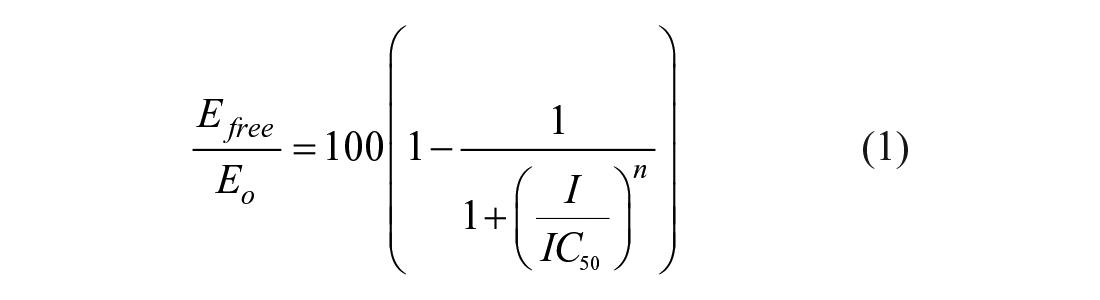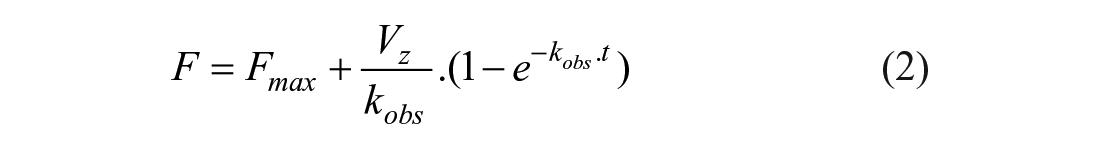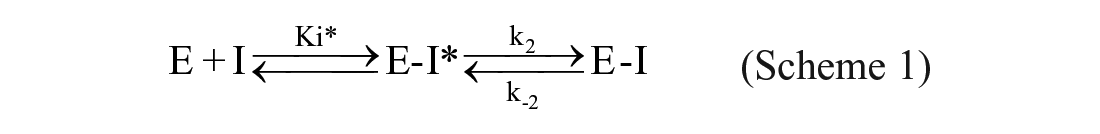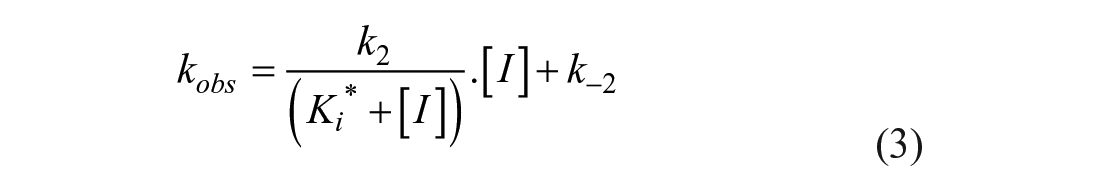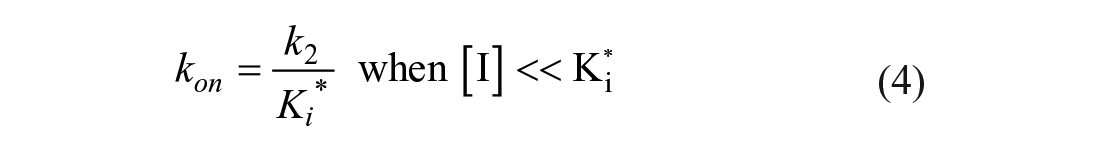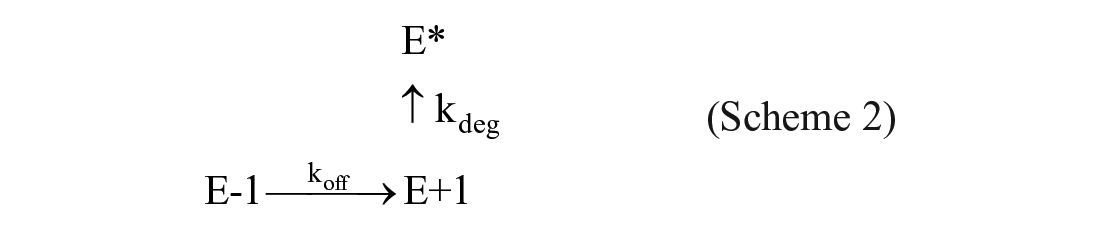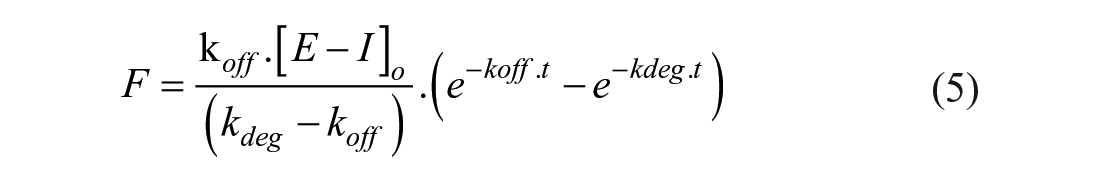Abstract
Two different signaling pathways lead to the activation of the transcription factor NF-κB, initiating distinct biological responses: The canonical NF-κB pathway activation has been implicated in host immunity and inflammatory responses, whereas the noncanonical pathway activation has been involved in lymphoid organ development and B-cell maturation, as well as in the development of chronic inflammatory diseases and some hematologic cancers. The NF-κB-inducing kinase (NIK) is a cytoplasmic Ser/Thr kinase and is a key regulator of the noncanonical pathway. NIK activation results in the processing of the p100 subunit to p52, leading to the formation of the RelB/p52 complex and noncanonical pathway activation. Because of its role in the development of lymphoid malignancies, this kinase has always been considered as an attractive target for the treatment of certain types of cancers and immune diseases. We at Takeda have pursued a drug discovery program to identify small-molecule inhibitors against NIK. This report provides an overview of the data generated from our screening campaign using a small fragment library. Most importantly, we also provide a kinetic analysis of published compounds and chemical series developed at Takeda that are associated with a slow tight-binding mechanism and excellent cellular potency.
Introduction
NF-κB proteins are a series of nuclear transcription factors that regulate a variety of physiological processes such as cell apoptosis, inflammation, and cell proliferation. This group includes RelA (also called P65), RelB, c-Rel, NF-ΚB1 p50, and NF-κB2 p52, which play key roles in the noncanonical and canonical NF-κB signaling pathways.
1
Each of these pathways is activated by different stimuli and involves subtle differences in their activation mechanisms that ultimately lead to distinct cellular responses. More specifically, the canonical pathway plays a key role in the innate and adaptive immune response. Its activation is rapid and transient, and will depend on the activation of transforming growth factor-β (TGFβ)-activated kinase-1 (TAK1) and the IkB kinase (IKK) complex. The IKK complex is an ensemble of three proteins, IKKα, IKKβ, and IKKγ, whose activation will lead to the phosphorylation of IkBα, a negative regulator of the NF-κB pathway. In its resting state, IkBα binds to the transcription factor RelA/p50, preventing this heterocomplex from migrating to the nucleus. Conversely, the activation of this pathway leads to the phosphorylation of IkBα, which dissociates from RelA/p50. Once free, IkBα undergoes proteolytic degradation by the proteasome. Its disappearance allows the RelA/p50 complex to translocate rapidly to the nucleus and to trigger the transcription of a variety of genes involved in the normal inflammation response. Alternatively, the activation of the noncanonical pathway plays a significant role in lymphoid organogenesis, B-cell survival and maturation, bone metabolism, and other cellular immune functions.2,3 Unlike the canonical pathways, the activation of the noncanonical signaling pathway is slow-paced, will last longer, and is activated by a variety of ligands such as TNF-like weak inducer of apoptosis (TWEAK), B-cell-activating factor (BAFF), CD40 ligand (CD40L), lymphotoxin beta (LTβ), and LIGHT, also known as tumor necrosis factor superfamily member 14 (TNFSF14). The binding of these ligands to their respective receptors at the surface of the cell activates the intracellular NF-κB-inducing kinase, also known as NIK or MAP3K14. When activated, NIK phosphorylates and activates IKKα kinase, which in turn will phosphorylate p100 protein. On phosphorylation, p100 is ubiquitinylated and undergoes limited proteolysis to form a smaller p52 protein. In return, p52 forms a complex with RelB and migrates to the nucleus to initiate transcription of specific genes distinct from the canonical pathway (
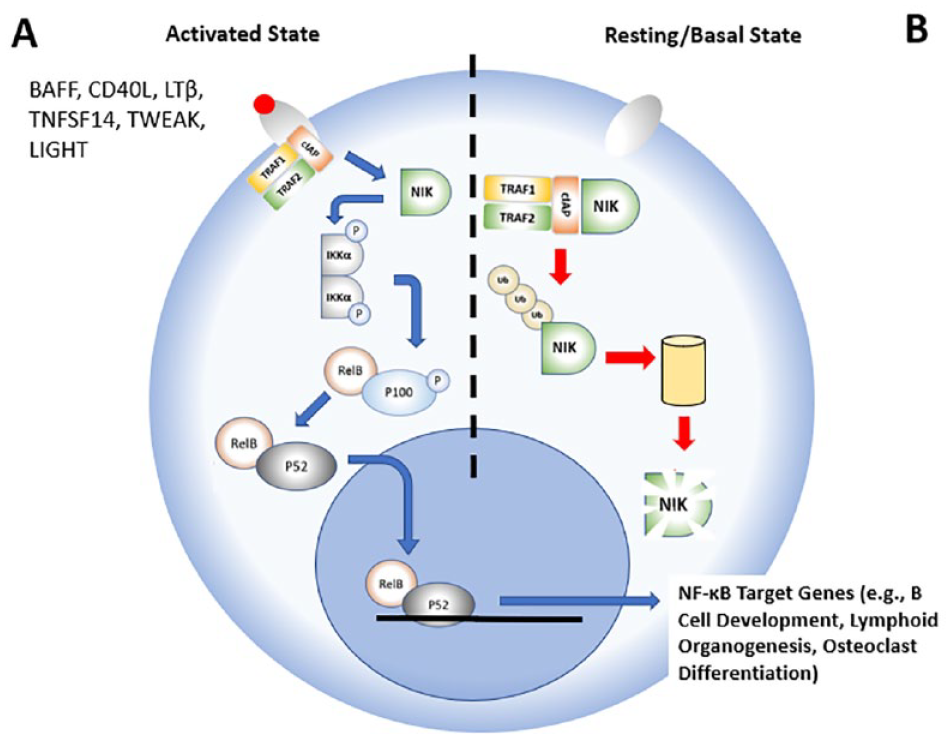
Overview of the noncanonical pathway in its activated and basal states, with emphasis on NF-κB-inducing kinase (NIK).
NIK is a key regulator of the noncanonical pathway whose role has been well described during lymphoid malignancy.6–8 As previously published, we also propose that NIK inhibition could be beneficial for the treatment of some cancers while preserving the ability of the canonical NF-κB pathway and the innate immune responses to fight against infection from external pathogens. In this study, we will highlight some of the results generated during our limited screening campaign, and exemplars of a few chemical lead series associated with the slow tight-binding mechanism and potent cellular data.
Materials and Methods
Materials
The NIK enzyme used in this study is a truncated recombinant protein (amino acid 319–947, 96 kDa) expressed in baculovirus with either an N-terminal glutathione-S-transferase (GST)-fusion or biotin tags. Both enzymes were purchased from Carna Biosciences (Kobe, Japan). The adenosine triphosphate (ATP)-binding site probe, tracer 236, and the europium (Eu)-labeled anti-GST antibody were purchased from Life Technologies/Thermo Fisher Scientific (Carlsbad, CA). The ADP-Glo kit was purchased from Promega (Madison, WI), and biotin full-length IKKα (dead mutant) was purchased from Carna Biosciences. Most of the studies described in the Methods section were conducted in 50 mM HEPES buffer containing 10 mM MgCl2, 1 mM ethylene glycol-bis(beta-aminoethyl ether) N, N, N’, N’-tetraacetic acid (EGTA), and 0.01% Brij-35 at pH 7.5 at room temperature (~22 °C), with the exception of the ADP-Glo assay that was conducted in 40 mM Tris, pH 7.5, 20 mM MgCl2, 0.1 mg/ml bovine serum albumin (BSA), and 50 µM dithiothreitol (DTT) at room temperature. The potencies of the NIK inhibitors [see the two-dimensional (2D) structures in
Methods
IC50 Determination
For the determination of IC50 values, increasing amounts of inhibitor were added (from 0 to 10 µM) to a fixed amount of enzyme (5 nM) with or without pre-incubation prior to the addition of the ATP-binding site probe (i.e., tracer 236, 100 nM) and the anti-GST antibody (2 nM). The decrease in Förster resonance energy transfer (FRET) signal [λex = 337 nm; λem = 620 and 665 nm; FRET signal = [(fluorescence signal recorded at 665 nm) / (fluorescence signal recorded at 620 nm) × 10,000] was recorded using a PHERAstar instrument from BMG LabTech (Cary, NC). The experimental data were fitted using Equation (1) for the determination of IC50:
where Efree and Eo are the free and total amounts, respectively, of NIK enzyme in the reaction mixture; n is the Hill coefficient; I is the free inhibitor concentration; and IC50 is the measure of the potency equivalent to the inhibitor concentration that leads to a 50% occupancy of the total enzyme. Each data were generated in duplicate; in addition, two IC50 values for each compound were determined, as illustrated in
To evaluate the time dependence mechanism, we used an ADP-Glo assay developed by Promega. Two sets of IC50 values were determined for each compound, with or without a 60 min pre-incubation period of mixing the enzyme (10 nM) and the inhibitor in the assay buffer prior to the addition of ATP (200 µM, ATP Km is 40 µM) and biotin–IKKα dead mutant (1 µM) as the protein substrate. The reaction was quenched after 120 min by adding the detection reagents from the ADP-Glo kit. The reagents were incubated for an extra 40 min at room temperature. The luminescence signal was then recorded using a PHERAstar instrument, and the experimental data were fitted using Equation (1) for the determination of IC50 values.
Thermal Shift Assay for the Determination of Compound Binding to NIK
The assay was conducted by adding 1 µM of recombinant NIK with 10 µM inhibitors to a 50 mM HEPES buffer containing 100 mM NaCl and SYPRO Orange (1×) at pH 7.4. Melting curves of the enzyme were recorded in the absence and presence of compound using the ViiA instrument from Life Technologies/Thermo Fisher Scientific. Briefly, the detection of a melting point (i.e., Tm) for a selected protein is achieved by gradually increasing the temperature of the medium, which causes the protein to switch from a well-folded state to a denatured form. The transition between these two states is measured by adding a fluorescent dye (SYPRO Orange) whose quantum yield increases when in contact with hydrophobic regions of the protein and whose enhanced fluorescence with time is recorded. A protein melting point is determined from the midpoint of the fluorescent curve. The binding of small molecules to the enzyme can be measured by their ability to affect the Tm of a protein. 10
Determination of On-Rate (kon) and Off-Rate (koff) Constants Using a Probe Displacement Assay
The same GST-tagged NIK kinase, Eu-labeled anti-GST antibody, and fluorescent probe as described above (see the IC50 Determination section) were used for the determination of kon. One potent inhibitor (compound 13) was selected for this study. Each of these inhibitors was available as a 10 mM stock solution and was used in two separate experiments for the determination of kon values. For the purpose of this study, compounds were diluted 100-fold in DMSO followed by a twofold serial dilution prior to being added to the assay buffer. In parallel, the enzyme, antibody, and active site probe were mixed together in the reaction buffer for a final concentration of 8, 100, and 8 nM, respectively. After a 5–10-min pre-incubation period, the inhibitor was added to this reaction mixture. The decrease in FRET signal was recorded with time using the same PHERAstar instrument as previously described. Initially, pseudo-first-order constants, kobs, were determined for each inhibitor concentration by fitting the experimental data using Equation (2):
where F is the FRET signal, Fmax is the FRET signal at t = 0, Vz is the initial rate for the displacement of the fluorescent probe from the enzyme, and kobs is the observed rate of association between the compound and enzyme.
The kobs values were replotted as a function of inhibitor concentration in a secondary replot, and the data were fitted using Equations (3) and (4) to determine kon, the overall on-rate; k2, the rate of isomerization of the intermediate complex E–I*; Ki*, the dissociation constant for the first step of the inhibition mechanism (see
A similar protocol was used for the determination of the off-rate constant, koff: An excess of Eu-labeled streptavidin (200 nM) was mixed with an equimolar amount of biotin-tagged NIK (200 nM) and inhibitor compounds (200 nM) in a 96-well plate format. After a 30 min pre-incubation, an aliquot of this mixture was diluted 100-fold into assay buffer containing an excess of active site tracer 236 (300 nM; 7.7-fold greater than its Kd; see
where F is the FRET signal, and kdeg is the constant describing the unfolding process of the free enzyme in the assay buffer. The detailed differential calculation process to generate Equation (5) is described by Capellos and Bielski. 11
Cell-Based Assay for the Detection of NIK-Dependent IKKα Phosphorylation
The assay was conducted using a stable double-transfected HEK293 Trex cell line (Invitrogen, Waltham, MA) that constitutively expresses the IKKα kinase domain and contains the full-length human NIK gene under control of the inducible doxycycline promoter. Prior to the addition of compound, cells are grown in a modified Eagle medium (MEM) with 10% fetal bovine serum (FBS) to reach 70–90% confluency. After a trypsinization step, cells are centrifuged and resuspended in MEM in the presence of doxycycline. Cells are added at a concentration of 5000 cells/well to a 384-well plate containing compounds in a final concentration of 0.5% DMSO. Negative control wells contain medium only. Plates are incubated overnight at 37 °C and 5% CO2. On the next day, the medium is replaced with a lysis buffer provided by the vendor. Plates are left at 4 °C for several hours in the dark. Detection of phosphorylated IKKα (p-Ser176/Ser180) is determined shortly after by adding donor and acceptor beads from the SureFire kit to each well according to the protocol provided by the vendor. Fluorescent signal is recorded using a PHERAstar instrument. Experimental data are normalized against DMSO-treated cells and after subtraction of background readings (i.e., wells containing medium only). Calculated values are fitted using a traditional IC50 curve; see Equation (1).
Results
Correlation between Biochemical, Biophysical, and Cellular Assay Data
A large subset of compounds was evaluated by measuring their potency (i.e., IC50) for binding NIK using the LanthaScreen FRET binding displacement assay and a thermal shift assay, which measures the ability of an inhibitor compound to stabilize a protein against thermal denaturation. Using this assay format, a large subset of compounds with a wide range of potency (i.e., IC50 values ranging from 10 to 0.01 µM) was evaluated in the presence of NIK. A plot of IC50 versus Tm indicates that the potency of these compounds is directly correlated to their ability to stabilize NIK under heat denaturation stress (
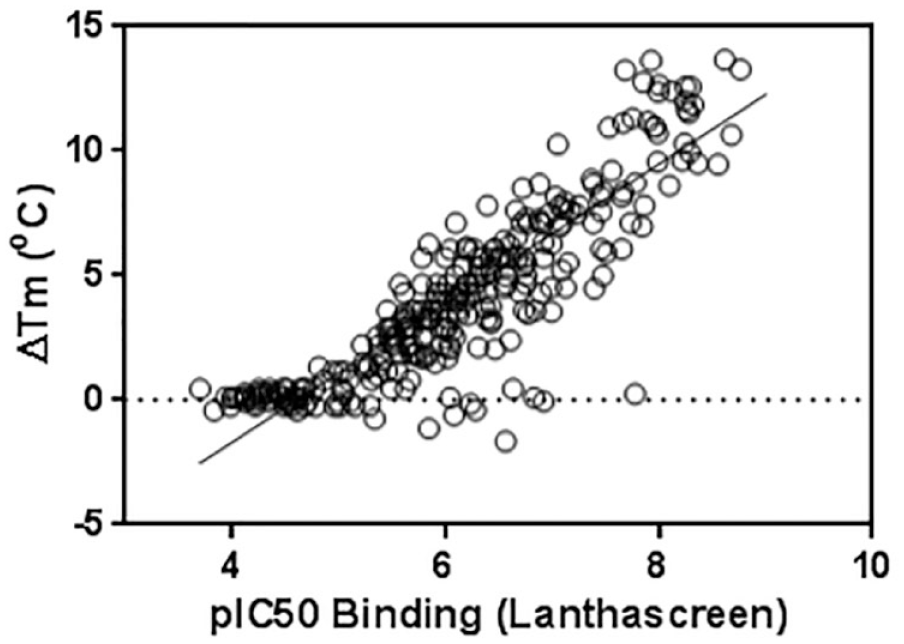
Correlation between data generated using thermal shift and Förster resonance energy transfer (FRET) binding assays (see Methods section). The data were fitted using a simple linear regression analysis for a final R2 of 0.81.
In a subsequent study, we also determined the cellular potency of these compounds in a stable-transfected HEK293 cellular model that monitors cellular NIK kinase activity. In the cytoplasm, NIK directly binds and phosphorylates IKKα (
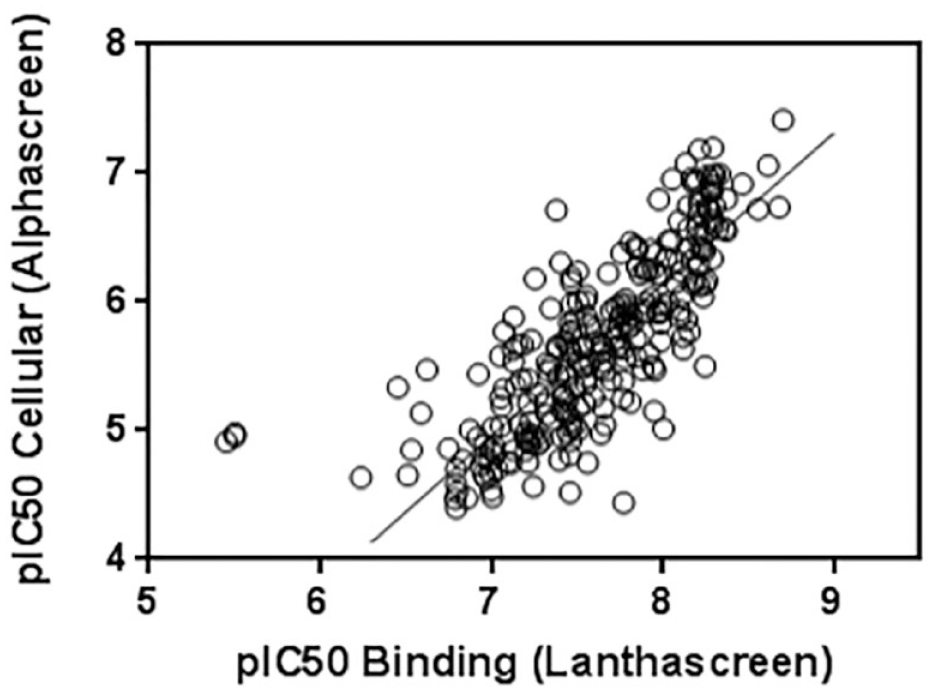
Correlation between data generated in the biochemical Förster resonance energy transfer (FRET) binding assay and cellular assay measuring NF-κB-inducing kinase (NIK)-dependent IkB kinase (IKKα) phosphorylation (see Methods section). The data were fitted using a simple linear regression analysis for a final R2 of 0.66. Despite a good correlation between these two sets of data, our compounds are, on average, ~100-fold less potent when evaluated using a cell assay. The reason behind this shift of potency remains unclear; nevertheless, we suggest that the presence of a high concentration of competing adenosine triphosphate (ATP) in the cell could explain to some extent these discrepancies. For the FRET biochemical assay, the Z’ factor for each plate varies between 0.7 and 0.8. For the cell assay, the Z’ factor varies between 0.5 and 0.7.
Kinase Selectivity Panel
The control compound 13 was submitted to the kinase panel to evaluate its inhibition profile against a panel of ~300 kinases (see
Time-Dependent Inhibition, Residence Time, Off-Rate and On-Rate Constants, and Binding Mechanism
Compounds used in this study were evaluated for their time-dependent inhibition mechanism against NIK. The study was conducted by measuring the potency of these compounds with and without pre-incubation with the enzyme. A few of these inhibitor compounds (I) showed an increase in potency after a pre-incubation period of 60 min with the enzyme (E), suggesting that formation of the E–I complex is slow to reach equilibrium (
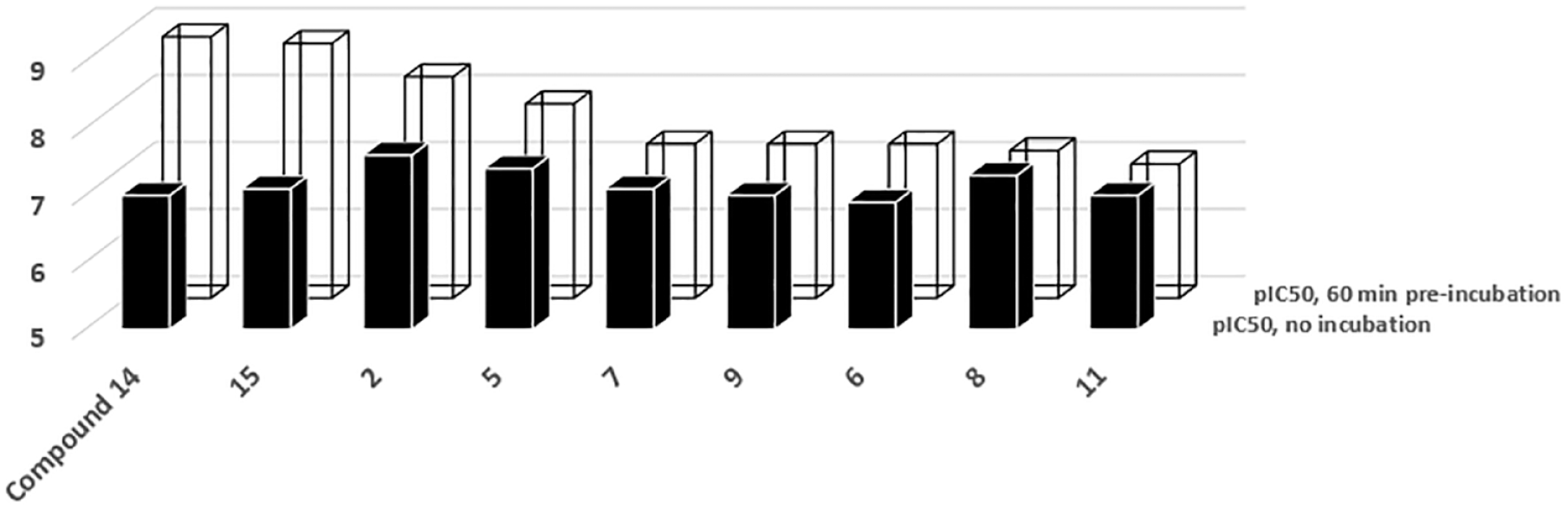
Time-dependent inhibition (TDI) of NF-κB-inducing kinase (NIK) inhibitors. Most of the compounds tested using the ADP-Glo activity assay gain potency (i.e., the IC50 value decreases) with time when pre-incubated in the presence of the enzyme. Each data were generated in duplicate and exhibit 20% or less in standard deviation.
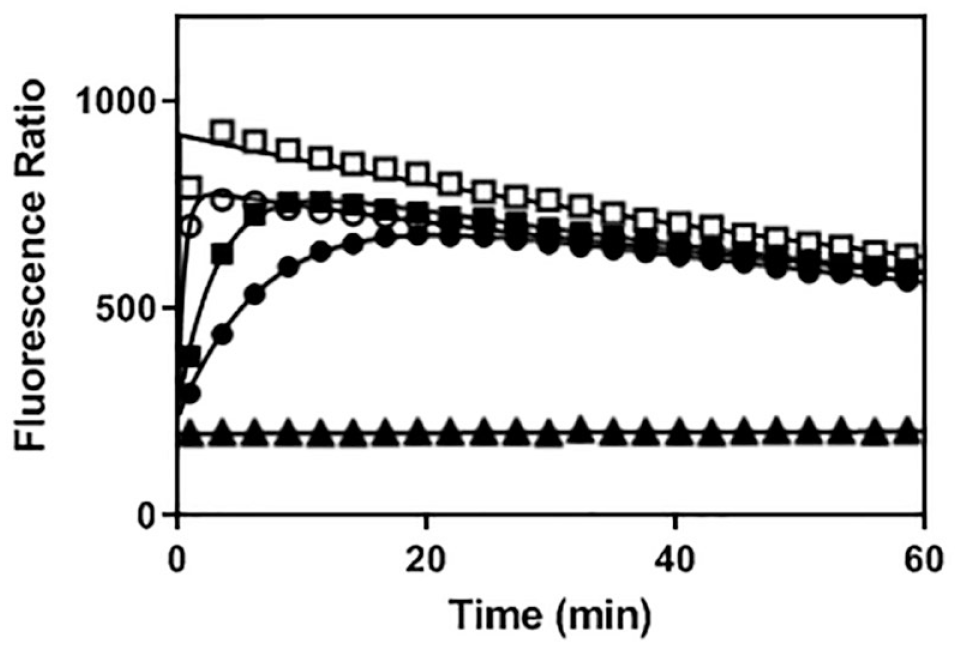
Off-rate determination of NF-κB-inducing kinase (NIK) inhibitors. As previously described in the Methods section, koff constants were determined by rapid dilution of the E–I (enzyme–inhibitor) complex in a large volume of buffer containing a high concentration of fluorescent-labeled adenosine triphosphate (ATP)-binding probe. As illustrated in this figure, compound 7 (•) is slowly released from the initial E–I complex as compared to compounds 9 (■) and 12 (○), respectively. Control experiments were conducted in the presence (□) and in the absence of enzyme (▲).
Off-Rate and Residence Time for Selected NF-κB-Inducing Kinase (NIK) Inhibitors.
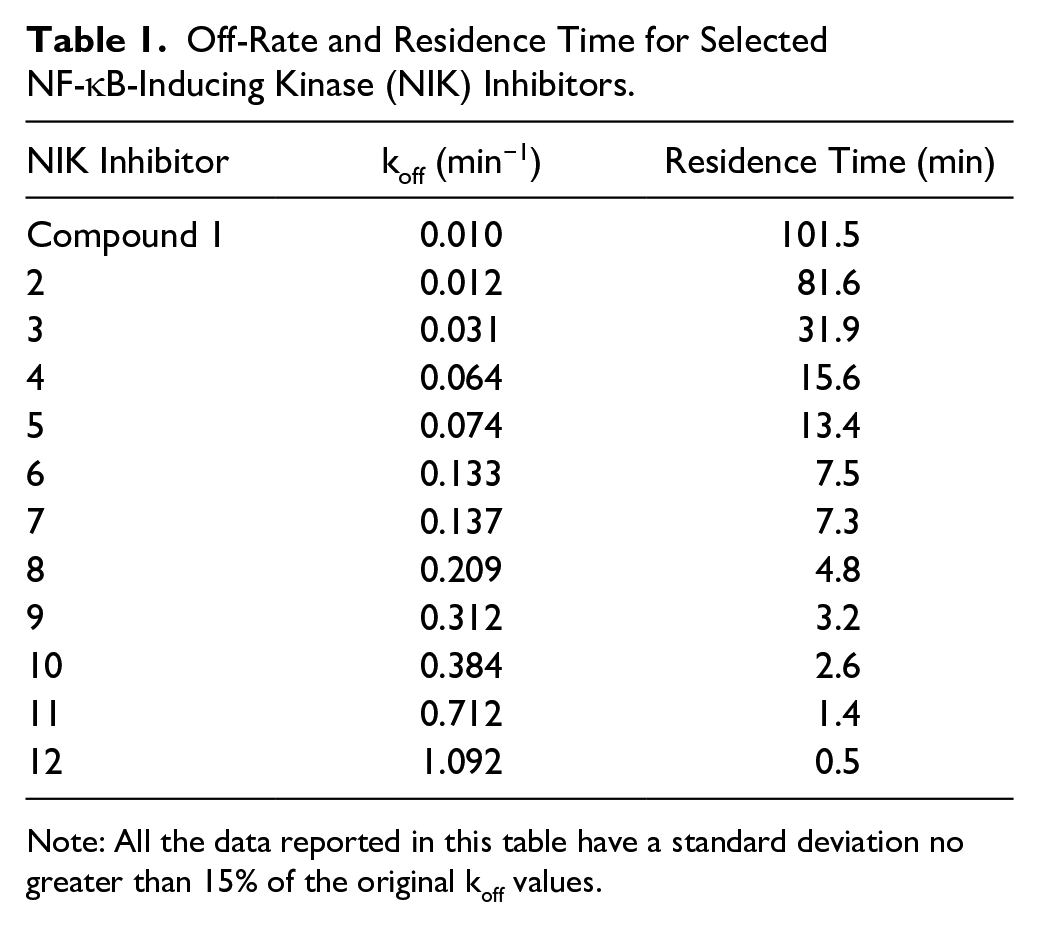
Note: All the data reported in this table have a standard deviation no greater than 15% of the original koff values.
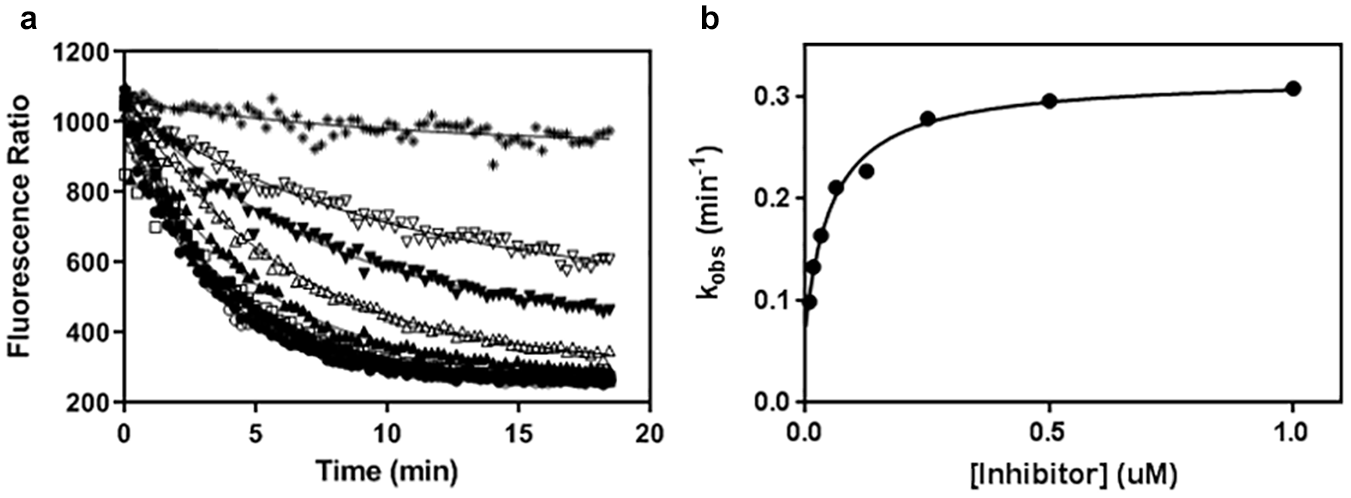
Mechanism of inhibition analysis for compound 13. The mechanism of inhibition and the kinetic constants of compound 13 were further analyzed. The rate of association of this compound to NF-κB-inducing kinase (NIK) is slow (see
Detailed Kinetic Profile of Compound 13.
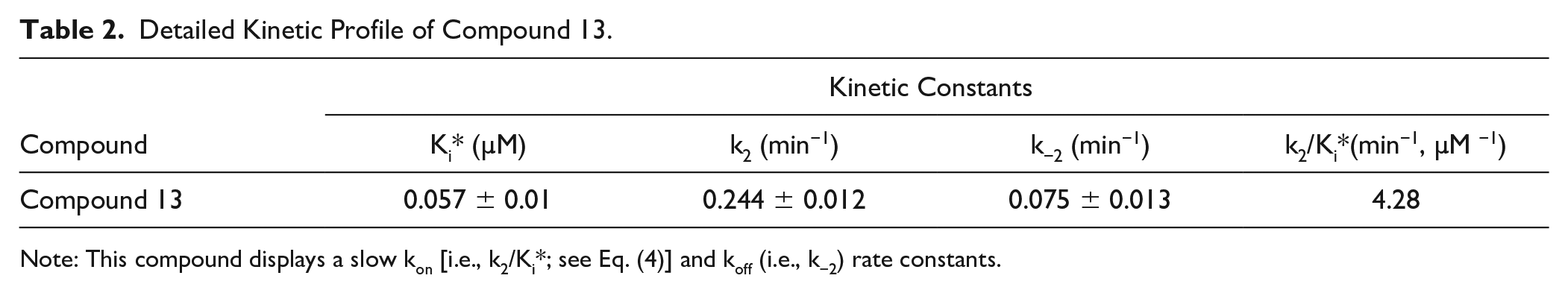

Note: This compound displays a slow kon [i.e., k2/Ki*; see Eq. (4)] and koff (i.e., k−2) rate constants.
Discussion
The noncanonical NF-kb pathway plays a vital role during the immune host response and the development of lymphoid organs, the maturation and survival of B plus dendritic cells, and the T-cell response to pathogens.
2
Conversely, the dysregulation of this pathway has been implicated in certain forms of cancers and severe autoimmune diseases such as lupus erythematosus, rheumatoid arthritis, inflammatory bone loss, and liver inflammation.14–18 A detailed inspection of this pathway suggests that NIK is the main gatekeeper responsible for its activation, which provides a unique therapeutic opportunity for the development of potent small-molecule inhibitors (see
Table 1
and the compound 2D structures in
In summary, we have described our in vitro pharmacology process to identify hit compounds against NIK. In addition to published inhibitors, we have also reported some of the compounds developed at Takeda and new methods to investigate the kinetic profile of these inhibitors. We hope that this study will provide useful information for the development of more efficacious inhibitors for this target against a variety of disease states from cancer to inflammation.
Supplemental Material
Kinase_selectivity_final_version – Supplemental material for Biochemical and Cellular Profile of NIK Inhibitors with Long Residence Times
Supplemental material, Kinase_selectivity_final_version for Biochemical and Cellular Profile of NIK Inhibitors with Long Residence Times by Petro Halkowycz, Charles E. Grimshaw, Walter Keung, Paul Tanis, Chris Proffitt, Kim Peacock, Ron de Jong, Mark Sabat, Urmi Banerjee and Jacques Ermolieff in SLAS Discovery
Supplemental Material
Supplemental_information_with_revisions_V2 – Supplemental material for Biochemical and Cellular Profile of NIK Inhibitors with Long Residence Times
Supplemental material, Supplemental_information_with_revisions_V2 for Biochemical and Cellular Profile of NIK Inhibitors with Long Residence Times by Petro Halkowycz, Charles E. Grimshaw, Walter Keung, Paul Tanis, Chris Proffitt, Kim Peacock, Ron de Jong, Mark Sabat, Urmi Banerjee and Jacques Ermolieff in SLAS Discovery
Footnotes
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.