Abstract
Affinity selection mass spectrometry (ASMS) has emerged as a powerful high-throughput screening tool used in drug discovery to identify novel ligands against therapeutic targets. This report describes the first high-throughput screen using a novel self-assembled monolayer desorption ionization (SAMDI)–ASMS methodology to reveal ligands for the human rhinovirus 3C (HRV3C) protease. The approach combines self-assembled monolayers of alkanethiolates on gold with matrix-assisted laser desorption ionization time-of-flight (MALDI TOF) mass spectrometry (MS), a technique termed SAMDI-ASMS. The primary screen of more than 100,000 compounds in pools of 8 compounds per well was completed in less than 8 h, and informs on the binding potential and selectivity of each compound. Initial hits were confirmed in follow-up SAMDI-ASMS experiments in single-concentration and dose–response curves. The ligands identified by SAMDI-ASMS were further validated using differential scanning fluorimetry (DSF) and in functional protease assays against HRV3C and the related SARS-CoV-2 3CLpro enzyme. SAMDI-ASMS offers key benefits for drug discovery over traditional ASMS approaches, including the high-throughput workflow and readout, minimizing compound misbehavior by using smaller compound pools, and up to a 50-fold reduction in reagent consumption. The flexibility of this novel technology opens avenues for high-throughput ASMS assays of any target, thereby accelerating drug discovery for diverse diseases.
Introduction
High-throughput screening remains one of the most popular, unbiased approaches used in drug discovery for identifying novel chemical matter against a target. Various technologies and high-throughput platforms are available for functional screening assays ranging from label-dependent approaches such as Förster resonance energy transfer (FRET) and radioactivity to label-free approaches such as mass spectrometry (MS). 1 However, not all therapeutic targets generate a functional readout; therefore, high-throughput target–ligand interaction assays are necessary to initiate drug discovery efforts.
Binding assays have historically relied on displacement formats utilizing fluorescent and radioactive probes.2,3 Affinity selection mass spectrometry (ASMS), first described more than 15 years ago, 4 has gained momentum as a powerful tool for identifying small molecules that engage protein and oligonucleotide targets. 5 The general ASMS workflow involves incubating a target with a single or collection of small molecules, isolating the complex, and then identifying the bound small molecule by its mass. This workflow has been adapted to various approaches for complex isolation and compound detection. Currently, a common option is to infer hit identification from MS detection following several steps, including mixing the target with up to 2500 compounds, 6 isolating the complexes under native conditions, dissociation of the bound compounds through denatured conditions, and MS analysis. A recent report described an alternative workflow where complex isolation is performed in a 384-well filter plate assay format, followed by compound detection using high-throughput matrix-assisted laser desorption ionization (MALDI) MS. 7 Our team previously highlighted how the combination of self-assembled monolayers of alkanethiolates on gold and MALDI time-of-flight (TOF) MS, a technique termed self-assembled monolayer desorption ionization (SAMDI)–ASMS, can be used to identify small-molecule noncovalent binders. 8 The original proof-of-concept study showcased how SAMDI-ASMS enables affinity capture of a target, followed by subsequent ionization to infer the identification of the noncovalent compound by its mass. The approach overcomes the shot-to-shot variability often experienced by MALDI matrix crystallization “hot spots” through uniform saturation of the surface with the captured protein, while reducing interference of small-molecule detection from various buffer components such as salts and detergents, along with the matrix.
Here we demonstrate, for the first time, the high-throughput capability of SAMDI-ASMS to screen more than 100,000 small molecules to identify novel binders to the human rhinovirus 3C (HRV3C) protease. This virus is responsible for the common cold, for which there remains a need for novel therapeutics, supporting the idea that alternative drug discovery strategies are necessary to identify chemical matter and drug leads. To accelerate the screen, the compounds were evaluated in pools of eight compounds, allowing the completion of the primary screen and data generation in less than 8 h. Hits identified by SAMDI-ASMS were validated in an orthogonal binding assay using differential scanning fluorimetry (DSF) and in a duplexed SAMDI-MS biochemical assay against HRV3C and SARS-CoV-2 3CLpro activity to shed light on potency and selectivity. The benefits of the SAMDI-ASMS assay over traditional ASMS approaches and implications for improved drug discovery efforts are discussed.
Materials and Methods
Proteins, Peptides, and Compounds
Biotinylated-Avitag-HRV3C was purchased from Sigma (St. Louis, MO) (SAE0110) and used in all assays. SARS-CoV-2 3CLpro was purchased from Kactus (Shanghai, China). Peptide substrates and internal standards were purchased from BioPeptide (San Diego, CA) at >95% purity—3CLpro substrate: Ac-TSAVLQSGFRKK(biotin)-NH2; 3CLpro internal standard: Ac-SAYRKK(biotin)-NH2; HRV3C substrate: biotin-REEVLFQGPG-NH2; HRV3C internal standard: biotin-REEVGFQ-NH2. All other chemicals were purchased from Sigma-Aldrich (St. Louis, MO). The SAMDI biochip arrays were prepared as previously reported 8 and quality controlled for self-assembled monolayer formation by assessing reproducible binding of a series of control peptides before use in assays.
HRV3C ASMS Assay
HRV3C binding assays were performed in a 5 µL volume in 384-well low-volume polypropylene microtiter plates (Greiner Bio-One, Kremsmünster, Austria; cat. 784201) at ambient temperature. The optimized assay buffer was 20 mM HEPES (pH 7.5), 10 mM NaCl, 1 mM EDTA, 0.005% Tween-20, 0.01% bovine skin gelatin (BSG), 1 mM DTT. For compound screening, HRV3C (final concentration 200 nM) was added using a Multidrop Combi (Thermo Scientific, Waltham, MA) and incubated for 30 min with a pool of eight small molecules (15 µM final concentration each) to allow for slow on rates. For SAMDI-MS analysis, 2 µL of each mixture was transferred using a 384-channel automated Thermo PlateMate liquid handler to SAMDI biochip arrays functionalized with a neutravidin-presenting self-assembled monolayer, as previously reported.8,9 The SAMDI arrays were incubated for 60 min in a humidified chamber to prevent evaporation and allow for specific immobilization of the biotinylated HRV3C protein, along with any bound small molecules. The SAMDI arrays were purified by a rapid <3 s gentle wash step with deionized ultra-filtered water (50 µL/spot) as previously reported 8 and dried with compressed air. The matrix solution comprising 20 mg/mL alpha-cyano cinnamic acid (CHCA), demonstrated in an earlier study to be optimal for the ionization of small molecules, 8 was prepared in 80% acetonitrile–20% aqueous ammonium citrate and 0.3% trifluoroacetic acid and was applied in an automated format using a Combi Nano by dispensing 50 nL to each spot in the array. SAMDI-ASMS was performed using the reflector positive mode on an AB Sciex TOF-TOF 5800 System (AB Sciex, Framingham, MA) with 400 shots/spot analyzed by a random raster sampling of the entire spot (20 shots/subspectrum with 20 subspectra pass acceptance), 400 Hz laser frequency, bin size of 1 ns, and detector voltage multiplier of 0.48. A mass window of m/z 230 to m/z 900 was used and a mass threshold of m/z 0.5 applied for peak identification.
ASMS Data Analysis
Internally developed custom software 8 relies on an input data sheet that includes the exact mass of each compound along with its corresponding plate ID and well location. MS detection of small molecules is dependent on ionization efficiency, which varies among different compounds. The >100,000 compounds screened here feature an average purity >95%, and >95% of the compounds can be observed by MALDI MS in the positive reflector mode (data not shown), in line with previous results using MALDI. 7 The area under the curve (AUC) of each peak that corresponds to the mass ID (mass tolerance m/z 0.5) in the provided input sheet is measured from the raw spectra for each well on the SAMDI array and reported along with the AUC of the tri(ethylene glycol) (EG3) terminated monolayer molecule, which serves as an internal comparator having an m/z 335.2. All masses generate a nonzero AUC value given that the baseline in the MS spectra is never zero. To calculate the relative signal value (RSV) for each compound, the AUC of each compound is divided by the sum of that same compound’s AUC and the AUC of the internal comparator. The extent of binding and ionization efficiency are two factors that may contribute to the overall RSV. The ratio of the RSV calculated for each compound analyzed in the presence of the target over the RSV from the absence of the target provides the signal-to-background (S/B) value. The S/B value informs on the selectivity of the compound to omit compounds that may bind to the surface. This is particularly important when using neutravidin for affinity immobilization as compounds may bind to the neutravidin protein. The average RSV across the screen was 0.06 with a standard deviation of 0.11. Importantly, the average RSV does not fluctuate more than 0.01 when analyzed on a plate-by-plate basis, supporting hit calling based on full-screen data rather than per plate. Hit calling with SAMDI-ASMS starts by setting thresholds for the RSV and S/B ratio. Based on these data, an RSV threshold of 0.2 was selected as it represents at least 1 standard deviation above the average, and an S/B threshold of 2.5 was selected, close to twofold above the average across the screen (1.3). Using these values, a 0.4% hit rate was achieved. These thresholds can be adjusted based on the desired hit rate, the library to be screened, and the concentration of the compound.
Duplexed SAMDI-MS Biochemical Assay for 3CLpro and HRV3C
Protease assays were performed in a 6 µL volume in 384-well plates. 3CLpro and HRV3C activities were measured using a recently described duplexed SAMDI-MS assay. 10 Assay robustness was determined by the Z factor, calculated using the following equation: Z factor = 1 – 3(σ+ + σ–)/(µ+ – µ–), where σ is the standard deviation and µ is the average conversion of positive (+) and negative (–) controls. 11 GraphPad Prism (San Diego, CA) was used to calculate IC50 values, and Hill slopes were generated using a four-parameter logistics fit. IC50 values were determined with a confined bottom at 0% inhibition and confined top at 100% inhibition, with the exception of STX32611319, which was not confined to 100% inhibition.
ASMS High-Throughput Screening
The 102,400 compounds included in this screen (SAMDI Tech collection) were derived from a diverse compound library devoid of pan-assay interference compounds (PAINS) synthesized in 2020 and dissolved in DMSO, with each compound achieving an average purity >95%. Compounds were screened in pools of eight (15 µM final concentration for each compound, 1.2% final DMSO concentration) by stamping 60 nL of 1.25 mM compound solutions into 384-well plates using a Mosquito HTS Automated Liquid Handler (SPT Labtech, Melbourn, Hartfordshire, UK). The compound pooling was conducted in a manner that reduced the overlap of compounds in the same well with the same mass. The assay was performed as described above. A selection of compounds were acquired as fresh powder and verified by liquid chromatography (LC)–MS (data not shown) before each compound was assayed individually in a duplicate hit confirmation step, followed by dose response in the ASMS assay and biochemical assays against 3CLpro and HRV3C.
DSF Assay
Thermally induced denaturation of HRV3C was evaluated in 384-well format in duplicate using a Bio-Rad C1000 thermal cycler over a temperature gradient from 25 to 80 °C, with steps every 0.5 °C, 30 s at each temperature. HRV3C was diluted to 5 µM in assay buffer in the presence of either 10 µM or 50 µM compound and 10 µM SYPRO Orange. Negative controls included DMSO only. Compounds exhibiting a shift in melting temperature (Tm) >0.5 °C were considered binders.
Results
Development of a High-Throughput ASMS Assay Using SAMDI Technology
The schematic of the workflow for an ASMS assay using SAMDI is presented in Figure 1 and described in detail in the Materials and Methods section. The reactions were initiated by incubating the compounds in the presence and absence of 200 nM biotinylated HRV3C in solution for 30 min in a 384-well plate. Next, the reaction solutions were transferred to neutravidin-presenting high-density SAMDI biochip arrays to selectively immobilize the biotinylated HRV3C along with any compounds in complex with the protein. The biochip arrays were then rapidly washed (<3 s) to remove unbound compounds and buffer components prior to matrix addition and SAMDI-ASMS analysis. This straightforward and general workflow can be adapted for screening virtually any target.
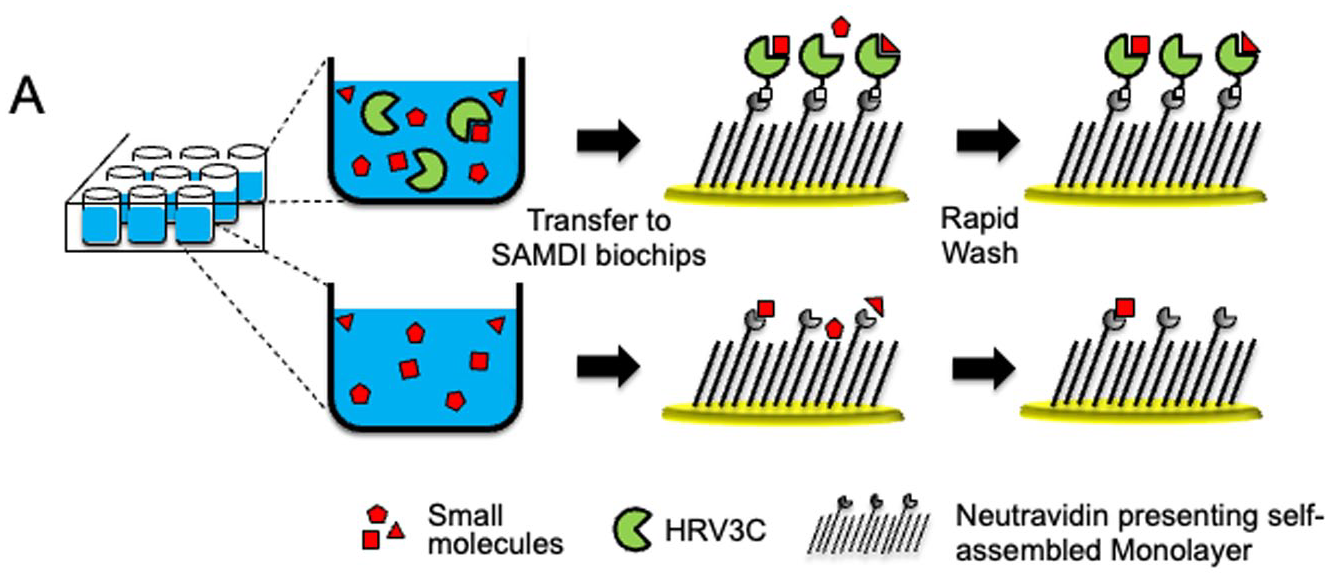
SAMDI-ASMS workflow. Pools of eight compounds are incubated in the presence (top) or absence (bottom) of the target of interest in a 384-well plate in buffer. The reactions are then transferred to high-density biochips to specifically immobilize the target of interest. The biochips are then washed to remove unbound compounds and analyzed by SAMDI-MS.
High-Throughput SAMDI-ASMS Screen of 102,400 Compounds against HRV3C
One of the foundations of the SAMDI technology is that upon laser irradiation, the self-assembled monolayer desorbs from the surface and can be detected by MALDI MS as an intact, singly charged species. 12 In the ASMS assay, the monolayer comprises a biotin terminated alkanethiolate and an EG3 terminated alkanethiolate, the latter of which is observed at m/z 335.2 ( Fig. 2A ). The EG3 alkanethiolate not only minimizes nonspecific adsorption of proteins to the surface, but also, since it is present at the same density on each spot, serves as an internal comparator to identify potential binders. 8 By comparing the AUC for a given compound peak to the AUC of the monolayer molecule, an RSV can be calculated for each compound (see Materials and Methods for more details). Given the immobilization step using affinity capture onto the SAMDI arrays, it is important to dismiss compounds with a propensity to bind to the background surface. To identify selective binders, the compounds are analyzed in the presence and absence of the target, essentially treating the neutravidin surface as a counterscreen. During analysis, the ratio of the RSV in the presence of the target to the RSV in the absence of the target represents the S/B value that ultimately informs on compound selectivity. In one example, the spectrum in Figure 2A displays a peak at m/z 460.3 that corresponds to compound A. The RSV is approximately 0.6 in the presence of the target and 0.05 in the absence of the target, with an S/B ratio >10. Conversely, the peak at m/z 419.4 (compound B) has an RSV of approximately 0.5 in the presence and absence of the target, generating an S/B ratio close to 1. The RSV and S/B value are the key data points that identify compound A as a selective binder (or designated a hit) and compound B as a nonselective binder (not a hit).
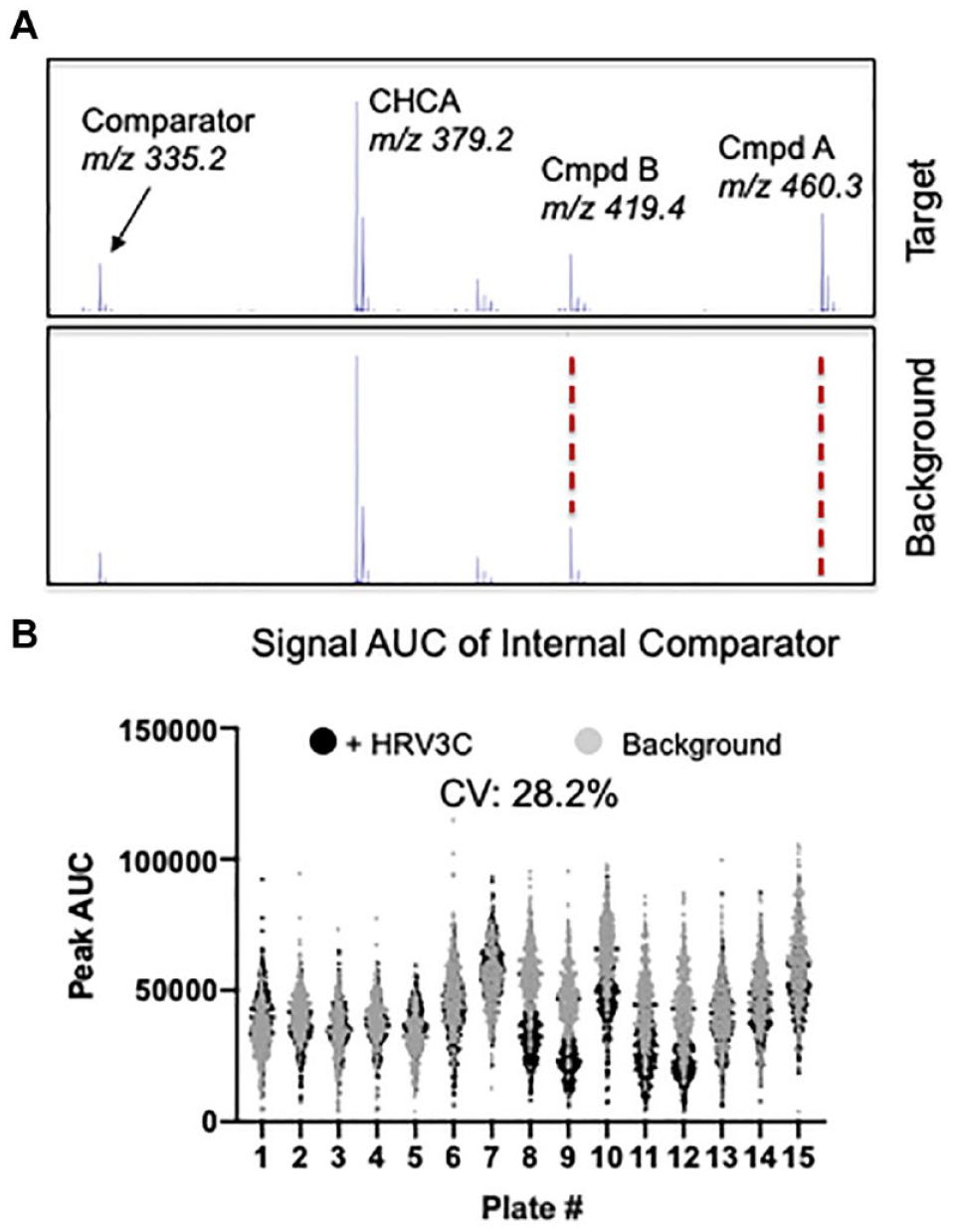
SAMDI-ASMS data acquisition and analysis. (
To demonstrate the high-throughput screening capability of SAMDI-ASMS, 102,400 compounds from a diverse small-molecule library were screened for binders to the HRV3C protease. The SAMDI-ASMS primary screening assay and data generation of 102,400 compounds were completed in <8 h, enabled by pooling 8 compounds per well and the high-throughput nature of the SAMDI technology. 13 The intensity of the internal comparator peak was evaluated throughout the screening process by measuring the AUC in the presence and absence of HRV3C ( Fig. 2B ). The coefficient of variation (CV) across all screening plates was determined to be 28.2%, consistent with the variability observed across other MALDI platforms for absolute signal. 7 These data underscore the importance of using the ratiometric analysis to account for the variability of a MALDI signal influenced by differences in matrix preparation and crystallization, for example. After completion of the screen, custom automated software was used to measure the AUC for peaks corresponding to the mass of each compound in the library. From these data, the RSV and S/B value can be determined. The average RSV across all compounds was 0.06 with a standard deviation of 0.11, and the average S/B ratio was 1.3. For hit calling, an RSV threshold of 0.2 was selected as it represents a value greater than 1 standard deviation above the average RSV, while the S/B threshold was set at nearly twice the average value with a cutoff of 2.5. Based on these thresholds, 414 compounds, corresponding to a 0.4% hit rate, were selected for further evaluation in follow-up assays as individual compounds.
SAMDI-ASMS Hit Confirmation and Concentration Response Analysis
The 414 compounds were reevaluated in a single concentration (15 µM) in duplicate as an individual compound (no longer pooled) in the presence and absence of 200 nM HRV3C. The RSVs achieved for each replicate support the reproducibility of the approach (R2 = 0.736 and 0.728 for plus and minus HRV3C, respectively) (
Fig. 3A
). Some variation is expected among independent replicates due to the dynamic nature of the target and for weaker binding compounds. These data confirmed 50 compounds that met the hit calling thresholds (12% validation rate), representing at least a fivefold greater rate than those achieved by other ASMS methods.7,14 Next, the 50 compounds were titrated starting at a 200 µM top concentration in a 6-point, threefold dilution series and analyzed in duplicate in the presence and absence of HRV3C. Of the 50 compounds tested, 15 generated a dose response. While the dose response analysis does not calculate a KD, it is well suited for rapidly rank ordering compounds for further evaluation, which highlighted six representative compounds (
Fig. 4
). Some compounds exhibit specific binding behavior relative to the background surface (
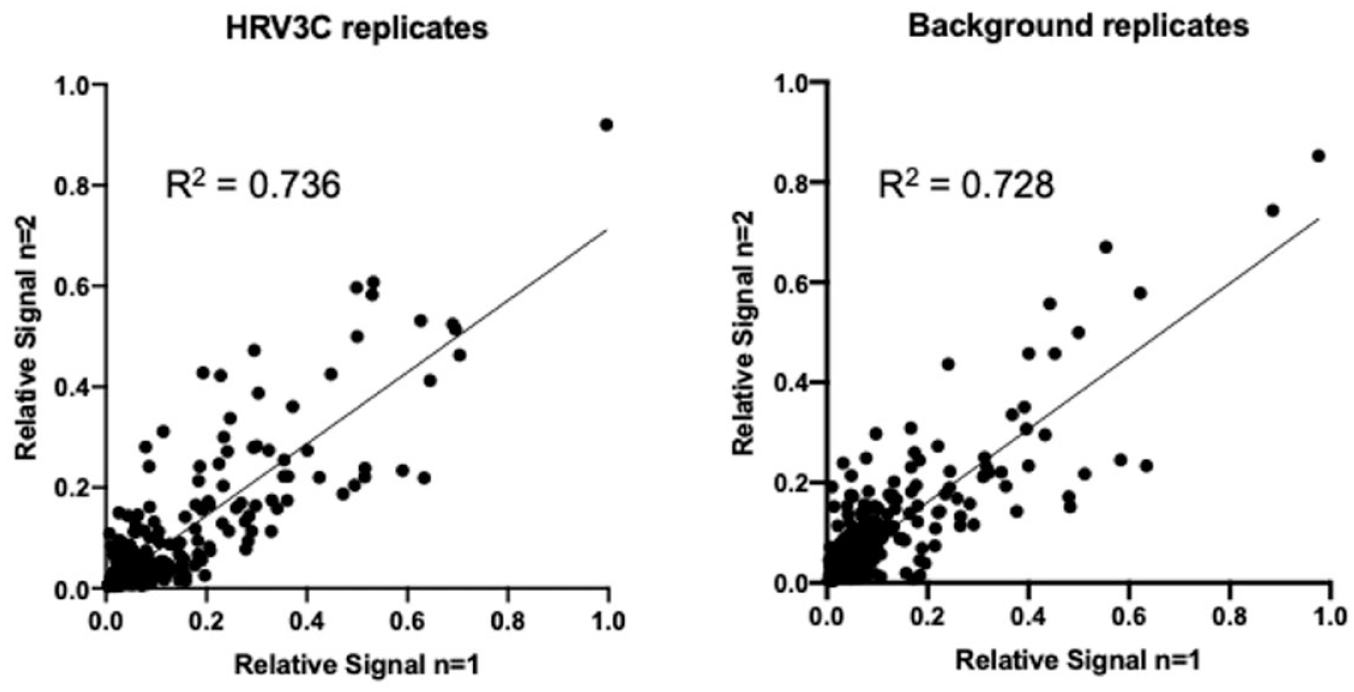
Confirming hits using SAMDI-ASMS. The relative signals for compounds tested for binding in the presence (left) and absence (right) of HRV3C are compared.
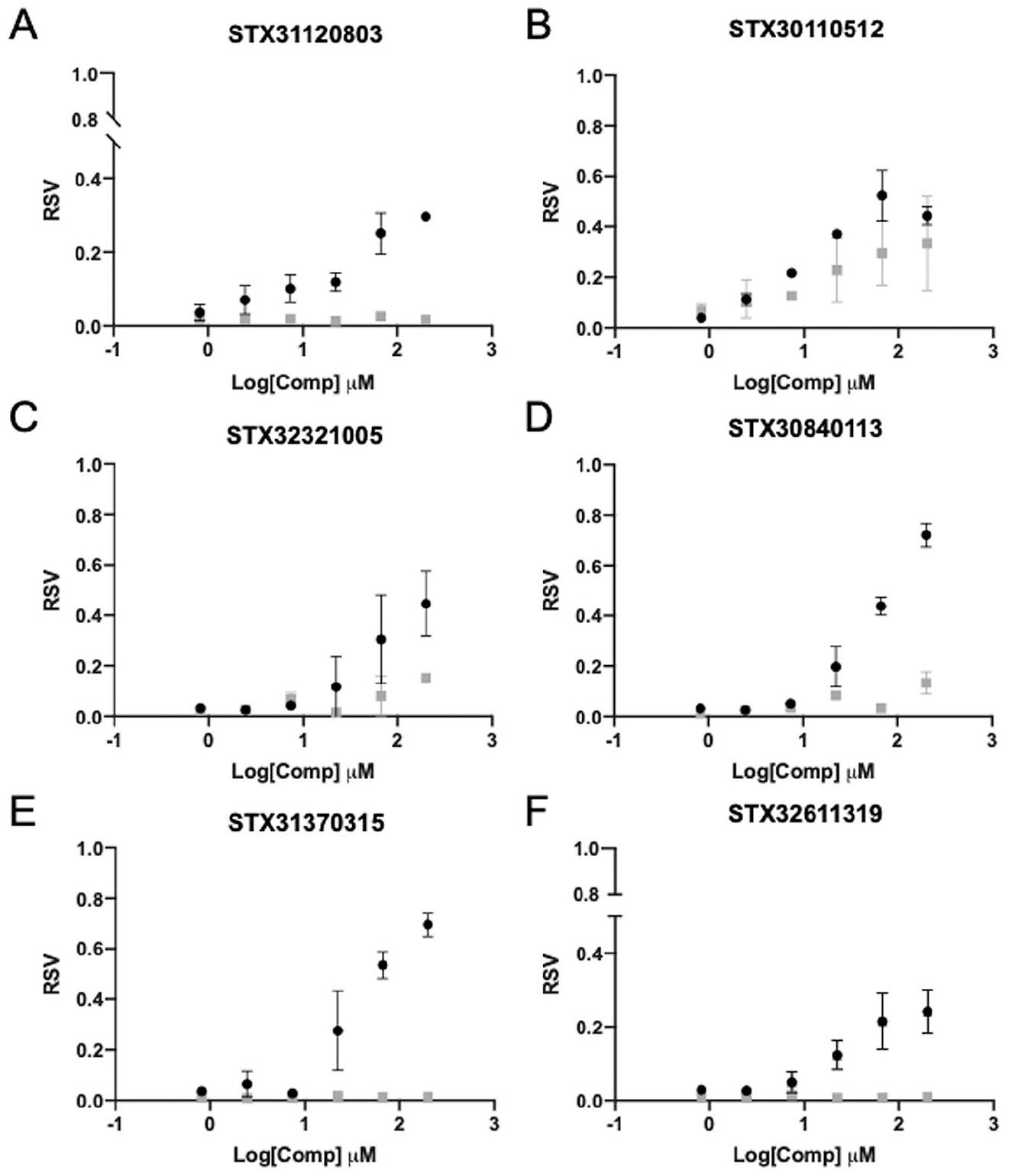
Concentration response analysis using SAMDI-ASMS. Representative concentration–response curves for six compounds shown in the presence (black circles) and absence (gray squares) of HRV3C. Points represent duplicate data.
Validation of HRV3C Binders in Biochemical Protease Assays
The six compounds were evaluated in functional assays to determine their ability to inhibit the protease activity of HRV3C along with the SARS-CoV-2 3CLpro enzyme as a counterscreen. For this, a recently described duplexed SAMDI-MS functional assay was used that simultaneously measures the protease activity of HRV3C and 3CLpro (also known as Mpro).
10
The assay was optimized and miniaturized with Z′ factors >0.9 for 3CLpro and 0.7 for HRV3C and validated with reported 3CLpro inhibitors.
10
The six lead compounds identified by the ASMS screen (
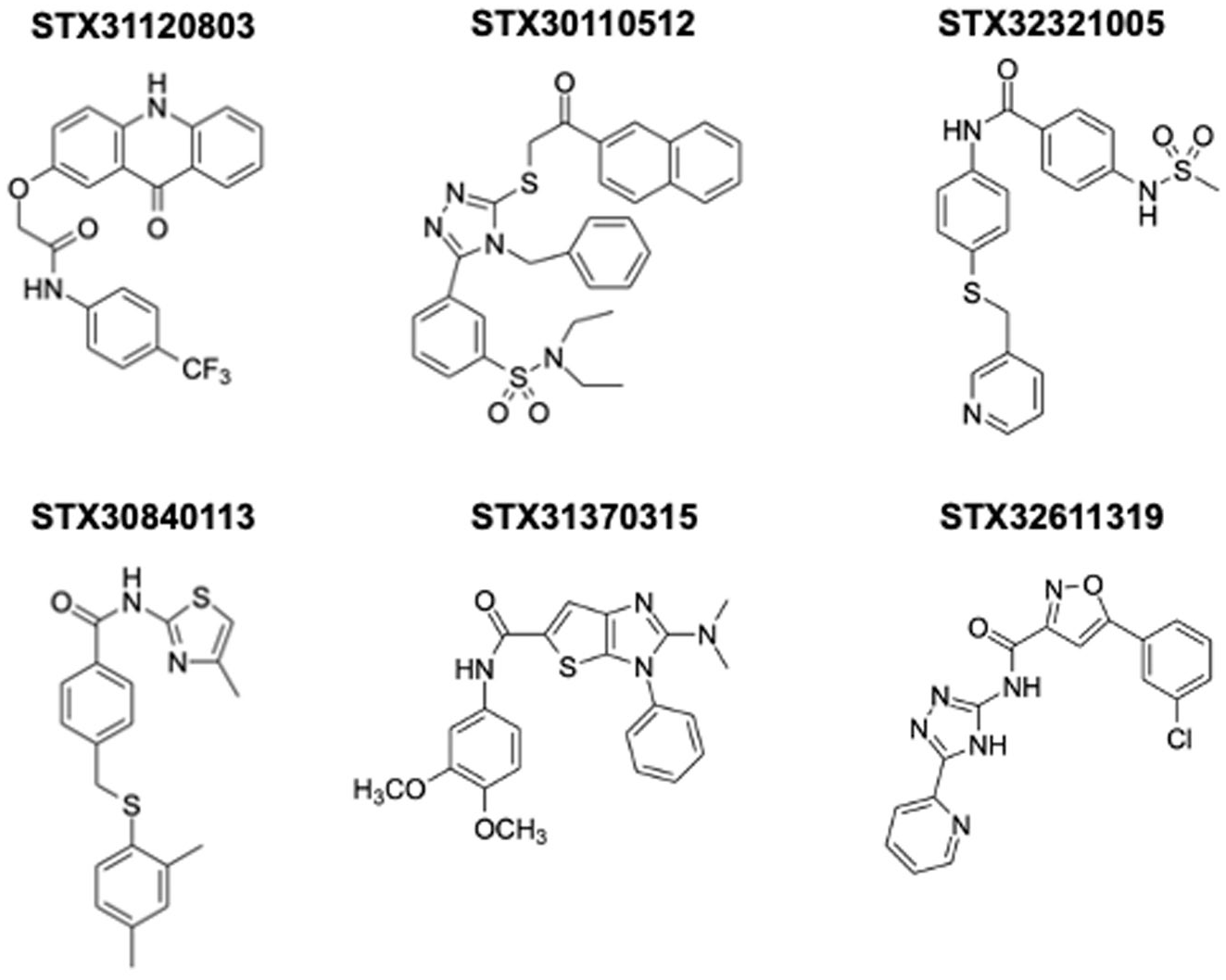
Structures of HRV3C binders identified by SAMDI-ASMS.
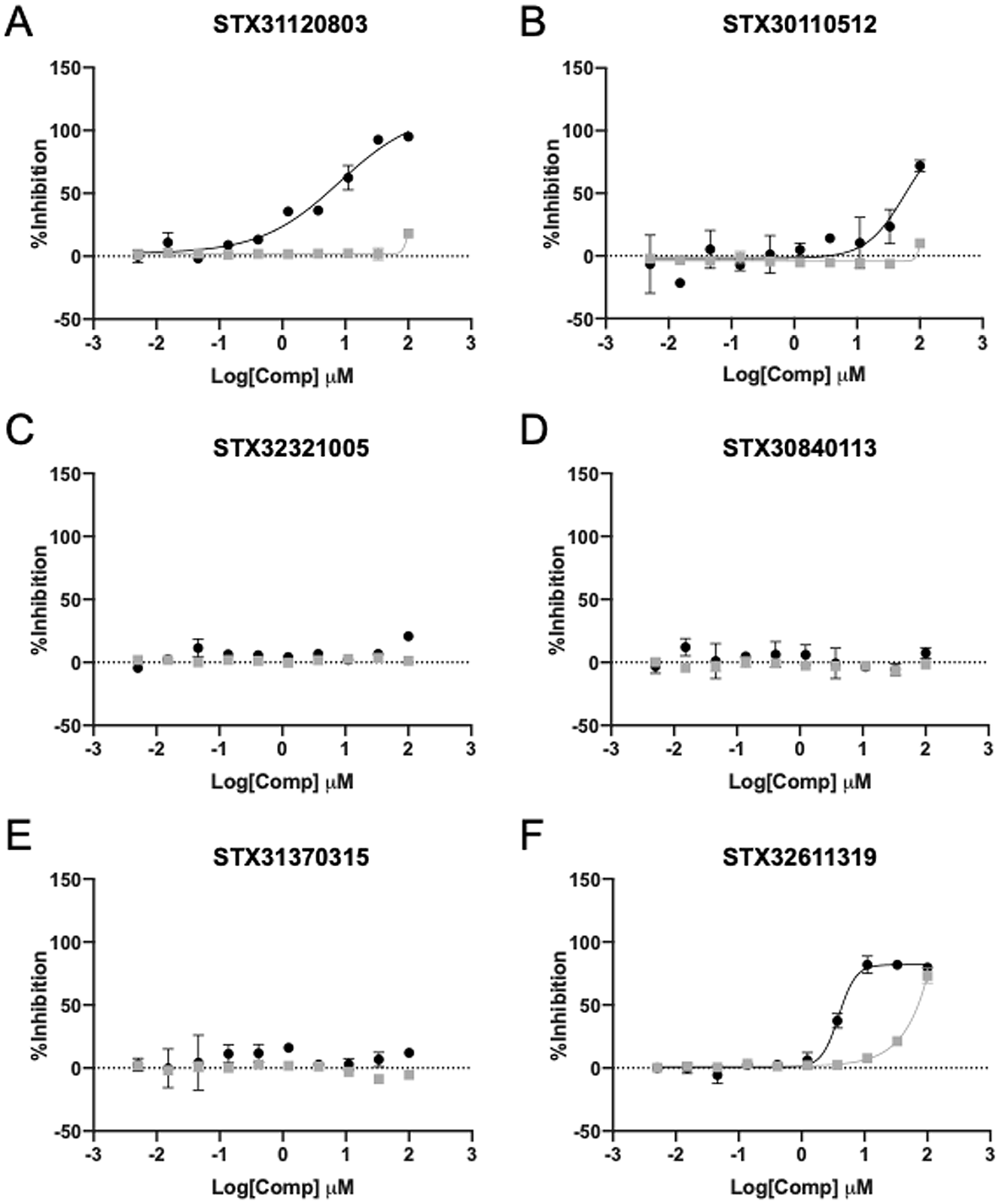
Biochemical analysis of HRV3C binders. Six compounds were evaluated in a duplex SAMDI-MS assay that reports on HRV3C (black circles) and 3CLpro (gray squares) activity.
DSF Analysis of HRV3C Binders
To assess the SAMDI-ASMS hits in an orthogonal binding assay, the six compounds from Figure 5 were analyzed by DSF. DSF is a technique that can infer interactions of small molecules with proteins by measuring compound-induced changes in the thermal stability, or melting temperature (Tm) of the protein. 15 The assay was set up by analyzing the same HRV3C protein (5 µM final concentration) in the presence of 10 µM and 50 µM of the six compounds in 5% DMSO final concentration. Three of the six compounds generated a ΔTm >0.5 C, while two others generated ΔTm values of 0.42 °C and 0.47 °C, near the general threshold used to indicate a weak binder. Table 1 summarizes the DSF results and SAMDI-MS functional assays. Interestingly, the compound that failed to show binding in the DSF assay is the most potent biochemical inhibitor of HRV3C. These data may be explained by a binding event that does not impact thermal stability, or an affinity beyond the limit of detection of DSF. Taken together, SAMDI-ASMS combined with orthogonal validation assays rapidly identified novel chemical matter that engages the HRV3C protease, offering new starting points for antiviral therapeutics.
Validation of HRV3C Binders Identified by SAMDI-ASMS.
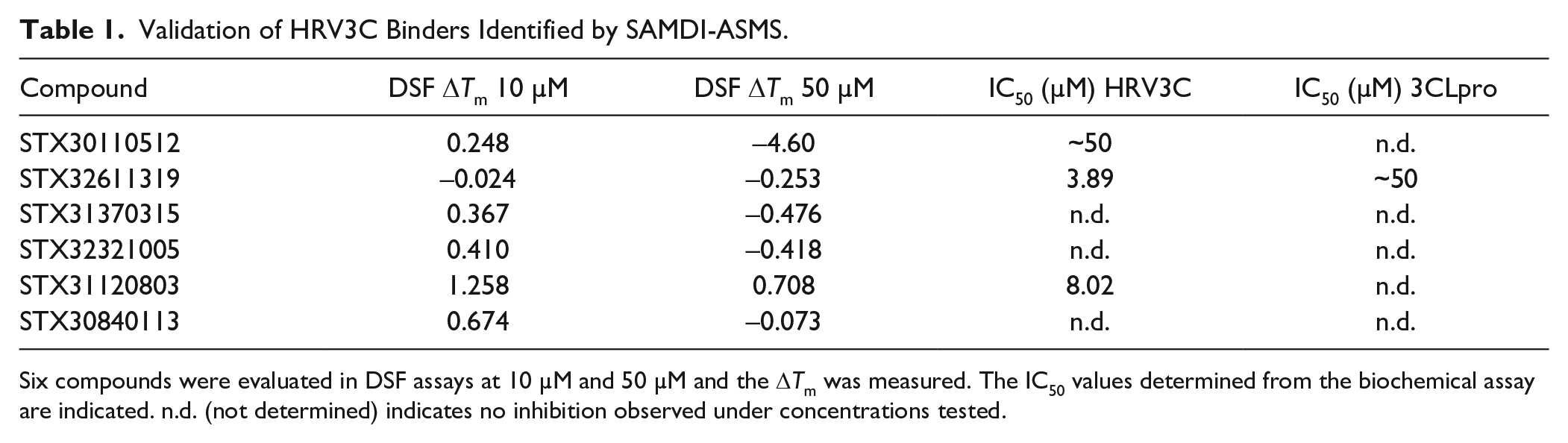
Six compounds were evaluated in DSF assays at 10 µM and 50 µM and the ΔTm was measured. The IC50 values determined from the biochemical assay are indicated. n.d. (not determined) indicates no inhibition observed under concentrations tested.
Discussion
This report is the first to employ SAMDI-ASMS as a high-throughput screening methodology by screening more than 100,000 compounds to identify small-molecule binders of the HRV3C protease. The significance of this study is twofold. First, to date there remains no approved small-molecule inhibitors for HRV3C, a viral protease implicated for function and replication of the common cold virus. Reported inhibitors have largely focused on peptidomimetics,16,17 many of which form a covalent bond with the active site cysteine. Covalent binders are not easily detected by SAMDI-ASMS since the resolution of MALDI is not sufficient to distinguish a small molecule bound to a large analyte. While this represented a challenge since well-characterized noncovalent binders to HRV3C were not available as controls, the pharmacophore-like small molecules identified here offer an attractive opportunity for therapeutic intervention, including through functional inhibition or targeted protein degradation strategies. Second, there remains a need for high-throughput and robust screening assays suitable for identifying ligand–target interactions. Here we expand the capabilities of the SAMDI technology, extensively reported as a powerful label-free and high-throughput assay for measuring biochemical and chemical reactions,9,18–28 to offer a high-throughput ASMS assay with key benefits over traditional ASMS approaches.
SAMDI-ASMS as a High-Throughput Binding Assay
The SAMDI-ASMS assay benefits from a straightforward, rapid workflow with automation integrated at each step to maximize efficiency and minimize human error. The use of self-assembled monolayers of alkanethiolates on gold offers key advantages for running ASMS assays. First, monolayers presenting oligomers of ethylene glycol are among the most effective at minimizing nonspecific adsorption of proteins to human-made materials. 29 Second, the surfaces can be tailored to immobilize a variety of analytes, 30 eliminating the need for tedious and slow chromatography steps. Currently, immobilization requires incorporating a handle, such as a biotinylated Avi-tag or His-tag for proteins, although opportunities for traceless immobilization strategies have been reported.27,28 It is important to ensure that the presence of a handle does not alter the protein structure and binding properties of the compounds. Third, the surfaces are efficient substrates for MALDI TOF MS analysis, 12 which represents the basis of the SAMDI technology.
Hits identified by SAMDI-ASMS are moved to validation assays as individual compounds followed by analysis in a dose response format. This strategy allows for a rapid rank ordering of the hits, rather than measuring a KD, which can be determined in downstream lower-throughput biophysical approaches. In this screen, performing the primary screen and data generation required less than 8 h. Generating data for the hit confirmation and dose response phases were each completed in a single day, highlighting the rapid hit identification enabled by SAMDI-ASMS. Choosing validation assays and understanding their benefits and limitations aid in properly assessing ASMS leads. Here, a functional assay was employed for HRV3C with the SARS-CoV-2 3CLpro enzyme as an off-target protease. Half of the SAMDI-ASMS binders tested showed selective enzymatic inhibition, supporting their engagement with HRV3C. The fact that some ASMS binders did not exhibit functional activity is not unexpected, as binding events may not elicit a functional response, but may offer a path toward developing compounds that operate through other mechanisms, such as targeted protein degradation. For an orthogonal binding assay, each compound was also assessed in DSF, which measures the ability of a compound to alter protein thermal stability. Several compounds generated a shift indicative of binding. It is important to consider that the DSF conditions are different than those used in the SAMDI-ASMS assay (25× higher protein concentration and no detergent in the buffer), which may impact binding. Moreover, compounds may bind but not elicit a change in thermal stability. Finally, the limit of detection of DSF is around 50 µM, suggesting that weak binders and fast off-rate compounds may be difficult to observe. Taken together, it is important to appreciate that each assay and methodology has advantages and potential caveats. Understanding functional aspects of each assay format and its limitations, while exploring multiple approaches, is critical in discerning the most promising compounds for hit-to-lead and preclinical phases.
A Comparison of SAMDI-ASMS with Other ASMS Methodologies
Integrating ASMS workflows in drug discovery continues to gain momentum, particularly as targets become more challenging and nonfunctional binders offer promising therapeutic avenues. When evaluating ASMS methodologies, it is important consider overall throughput, reagent consumption, and data quality. Traditional ASMS approaches rely on tedious column chromatography steps that limit throughput. To compensate, hundreds or thousands of compounds are pooled in a single reaction. 6 One example, termed automated ligand identification system (ALIS), relies on size exclusion chromatography (SEC) to first isolate the ligand–target complex coupled to an electrospray ionization LC-MS backend.6,31,32 Even with recent improvements in throughput for LC systems, 33 the approach requires the processing of each sample individually. The recent report of a plate-based filter assay coupled with compound detection by MALDI MS benefits from a higher throughput, evidenced by screening 23,000 compounds in 24 h. 7 SAMDI-ASMS also benefits from the high-throughput capability of the MALDI instrument13,34 and the purification of up to 1536 samples in seconds through the immobilization approach. Another limitation of ASMS is the reagent requirement, with various approaches using 5–10 µM of the target.6,7,14,35 Conversely, SAMDI-ASMS technology benefits from up to 50-fold less concentration of the analyte, opening avenues to screen previously intractable targets that may suffer from low expression. Lastly, go or no-go decisions rely on the quality of data generated. Although one may qualify a screening campaign as successful even if only a single hit is identified, validation rates from initial hit finding through dose response analysis inform on robustness and the quality of the library. In this screen, of the 414 initial hits (0.4% hit rate), 50 were validated when assessed as individual compounds, a validation rate of 12%, a rate over five times higher than traditional ASMS approaches.7,14 There are several factors to consider when assessing the validation rate. First, multiple compounds may be called a hit from the primary screen if they have an exact mass (within m/z 0.5 for SAMDI-ASMS) of another compound in the pool. This rare occurrence will lead to two or more compounds being called hits, where based on probability, only one is likely a true binder, thereby decreasing the validation rate. Second, some compounds may appear as hits in the primary screen due to misbehavior of another compound in that well and will therefore not appear as a hit when assessed individually. The SAMDI-ASMS approach achieves higher validation rates due to utilization of smaller compound pools, and testing compounds against the background surface further assists in omitting misbehaving compounds early.
In conclusion, this study showcases the first application of SAMDI-ASMS as a high-throughput ASMS screening tool, evaluating 102,400 compounds in <8 h (in the presence and absence of the target) to reveal novel ligands for the HRV3C protease. The SAMDI-ASMS assay described here offers several solutions over traditional ASMS approaches, including faster throughput, higher validation rates, and a straightforward and streamlined approach amenable to screening virtually any target. Moreover, the compounds identified in this screen offer novel leads to identify specific and pan-viral inhibitors and will be the focus of future work in hit-to-lead optimization efforts.
Footnotes
Acknowledgements
The authors wish to thank Sandra Dib for assistance with compound handling and the SAMDI Tech team for critical discussions.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
