Abstract
The COVID-19 pandemic, caused by the SARS-CoV-2 virus, has a huge impact on the world. Although several vaccines have recently reached the market, the development of specific antiviral drugs against SARS-CoV-2 is an important additional strategy in fighting the pandemic. One of the most promising pharmacological targets is the viral main protease (Mpro). Here, we present an optimized biochemical assay procedure for SARS-CoV-2 Mpro. We have comprehensively investigated the influence of different buffer components and conditions on the assay performance and characterized Förster resonance energy transfer (FRET) substrates with a preference for 2-Abz/Tyr(3-NO2) FRET pairs. The substrates 2-AbzSAVLQSGTyr(3-NO2)R-OH, a truncated version of the established DABCYL/EDANS FRET substrate, and 2-AbzVVTLQSGTyr(3-NO2)R-OH are promising candidates for screening and inhibitor characterization. In the latter substrate, the incorporation of Val at position P5 improved the catalytic efficiency. Based on the obtained results, we present here a reproducible, reliable assay protocol using highly affordable buffer components.
Introduction
The coronaviruses are a group of positive-strand RNA viruses that infect mammals, birds, reptiles, and amphibians. In humans, several species of the group are important as pathogens causing various pathologies. These range from mild respiratory infections to life-threatening diseases such as severe acute respiratory syndrome (SARS). Coronaviruses are responsible for the SARS pandemic in 2002–2003, the Middle East respiratory syndrome (MERS) epidemic (ongoing since 2012), and the present COVID-19 pandemic (from 2019).1–3 The viral disease COVID-19 is caused by SARS-CoV-2, which was first described in Wuhan, China, in December 2019. 4 The present worldwide pandemic has, as of January 25, 2021, affected more than 99 million persons, with more than 2.1 million reported deaths. 5 In around 80% of the symptomatic infections, a mild disease with fever or mild pneumonia can be observed. Fourteen percent of the cases are more severe, and about 5% of the patients have to be treated in intensive care units. 6 The infection appears to be mainly transmitted by aerosols during social interactions. 7
Specific antiviral drugs against infections with SARS-CoV-2 and other coronaviruses have not yet been approved. The main protease, Mpro (3CLpro, nsp5), of coronaviruses has no closely related homologs in humans, is involved in an essential step of the viral life cycle, and is therefore considered a promising drug target. 8 Mpro is a chymotrypsin-like cysteine protease that cleaves the viral polyprotein into several functional proteins. The relevant proteolytic species is the dimer of the protease. 9 Mpro cleaves the polyproteins exclusively after glutamine residues at 11 sites.8,10,11 A number of compounds have been described to act as inhibitors of coronaviral Mpro. These frequently have a peptidic structure and contain an electrophilic group to promote covalent binding to the catalytic cysteine.12–17
Biochemical assays are valuable tools to study viral proteases and to identify, develop, and optimize protease inhibitors. A frequently used, high-throughput-capable technique to investigate the activity of proteases is Förster resonance energy transfer (FRET)-based assays. In FRET, the energy of an excited donor group is transferred to an acceptor moiety in close proximity, the “quencher.” The energy transfer is radiation-free and does not involve intramolecular emission and absorption of photons. By incorporation of a FRET pair into an enzyme substrate, the cleavage rate of the substrate can be investigated.18,19
We herein present the development and optimization of a biochemical FRET-based SARS-CoV-2 Mpro assay. Several assay conditions and additives such as salts, polyols, and detergents were studied. Furthermore, several established and new FRET substrates were synthesized, compared, and tested, aiming to provide a robust and cost-effective protocol that can be carried out under a wide range of infrastructural conditions.
Materials and Methods
General Comments
FRET substrates were synthesized via the Fmoc (fluorenylmethyloxycarbonyl protecting group) solid-phase peptide synthesis (SPPS) protocol, as described in the supplemental information. Other chemicals were purchased from commercial suppliers. Boceprevir was obtained from Biosynth-Carbosynth (Bratislava, Slovak Republic). The measurements were performed in black 96-well V-bottom plates (Greiner Bio-One, Germany) using a BMG Labtech Fluostar OPTIMA microtiter fluorescence plate reader at an excitation wavelength of 330 nm and an emission wavelength of 430 nm. All measurements were performed at room temperature.
Construct Design
The expression construct for the SARS-CoV-2 Mpro was designed by multiple sequence alignment of the Wuhan seafood market pneumonia virus isolates 2019-nCoV (accession numbers MN938384.1, MN975262.1, MN988713.1, and MN985325.1). The reading frame for the SARS-CoV-2 Mpro was determined by alignment with previous SARS-CoV Mpro expression constructs. 20 Codon usage was optimized individually for optimized expression in both Escherichia coli and eukaryotic systems. The gene sequence coding for the SARS-CoV-2 Mpro protease was ordered at Eurofins with NotI and BamHI restriction sites at the 3′ and 5′ ends, respectively. The gene sequence encoding the SARS-CoV-2 Mpro was inserted by restriction-based cloning into a pET28a(+) expression vector to obtain a C-terminal His-tag. The C-terminal His-tag was chosen because a structural analysis of SARS-CoV Mpro indicated less disruption of dimerization in the presence of a C-terminal tag compared with an N-terminal tag. The cleavage inactive mutant C145A was cloned from the wild-type construct by molecular assembly with two sets of primers. The identity of the constructs was determined by agarose gel electrophoresis, colony PCR, and sequencing.
Expression and Purification of SARS-CoV-2 Mpro
The plasmid encoding the SARS-CoV-2 Mpro was transformed into E. coli BL21 DE3 cells for expression. Overnight culture from a single colony was grown by shaking at 37 °C in LB medium supplemented with 50 mg/mL kanamycin. On the next day, prewarmed LB medium with kanamycin was mixed 1:40 with overnight culture and bacteria were grown until the optical density (OD) at 600 nm reached 0.3–0.4. Subsequently, the temperature was reduced to 25 °C. Protein expression was induced by addition of 1 mM isopropyl-
Purification of the SARS-CoV-2 Mpro was performed on ice or at 4 °C. One gram of cell pellet was thawed and resuspended in 10 mL of ice-cold buffer A (20 mM Tris, 200 mM NaCl, 10 mM imidazole, pH 7.6). The bacteria were lysed using a high-pressure cell disrupter (One Shot, Constant Systems LTD). Cell debris was centrifuged 2 h at 50,000g to remove insoluble aggregates and inclusion bodies. The supernatant was mixed after centrifugation with 1 mL of preequilibrated (buffer A) Ni-NTA-beads and incubated on a rolling shaker for 30 min at 4 °C. Subsequently, the Ni-NTA-beads were washed with buffer A, containing increasing concentrations of imidazole (10 mM, 20 mM, 50 mM, pH 7.6; 10–20 mL buffer per step). The protein was eluted by multiple elution steps, each with 1 mL of buffer B (20 mM Tris, 200 mM NaCl, 500 mM imidazole). Elution fractions were analyzed by OD measurement and sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE). Fractions containing the SARS-CoV-2 Mpro were concentrated and buffer was exchanged with buffer C (20 mM Tris, 200 mM NaCl, 1 mM DTT, 1 mM EDTA, pH 7.6) using Amicon centrifugation filters. Protein was further purified by size-exclusion chromatography using an S200 column and buffer C as running buffer. Fractions with pure protein were concentrated and mixed with 50% sterile glycerol, and aliquots at 8 mg/mL (~228 µM) were flash frozen in liquid nitrogen and stored at –80 °C until further use.
Evaluation of Assay Buffer Composition
Buffering compounds, salts, additives, polyols, and detergents were evaluated by preparing a buffer solution with corresponding concentrations of the components. In the experiment evaluating the buffer components, 50 mM Tris-HCl, phosphate, and HEPES buffer were prepared without the addition of other components at pH 7.5. In the experiment with salts and additives, 50 mM Tris-HCl buffer, pH 7.4, was used with the addition of NaCl at various concentrations (50 mM, 100 mM, or 150 mM), 1 mM DTT, 1 mM TCEP, or 1 mM EDTA. For the evaluation of polyols, ethylene glycol (10%, 20%, or 30% v/v) or glycerol (10%, 20%, or 30% v/v) was added to the 50 mM Tris-HCl buffer, pH 7.4. The influence of detergents was evaluated using 50 mM Tris-HCl buffer, pH 7.4, with the addition of 0.01% detergent. In all experiments, the prediluted solution of Mpro was prepared by pipetting the corresponding volume of the enzyme storage buffer (enzyme concentration 228 µM) and adding the tested buffer to obtain an enzyme concentration of 5 µM. Substrate
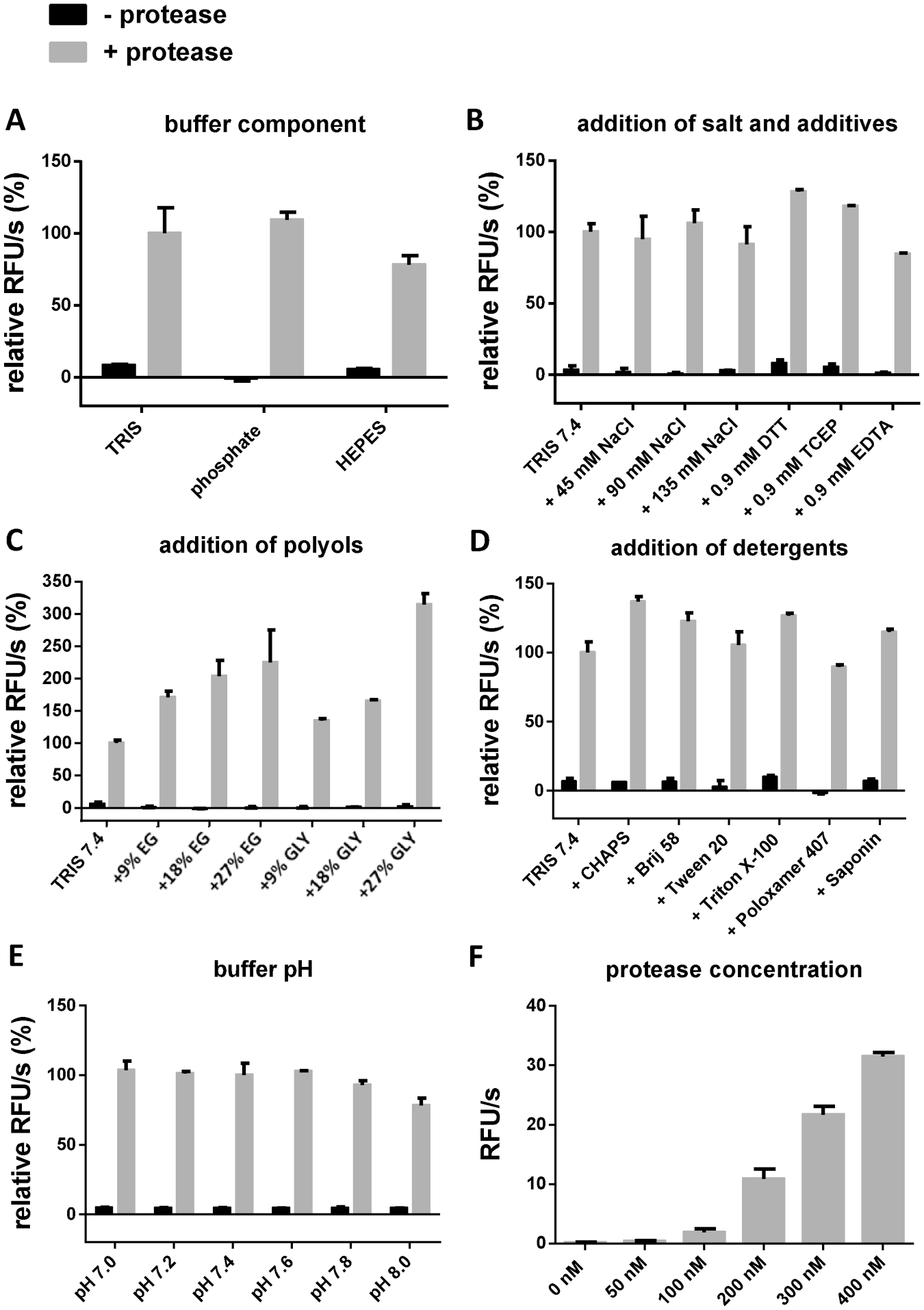
Buffer optimization of the FRET assay. The measurements were performed with substrate
Evaluation of pH Value
The assay buffer (50 mM Tris-HCl, 100 mM NaCl, ethylene glycol [20% v/v], 0.0016% Brij 58) was prepared with the pH value varying by 0.2 units in the range 7.0–8.0. The prediluted solution of the protease was prepared by pipetting the required volume of the enzyme in storage buffer (enzyme concentration 228 µM) and adding the assay buffer, pH 7.4, to obtain an enzyme concentration of 10 µM. The prediluted solution of substrate
Evaluation of Protease Concentration
The prediluted solution of protease was prepared by pipetting the corresponding volume of the enzyme storage buffer (enzyme concentration 228 µM) and adding the assay buffer (50 mM Tris-HCl, pH 7.4, 100 mM NaCl, ethylene glycol [20% v/v], 0.0016% Brij 58) to obtain an enzyme concentration of 5 µM. The prediluted solution of substrate
Evaluation of Substrates
The Mpro prediluted solution (prepared in assay buffer) was pipetted in the wells (10 μL), followed by assay buffer (80 μL, 50 mM Tris-HCl, pH 7.4, 100 mM NaCl, ethylene glycol [20% v/v], 0.0016% Brij 58). Reactions were initiated by the addition of 10 μL of the corresponding substrates (prepared in the assay buffer). The final protease concentration in the assay was 300 nM, whereas the concentration of the substrate was 50 µM. The final volume was 100 µL per well. The enzymatic activity was monitored for 15 min and determined as a slope of relative fluorescence units per second for each substrate. The results are expressed as relative fluorescence units per second, as a mean of the triplicates and the respective standard deviation. In each measurement, a corresponding solution without the addition of the protease was used as a negative control.
Kinetic Measurements
The required volumes of prediluted selected substrates, dissolved in assay buffer, to yield final concentrations in the range of 0–400 μM were pipetted into the wells, followed by a corresponding volume of the assay buffer (50 mM Tris-HCl, pH 7.4, 100 mM NaCl, ethylene glycol [20% v/v], 0.0016% Brij 58). Reactions were initiated by the addition of 10 μL of the prediluted protease (prepared in assay buffer) to obtain a final enzyme concentration of 300 nM. The final volume was 100 μL per well. The enzymatic activity was monitored for 15 min and determined for each concentration as a slope of relative fluorescence units per second. The obtained values were divided by the correction factor for each concentration. Additional details are provided in the supplemental information (see
Z′ Value Determination
The Mpro prediluted solution (final concentration 300 nM) together with the assay buffer (50 mM Tris-HCl, pH 7.4, 100 mM NaCl, ethylene glycol [20% v/v], 0.0016% Brij 58) was pipetted in 47 wells. The same volume of the assay buffer without the protease was pipetted in 48 wells. After 15 min of preincubation, substrate
SARS-CoV-2 Main Protease Relative Inhibition Assay
The prediluted solutions of the reference compounds (500 μM) were obtained by diluting the compound stock solution (10 mM in DMSO) in the assay buffer (50 mM Tris-HCl, pH 7.4, 100 mM NaCl, ethylene glycol [20% v/v], 0.0016% Brij 58). The prediluted solution of protease (3 µM) was prepared by adding the corresponding volume of the assay buffer to the enzyme storage buffer (enzyme concentration 228 µM). The prediluted solution of the protease (10 µL), the assay buffer (60 µL), and the prediluted solution of the inhibitor (10 µL) were incubated for 15 min. The reaction was initiated by the addition of 20 µL of the prediluted solution of FRET substrate
SARS-CoV-2 Mpro Assay for IC50 Determination
Eight inhibitor concentrations with 1:2 serial dilutions and covering the range 0–10 or 0–200 μM were studied. The prediluted inhibitor solutions were prepared by diluting the compound stock solution (10 mM in DMSO) in the assay buffer (50 mM Tris-HCl, pH 7.4, 100 mM NaCl, ethylene glycol [20% v/v], 0.0016% Brij 58). The prediluted solution of protease (3 µM) was prepared by adding the corresponding volume of the assay buffer to the enzyme storage buffer (enzyme concentration 228 µM). The prediluted solution of the Mpro (10 µL), the assay buffer (60 µL), and the prediluted solution of the inhibitor (10 µL) were incubated for 15 min. The reaction was initiated by the addition of 20 µL of the prediluted solution of FRET substrate
Results and Discussion
Assay Conditions
In order to find the optimal assay conditions, measurements with substrate
The chelating agent EDTA reduced the enzyme activity slightly. Using polyols as additives, we observed the highest enzymatic activities at high concentrations (
Furthermore, we tested the influence of different pH values on substrate cleavage velocity and found no significant influence in the pH range from 7.0 to 8.0 (
For further investigations, we used the combination of 300 nM enzyme, 50 mM Tris-HCl, pH 7.4, 100 mM NaCl, 20% ethylene glycol, and 0.0016% Brij 58 because this ensures a high activity of the protease as well as reliability and easy handling.
Substrate Characterization
After establishing the assay conditions, we proceeded with the investigation of the substrates. Eight FRET substrates were designed and synthesized, with N-terminal 2-aminobenzoic acid as a fluorophore (
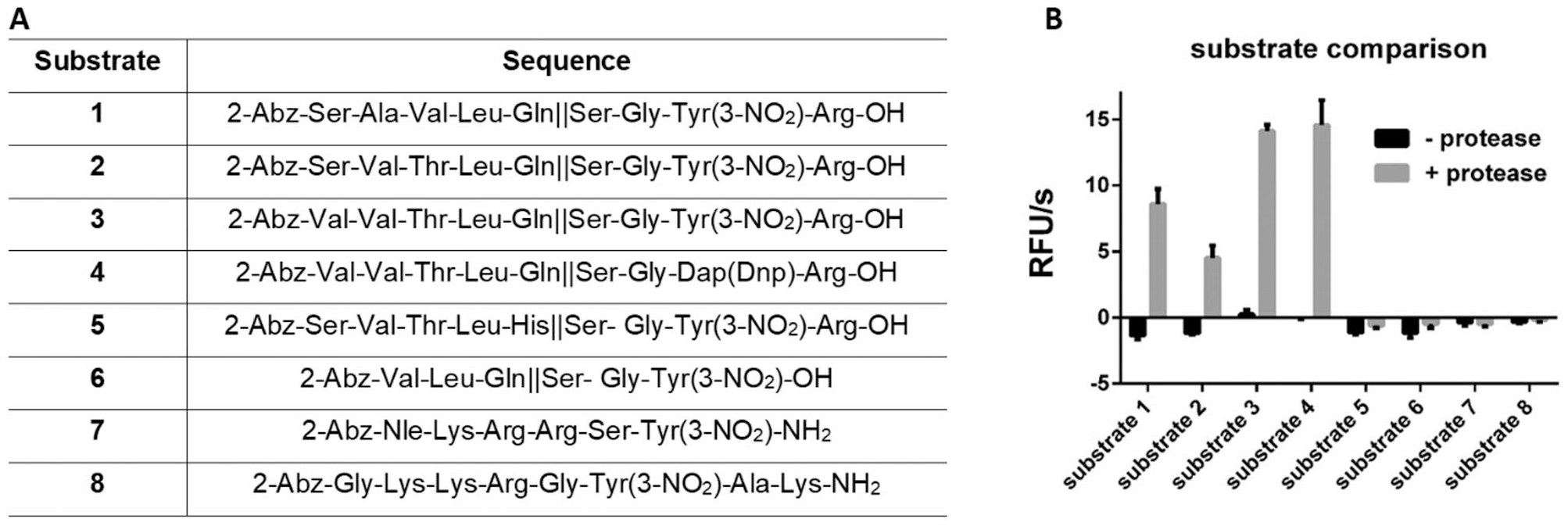

(
We first screened the activity of the protease at a substrate concentration of 50 µM. The highest cleavage was observed for substrates
By determining Km, Vmax, Kcat, and Kcat/Km values, we further characterized four substrates that displayed cleavage in the initial screening (
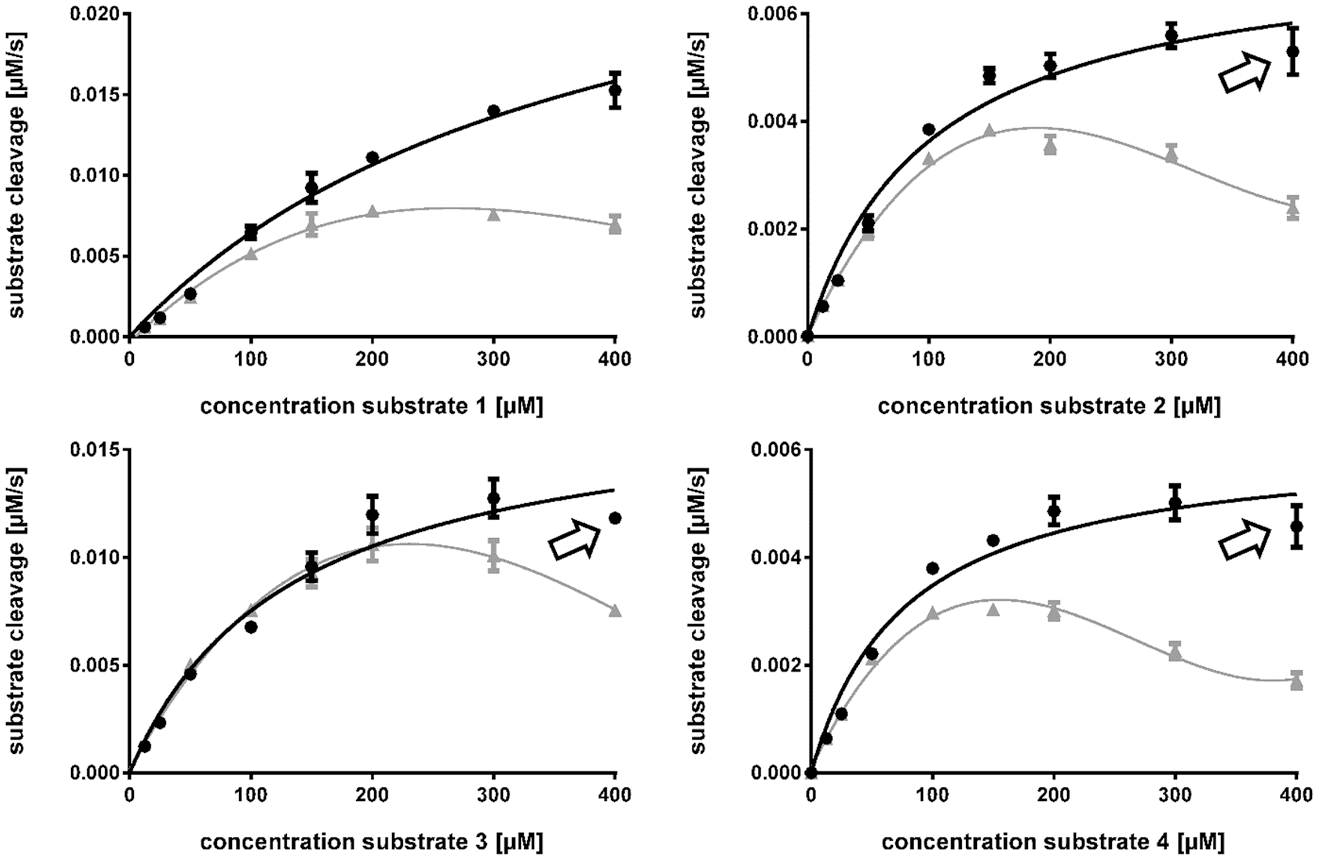
Michaelis–Menten curves for substrates
Km, Vmax, Kcat, and Kcat/Km Values for Different Substrates Determined by the FRET Assay, with Their Respective Errors (Kcat/Km) and Standard Deviations (Other Quantities).
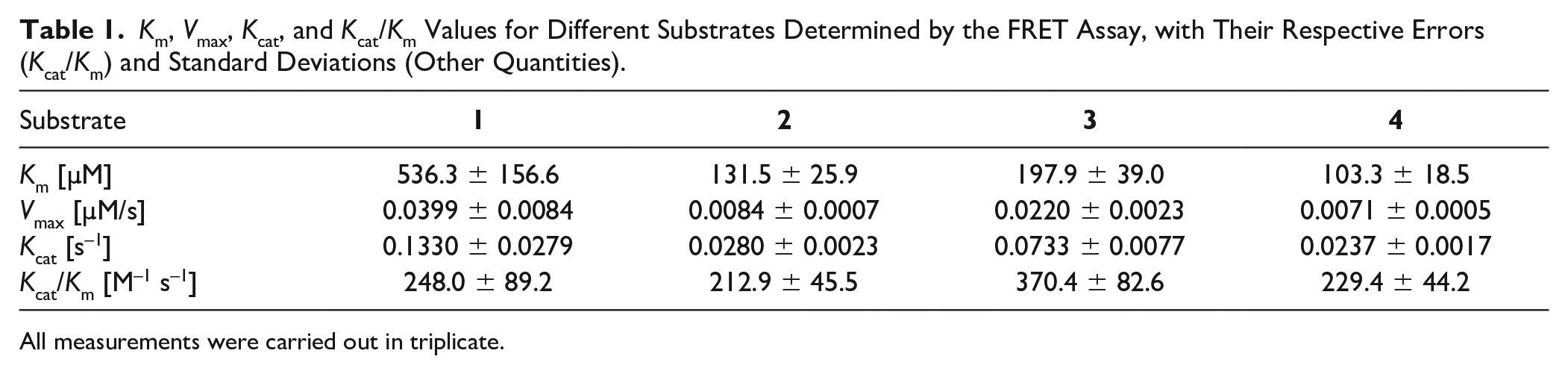

All measurements were carried out in triplicate.
Substrate
However, the full-sequence substrate showed comparable catalytic efficacy (214 M–1 s–1) to our substrate
Z′ Score
The Z′-score determination for the assay was performed with substrate
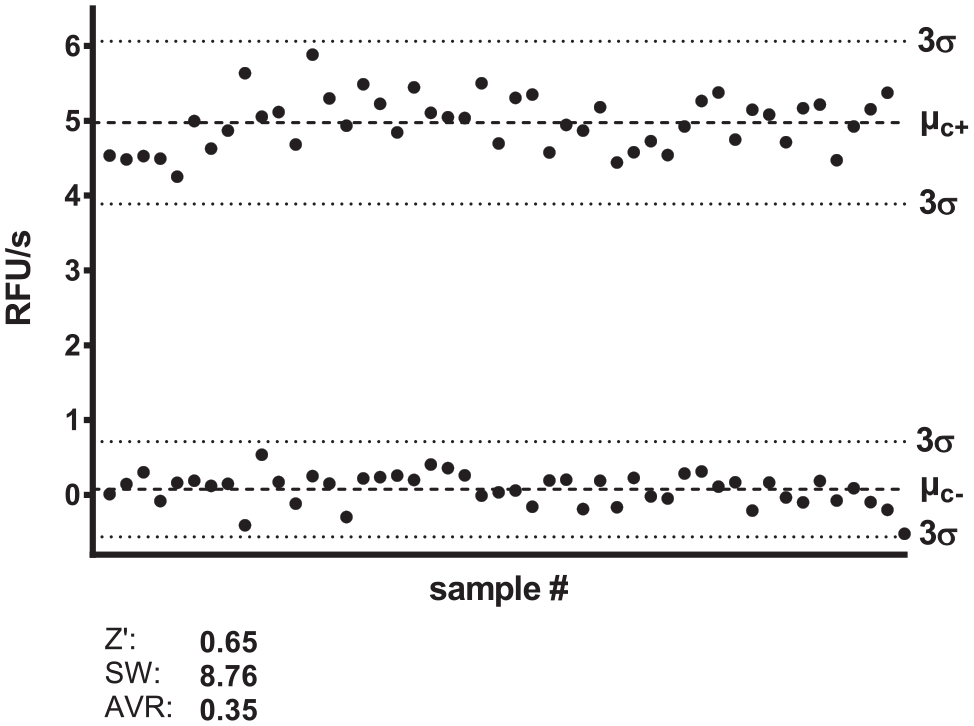
Z′ factor and signal window determination.
Reference Inhibitors
Substrate
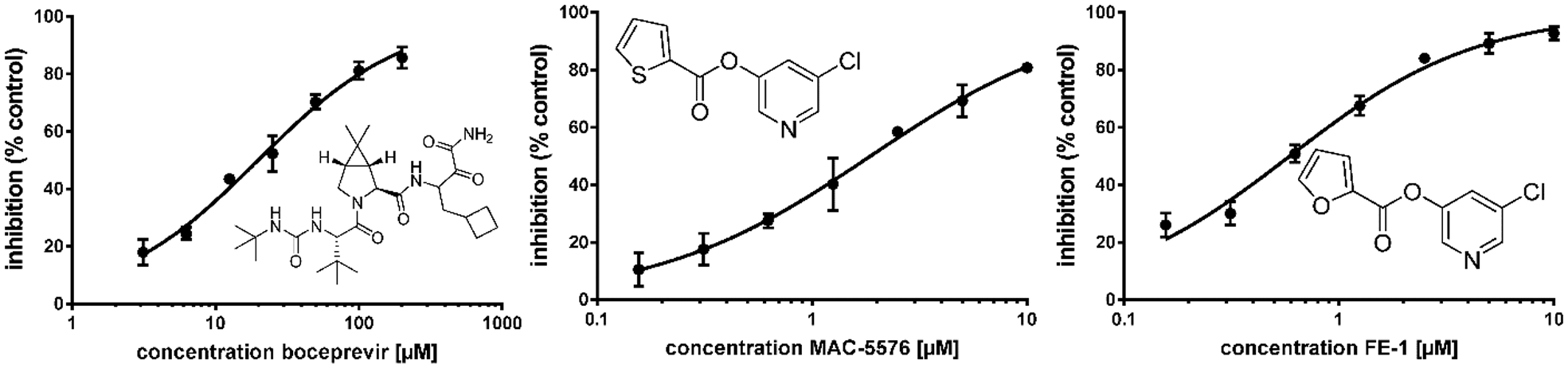
Dose–response curves and structures of reference compounds boceprevir (IC50 19.9 ± 1.1 µM), MAC-5576 (IC50 1.9 ± 0.05 µM), and FE-1 (IC50 0.59 ± 0.04 µM). R2 values for all curves are > 0.99. All measurements were performed in triplicate. Hill slopes: boceprevir, 0.85 ± 0.05; FE-1, 0.99 ± 0.07; MAC-5576, 0.87 ± 0.02.
Inhibitory Activity of Reference Compounds against Isolated SARS-CoV-2 Mpro and Published IC50 Values against SARS-CoV and SARS-CoV-2 Mpro.
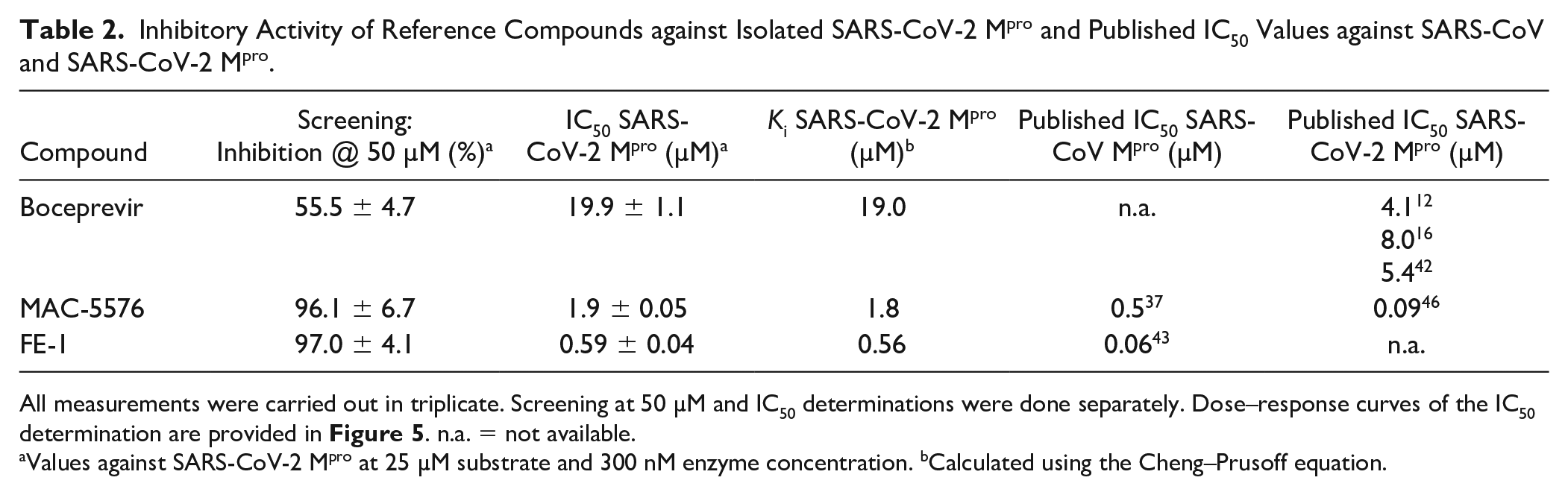

All measurements were carried out in triplicate. Screening at 50 µM and IC50 determinations were done separately. Dose–response curves of the IC50 determination are provided in Figure 5 . n.a. = not available.
Values against SARS-CoV-2 Mpro at 25 µM substrate and 300 nM enzyme concentration. bCalculated using the Cheng–Prusoff equation.
To summarize, a systematic evaluation of assay conditions for the SARS-CoV-2 protease was performed in this work. It was shown that the assay performance with Tris and phosphate buffer is improved in comparison with HEPES buffer. The addition of salts had no influence on the protease activity, whereas polyols, as well as most of the tested detergents, improved activity. Six substrates were designed and tested under the newly established assay conditions. Substrate
Conclusion
We have developed a reliable and reproducible biochemical assay for the SARS-CoV-2 main protease, which can be applied in high-throughput screenings and focused characterization of inhibitors. We hope that the newly discovered conditions and substrates will aid in the development of potent antiviral compounds against SARS-CoV-2.
Supplemental Material
sj-pdf-1-jbx-10.1177_24725552211020681 – Supplemental material for Efficiency Improvements and Discovery of New Substrates for a SARS-CoV-2 Main Protease FRET Assay
Supplemental material, sj-pdf-1-jbx-10.1177_24725552211020681 for Efficiency Improvements and Discovery of New Substrates for a SARS-CoV-2 Main Protease FRET Assay by Tonko Dražić, Nikos Kühl, Mila M. Leuthold, Mira A. M. Behnam and Christian D. Klein in SLAS Discovery
Footnotes
Acknowledgements
We thank Malte Hermes, Max Schmitt, Katharina Eckstein, and Aline Schöllkopf for their involvement in SARS-CoV-2 Mpro production. We also thank Tobias Haupt, Mathias Büchel, and Marie Hermann for preparing substrates
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material is available online with this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
