Abstract
High-throughput matrix-assisted laser desorption/ionization mass spectrometry (HT-MALDI-MS) has garnered considerable attention within the drug discovery industry as an information-rich alternative to assays using light-based detection methods. To date, these efforts have been primarily focused on assays using protein or peptide substrates. Methods for RNA or DNA analysis by HT-MALDI-MS have not been extensively reported due to the challenges associated with MALDI-MS of oligonucleotides, including the propensity to form multiple salt adducts, low ionization potential, and ease of fragmentation. The objective of this work was to develop a platform suitable for HT-MS analysis of RNA and DNA substrates that overcomes these hurdles by combining on-surface sample preparation with soft ionization. This has been accomplished through the selective immobilization of fluorous-tagged oligonucleotides on a fluorous-modified MS target plate, followed by on-surface enrichment, matrix addition, and direct laser desorption/ionization, a process dubbed fluorous HT-MS (F-HT-MS). The work has resulted in methods by which RNA and DNA substrates can be detected at nanomolar concentrations from a typical assay buffer system using procedures that are amenable to full automation. The protocols were applied to an miRNA biogenesis assay, demonstrating its potential for RNA processes and thereby filling a prominent gap in RNA drug discovery: the paucity of in vitro functional assays.
Introduction
Mass spectrometry (MS) has received a high level of interest as an alternative to light-based detection in high-throughput screening (HTS) since MS provides structural information, can detect multiple analytes simultaneously, lowers false signal rates, and eliminates the use of antibodies.1–6, By combining the information density of MS-based analysis with the throughput, a high level of data, in terms of both quantity and quality, can be achieved that is not available by other detection methods.
The primary ionization methods for MS detection are electrospray ionization (ESI) and laser desorption/ionization (LDI), 7 with the latter in the form of either matrix-assisted LDI (MALDI) or surface-assisted LDI (SALDI). 8 Regardless of the ionization method used, the assay buffer composition and sample preparation are important factors, as salts, buffers, and detergents can lead to significant signal suppression, lower ionization efficiency, and degradation of signal quality.9,10 This has led to a number of different approaches toward optimizing MS signal by analyte enrichment, including in-line solid-phase extraction or chromatography for ESI methods11–13 or off-surface ion exchange or solid-phase analyte capture for LDI methods.14,15 The majority of these sample preparation methods, however, limit throughput, thereby precluding these methods from being used in primary high-throughput screens.
One solution to this issue is to incorporate on-surface analyte enrichment and cleanup directly on LDI surfaces.8,16 Methods developed include surface modifications or affinity capture by various chemistries, followed by on-surface sample cleanup, and they have been applied to a number of MS surfaces, including DIOS, 17 SELDI, 18 SAMDI, 19 and NIMS. 20 On-surface capture chemistries that have been utilized include hydrophobic interactions, antibody–antigen, cysteine–maleimide, His6-Ni2+, biotin–streptavidin, and click chemistry. 21 For example, SAMDI-MS uses a thiol-modified maleimide immobilized on a gold surface to capture cysteinyl-tagged analytes that are then ionized from the gold surface with matrix assistance, 19 although for oligonucleotides it was found that a biotin–streptavidin-modified gold surface was required for immobilization and ionization biotin-tagged RNA and DNA.22,23 These on-surface sample preparations simplify workflows, allow for automation, and are amenable to high-throughput primary screens to varying degrees and have been used in a number of assays. 16
For assays using oligonucleotide substrates, sample preparation and desalting take on increased importance since RNA and DNA analysis by MS is less tolerant of salts, buffers, and detergents than peptide analysis. 24 MS analysis of oligonucleotides is inherently more difficult due to their lower ionization potential, the formation of multiple cationic adducts, and a higher propensity to fragment. 25 Given both the relative paucity of HT-MS oligonucleotide assays 16 and the current interest in RNA processes as therapeutic targets, 26 the development of a general platform for functional HTS would be a valuable addition to the discovery toolbox. 27
One interesting capture chemistry that has been used in various applications across multiple molecular classes is fluorous partitioning where perfluoroalkyl-tagged molecules selectively partition into a perfluoroalkyl liquid or solid phase in preference to an aqueous or organic phase.28,29 Fluorous-based immobilization for array formation has been utilized with small molecules,30,31 peptides, 32 and recently DNA. 33 In 2005 Brittain et al. provided the first example of using fluorous partitioning for analyte enrichment of peptides prior to ESI-MS analysis and found the tags to be nonfragmentary and non-ion suppressive. 34 Since then, three fluorous surfaces for selective capture of fluorous-tagged peptides and carbohydrates, followed by desorption/ionization from the surface, have been reported: fluorous-modified DIOS, 35 NIMS, 20 and fluorous-modified aluminum oxide-coated slides. 36 These surfaces, however, have proven to be difficult to produce consistently and none of these surfaces are currently commercially available. There have also been no reports to date of fluorous-tagged RNA (f-RNA) or fluorous-tagged DNA (f-DNA) being enriched and detected by MS from these surfaces. Herein we report the fluorous partitioning-based enrichment and MS detection of f-RNA and f-DNA substrates from a fluorous-modified stainless steel plate amenable to high-throughput screening, a platform dubbed fluorous high-throughput MS (F-HT-MS). The platform was then applied to an miRNA biogenesis assay to detect Dicer processing of fluorous-tagged precursor microRNAs (pre-miRNAs).
Materials and Methods
Materials
All RNA and DNA compounds were synthesized and prepared by Dharmacon Inc. (Lafayette, CO) containing either a 5′-terminal fluorous tag or an internal amino allyl-modified uridine. Fluorous phosphoramidite
Methods
Synthesized f-RNA and f-DNA from amine-modified RNA and DNA (pre-miR-21 and pre-miR-155 RNA) were purchased from Dharmacon with an internal 5-aminoallyl-uridine as a lyophilized solid. The full-length pre-miRNA hairpins were annealed by heating to 95 °C and cooling to 12 °C over 10 min in a thermal cycler (Bio-Rad, Hercules, CA). For coupling reactions, 10 μL of RNA (1 mM stock) was diluted in 35 μL of sodium bicarbonate buffer (100 mM, pH 8.0). DMSO (45 μL) was added followed by N-succinimidyl 3-(perfluorooctyl)propionate (10 mM) that was added in five 10 μL aliquots every 30−60 min. After the last addition, the reaction was stirred overnight at room temperature. Ethanol precipitation was performed at −80 °C for 1 h and then centrifuged at 16,000g and 4 °C for 30 min. The supernatant was removed, and the pellet was washed with 500 μL of ethanol (70% v/v in water). The solution was centrifuged a second time at 16,000g and 4 °C for 30 min. The fluorous-modified oligonucleotide product was isolated by high-performance liquid chromatography using a Shimadzu LC-20 A equipped with an ACE Equivalence C18 column. The mobile phases consisted of 100 mM triethylammonium acetate and acetonitrile. A gradient of 5%−95% acetonitrile for 30 min was used with a flow rate of 0.75 mL/min, and the absorbance at 260 nm was monitored. The purified oligonucleotide was desalted on a 10 kDa molecular weight cutoff (MWCO) column (MilliporeSigma cat. UFC500324) following the manufacturer’s protocols. The purity of the modified oligonucleotides was assessed by gel electrophoresis, and the correct mass was verified using MALDI-MS.
Spotting Solutions
All RNA and DNA spotting solutions were freshly made from ~50 µM stock solutions of fluorous-tagged and untagged RNA and DNA by the addition of the appropriate amounts of nuclease-free water or miRNA assay buffer solution (Tris [10 mM, pH 7.5], MgCl2 [5 mM], ATP [1 mM], and DTT [1 mM]). The matrix solution was a solution of recrystallized 5-methoxy-salicyclic acid (26.7 mg/mL) and ammonium citrate dibasic (16.9 mg/mL) in a 4:3:1 isopropanol−water−2,2,2-trifluoroethanol mixture.
Dicer Expression and Purification
HEK293T cells were plated (2.5 × 106 cells/plate) in a 10 cm plate and incubated overnight. Cells were transfected with pFRT/TO/FLAG/HA-DEST Dicer plasmid (8 µg; Addgene, Wastertown, MA, cat. 19881) using Lipofectamine 2000 (Thermo Fisher, Pittsburgh, PA) for 5 h. Cells were grown to confluence (72 h) and then lysed (Tris-HCl [50 mM, pH 7.5], NaCl [150 mM], Triton X-100 [1% v/v], and sodium dodecyl sulfate [SDS; 0.1% w/v]). Lysate was centrifuged and the supernatant was incubated with 25 μL of anti-FLAG antibody (Proteintech, Rosemont, IL, cat. 20543-1-AP; 6 μg) overnight at 4 °C. After incubation, protein A/G resin (Pierce, Waltham, MA, cat. 20421; 100 μL) was washed with immunoprecipitation buffer (IP; Tris-HCl [25 mM, pH 7.2] and NaCl [150 mM]) three times and added to the lysate and incubated for 3 h. The Dicer-bound resin was then washed three times with IP buffer and stored in 50% glycerol at −20 °C until use.
Pre-miRNA Cleavage by Dicer
Preannealed and desalted pre-miRNA was diluted to the appropriate concentration in Dicer assay buffer (Tris [10 mM, pH 7.5], MgCl2 [5 mM], ATP [1 mM], and DTT [1 mM]). Immobilized Dicer was washed twice with assay buffer and then resuspended in the appropriate amount of assay buffer to prepare a working stock of immobilized Dicer consisting of a 50% slurry (e.g., 50 μL of bedded resin and 50 μL of assay buffer). Immobilized Dicer (10%−50% v/v) was added to the reaction mixture (final volume, 10−15 μL), and the reaction was incubated at 37 °C for 24 h.
Gel Analysis
Reaction mixtures were analyzed by 16% Tris-borate-EDTA polyacrylamide gel electrophoresis gel. A 3× gel loading solution consisting of urea (6 M), EDTA (250 mM), glycerol (10% w/v), and bromophenol blue (0.1% w/v) was added to the reaction mixture and then separated by gel electrophoresis (140 V, 40 min). The gel was stained using 1× SYBR Gold stain (Invitrogen, Carlsbad, CA) and imaged using on a ChemiDoc XRS+ (Bio-Rad).
The on-target sample preparation protocol is as follows:
a. Spot a 1 µL aliquot of the sample onto the fluorous-modified plate.
b. Evaporate to dryness (protocol A) or allow to stand for 4 min and then remove (protocol B). For protocol A, proceed to steps c and d. For protocol B, proceed to step e.
c. Add 1.4 µL of 0.01−0.1 M aqueous citrate solution.
d. Let stand for 2 min and then remove.
e. Add matrix solution (800 nL) and allow to dry.
MS spectra were then collected on a Bruker Ultraflextreme MALDI-MS equipped with a Smartbeam IITM (Nd:YAG) laser in the linear-positive ion mode. Two thousand laser shots were accumulated at a 1000 Hz acquisition rate. The diameter of the acquisition area was restricted to 1250 μm using a random walk path. The primary readout was the ratio of the RNA-DNA internal standard signal intensity.
Results and Discussion
Since MALDI-MS requires a conductive surface, a proprietary process for the fluorous modification of standard stainless steel MALDI-MS target plates was developed based on published methods for the modification of oxide surfaces using a perfluoroalkyl phosphonic acid. 37 These F-HT-MS plates are manufacturable in a consistent and scalable manner and represent the first commercializable fluorous MS plate. The plates are shelf stable for more than 18 months with no special storage conditions required, and they were also found to be reusable up to eight times each (four times on each side) after cleaning with water, 50:50 water−isopropanol, and 100% isopropanol between each use. The reusability of the F-HT-MS surface is consistent with that observed with other fluorous surfaces,31,33 despite the extra burden of exposure to laser ablation at fixed positions. Reusability of the F-HT-MS plates is highly desirable to reduce the overall cost of an HTS campaign, particularly in comparison with other capture surfaces that are generally single use only. 38
The synthesis of fluorous-tagged oligonucleotides is straightforward as fluorous tags have been used in the synthesis and purification of oligonucleotides.
39
Fluorous tagging was conducted using standard chemistries similar to other tagging methods, for example, biotin, with minor adjustments using either (1) commercially available fluorous phosphoramidite
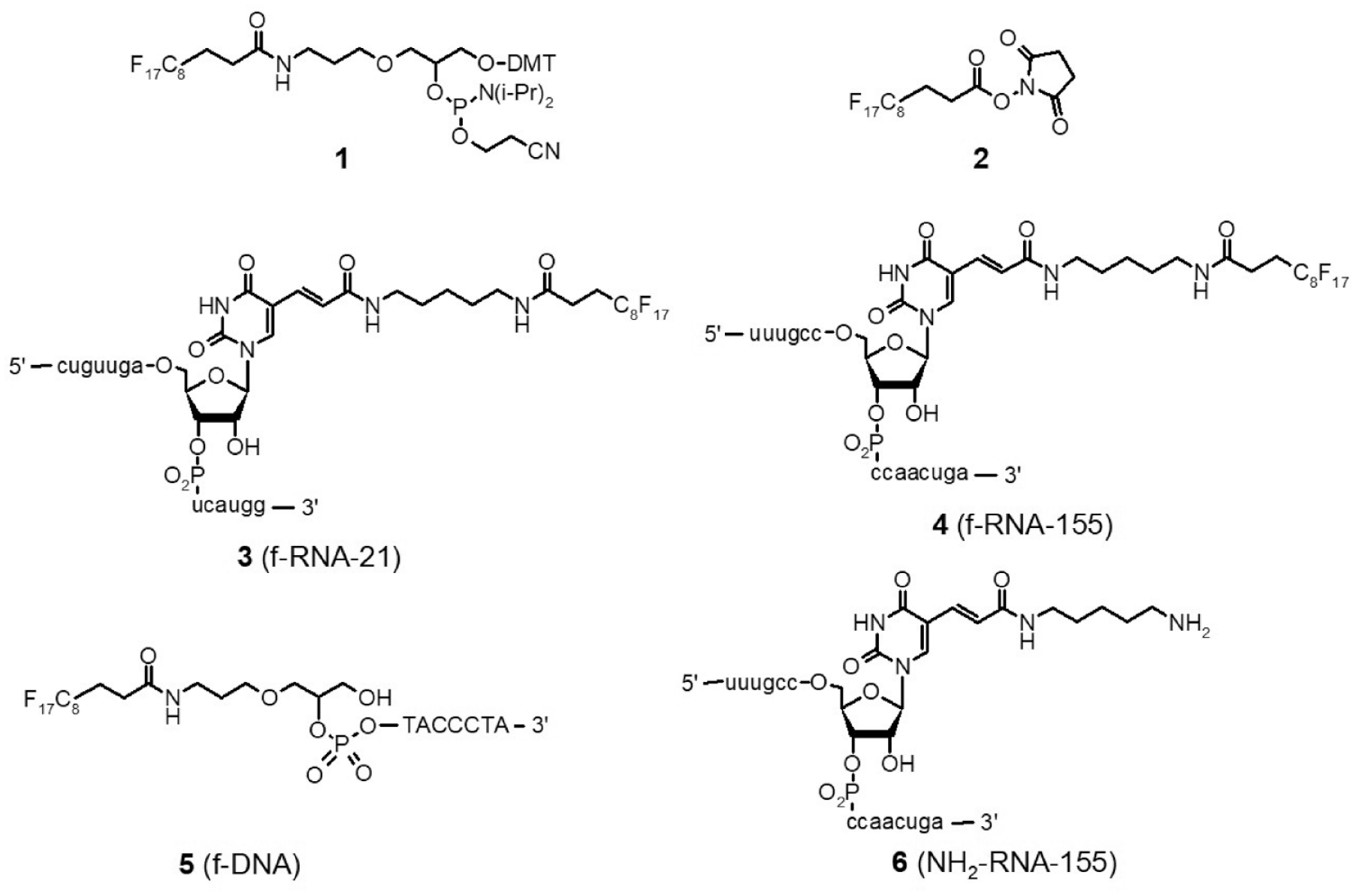
Fluorous tagging reagents (
The protocol first developed (protocol A) involved spotting an aliquot (1 µL) of the analyte solution, evaporation to dryness, and washing the spot area with an aqueous citrate solution (1.4 µL) that is drawn off after 2 min, followed by addition of the matrix solution (0.8 µL). The fluorous-tagged analytes retained on the F-HT-MS plates are then detected by direct laser ionization from the surface. These relatively large spot volumes are possible due to the high solvophobicity of the F-HT-MS plate that allow greater flexibility in solvent composition and volume while still maintaining spot diameters suitable for 384 and 1536 spot configurations. A number of wash solutions were examined before settling on a simple 0.1 M aqueous citrate wash. An investigation of many of the matrices that have previously been reported for oligonucleotide ionization was undertaken, with optimal ionization found using a matrix consisting of 5-methoxy-salicylic acid (5-MSA) and ammonium citrate in 4:3:1 isopropanol−water−trifluoroethanol.
The selectivity of f-RNA and f-DNA capture was confirmed using an aqueous solution of
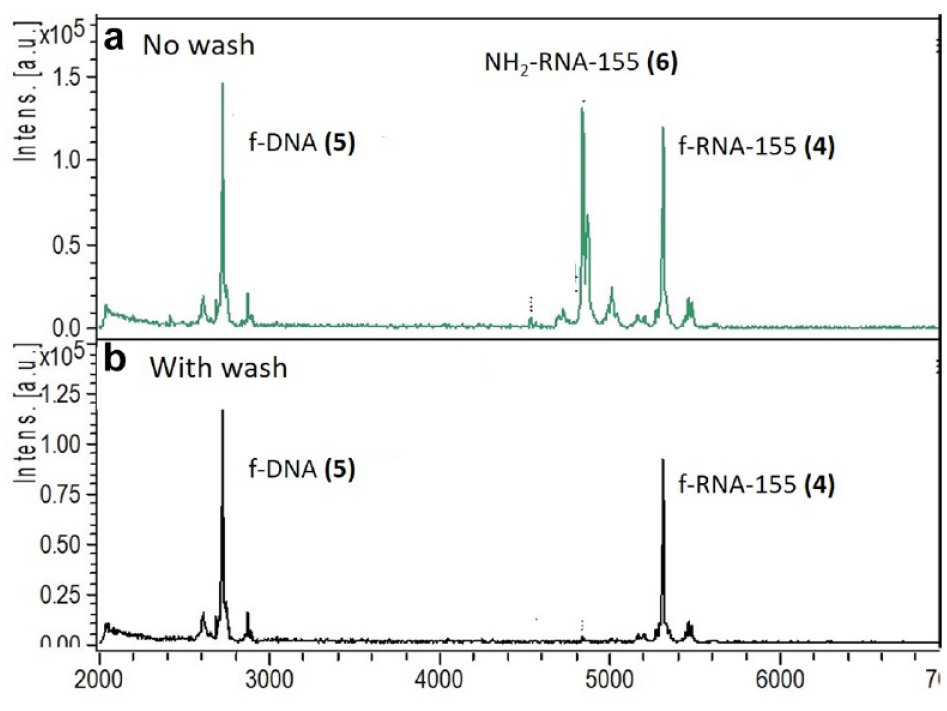
F-HT-MS of RNA and DNA: (
As with MALDI-MS, proper matrix crystallization is critical to obtaining high-quality consistent spectra. Ideally, the matrix and analytes are co-crystallized to achieve maximal energy transfer, greater spot heterogeneity, and increased reproducibility.
9
Methods such as the dried-droplet and thin-layer methods, however, are difficult to employ in conjunction with an on-surface wash step for analyte enrichment. F-HT-MS matrix solution, however, promotes co-crystallization due to the addition of 2,2,2-trifluoroethanol, which partially desorbs the fluorous oligonucleotides from the surface, allowing co-crystallization of the matrix and analytes. This was confirmed by employing a wash using the matrix solvent system of 4:3:1 isopropanol−water−TFE after spot drying, resulting in an average reduction in signal intensity for both the f-RNA and f-DNA of 60%−70% in comparison to the 0.1 M citrate wash (
Attention was then turned to applying the platform to a relevant RNA assay, the Dicer-mediated cleavage of pre-miRNAs. miRNAs are important regulators of gene expression associated with abnormal cell states and viewed as potential drug targets.
40
The repression of aberrantly upregulated miRNAs by inhibiting Dicer cleavage of targeted pre-miRNAs has been postulated as a possible intervention point in miRNA biogenesis, and several assays to identify putative inhibitors have been described.
41
These assays have all employed light-based detection methods, that is, fluorescence or chemiluminescence. A functional MS-based assay would provide several notable advantages, including the elimination of fluorescence interference, the detection of noncanonical cleavage by Dicer, and the ability to multiplex through the addition of several pre-miRNAs. The general scheme for the F-HT-MS miRNA maturation assay is depicted in
Figure 3
. Cleavage of the f-pre-miRNA by Dicer results in formation of the miRNA duplex along with the fluorous-tagged cleaved loop RNA fragment that is detected by F-HT-MS. Inclusion of f-DNA
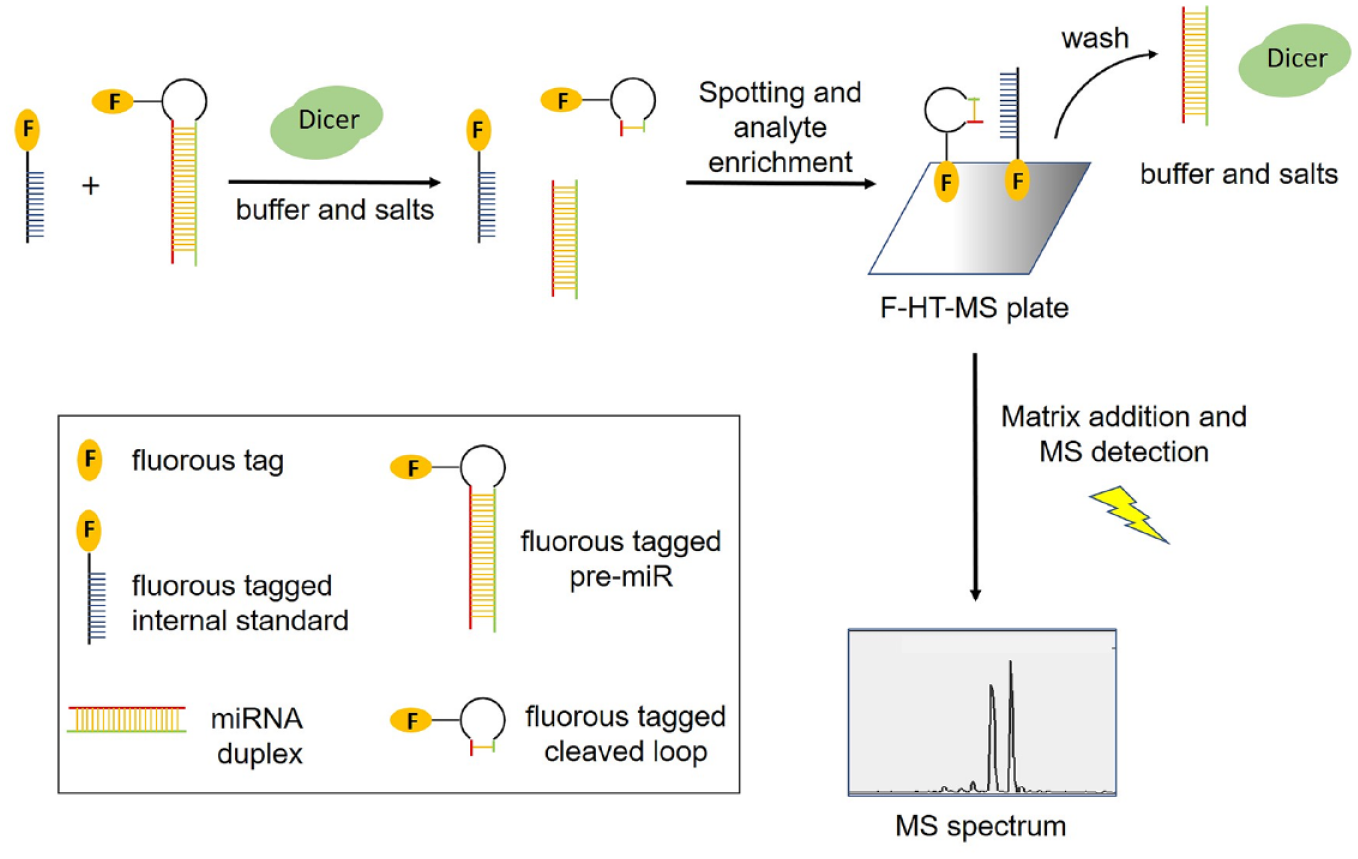
General scheme for f-pre-miRNA processing assay using F-HT-MS.
The F-HT-MS detection of compounds
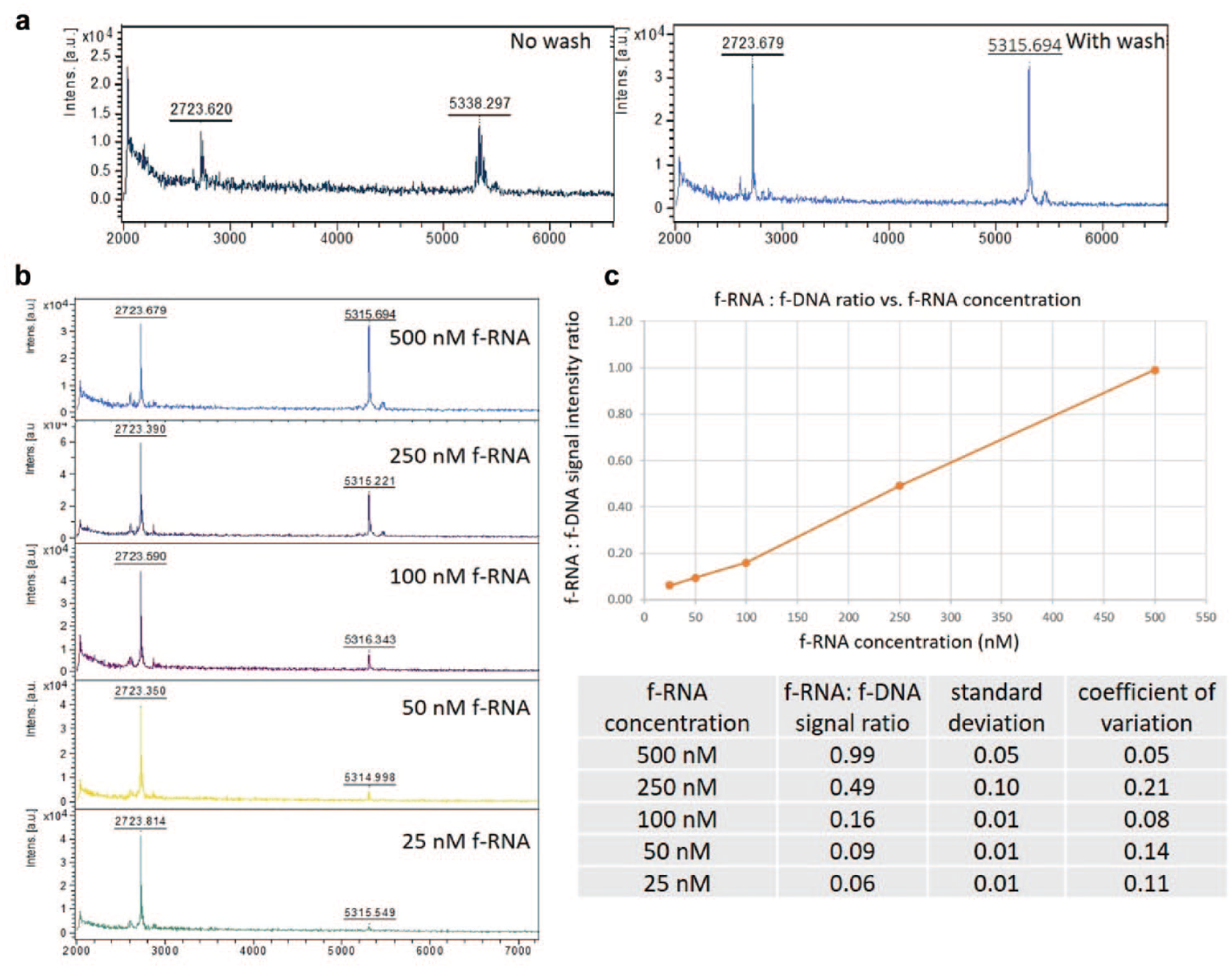
F-HT-MS of f-RNA and f-DNA from Dicer assay buffer. (
With the feasibility of an F-HT-MS-based assay established, the detection of Dicer cleavage of fluorous-tagged pre-miRNAs was pursued. Pre-miR-21 and pre-miR-155, both containing an amine-modified uridine in the loop section, were fluorous modified through carboxylation with
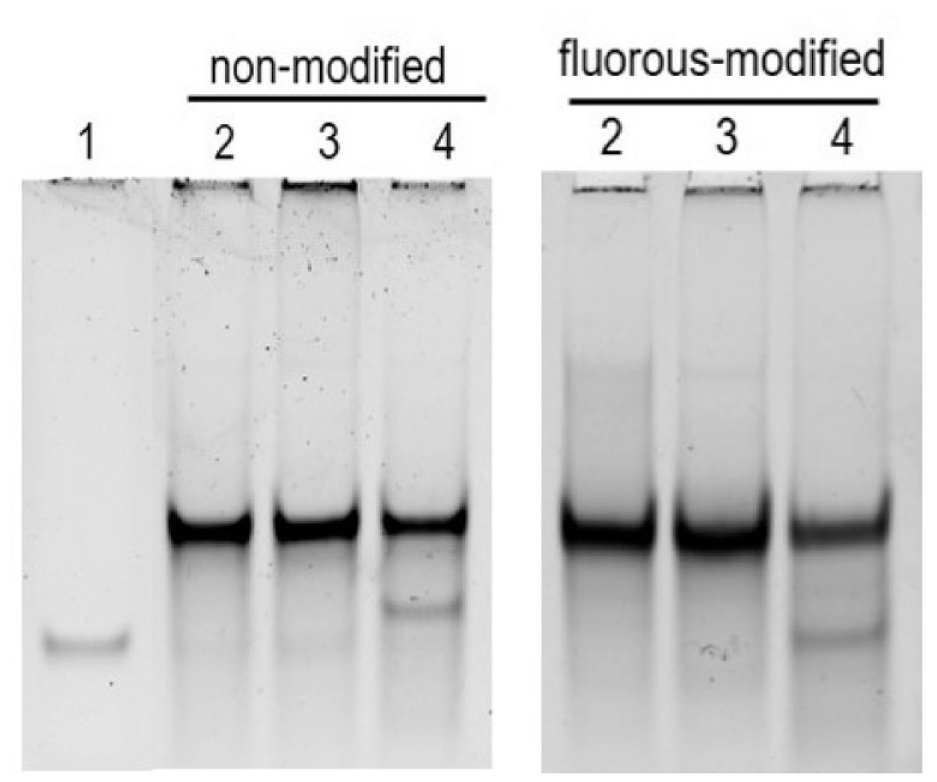
Dicer activity with differently modified RNAs. Immobilized Dicer (50% v/v) incubated with nonmodified or fluorous-modified pre-miR-21 (100 nM) for 24 h. Lane 1: Mature miR-21. Lane 2: Pre-miR-21. Lane 3: Pre-miR-21 + immobilized Dicer + EDTA. Lane 4: Pre-miR-21 + immobilized Dicer.
F-HT-MS detection was then used to detect f-pre-miRNA cleavage by the immobilized Dicer. Negative controls included incubation in the absence of Dicer and in the presence of EDTA, which inhibits Dicer activity through magnesium chelation.
45
The cleavage of both f-pre-miR-21 and 155 was observed by the presence of the fluorous-tagged cleaved loop the 5′-phosphorylated segment (
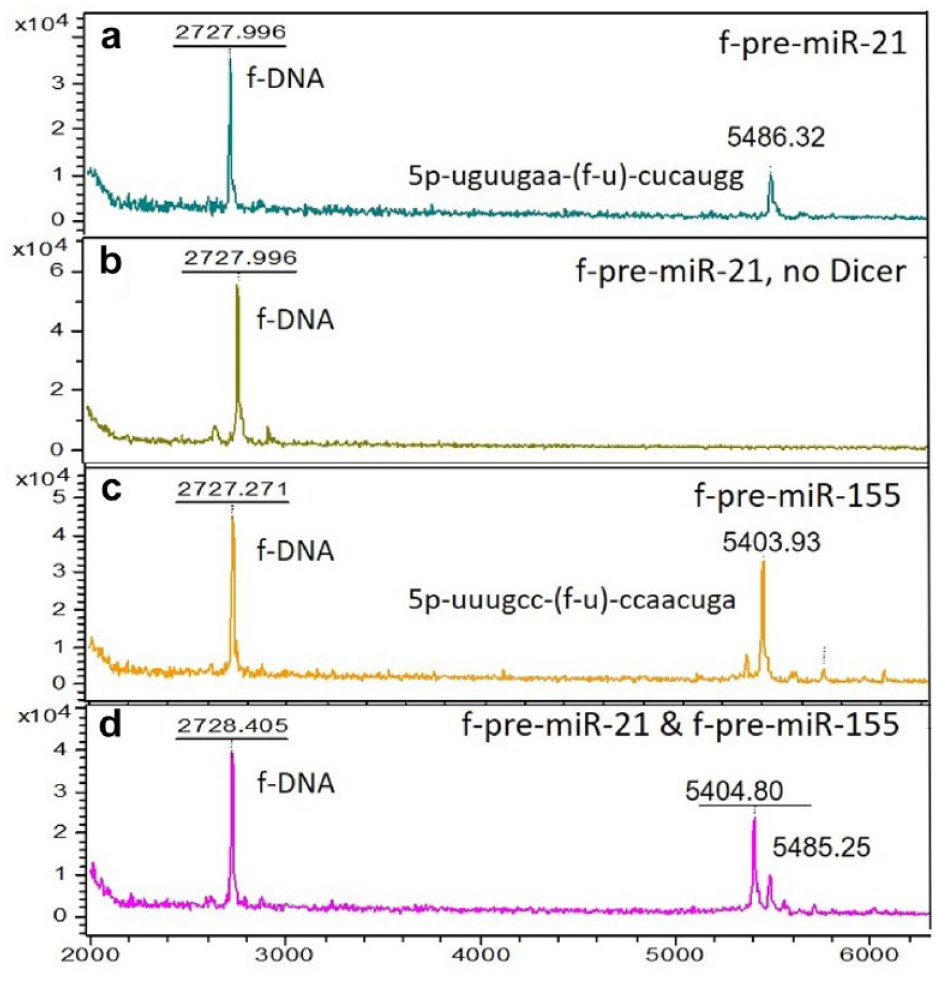
Dicer activity with f-pre-miRs 21 and 155: (
During the course of these studies, it was also found that the initial reaction aliquot need not be evaporated completely to dryness, but that a simple “stand and remove” method (protocol B) was an equally effective analyte enrichment procedure. In this revised protocol, the reaction aliquot is allowed to stand for 4 min and the droplet is then removed prior to the application of the matrix solution. This revised protocol significantly simplifies and shortens the entire process, increasing its suitability for HTS, particularly relative to other on-surface enrichment processes. Importantly, all of these steps have been demonstrated using a Mosquito TPP liquid handler and Bruker MALDI-MS, ensuring that either protocol is automation capable and amenable to a large high-throughput screen.
The ability to detect Dicer inhibition by F-HT-MS assay using protocol B was demonstrated with two compounds: (1) EDTA, which sequesters the Mg2+ required for Dicer activity, and (2) methacycline, which has been reported by Garner et al. as an inhibitor of pre-miRNA cleavage by Dicer, although the exact mechanism is not clear.
47
The inhibition of Dicer activity as a function of EDTA concentration was readily observed as the EDTA concentration approached the MgCl2 concentration of the assay buffer, but the effect of methacycline on miRNA maturation using the F-HT-MS assay required some modification as the signal intensity for both the internal standard
A major advantage of MS-based assays is the ability to analyze multiple analytes simultaneously. The multiplexing capability of the F-HT-MS miRNA biogenesis assay was tested by adding both f-pre-miR-21 and f-pre-miR-155 in a single reaction. Both were digested by Dicer, and each fluorous-tagged loop segment was observed as in the single-substrate reactions, thereby demonstrating the ability to multiplex ( Fig. 6d ). We extended the multiplexing to inhibition by EDTA and methacycline and only observed complete inhibition of both f-pre-miR-21 and f-pre-miR-155 processing, indicating that methacyline is a nonselective inhibitor. While we utilized only two pre-miRNAs in this demonstration, the assay should be applicable to analyzing multiple f-pre-miRs limited only by redundant molecular masses, an issue that can be easily overcome through modification of the linker and fluorous tag in order to produce distinctive m/z peaks corresponding to each particular cleaved loop segment. The ability to use multiple substrates provides a platform that can not only detect potential inhibitors of each of the pre-miRNAs, but also provide an initial indication of selectivity of those inhibitors, thereby focusing HTS triage and hit validation activities on the most promising hits only based on activity and some initial indication of selectivity.
Given the recent surge of interest in RNA processes as drug targets, the development of new high-throughput screening methods to identify modulators of RNA structure and function is of high interest to the drug discovery community, particularly those investigating epitranscriptomics, pre-mRNA splicing, or miRNA processing, all of which result in a structure and mass change in the RNA substrate. The results reported herein provide the basis for the development of new in vitro functional RNA assays, a prominent gap within the field,27,47 since a common feature of small molecule−RNA interactions is that binding does not necessarily equate to function. 50 Noteworthy aspects of the F-HT-MS platform include an easy-to-manufacture, stable, and reusable capture surface; compatibility with existing instrumentation; and multiplexing capability. Substantial benefits compared with existing methods include simpler protocols, reduced costs, lower false signal rates, and the ability to screen against multiple assay substrates in a single screen. The application of the miRNA maturation assay to an HTS campaign and the development of additional functional assays probing other RNA processes will be the subject of future research.
Supplemental Material
Yu_202006_Supplemental_information – Supplemental material for High-Throughput Amenable MALDI-MS Detection of RNA and DNA with On-Surface Analyte Enrichment Using Fluorous Partitioning
Supplemental material, Yu_202006_Supplemental_information for High-Throughput Amenable MALDI-MS Detection of RNA and DNA with On-Surface Analyte Enrichment Using Fluorous Partitioning by Cole Emanuelson, Nicholas Ankenbruck, Alexander Deiters and Marvin S. Yu in SLAS Discovery
Footnotes
Acknowledgements
The authors would like to thank Drs. Amanda Garner and John Schneekloth for helpful discussions regarding Dicer, Dr. Scott Busby for access to HT-MS instrumentation, Dr. Erin Seeley for mass spectrometry guidance, and New River Labs, Helomics Inc., Boron Specialties, and Sharp Edge Labs for laboratory and instrumentation access.
Declaration of Conflicting Interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: Dr. Yu has an equity stake in MS2 Array LLC.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Funding was provided by NIH SBIR grant R43 GM125398-01 and NSF grant CCF-1617041.
Supplemental material is available online with this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
