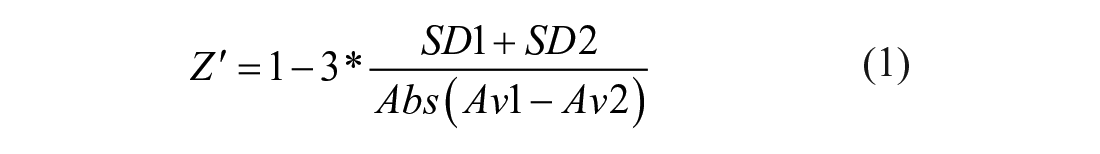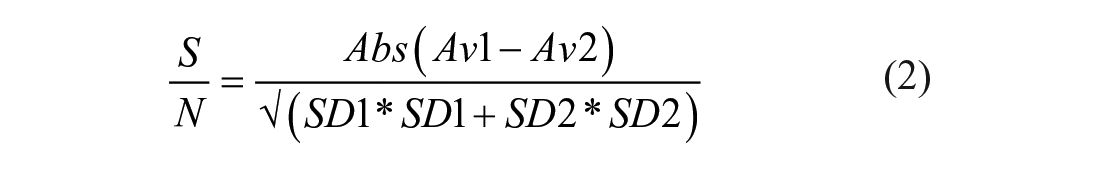Abstract
With improving survival rates for cancer patients, the side effects of radiation therapy, especially for pediatric or more sensitive adult patients, have raised interest in preventive or rescue treatment to overcome the detrimental effects of efficient radiation therapies. For the discovery of rescuing small molecules for radiation damage to the endothelium, we have been developing a 96-well microplate-based in vitro assay for high-throughput compatible measurement of radiation-induced cell damage and its rescue by phenotypic high-content imaging. In contrast to traditional radiation assays with detached cells for clonogenic formation, we observed cells with live-cell imaging in two different kinds of endothelial cells, up to three different cell densities, two gamma-infrared radiation dose rates, more than four different radiation doses, and acute (within 24 h with one to two h intervals) and chronic (up to 7 days) responses by phenotypic changes (digital phase contrast) and functional assays (nuclear, live-cell, and dead-cell staining) at the end of the assay. Multiple potential small molecules, which have been reported for rescuing radiation damage, have been tested as assay controls with dose responses. At the end, we did not move ahead with the pilot screening. The lessons learned from this “failed” assay development are shared.
Keywords
Introduction
Assay development, especially for high-throughput screening, is a tough business. 1 Usually the challenges, known as the “valley of death” of the drug discovery process,2,3 are attributed to the nonperfect in vitro assays for early drug discovery. The robustness, biology/physiology relevance, and automation compatibility (microplate based) of the assays are to be optimized to generate hits that ultimately can be used/developed for in vivo and clinical application. 4 These three factors also greatly limit the available in vitro models5,6 and the windows of the signals7,8 in assay development, despite there being a plethora of target-based and phenotypic assays that have been developed for early drug discovery and systems biology research.9,10 Here we present a “failed” assay development for the discovery of small molecules that rescue or mitigate radiation damage in noncancerous tissue. This example, in contrast, demonstrates the complications and difficulties that can arise during assay development.
Radiation therapy for cancer 11 has been part of standard treatment for 50% of cancer patients, in addition to surgery, chemotherapy, and, more recently, immunotherapy. 12 Because of the continuingly improving survival rate of cancer patients, the long-term consequences of this primary treatment have emerged as a significant risk factor, 13 especially on the cardiovascular-related side effects.14–16 Small-molecule drugs have been explored for rescuing radiation damage therapies for their transient, efficient, and biologically available characteristics. Recilisib sodium 17 (Ex-Rad) has been in a phase I trial as the only known oral radioprotectant. There are five major categories of reported irradiation rescue effects: (1) antioxidants or anti-inflammatory (e.g., auranofin, 18 GC1149, 19 or Mito-TEMPO 20 ); (2) 8-oxoguanine glycosylase (OGG) activators (e.g., V028-5832 21 or melatonin 22 ); (3) GPX4 activators through ferroptosis (Y600-0815 23 or (±)-α-tocopherol acetate 24 as a vitamin E analog); (4) hits/leads from whole-animal irradiation experiments (e.g., recilisib sodium, 17 also known as Ex-Rad, and γ-tocotrienol 25 ); and (5) others, for example, histone deacetylase (HDAC) inhibitors (trichostatin A 26 ). However, there are no systematic studies on the rescuing molecules, especially on specific tissues/organs, for example, cardiac endothelium or brain microvascular endothelium. A high-throughput plate-based in vitro assay would greatly facilitate the discovery of rescuing small molecules from radiation damage. The abovementioned small-molecule compounds will be utilized as positive controls for the assay development.
Currently, most of the assays for the discovery of rescuing small molecules from radiation damage are (1) clonogenic formation assays of radiated detached cells and (2) in vivo (mostly through mouse) experiments. Both are limited by their throughput to do systematic unbiased study on radiation effects. A high-throughput plate-based in vitro assay would attest not only hypothesis-based (e.g., OGG activators) but also unbiased phenotypic cell-based (e.g., synergistic effects of two or more targets) discovery of small molecules for rescuing radiation damage.
In this report, we describe how we developed a 96-well microplate-based in vitro assay for the high-throughput measurement of radiation-induced cell damage and its rescue by phenotypic high-content imaging. In contrast to traditional radiation assays with detached cells for clonogenic formation, we observed cells with live-cell imaging in two different kinds of endothelial cells, with up to three different cell densities, at two gamma-infrared radiation dose rates, and at more than four different radiation doses, for acute (within 24 h with one to two h intervals) and chronic (up to 7 days) responses by phenotypic changes (digital phase contrast [DPC]) and functional assays (nuclear, live-cell, and dead-cell staining) at the end of the assay. Eleven reported small molecules have been tested with dose responses as controls. In the end, we did not move ahead with the pilot screening because we could not identify any potential radiation rescuer controls. The lessons learned from this failed assay development are discussed.
Materials and Methods
Chemicals and Reagents
Dactinomycin and piplartine were purchased from Selleck Chemicals (Houston, TX); melatonin, 22 trichostatin A, 26 γ-tocotrienol, 25 auranofin, 18 genistein, 27 and Mito-TEMPO 20 from Cayman Chemicals (Ann Arbor, MI); GC1149 19 from Galera Therapeutics (Malvern, PA); recilisib sodium (also named Ex-Rad) 17 from MedKoo Biosciences (Morrisville, NC); and V028-5832 (also named compound C) 21 and Y600-0815(also named PKUMDL-LC-101) 23 from Enamine (Monmouth Junction, NJ). The stock solutions of all these chemicals at 10 mM DMSO were prepared, unless specified otherwise.
Cell-permeable Hoechst 33342 dye and Calcein-AM were from Thermo Fisher Scientific (Waltham, MA). Stock solutions (1 and 2 mM; 1000×) were prepared according to the vendor’s instructions, respectively.
Cells and Cell Culture
All cells were incubated at 37 °C in a humidified atmosphere of 5% CO2. An hCMEC/D3 BBB cell line from Millipore Sigma (Burlington, MA) was cultured in DMEM:F12 (Dulbecco’s Modified Eagle’s Medium nutrient mixture Ham’s F-121 in a 1:1 mixture) supplemented with 10% filtered fetal bovine serum (FBS; 10:100 media), 100 U/mL penicillin/100 µg/mL streptomycin (1:100 media), sodium pyruvate (1:100 media),
Radiation and Drug Treatment
Before radiation exposure, the cells were cultured in T75 flasks in less than 90% confluency before cell seeding with routine cell detachment/splitting and counting for each cell line. In most cases, unless otherwise specified, the cells were seeded onto 96-well microplates (PerkinElmer CellCarrier-96 Ultra microplates, tissue culture treated, black, 96-well with lid, Waltham, MA) at a cell density of either 2500 or 10,000 cells/well and incubated for 48 h to ensure that the cells formed a high-density, confluent monolayer, but still separated at a low cell density.
The control compounds were prepared in a 96-well drug plate (Greiner, Monroe, NC, microplate, 96-well, polypropylene, U-bottom, natural) for the serial dilution (1:3 for 8 wells or 1:2 for 12 wells) of each control compound. The drugs were added 24 h, 4 h, or immediately before the irradiation treatment (unless otherwise specified) with a change of culture media using the Hamilton MicroLab Star liquid handling system (Reno, NV).
Irradiation was performed on the abovementioned 96-well microplates with the culture media, with or without control compound treatment in serial dilutions. Cells were irradiated from 0 up to 440 Gy x-rays in the Radiation and Free Radical Research Core, delivered with either a PANTAK HF-320 ortho volt x-ray unit or an Xstrahl small-animal radiation research platform (SARRP) unit with different dose rates and doses (Gy) as specified. Cells in complete culture medium in microplates were exposed to irradiation at 22 °C, usually within half an hour. Two approaches of irradiation were applied on the microplate irradiation: one was covered with lead metal plates so that one single SBS standard 96-well plate can be divided into three or four blocks with different doses by controls of the irradiation time; the other microplate was set up using one dose per plate.
The PerkinElmer Operetta High Content Imaging System was used to monitor microplates using DPC imaging at 1 h intervals during the first 12 h, followed by either 2 or 8 h intervals over 24 h, unless otherwise specified. Usually, each plate was imaged before irradiation as well (denoted as –1 or –2; negative here indicates hour[s] before irradiation). After 24 h, plates were imaged every 24 h, after the daily media change, up to 7 days. At the end of the experiment, all the plates were stained with Hoechst for nuclear staining, calcein-AM for live-cell staining, and/or ethidium homodimer for dead-cell staining. Three additional channels of excitation (360–400 nm) and emission (500–550 nm) for Hoechst, excitation (460–490 nm) and emission (500–550 nm) for calcein-AM, and excitation (520–550 nm) and emission (580–650 nm) for ethidium homodimer were imaged.
Image analysis was done by instrument-accompanying Harmony software for a single DPC channel or four (DPC, Hoechst, calcein-AM, and ethidium homodimer) channels. Nuclei and cells were selected for both the intensity of all channels and morphology studies. The responses of radiation doses, control compound doses, and time-courses were analyzed by individual wells and/or individual cells within the wells using Spotfire (TIBCO and PerkinElmer) software for the visualization of the data sets. In cases where there was a dose response, additional analysis was performed with GraphPad Prism 9 (San Diego, CA).
Z′ factor and signal-to-noise (S/N) ratio are calculated according to the following formulas:
Results and Discussion
Assay Development Scheme: How and Why?
The objective of this assay development was to develop a microplate-based in vitro high-throughput assay that can be used for the discovery of and screening for small-molecule radiation protectors and radiation mitigators in endothelium upon radiation damage.
Current epithelial cell-based models for radiation damage detection use hCMEC/D3 and HUVECs, and in some cases, bovine aortic endothelial cells (BAOECs) are also used to evaluate radiation damage. In this report, hCMEC/D3 cells and HUVECs were used as cell-based in vitro models; the hCMEC/D3 cell line was used for its availability and ease of use, and primary HUVECs were used for their human relevance.
The phenotypic assays were intentionally picked for major assay development based on the earlier observation of the transient cell size changes when irradiated. The choice of the phenotypic assays also reflects the uncertainty of molecular mechanisms of action, especially the dose-dependent and time-related mechanisms and responses. There have been reports on target-based assays for radiation responses, for example, DNA damage based; mitotic catastrophe and mitotic death; apoptosis, necrosis, and senescence; necroptosis and ferroptosis; and bystander effect (e.g., immunogenic cell death). However, no dominant factor has yet been identified that correlates well with the radiation cellular effects. Thus, for this project, the phenotypic assay development was the major direction we chose.
Our hypothesis for working with a phenotypic assay (
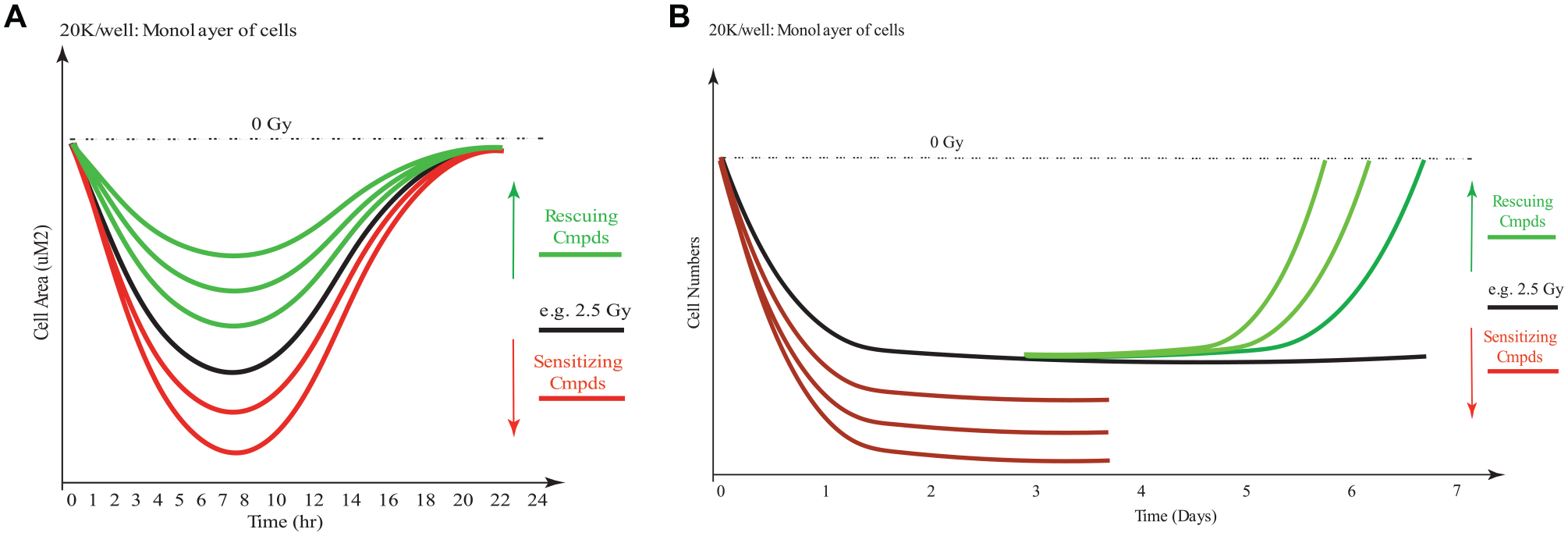
Hypothesis of acute and chronic responses of cells upon irradiation. (
To validate our hypothesis of the phenotypic assay, we designed the experiment as summarized in
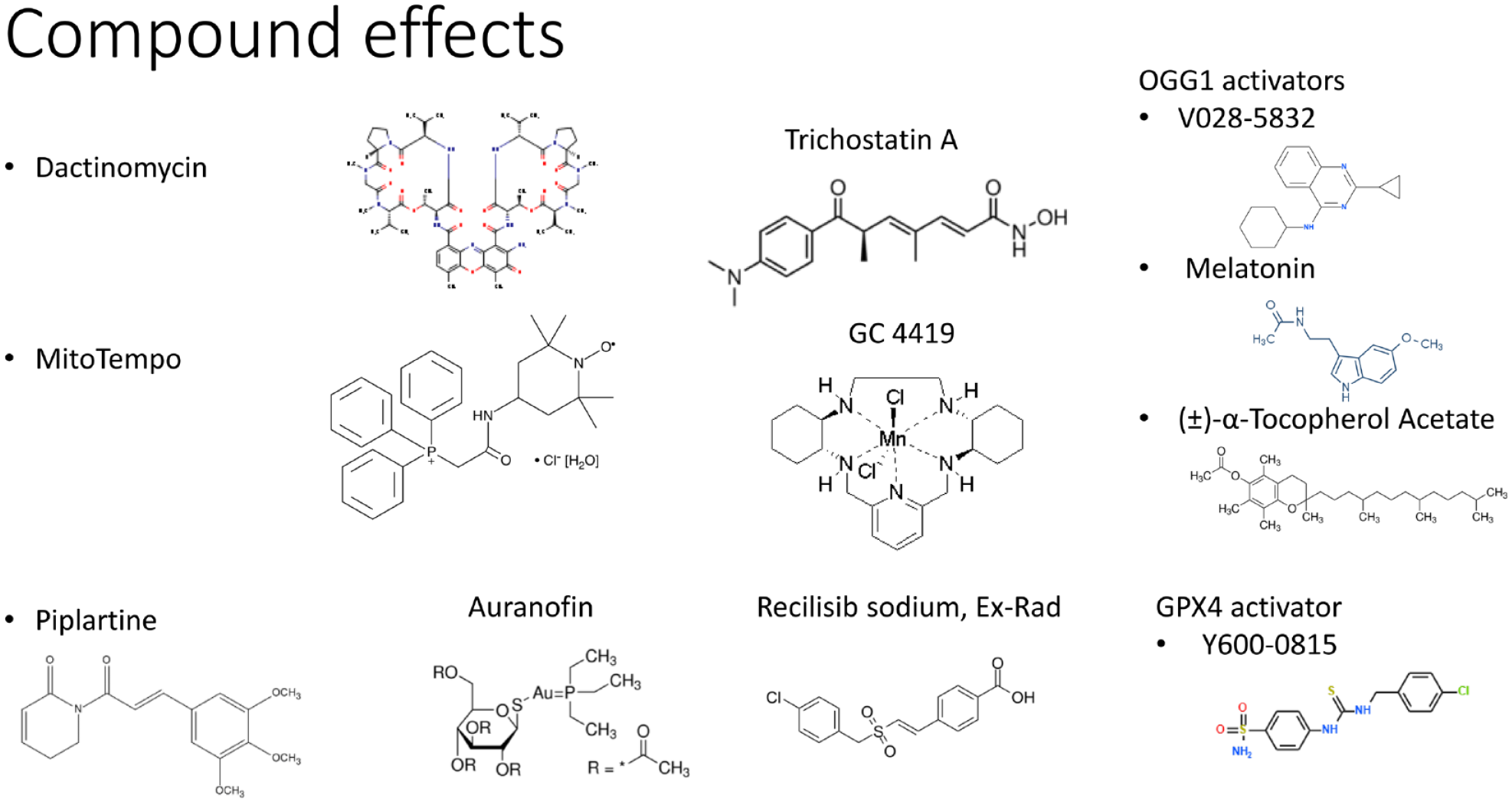
Representative control compounds tested in dose responses in the phenotypic assay development as positive or negative controls to evaluate the assay responses.
The control compounds (
The assay development workflow (
The results from this workflow, experimental design, and timeline are discussed separately in the following sections.
Radiation Doses and Cellular Responses
For the development of our in vitro plate-based assay, the maximum radiation dose up to 444 Gy at 22.2 Gy/min was applied to the hCMEC/D3 cells. Representative live-cell imaging of the time-course responses upon irradiation is shown in Figure 3 . The label-free DPC imaging of the plate provided the time-dependent phenotypical changes in cell numbers, cell area, cell roundness, and cell contrast of DPC intensity.
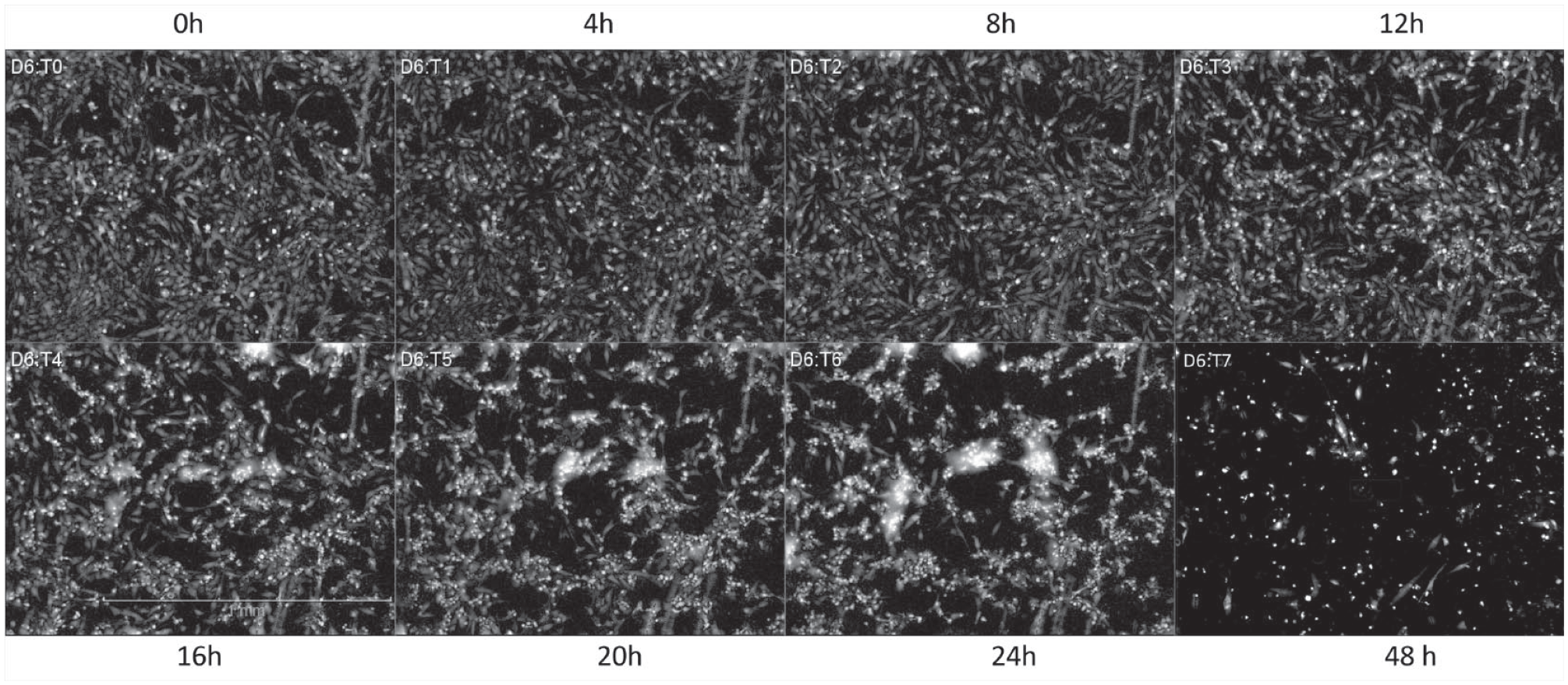

Images (DPC) for the time-course of hCMEC/D3 cells with a 444 Gy dose of irradiation at 22.2 Gy/min. Seeding in 20K/well, monitored for 2 days postirradiation.
The radiation dose effects on the cells were further studies at four different doses at the same dose rate of 22.2 Gy/min at a seeding density of 20K/well (monolayer formation). The postirradiation cell response over 72 h is summarized in
Figure 4
. The cell number responses postirradiation are shown in
Figure 4A
; as the irradiation increased from 0 Gy to 111 Gy, the cell number decreased accordingly up to 72 h. The cell area (µm2), cell roundness, and cell DPC intensity contrast upon irradiation at 0, 25, 55, and 111 Gy are shown in
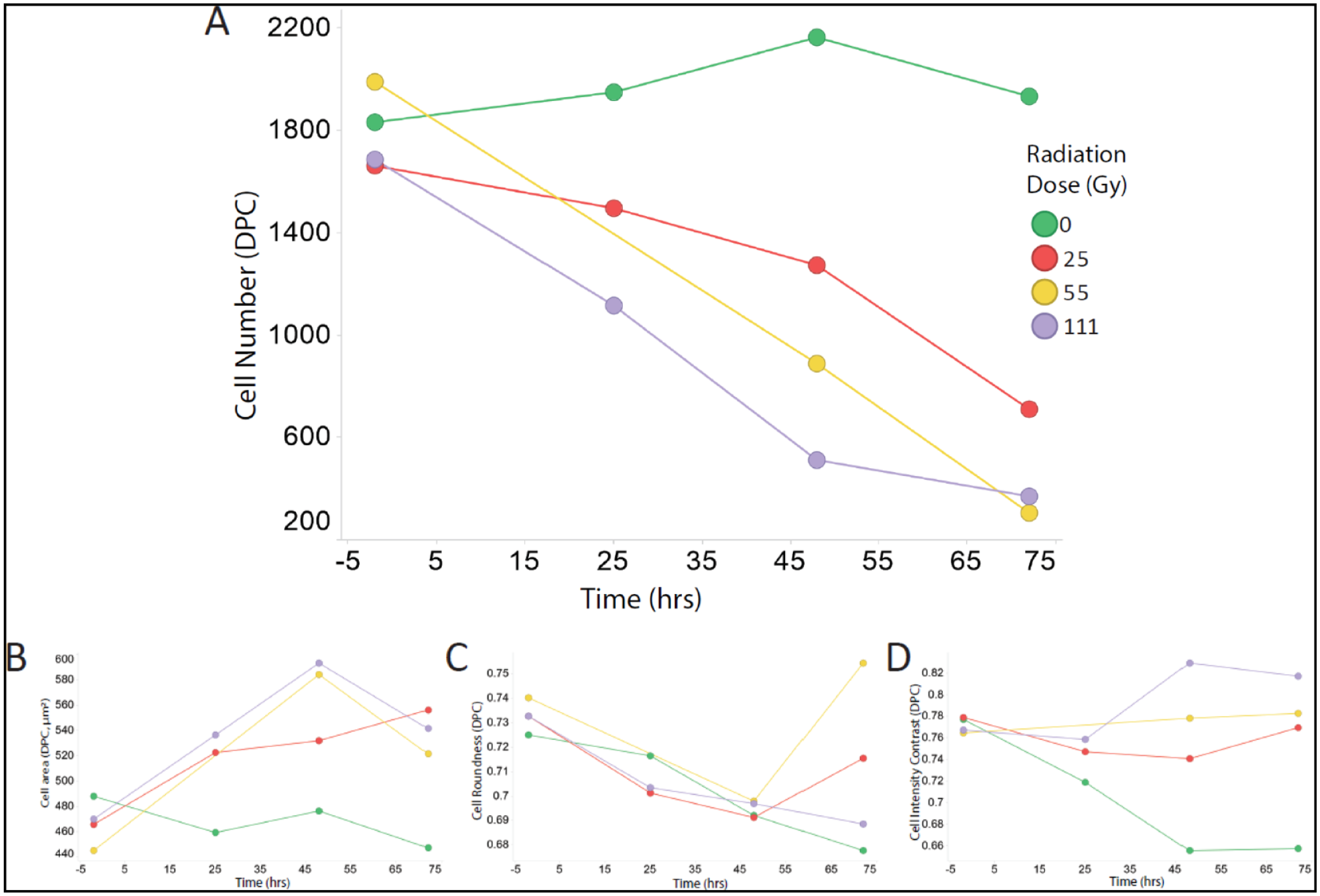
Radiation dose responses of the endothelial cells in a 72 h time-course. (
Radiation Doses and Control Drug Dose Responses
The effects of six control compounds (auranofin, dactinomycin, GC1149, Mito-TEMPO, recilisib sodium, and trichostatin A) in serial dilutions were tested at four different radiation doses at the same dose rate of 22.2 Gy/min (
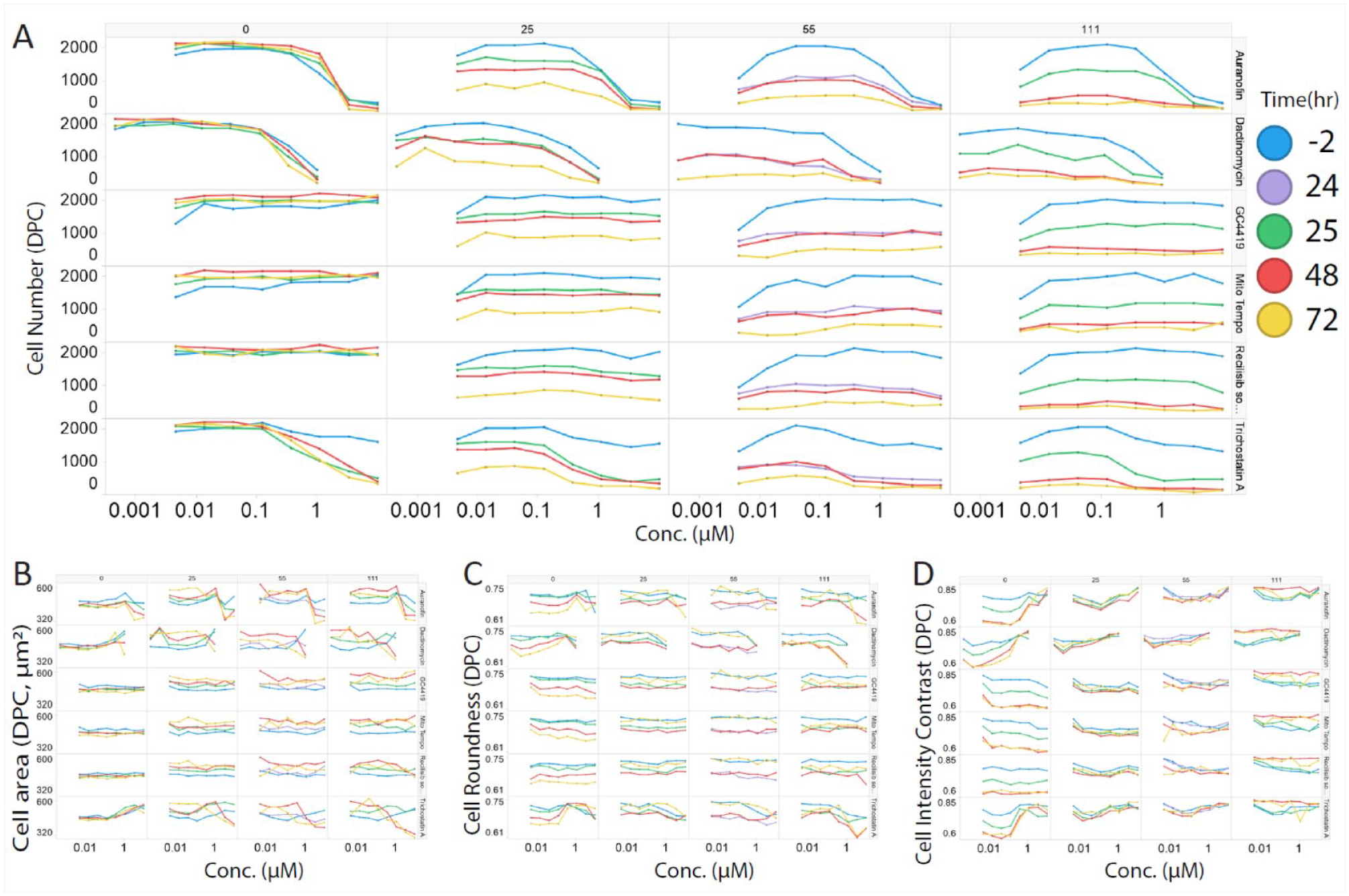
Compound effects (dose responses) in different radiation doses. (
The postirradiation effects of these six control compounds on other phenotypes, including cell area (µm2), cell roundness, and cell DPC intensity contrast, are shown in
The effect of the radiation doses on the cell numbers for control compounds (
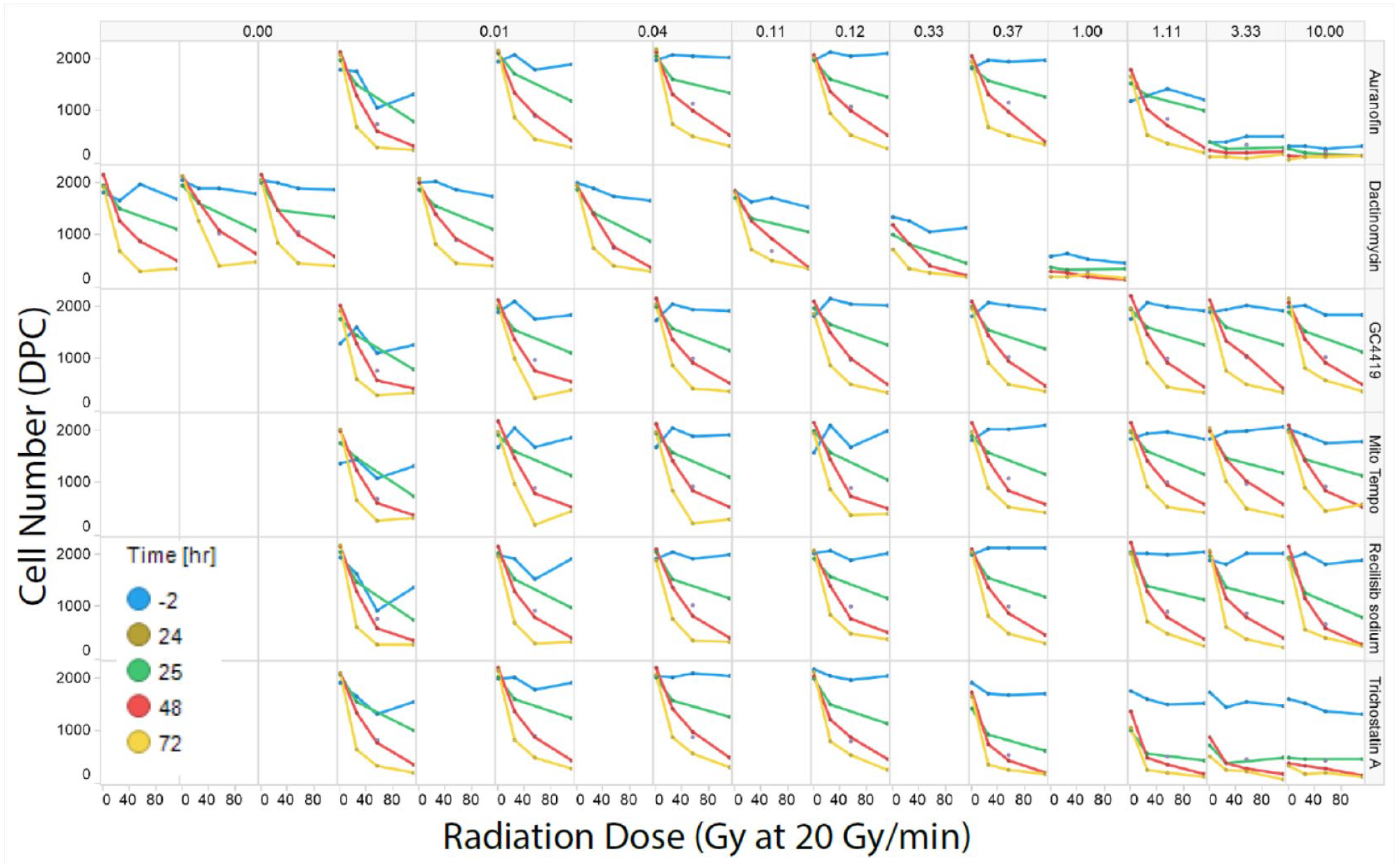
Radiation dose effects with different drug controls and time-course responses. Top: Control drug concentrations (µM). Right vertical: Compound names: auranofin, dactinomycin, GC1149, Mito-TEMPO, recilisib sodium, and trichostatin A. hCMED/D3 cells in 20K/well, monitored for 2 days after irradiation. See article online for color figures.
The timing of the application of the small-molecule rescuers/mitigators in all experiments is intentionally set at 24 h before irradiation. This timing could maximize the drug effect (protection or sensitization) in the cells in combination with serial doses of the drugs. This could minimize the effect of timing by covering all possible scenarios with diverse concentrations instead of a single concentration. There was no drug in the cell anymore after irradiation. All the effects are due to the downstream effects of the drugs before irradiation. This is also intentionally correlated with potential clinical application to eliminate radiation side effects before treatment. Application after irradiation was not explored at this time because the different drugs might take different lengths of time to be effective and might induce further compound effects for result interpretation.
Nonlinear Cell Responses and Irradiation Doses
It has been established that the postirradiation noxious effects to cells and tissue are nonlinear at low and high doses,
28
arbitrarily separated at around 0.1 Gy, based on epidemiology studies (
Furthermore, the bystander effect (
The postirradiation effects of the cell density and different cell monocultures (hCMEC/D3 and HEC-50) were tested (
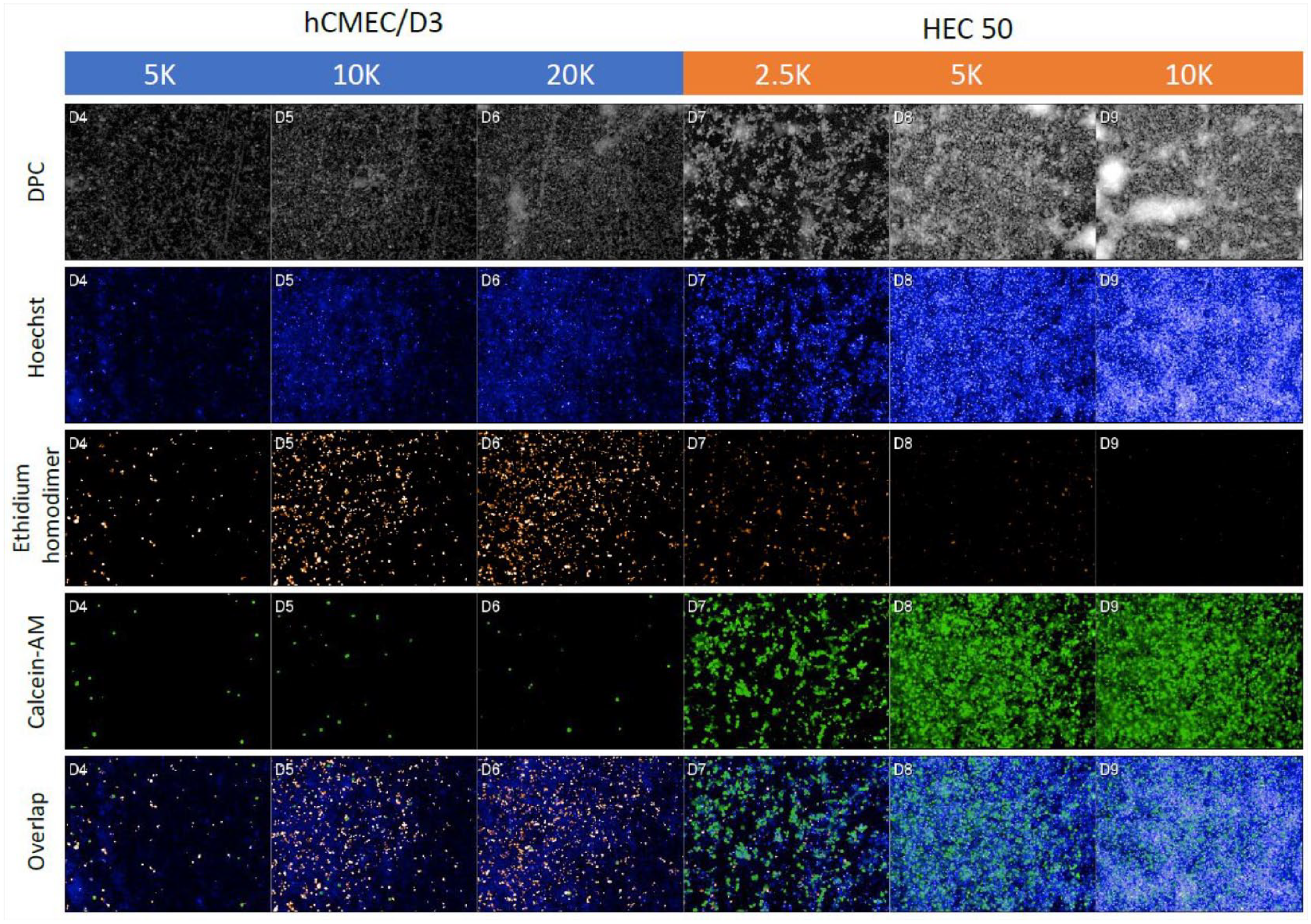
Cell density effects of hCMEC/D3 and HEC-50 cells upon irradiation by 444 Gy at 22 Gy/min at the 72 h time point. The cells at different cell densities were stained with Hoechst for nuclei, calcein-AM for live-cell staining, and ethidium homodimer for dead-cell staining. DPC was used for label-free cell staining.
Irradiation Rescuing/Mitigating Molecule Assays
Based on the abovementioned optimization of radiation doses, cell density, and time-course responses from acute (<24 h) and chronic (up to 7 days) phases, we further tested irradiation doses of less than 10 Gy with reported radiation rescuing control compounds and live-cell, time-course imaging.
As for the acute radiation effect (<24 h) at 2.5 Gy, there were no significant effects on the cell number (
As for the chronic radiation effect (up to 7 days) with the same plate map with 0, 2.5, 5, and 10 Gy/plate of irradiation, only dactinomycin demonstrated dose responses with increasing efficacy in cell number changes with increasing irradiation doses (
No dose responses were observed in both acute and chronic irradiation effects, with five reported rescuing control compounds in serial dilution and four different doses of “low-dose” irradiation.
Based on these results, we decided not to proceed with the pilot screening for the following considerations: (1) With the current phenotypic assay approach, we could not generate a dose response with the control compounds, even though some reported rescuing molecules have already been tested in vivo and in some clinical trial cases. 35 (2) Multiple mechanisms of action, especially in the low-dose irradiation condition, might not generate enough of a signal window to produce the necessary cellular phenotypes for irradiation rescuing/mitigating compounds. (3) For low-dose irradiation, the signal window is too small to be considered for physiological relevance. Despite our efforts, we were not able to determine the optimal radiation dose. Radiation doses that were too low resulted in a small signal, and those that were too high (or even at a medium-high level) raised concerns about the relevance by extrapolation and difficulty of the rescue.
We are reporting this failed assay to demonstrate that even with much effort and a strong rationale supporting the different conditions tested, the signal window can still be limited to the extent that it does not generate a robust assay for potential screening effects. These lessons learned fall into the category of the proverbial valley of death often encountered in assay development, which can also be similar to the challenges experienced during clinical trials: (1) phenotypic assays are not exactly physiologically relevance, and (2) target-based assays are not necessarily correlated to phenotypic observations. The different kinds of targets discussed above can only be considered as related, but not correlated, especially when the kinetic time-course measurement is used. To overcome these valleys of death in assay development, more thorough systematic approaches are needed to bridge the gap between assays and phenotypes, similar to the efforts in this failed assay. In addition, there is a need to target the responses required to facilitate screening efforts that allow for the discovery of in vivo and preferably clinical trial-relevant hits, which can eventually lead to therapeutics.
Despite the “failed” development of phenotypic assays for irradiation rescue of compounds in this report, we are still working on alternative assays: (1) target-based assays based on the low-dose mechanisms of action, for example, irradiation-induced direct DNA damage level and recovery, mitochondrial DNA damage and recovery, and reactive oxygen species and nitric oxide synthase as cellular signaling pathways for both direct-hit cells and bystander cells; (2) better in vitro cellular models for irradiation effects, such as the transwell systems for irradiation-induced leaking of epithelial monolayers, wound-healing functional models for epithelial cells, and 3D organ-on-a-chip models for epithelium (e.g., MIMETAS system); and (3) innate immune responses upon irradiation as well as co-culture immune responses for irradiation effects.
Supplemental Material
sj-pdf-1-jbx-10.1177_24725552211020678 – Supplemental material for A “Failed” Assay Development for the Discovery of Rescuing Small Molecules from the Radiation Damage
Supplemental material, sj-pdf-1-jbx-10.1177_24725552211020678 for A “Failed” Assay Development for the Discovery of Rescuing Small Molecules from the Radiation Damage by Kuo-Kuang Wen, Stephen Roy, Isabella M. Grumbach and Meng Wu in SLAS Discovery
Footnotes
Acknowledgements
We are deeply grateful to Dr. Douglas Spitz and Dr. Amanda Baker with the Free Radical and Radiation Biology program, for both the irradiation and their helpful comments and discussions. Dr. Kim Leslie and Dr. Xiangbing Meng are greatly appreciated for sharing HEC-50 cells.
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: We appreciatively acknowledge research grants R50CA243786, P30CA086862, and S10 RR029274, which provide funding to Dr. Meng Wu and the University of Iowa High Throughput Screening (UIHTS) Core, and R01HL108932, I01BX000163, and R01EY031544, which fund Dr. Isabella M. Grumbach.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.