Abstract
Organoids are three-dimensional, functional structures that mimic in vivo organs. They offer new opportunities for the modeling of cancer and infectious and rare hereditary diseases. Furthermore, the advent of organoid biobanks opens new avenues for drug screening in a personalized fashion and holds much promise for personalized regenerative medicine. Thus, there is a need for reproducible, large-scale organoid generation with minimal variability, making manual approaches impracticable. Here, we review the current use of automation in organoid culture and analysis, using cerebral and retinal organoids as illustrations of current applications. An increased demand for automated organoid platforms is anticipated.
Graphical Abstract
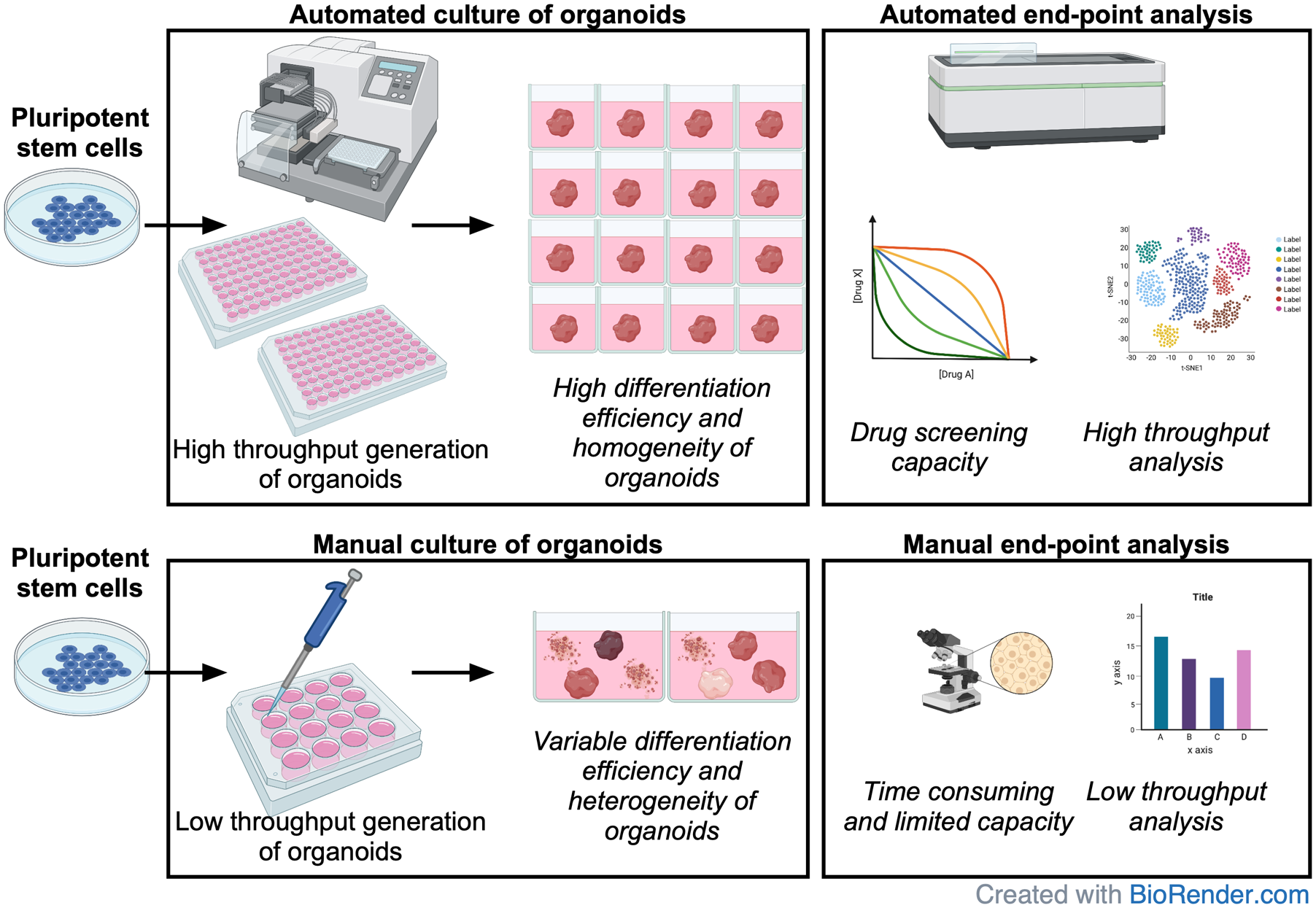
Introduction
Organoids are self-organized, three-dimensional (3D) structures derived from pluripotent or adult stem cells containing multiple functional, organ-specific cell types.1–4 Early examples of pluripotent stem cell (PSC)-derived organoids were established by Eiraku and colleagues, who generated 3D cortical tissues from both mouse and human embryonic stem cells (ESCs). 5 Since then, organoids have been developed for multiple healthy and cancerous tissues.1,6 Organoids are more structurally and functionally complex compared with conventional two-dimensional (2D) systems, and thus are likely to mimic organ responses to external stimuli more accurately than monolayer cell cultures would.7,8 Organoids can also be genetically engineered with CRISPR/Cas9. A landmark paper reported genetic correction of CFTR mutation in intestinal organoids that restored its function. 9 Others have introduced mutations to study cancer in intestinal,10–12 breast, 13 and liver organoids,14,15 as well as polycystic kidney disease in kidney organoids. 16 This exemplifies how organoids are a valuable tool for studying biological processes associated with development and disease that cannot be replicated in other model organisms. 6 Furthermore, they can be incorporated into drug development and potentially cell replacement therapy pipelines. Nevertheless, despite their great potential, several limitations remain. First, organoid generation is usually performed manually. Although successfully implemented into numerous protocols, these steps are time-consuming and susceptible to technician variability and human error. 17 This contributes to significant heterogeneity of cell composition between individuals and protocols6,18 and restricts the scalability and reproducibility of these approaches. The use of automation might offer a solution for some of these issues. The automated approach is likely to significantly improve the consistency of generated organoids and improve differentiation reproducibility. This in turn will allow processing of much larger numbers of samples, opening new avenues in disease modeling and drug discovery. 19 Second, lack of vascularization reduces nutrient supply and may impact growth potential; however, recently reported vascular organoids may help to address this problem. 20 Yet, transition of organoid culture into automation will not improve vascularization. Automation combined with automated imaging systems will facilitate the optimization of differentiation protocols by assessing the impact of modifying the initial number of cells used for organoid generation and the timing and concentration of small molecules.21,22 Third, many protocols rely on Matrigel, significantly increasing the costs of organoid cultures. Furthermore, its variable composition introduces batch-to-batch variability and poses a risk of immunogen transfer, prohibiting potential clinical applications. Synthetic, chemically defined hydrogels are a promising alternative to Matrigel.17,22–24 Fourth, long-term tracking of organoid cultures typically involves imaging systems and algorithms. In situ monitoring is unfeasible and requires organoid dissociation, limiting the analysis to a single time point. Incorporation of microfluidics for organoid research may help to address this problem, and several multiorgan devices coupled with automation have already been reported in the literature.25–28 Here, we review the current research surrounding the automation of organoid generation, cell tracking, phenotypic analysis, and drug screening, with a particular focus on both cerebral and retinal organoids.
Automated Applications for Organoids
Biobanking
The rapid progress in the induced PSC (iPSC) technology prompted the establishment of large-scale banks of patient-specific iPSC lines derived from both disease-affected individuals and healthy controls (reviewed in Daniszewski et al. 19 and McKernan and Watt 29 ). Their overarching goal is to provide researchers with a reliable source of samples for downstream analyses. It is thus imperative that processing steps are standardized to maintain the biospecimen integrity during the procedure. This issue was addressed by the International Organization for Standardization (ISO), which developed standards (ISO 20387:2018 and ISO 22758:2020) to ensure quality and reproducibility in biobanking. 30 The organoid banking has previously been reported;31,32 however, the procedure was performed manually. To the best of our knowledge, automated organoid freezing has not been reported; however, Linsen et al. have recently described the automated system for storage and sample retrieval, which allowed significant improvements to hands-on sample handling times. 33 Nevertheless, the overall process to install and operate the system took more than 4 years, underlining the substantial effort needed for their adaptation to site-specific requirements.
Phenotypic Analysis and Cell Tracking
The development of organoids has been accompanied by rapid progress in 3D imaging methods to visualize their structural complexity. These new techniques combined with improved clearing procedures 34 allow researchers to observe cellular architecture at an unprecedented level and without compromising tissue integrity. 35 Furthermore, organoids are amenable to live imaging and studying spatiotemporal changes in biologic processes at high resolution.11,36,37 Nevertheless, the automation of organoid imaging is limited due to high dimensionality of the data, acquisition artifacts, low contrast, and bright-field noise.38,39 To address this bottleneck, several cell segmentation and tracking algorithms have been developed. For example, Borten et al. described OrganoSeg, an automated package for analysis of bright-field images. The morphometric profiling could be correlated with transcriptomic profiles from bulk RNA sequencing to identify genes associated with organoid size. 39 Others focused on algorithms for fluorescent microscopy designed for high-throughput applications. Brandenberg et al. used a semiautomated method based on Fiji (ImageJ) 40 software to compare the responses of colon tumoroids derived from patient biopsies and the human colon carcinoma HTC 116 cell line to FDA-approved and experimental chemotherapeutics. 22 The imaging system was able to extract more than 250 features from each organoid and distinguish their sensitivity to epidermal growth factor receptor (EGFR) inhibitors. Responsiveness to EGFR inhibitors depended on the presence of KRAS mutations and was in line with previous findings.41,42 Furthermore, Czerniecki et al. developed a custom algorithm to identify subcompartments of kidney organoids and optimize their automated differentiation based on dose response to the GSK3 inhibitor, CHIR99021. The proportion of proximal and distal tubules as well as podocytes varied in a CHI99021 dose-dependent manner. 21 Bulin et al. described CALYPSO (comprehensive image analysis procedure for structurally complex organotypic cultures) to analyze the impact of combinations of photodynamic therapy, chemotherapy, and radiotherapy on the viability of pancreatic tumoroids. The software was able to analyze tumoroids grown alone and in co-culture and find correlations between their size, residual viable area, and viability in a dose-dependent manner. 43
The introduction of life imaging has facilitated the investigation of spatiotemporal dynamics at the cellular level; however, it also presented new sets of challenges (elegantly reviewed in Skylaki et al. 44 ). These include the choice of molecular markers for imaging, which are often conjugated to fluorescent proteins, to prevent artifacts. Optical monitoring of organoids is routinely used due to the destructive nature of currently available techniques for the functionality assessment. 17 Furthermore, high-power lasers used for image acquisition in traditional 3D confocal microscopy significantly restrict observation times without inducing photobleaching and phototoxicity.45,46 This limitation can be overcome with the use of nondestructive imaging techniques, such as multiphoton (MP) and light-sheet microscopy (LSM), where only the in-focus part of the sample is illuminated by laser, decreasing phototoxicity and improving imaging speeds.47,48 Interestingly, use of optical coherence tomography for live 3D imaging of retinal organoids has recently been reported. 49 Both MP and LSM allowed researchers to combine 3D imaging with time-lapse imaging to generate four-dimensional (4D) imaging and observe breast tumor mobility 50 and Drosophila embryogenesis at single-cell resolution,47,51 respectively. An advancement of LSM was described by Chen et al., who integrated optical lattices and LSM, creating lattice LSM (LLSM) with improved acquisition speeds and reduced invasiveness. 36 The method enabled imaging of a wide range of samples from molecules to developing embryos; however, imaging depth was limited due to aberrations caused by heterogeneous refractive indexes in multicellular samples. 36 The aberrations can be corrected with the use of adaptive optics (AO) to increase resolution. 52 Combining AO with LLSM into AO-LLSM retains benefits of both techniques for noninvasive, high-resolution imaging in a physiologically relevant manner. 53 This allowed observation of clathrin-mediated endocytosis in human intestinal organoids as well as organ development and cell migration during zebrafish embryogenesis.53,54 AO-MP microscopes have been used for in vivo imaging of the retina 55 and lymph nodes; 56 however, to the best of our knowledge, AO-MP has not been used for organoid imaging yet.
Furthermore, the vast amount of data generated during imaging is a significant problem for postacquisition processing. Over the years, different computational approaches have been developed for cell segmentation, tracking, and quantitative data analysis, 44 including TrackMate. 57 TrackMate is an open-source Fiji plugin that was successfully used for cell tracking in embryos,57–59 as well as kidney 60 or endometrial organoids. 61 Many other algorithms for cell tracking purposes were generated, including those created as part of the Particle and Cell Tracking Challenge.62,63 Interestingly, algorithms using machine learning performed best in most segmentation scenarios, 63 with several others also reported in the literature.64–67 The OrganoidTracker developed by Kok et al. employs neural networks in machine learning to detect cell nuclei and form cell trajectories with high accuracy. A built-in error checker flags potential tracking errors and marks them for manual verification, improving the performance of the neural network. 68 Furthermore, other tools for automated time-lapse analysis in 3D69–74 and 4D54,75 have also been described. The combination of AO-LLSM with OrganoidTracker would allow noninvasive, real-time assessment of organoid behavior following drug exposure, opening new avenues for drug screening.
Drug Screening
Despite the significant cost associated with drug development, the expenditure does not translate into the number of new drugs being approved every year. In 2020, only 53 new molecular entities (NMEs) were approved in the United States, 76 which is in part due to the failure to translate the results of preclinical research into clinically effective compounds. According to the R&D productivity model, only 54% of new drugs reach phase I of clinical development; this accounts for 32% of the cost of their development. 77 The percentage of attrition is even higher at later stages, with 66% and 30% in phases II and III, respectively, leading to a major loss of time and resources. 77 iPSC-derived kidney and liver organoids may pave the way for predictive toxicology assessment. This would allow implementation of the “fail early, fail cheaply” framework, 78 where compounds would be withdrawn from the drug development process if they are found to be hepato- and/r nephrotoxic. It would decrease the attrition rate of NMEs at early stages of clinical trials and potentially lead to a vital reduction of animal testing at the preclinical stage. 79
The first automated system for high-throughput organoid generation was described in 2014. 80 The authors used breast, prostate, and colorectal organoids and identified nine compounds targeting the respiratory chain that are effective against dormant cells in the core tumor regions. 80 On the other hand, Boehnke et al. described a platform for screening chemotherapeutics on colon cancer organoids. They validated the robustness and reproducibility of the platform, which passed the criteria of replicate-experiment study and achieved pharmacologic relevance.81,82 The subsequent systems examined nephrotoxicity in kidney organoids generated from primary cells and human iPSCs by assessing the expression of heme oxygenase 1 (HO-1) and kidney injury molecule 1 (KIM-1), respectively.21,83 Both studies observed dose-dependent upregulation of HO-1 and KIM-1 following exposure to nephrotoxic compounds, for example, cisplatin, in line with previous data.84,85 Furthermore, Czerniecki et al. attempted to optimize vascularization of iPSC-derived kidney organoids with increasing concentrations of VEGF. The study observed a 10-fold increase in the number of endothelial cells (ECs) expressing CD31 and vascular endothelial (VE) cadherin; however, ECs failed to invade podocytes to establish the glomerular basement membrane. 21
Limited vascularization has been a persistent problem in organoid research. Interestingly, this limitation may be addressed by moving into microfluidics, which could also offer other improvements over standard-size cell culture. These include precise control of the microenvironment in a spatiotemporal manner and low reagent consumption. The former allows introducing flow and regulating shear stress, which improves maturation and vascularization of kidney organoids 86 and can also be used to mimic peristaltic activity in gastric organoids. 87 The latter can significantly reduce costs, delivering liquids at femtoliter volumes. 88 Furthermore, microfluidic platforms are amenable to automation requiring minimal human intervention and in turn reducing variability. Schuster et al. developed an automated microfluidic platform for screening chemotherapeutics on primary pancreatic tumoroids at clinically relevant doses. Interestingly, sequential delivery of chemotherapeutics significantly increased apoptosis in tumoroids in a patient-specific fashion and allowed the authors to predict the most effective combination of drugs in a personalized fashion. 89 The microfluidic systems, also known as organs-on-chips, may be connected with each other with endothelium-lined vascular channels to form a body-on-a-chip (reviewed in Sung et al. 90 and Bhatia and Ingber 91 ). Such a setup would allow researchers to examine interactions between different tissues following the drug exposure and conduct ADMET (absorption, distribution, metabolism, elimination, and toxicity) analysis. This is crucial for the assessment of pharmacokinetics and pharmacodynamics (PKPD) to determine the pharmacological effect of the drug. Zhang et al. reported two two-tissue systems (heart, liver and heart, liver cancer) to test cardio- and hepatotoxicity. 25 Skardal et al. developed a three-tissue system (heart, lung, liver) and identified the mechanism underlying cardiotoxic side effects of the anticancer drug bleomycin. 26 Novak et al. recently described an eight-organ system (intestine, liver, kidney, heart, lung, skin, blood–brain barrier, and brain) combined with automated imaging and sampling. 27 A derivation of this system containing bone marrow, gut, liver, and kidney was used to analyze drug PKPD of cisplatin, an anticancer drug with known myeloid and renal toxicity. 28 The system was able to recapitulate those side effects and provided better prediction of drug liver clearance compared with rodent data. 28 Interestingly, similar findings were reported in 3D printed kidney 92 and liver organoids. 93 The advantage of 3D printing is the ability to assemble tissue in automated and spatially controlled patterns, improving throughput and reproducibility. This approach also elucidated the mechanism of hepatotoxicity caused by an antibiotic trovafloxacin that was not identified during clinical trials. 94 Intriguingly, the platform described by Jiang et al. combined microfluidics and 3D printing of Matrigel droplets to generate liver, lung, kidney, and colon tumoroids. Thanks to high cell density in Matrigel droplets, they observed significant improvement of growth time as well as high interorganoid homogeneity. Furthermore, liver and colon tumoroids recapitulated the patient’s response to anticancer drugs with 81% accuracy. 95 This predictive ability was consistent with previous reports,96–99 further supporting the use of organoids for drug screening in the future.
Cerebral Organoids
Although animal models and 2D culture systems have significantly advanced our understanding of human brain development and neurological disorders, they have limitations.100,101 Consequently, the generation of cerebral organoids that contain brain-specific cells and resemble the developing human brain has provided a novel tool for brain research. 102 However, as with all organoids, cerebral organoids do not perfectly model the human brain and face considerable limitations, such as the lack of reproducibility and long culturing times. These could potentially be addressed by automation. Due to high heterogeneity in cell distribution and morphology within organoids, quantifying data sets for analysis is a common challenge. 103 Automated protocols for differentiation have significantly reduced this heterogeneity; however, organoids still show heterogeneous spatial arrangement. This makes it difficult to interpret the cellular phenotype without being able to assess the organoids in a larger spatial context and remains a major limitation to gaining a deeper understanding of human pathology. 100 Promisingly, RNA sequencing analysis has suggested that cell developmental trajectories are largely consistent and reproducible within organoids. 104 Although this provides promise for future developments that decrease batch variation, protocols remain strict and labor-intensive. Renner et al. recently described the medium- to high-throughput generation of human automated midbrain organoids (AMOs) from neural progenitor cells, previously originated from iPSCs, using a 96-channel pipetting robotic attachment (Biomek FXP, Beckman Coulter, Brea, CA). 105 AMOs were generated via a guided differentiation protocol, as opposed to a self-patterning differentiation method followed for organoid generation. The workflow permits the generation and maintenance of AMOs, as well as their fixation, staining, and clearing for optical analysis, followed by fixation, whole-mount staining, and clearing steps that were fully automated. 105 This automated protocol was combined with cell culture modifications to improve consistency of cultures. For instance, there was no Matrigel embedding of organoids to reduce batch-to-batch variability. Further, as each organoid was set up in an individual well, the setup also reduced the impact of potential paracrine signaling, which could also play a role in batch variations. Additionally, the culture conditions were optimized to reduce the formation of local subdomains, such as rosettes, within the organoids. This was done to overcome the complication of heterogeneity of structures within the organoids, allowing a clearer screen and analysis of organoids. The resulting homogeneity maximized the ability to extrapolate information from a single image from a medial confocal slide without having to image the whole organoids and hence increasing the power on the number of organoids that can be analyzed. Further characterization of the AMOs using RNA sequencing and functional assays such as calcium imaging and electrophysiology confirmed that this method produces midbrain organoids with a pattern of gene expression similar to previously published midbrain organoids and primary human midbrain tissues, and with spontaneous and synchronized neuronal activity. Interestingly, this workflow was proven efficacious for the generation of neural progenitor cell-derived AMOs as well as for iPSC-derived automated cortical organoids, but differences in levels of homogeneity were significant; the automated cortical iPSC differentiation method producing organoids showed much higher levels of heterogeneity, both structurally and functionally. Further, the AMOs showed faster maturation than iPSC-derived automated organoids, with characteristics of the midbrain, including mature functional dopaminergic neurons. This platform was subsequently used for proof of concept of high-throughput drug and toxicity screening, treating AMOs with 6-hydroxydopamine and 1-methyl-4-phenylpyridinium, known to be toxic to dopaminergic neurons, and using cleaved caspase 3 to monitor apoptosis as a robust readout. 105 Overall, this work highlights the importance of automation and cell culture processes to reduce heterogeneity, as well as the intrinsic variabilities associated with PSCs, which cannot be fully controlled despite the rigor and consistency of automation as well as the compatibility of different readouts that can be applied to large-scale screening models. Another automated workflow for the scaling and phenotyping of automated organoids has been recently described, although not yet peer-reviewed. 106 The automated Organoid Culture and Assay (ORCA) platform uses automated liquid handlers (PA PF-400) to standardize, scale, and automate the culture of forebrain organoids from multiple patient lines, and subsequently permits their phenotyping in an unbiased fashion. This setup can use 24- or 96-well plates to cultivate and harvest organoid cultures without manual manipulations or corrections. It is able to culture more than 40,000 organoids simultaneously while also being individually tracked. The ORCA-cultured organoids displayed characteristics superior to manually cultured cortical organoids, including reduced size variability and endogenous neural network activity. This study subsequently applied machine learning methods to characterize the organoid variability and found that the donor, clone of the cells, and batch of differentiation are significant factors that contribute to the variability of the organoids, measured by gene expression, cell composition, and morphology. The study suggests that diseased phenotype may be concealed or influenced by these factors (donor, cell, and batch) if not properly verified. In fact, using the machine learning methodology, the ORCA-cultured forebrain organoids were used to study tuberous sclerosis, a genetic disease known to cause tumor formation in different organs, including in the brain, and for the first time they were able to report a highly reproducible disease phenotype in organoids with heterozygous mutation in the TSC2 gene at early stages of differentiation (at day 35 compared with >100 days for other studies). Interestingly, the structure variability was matched across multiple assays and proposed that an automated nondestructive assay (such as bright-field imaging) could be used to assess the organoid variability as well as destructive time-consuming assays would (such as single-cell RNA analysis). Additionally, the Single-cell and Cytoarchitecture analysis of Organoids using Unbiased Techniques (SCOUT) pipeline also permits an automated multiscale analysis of cerebral organoids, capable of phenotyping organoids at a single-cell and whole-organoid level while also analyzing their cytoarchitecture by rapidly clearing, staining, imaging, and analyzing intact organoids. 103 As a proof-of-concept experiment to assess the efficacy and accuracy of SCOUT phenotyping abilities, the pipeline was used to distinguish between organoids obtained via separate differentiation protocols (unguided cerebral organoids vs guided forebrain organoids). Upon comparison from around 120 organoids, the pipeline detected 29 major differences between the two culture methods, demonstrating the robustness of this approach. Interestingly, the SCOUT pipeline also provided novel information on Zika infection in the developing brain. 103 Human cerebral organoids infected with Zika virus were subjected to the SCOUT pipeline to assess if phenotypes could be observed and quantified, concentrating on cell number, size of ventricles, and general cytoarchitecture. The SCOUT pipeline could indeed robustly characterize Zika-associated phenotypes, including reduced organoid size, increased ventricle size, and an increase in neural progenitor cells and neurons, consistent with previous reports.107–109 However, the SCOUT pipeline is limited to the analysis of only two nuclear markers at one time, and further implementation should be adapted to fully unveil its potential for large-scale analysis.
Retinal Organoids
The development of new therapeutics for retinal diseases has been hindered by the lack of good in vitro models. Retinal organoids recapitulate early retinogenesis and can display some functional properties, such as a photoreceptor response to light.31,110–113 Although retinal organoids have improved our understanding of some retinal diseases, including retinitis pigmentosa114–117 and Leber congenital amaurosis,118,119 challenges still remain limiting their use as in vitro models. These include batch-to-batch and interprotocol variabilities, as well as low-throughput culturing associated with cell immaturity. So far, only a handful of groups have attempted incorporating automation into their organoid studies. Kegeles et al. recently described a semiautomated approach for the differentiation of mouse and human retinal organoids. The workflow included automated media exchange (Thermo WellWash Versa and Thermo RapidStack robot) that performed on par with human operators. 120 Furthermore, an automated imaging system was used to assess differentiation quality and confirmed that the addition of forskolin enhances retinal differentiation. The other automated system is the 3D automated reporter quantification (3D-ARQ) platform for fluorescence-based analysis of retinal organoids. 121 The platform allowed researchers to quantify developmental progression by examining Brn3b and Otx2, which are expressed in retinal ganglion cells (RGC) and photoreceptor progenitors, respectively. Subsequent proof-of-principle experiments measured organoid responses to oxidative stress and mitochondrial uncoupling. In both instances, 3D-ARQ detected significant differences in fluorescent intensities between conditions, supporting its use in therapeutic screening. Yet, both systems relied on manual organoid generation, significantly reducing the possibility of scaling up. Moreover, fluorescent reporter lines could not be used to monitor organoid differentiation outside of research laboratories. Interestingly, such quantitative assessment may also be conducted using machine learning to predict retinal differentiation and functional maturity in a noninvasive manner.122,123 This would ensure that only “good” retinal organoids proceed in differentiation to generate fully mature retinal cell types, which often require 200+ days, mimicking the development of human retina.110,124 Nevertheless, the cost of upscaling such a process would not be commercially feasible. DiStefano et al. attempted to accelerate the differentiation by incorporating rotating-wall vessel bioreactors and observed slight improvement in speed and organoid size compared with static culture. 125 Thus, further research is required to generate functionally mature organoids in a shorter time frame to progress their use in high-throughput drug screening studies.
Conclusion
Human PSC-derived organoids provide new opportunities for disease modeling, for drug screening, and to improve our understanding of many developmental processes. Coupling them with automation could significantly impact the medical research and pharmaceutical industry by reducing or even eliminating human error and batch-to-batch variability from experiments. Nevertheless, the automation will not improve the inherent characteristics of the model on its own; thus, model optimization should precede transition into automation to achieve optimal results. Furthermore, thanks to advances in microscopy, it is also possible to image organoids with unprecedented resolution to gain insight into physiological processes. Yet, challenges remain, including improvements to culture conditions and co-cultures of organoids with other cell types to better mimic native microenvironment and cellular interactions that could potentially accelerate the speed of organoid maturation. Also, data storage may still be problematic, while data analysis requires sophisticated algorithms that often lack flexibility in their applications. Nevertheless, with the technological progress, automated platforms will become more versatile, allowing researchers to generate and analyze different organoid types. It will also allow standardization of protocols to ensure scalable supply of high-quality, mature organoids for research and clinical applications.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by a National Health and Medical Research Council Senior Research Fellowship (A.P., 1154389), the Yulgilbar Alzheimer’s Research Program (D.H., A.P.), and a University of Melbourne Early Career Grant (M.D.).
