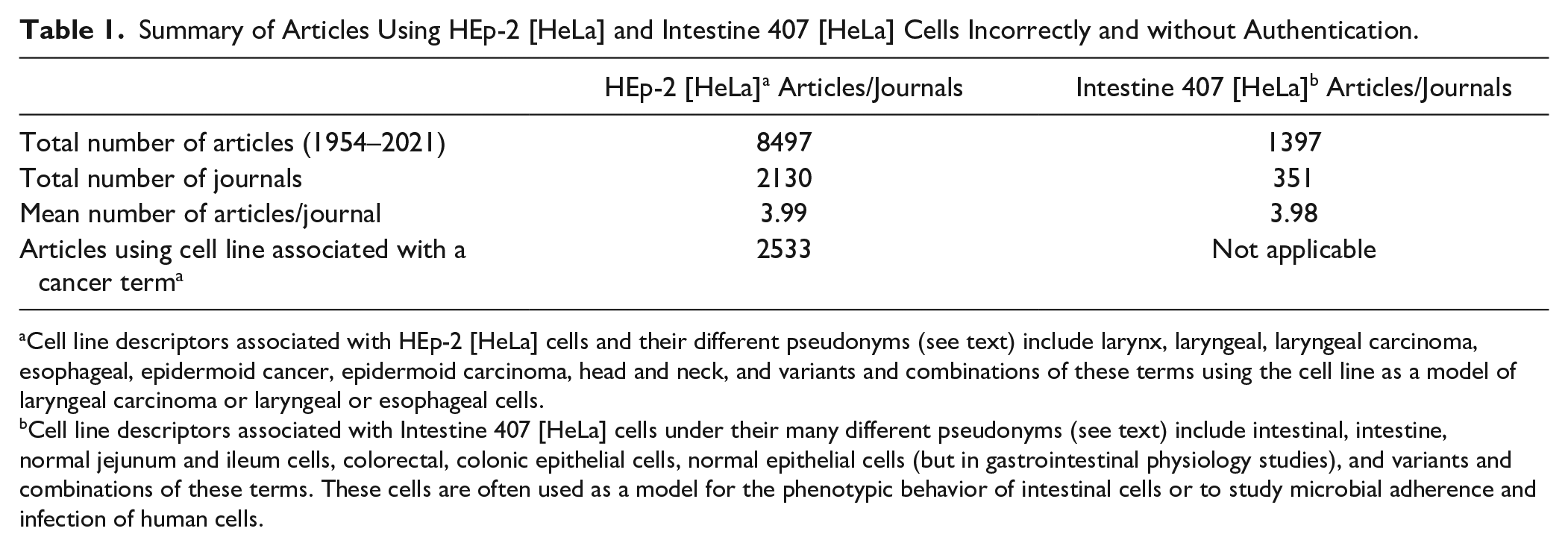
Cell line descriptors associated with HEp-2 [HeLa] cells and their different pseudonyms (see text) include larynx, laryngeal, laryngeal carcinoma, esophageal, epidermoid cancer, epidermoid carcinoma, head and neck, and variants and combinations of these terms using the cell line as a model of laryngeal carcinoma or laryngeal or esophageal cells.
Cell line descriptors associated with Intestine 407 [HeLa] cells under their many different pseudonyms (see text) include intestinal, intestine, normal jejunum and ileum cells, colorectal, colonic epithelial cells, normal epithelial cells (but in gastrointestinal physiology studies), and variants and combinations of these terms. These cells are often used as a model for the phenotypic behavior of intestinal cells or to study microbial adherence and infection of human cells.
Publications Using the HeLa Derivative Intestine 407 [HeLa]
Intestine 407 [HeLa] was established in 1955 by Henle and Deinhardt, who described its history and behavior in 1957, 37 claiming it was derived from the “jejunum and ileum of a human embryo of approximately 2 months gestation.” After subculturing, epithelial-like “cells eventually overgrew the cultures.” that resembled “HeLa cells morphologically.” 49 These authors distributed this cell line prior to publishing their report, and some of its characteristics were described in 1956–1957 by three other groups,34–36 in which the morphological similarities to HeLa cells were also noted. Intestine 407 [HeLa] has been widely used to study and model intestinal diseases (colorectal cancer and microbial infections causing intestinal diseases). In 1967, as described for the HEp-2 cell line, these cells were analyzed by Stanley Gartler using isozyme genetics and shown to be derived from HeLa cells.7,8 The true identity of the cell line was confirmed using multiple techniques, and users of the 1975 ATCC catalog were warned that Intestine 407 was actually HeLa.
Our searches for Intestine 407 [HeLa] were hampered by the use of multiple names—variants of either Intestine 407 [HeLa] (e.g., Intestine 407, Intestine-407, Intestine407, Int-407, Int407, Int 407, I-407, I407) or Henle (e.g., Henle 407, Henle-407; not to be confused with Henle cells in the kidney structure loop or limb of Henle). Because of the limitations of the different databases mentioned earlier, several databases were searched using variants of the complex Boolean search strings shown in the Supplemental Information. Results from a PubMed search were combined with searches of Highwire + PubMed, the Embase database, the Web of Science database, the PLoS website, and the ASM website. Once imported into EndNote, duplicates were removed and the abstracts and (in many cases) the Materials and Methods of some 1500+ articles were reviewed to identify those that used Intestine 407 [HeLa] cells. Most are peer-reviewed research articles; a few results are reports from meetings (which are usually not peer-reviewed) and these, if identified, were deleted. As of June 2021, we have found 1397 articles that used Intestine 407 [HeLa] inappropriately, published in 351 journals (see
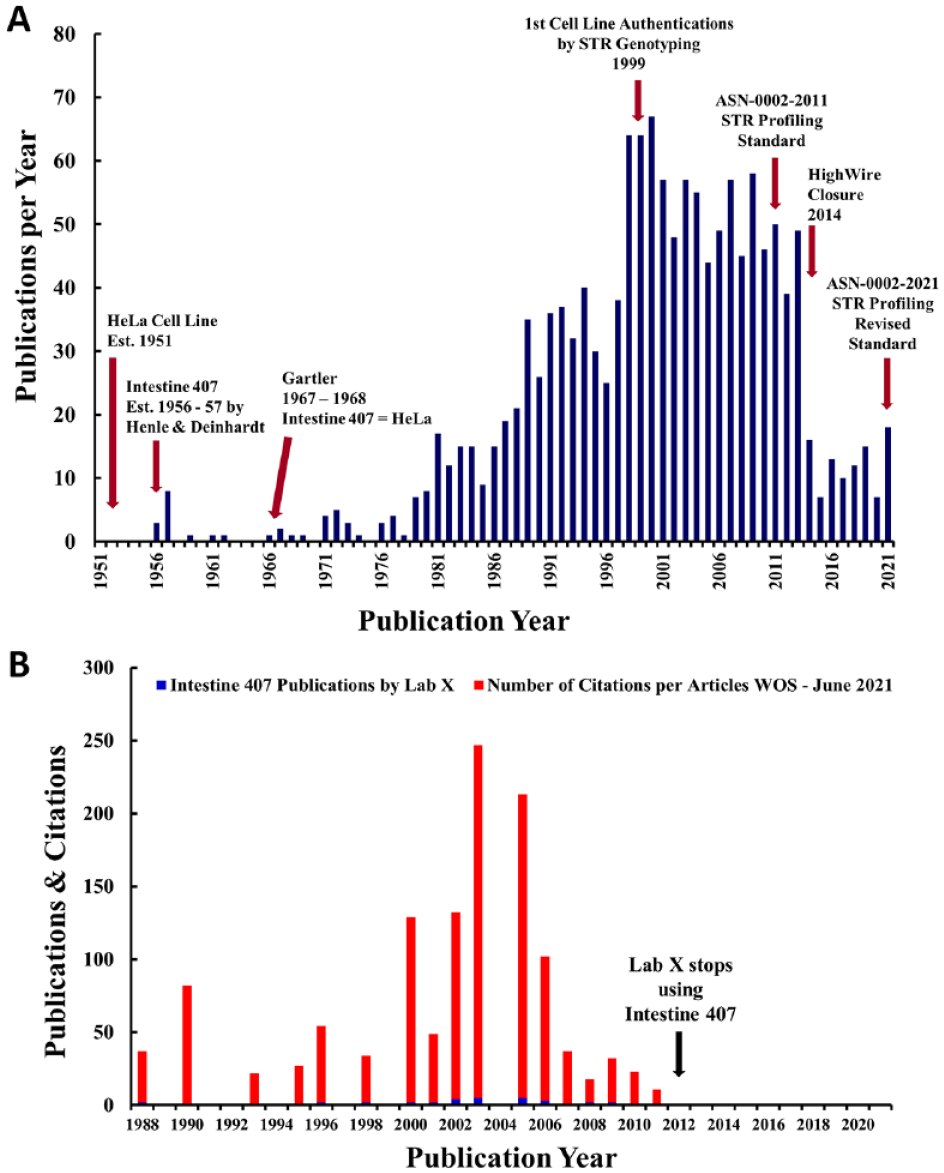
Top panel: Annual incidence of publications using Intestine 407 [HeLa]. Note that for some reason, the use of this cell line appears to have dropped drastically in 2014. This probably relates to some data for the period of 2014–2021 not being available after closure of the HighWire database in 2014. Key events in the use of this cell line and the authentication of cell lines are noted in the figure. Bottom panel: Annual incidence of publications from Lab X using Intestine 407 [HeLa] (blue bars) versus their citations (red bars). The total number of citations of the one or more articles published by Lab X are indicated in the year that each original article was published.
In summary, between 1954 and mid-June 2021, HEp-2 [HeLa] has been published in at least 2130 different journals and Intestine 407 [HeLa] in 351 journals, with a mean number of about four articles per journal for each cell line ( Table 1 ). Clearly, this problem is widespread, and it would appear that the best target for tackling the publication of articles using false cell lines would be the large publishing house that publishes multiple biomedical journals.
One Laboratory’s Use of Intestine 407 [HeLa] Leads to 1212 Citations
To further illustrate the consequences of using false cell lines, we examined a set of publications from a research group (herein identified as “Lab X”) that used Intestine 407 [HeLa] as a model for intestinal cells. Of 1397 publications that were found to use this cell line (
In 2014 we approached the principal investigator of Lab X (Lab X-PI) regarding the usage of Intestine 407 [HeLa] as normal intestinal cells in their publications instead of its actual identity. Lab X-PI was kind enough to discuss and share some STR genotyping data with us. After thorough discussion and evaluation of discrepancies in these data, they were convinced that Intestine 407 was indeed a derivative of HeLa cells. From then on, Lab X discontinued publishing on this misidentified cell line. However, we have not seen any of these articles retracted nor any corrigenda or letters of concern issued for them in PubMed or PubPeer. In fact, we could not find in PubMed a corrigendum for any article using Intestine 407 [HeLa] and very few for articles using any other misidentified cell lines. 24
Although Lab X has ceased to use Intestine 407 [HeLa], their publications continue to be cited by others. A citation of earlier work can indicate that it was used in some way as a basis for the newer work; for example, using a previously described reagent (cell line) or technique, or referring to an observation or conclusion made in the earlier report. If one attempts to reproduce a published work using a false cell line, like Intestine 407 [HeLa], without confirming its identity by genotyping, then there is a risk that the sample of this cell line used by one group comes from a different donor than a sample used by another group. Furthermore, scientists reading such articles may interpret that the findings apply to the physiology and genetics of the falsely associated tissue or disease and will proceed to base new research on this premise.
As of July 2021, Lab X’s 37 publications have been cited a total of 1212 times according to Web of Science (Thomson Reuters). This group has been the source of Intestine 407 [HeLa] cells for other groups, who have acknowledged the group’s leader for the cell line they used in their research. Based on the Web of Science citation rates, this lab’s publications are cited on average 32.7 times per article. Taking the 1397 publications that have used Intestine 407 [HeLa] and multiplying by this average citation rate per article indicates that there could be as many as 45,682 citations referring to reports of questionable significance based on this cell line. Similarly, as many as 277,852 citations could refer to the 8497 articles using the HEp-2 [HeLa] cell line during their citation life span, which can be as long as 40 years. 50
Discussion
Horbach and Halffman 50 reported finding 32,755 articles for 255 of the cell lines listed in the ICLAC Register of Misidentified Cell Lines. For the false cell lines HEp-2 [HeLa] and Intestine 407 [HeLa], they found 3571 and 201 publications in which these lines were used, respectively. Gorphe 51 reported finding 5461 publications using HEp-2 [HeLa], of which 1036 articles refer to its purported laryngeal origin. In contrast, we found 8497 publications using HEp-2 [HeLa], of which 3163 used a laryngeal cancer description, and 1397 publications using Intestine 407 [HeLa] cells. We were able to identity a larger number of publications probably due to our better naming and more extensive description variations for the two cell lines.
The large numbers of research articles that have used the false cell lines HEp-2 [HeLa] and Intestine 407 [HeLa] make up a tangled web of reports, based on assumptions with different degrees of validity. We envision that fields of scientific endeavor that are based on false cell lines can be represented as inverted pyramids, emanating from a single publication at its bottom apex and growing upward through widespread use of that cell line—with each subsequent publication being cited in turn. Such bodies of research may not be based on solid, reproducible footings and be only deceptively held together, and in this case, solely on the basis of the two original articles describing the false cell lines Intestine 407 [HeLa] and HEp-2 [HeLa], which were published in the mid-1950s.
Authentication Testing Is Needed to Address Ongoing Usage
The work by Lab X exemplifies how science, when based on imposter reagents, contributes to a precarious situation. Many studies of intestinal and laryngeal cancer have used these two cell lines to model the physiology of intestinal and laryngeal cells, to study their diseases, and to develop treatments for bacterial and viral pathogens that adhere to and invade cells along the human alimentary canal, as illustrated in Figure 1 . Using these cell lines has fostered an understanding of how pathogens can adhere to and invade human cells. However, using them as a model of intestinal or nasal pharyngeal cells raises the question of how useful these studies are for the development of treatments of these diseases. Furthermore, readers of articles ascribing the origin of cell lines incorrectly or falsely can lead them to base future research on such false models. This case emphasizes that to address the ongoing usage of false cell lines, it is essential for journals and publishers to develop policies requiring authentication testing as a mandatory step prior to publication.
What Is the Cost of This Type of Research?
How much research funding has supported publications and subsequent citations that use Intestine 407 [HeLa] and HEp-2 [HeLa]? Stern et al. studied the cost of retracting articles using grant funding data and found that the average cost of a retraction was between $300,000 and $400,000 per article. 52 If we assume that each publication conservatively costs circa US$100,000 in today’s dollars—an estimate that includes labor (one or more authors), equipment, materials and reagents, facility infrastructure, journal charges, and so forth—then the cost for all original research publications using just these two imposter cell lines (1397 + 8497) ( Table 2 ) would be approximately US$990 million.
Estimates of the Costs of Using Misidentified Cell Lines.
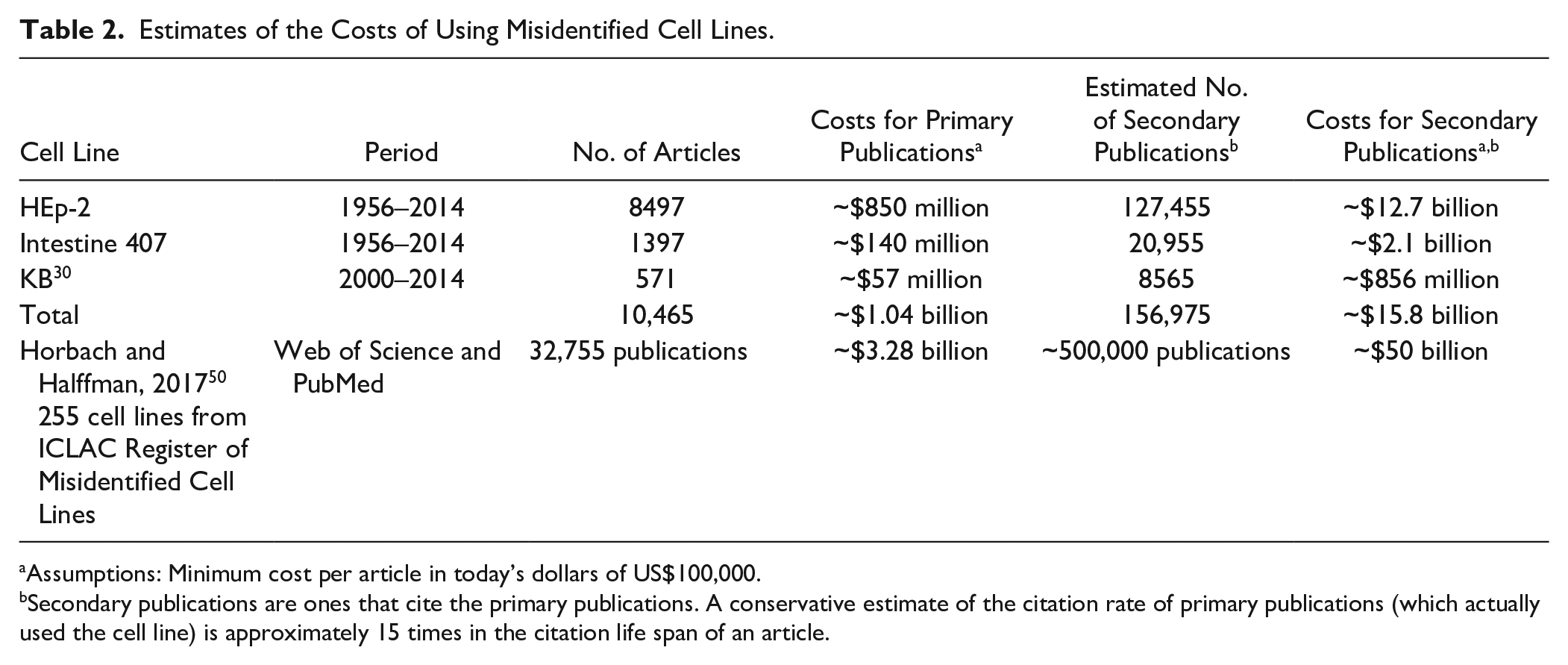
Assumptions: Minimum cost per article in today’s dollars of US$100,000.
Secondary publications are ones that cite the primary publications. A conservative estimate of the citation rate of primary publications (which actually used the cell line) is approximately 15 times in the citation life span of an article.
Assuming each article using these two cell lines was cited on average by 15 articles, the cost of these publications could be as high US$14.8 billion ( Table 2 ). If we take an even more conservative approach and estimate that the combined 9894 original publications that have used Intestine 407 [HeLa] and HEp-2 [HeLa] result in only five citations each, then more than US$4.9 billion may have been spent to support subsequent research based on these two unauthenticated, imposter cell lines.
How much research funding has supported publications and subsequent citations that use any of the 531 false cell lines listed in the ICLAC Register of Misidentified Cell Lines that do not have authentic stocks? Cell lines will vary in their usage, with some used extensively and others rarely. Horbach and Halffman estimated the number of primary publications that used 255 of the cell lines in the ICLAC list to be 32,755, which in turn were cited by approximately a half million other publications, that is, a citation frequency of 15. 50
For Intestine 407 [HeLa], there were 1397 publications over 66 years, averaging about 21 publications per year. If we use the above cost estimates for each publication, and multiply them by a very conservative estimate of 5 publications per year that use each imposter cell line, this yields an estimated 2655 publications annually that use one or more of the 531 imposter cell lines—at a cost of well over US$265 million per year. If we continue to take a conservative approach and estimate that each of these publications is cited five times, an additional US$1.33 billion research funding may have been used to support work using unauthenticated, imposter cell lines. In addition to the financial costs of retracting an article, there are the other costs, including tarnished reputations of the principal investigator and the laboratory, the laboratory staff, and the research institution; the potential loss of current funding and employment; and possibly reduced funding of future grant applications. 52
Has This Research and Associated Funding Been Wasted?
Each publication that uses false cell lines will have questionable value and validity. Although the cost estimates above are very high, we tried to be very conservative in our assumptions. Even while being very conservative in estimating how many publications use imposter cell lines and how many times these publications are subsequently cited, it is clear from this exercise that the use of imposter cell lines can be quite costly for research. However, it does not mean that the resources have been wasted or that all research using misidentified cell lines has been meaningless. The two cell lines used for this study have been employed in microbiology and yielded significant understanding of microbial attachment and invasion of mammalian epithelial cells. Correctly identified HeLa derivatives have served many purposes, including the development of the polio vaccine in the 1950s 3 and the discovery of a new papillomavirus in the 1980s, 53 leading to the development of a vaccine against cervical cancer.
How Can the Questionable Literature Be Appraised and the Losses Avoided?
Very few publications using false cell lines have been retracted or had corrigenda issued about them. 24 Simplistically, some might argue that all these publications should be retracted and thus the cell biology literature would be cleansed of this pollution. However, that is not realistically achievable and any valuable knowledge in these reports would be discarded. Gorphe presented two scenarios for evaluating the value of the numerous publications using misidentified cell lines and rescuing the scientific merit of many publications. 51 To develop the two scenarios, one must first distinguish between publications that used a cell line to represent a specific tissue or disease and those that did not use a specific cell line to model a specific biological process. For the first scenario, Gorphe argued that salvaging the results cannot be performed by simply changing the emphasis of the text and examining the results; in this scenario, the model was incorrect from the beginning. For the second scenario, removing the reference to the false identity of the cell line would not change the importance of the finding. These scenarios are useful when assessing the impact of a false cell line in a specific publication.
An example of postpublication review that may provide useful findings is the study of St. Geme et al. 16 we discussed in the introduction and illustrated in Figure 1 . Recently, Zaaijer and Capes-Davis argued that cell line ancestry is also important when using cell lines for biomedical research and eventual development of clinical treatments in order to understand whether findings are applicable to all people or only those of a specific ancestry. 54 On examining the ancestry of the cell lines used Figure 1 in the Cellosaurus database ( Table 3 ), it is curious that the cervical cancer cell line ME-180 has 97% European ancestry and the HEC-1-B endometrial cell line has 99% Asian ancestry, while the HeLa cell and its derivatives have 65% African and 32% European ancestry. H. influenzae can cause genital infections, so one might expect that this bacterium would be able to bind to cervical carcinoma lines. The fact that hia expressing E. coli does not bind to ME-180 or HEC-1-B cells is intriguing. Prior to the introduction of the vaccines against H. influenzae type B, the incidence of meningitis caused by this bacterium among African American children was significantly greater than in White children, even after correction for socioeconomic factors.55,56 Is it plausible that because of their African ancestry, HeLa cells express a surface factor to which E. coli cells expressing the hia fibril gene of H. influenzae bind preferentially? Similarly, reanalyses of studies of publications using false cell lines may provide new useful findings.
Information regarding cell line ancestry comes from the Cellosaurus information resource and suggests a possible reason for the differences in biology (see Discussion).
Postpublication peer-review platforms are also useful to flag specific instances of misidentified cell lines and provide a forum for addressing questionable and irreproducible findings and data. PubMed Commons was discontinued in February–March 2018, leaving only PubPeer to act in this capacity. When a suitable browser extension is installed, PubMed users can see links to comments that have been posted on PubPeer, offering a mechanism to improve transparency and raise awareness regarding problems. Scientists concerned about misidentified cell lines could tackle the problem by posting comments on PubPeer. The onerous aspects of this task could be ameliorated if the labor is shared by the members of a group.
Mechanisms for Tackling Cell Line “Imposter Syndrome”
The high number of journals publishing articles using HEp-2 [HeLa] or Intestine 407 [HeLa] (2130 and 351, respectively) demonstrates that global policies are needed to address false cell lines in an efficient way. Publishers, editors, reviewers, and editorial staff of journals act as gatekeepers who are best placed to reduce the usage of false cell lines at publication. Global policies are already in place at some publishers; for example, Nature Portfolio journals have a reporting checklist for authors to complete prior to publication. This checklist, which includes cell line questions, is reported by authors to be useful and effective (https://media.nature.com/original/magazine-assets/d41586-018-04590-7/15675426).
We also urge that all journals, as a prerequisite for article submission, mandate cell line authentication using STR genotyping or other means appropriate for the species of the cell line. The International Journal of Cancer has an admirably stringent requirement for providing this journal’s editors and reviewers with valid and meaningful genotyping data to accompany manuscripts that use cell lines. The journal editors have reported that since their implementation of such a policy, they have seen an increase in manuscript submissions. 57 Similarly, granting agencies need to stringently enforce their cell line authentication policies, such as those promulgated by the NIH (Authentication of Key Biological and/or Chemical Resources, NOT-OD-17-068 [https://grants.nih.gov/grants/guide/notice-files/NOT-OD-17-068.html] and earlier versions).
The ICLAC Register of Misidentified Cell Lines is linked to the Cellosaurus knowledge resource that attempts to describe all cell lines used in biomedical research. As of June 2021, it has information on 128,806 metazoan cell lines, which include 96,820 human, 21,791 mouse, and 2444 rat cell lines. The Cellosaurus website provides an incredible wealth of information and an STR search tool (CLASTR) that allows the comparison of an STR genotype to the STR profiles of 7647 human cell line STR profiles, 74 mouse STR profiles, and 36 dog STR profiles.
There are several technical manuals for the authentication of (primarily) human cell lines. These include a written standard for the authentication of human cell lines, published by the American National Standards Institute together with ATCC. 58 The initial document from 2011 has been greatly revised and expanded and was republished in April 2021. 59 It is available for purchase online (https://webstore.ansi.org/standards/atcc/ansiatccasn00022021) and described in an ANSI blog (https://blog.ansi.org/authentication-human-cell-lines-str-atcc-asn-0002/#gref), with a printed hard copy version being planned. The U.S. National Library of Medicine published in 2013 a chapter in the Assay Guidance Manual (AGM) e-book on the authentication of human cell lines by STR profiling, 60 which is planned to be revised for republication.
Anyone who implements a policy to address false cell lines needs support and training in order to do so effectively. This is particularly true for journal staff members who facilitate mandatory presentation and review of authentication data. The research community (principal investigators, technicians, students, departmental chairs, institutional policy boards) needs similar training on the proper handling of cell lines. ICLAC provides useful information and training modules on their website, including slide decks for educational seminars (https://iclac.org/education/). An institutional cell line authentication policy, such as the example provided on the ICLAC website (https://iclac.org/resources/cell-line-policy/), is recommended to provide further guidance on how to avoid cell line cross-contamination in the laboratory and bring about a change of work culture that may reduce cell line problems. Besides the resources mentioned above, good cell culture practice is clearly set out in textbooks 61 and published guidelines for the use of cell lines in biomedical research. 22
Conclusions
Freedman et al. 62 reported that more than 50% of surveyed researchers never authenticated their cell lines; some even saw no need to do so since “they never made mistakes.” Although checking lists and testing cell lines may be considered cumbersome by some researchers 50 —such as when a cell line is renamed by a group—doing nothing is not an option. The best approach to address false cell lines is to provide definitive authentication data, such as STR genotypes, in publications for future reference. Misidentified cell lines have contributed to the last half-century of biomedical research, but we need to ensure they are identified correctly if they are to become more useful tools in the future.
Supplemental Material
sj-pdf-1-jbx-10.1177_24725552211051963 – Supplemental material for The Extensive and Expensive Impacts of HEp-2 [HeLa], Intestine 407 [HeLa], and Other False Cell Lines in Journal Publications
Supplemental material, sj-pdf-1-jbx-10.1177_24725552211051963 for The Extensive and Expensive Impacts of HEp-2 [HeLa], Intestine 407 [HeLa], and Other False Cell Lines in Journal Publications by Christopher T. Korch and Amanda Capes-Davis in SLAS Discovery
Footnotes
Acknowledgements
We are grateful to Dr. John Masters for providing us with information about publications from Lab X describing their use of Intestine 407 [HeLa], and to Ms. Jill Neimark for her persistent interest in this project and for facilitating contact with the principal investigator of Lab X. We wish to dedicate this article to the memory of Drs. Roland M. Nardone and R. Ian Freshney, who strongly advocated for reproducible research by authentication of cell lines.
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was not funded by any sources, other than our use of the libraries in the organizations with which we are affiliated as retired or honorary faculty.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.