Abstract
The identification of novel peptide hormones by functional screening is challenging because posttranslational processing is frequently required to generate biologically active hormones from inactive precursors. We developed an approach for functional screening of novel potential hormones by expressing them in endocrine host cells competent for posttranslational processing. Candidate preprohormones were selected by bioinformatics analysis, and stable endocrine host cell lines were engineered to express the preprohormones. The production of mature hormones was demonstrated by including the preprohormones insulin and glucagon, which require the regulated secretory pathway for production of the active forms. As proof of concept, we screened a set of G-protein-coupled receptors (GPCRs) and identified protein FAM237A as a specific activator of GPR83, a GPCR implicated in central nervous system and regulatory T-cell function. We identified the active form of FAM237A as a C-terminally cleaved, amidated 9 kDa secreted protein. The related protein FAM237B, which is 64% homologous to FAM237A, demonstrated similar posttranslational modification and activation of GPR83, albeit with reduced potency. These results demonstrate that our approach is capable of identifying and characterizing novel hormones that require processing for activity.
Introduction
There are more than 80 known human peptide hormones that mediate a wide variety of physiological processes, including reproduction, appetite, metabolism, growth, behavior, cardiovascular function, and electrolyte balance.1,2 The diversity and importance of hormone function in health and disease have prompted searches for new peptides that may offer novel insights into homeostatic mechanisms and novel targets for therapeutics.
The first peptide hormones were discovered more than 75 years ago by arduous biochemical strategies involving stepwise purification from large amounts of tissues, guided by activity in cell-based assays or animal models. The biologic activity of hormones often required extensive posttranslational processing from inactive precursor proteins. One critical modification is the sequential cleavage of precursors in a sequence and tissue specific manner by a family of endoproteases known as the prohormone convertases. 3 A well-known example is preproopiomelanocortin, 4 which contains as many as eight cleavage sites that are used by convertases to generate at least 10 different biologically active peptides, depending on the tissue of origin. In addition to proteolytic cleavage, hormones may also require a variety of other posttranslational modifications for activity, including N-terminal acetylation, C-terminal amidation, formation of N-terminal pyroglutamyl residues (pyrrolation), tyrosine sulfation, phosphorylation, glycosylation, disulfide bond formation, and lipidation. 4 Methods for predicting cleavage and modification sites from sequence information have been described and used successfully to identify novel peptide hormones,5–7 but the challenge of producing the novel hormones in the correctly processed form at scale means that characterization is limited to individual or a small number of candidate hormones.
Our approach to the challenge of assessing predicted candidate hormones at scale was to develop a process to discover novel hormones from a library of predicted preprohormone genes that were identified using an algorithm. Host cell lines competent for regulated secretion were engineered to express the candidate peptide hormones, which were then screened for function in cell-based assays along with known controls. As proof of concept for our approach, we screened the novel hormone library for the ability to activate a set of G-protein-coupled receptors (GPCRs), which were selected by giving preference to orphan receptors as determined by IUPHAR, 8 because the receptors for many known hormones are GPCRs. 9 We successfully identified and characterized two novel, related peptides as potential activating ligands for GPR83.
Materials and Methods
Identification of Candidate Novel Preprohormone Genes
A supervised classifier based on Random Forests of Decision Trees
10
was used to identify potential prohormone convertase substrates. Two training sets of 350 different sequences each were constructed, consisting of 8-mer amino acid sequences derived from unique human proteins from the UniProt database.
11
Each 8mer contained centrally positioned residues known to be cleaved by convertases (GRR, KK, KR, RR, RXKR, RXRR, GKR, GKR, and/or RXXR, where X denotes any residue). The positive control set contained sequences that are known to be cleaved, while the negative control set contained sequences that have the characteristic residues but are known not to be cleaved. Associated with each 8-mer were 20 different attributes derived from the following: two Hidden Markov models representing 108 dibasic sites cleaved by the convertases PC1/3 and PC2; nine Hidden Markov models representing each of the characteristic cleavage sites listed above; residue statistics on the four residues immediately surrounding the cleavage sites; and secondary structure analysis of the full-length protein in the region around the cleavage site. The test set consisted of a random selection of 10% of the sequences removed from the two training sets. This out-of-sample set was used to test the overall performance of the classifier, and the algorithm was modified as necessary. The process was repeated until there was an insignificant change in the classification error rate. All sequences were then classified with an overall score ranging from 0 to 1, where 0 is not cleaved and 1 is cleaved. Based on the out-of-sample test set, a baseline score of 0.4 was set and all genes with a score of ≥0.4 and not previously characterized as preprohormones were chosen for further analysis. We identified 56 genes from this set that were additionally conserved across species and co-expressed in tissues with the prohormone processing enzymes carboxypeptidase E, PC1/3, or PC2 as determined by using the GTEx portal (https://www.gtexportal.org/home/). The complete set of 56 candidate preprohormone genes identified, along with the corresponding cloned nucleotide sequences, are shown in
Candidate Preprohormone Genes Cloned into STC1 and AtT20-PC2 Comprising the Peptide Hormone Library.
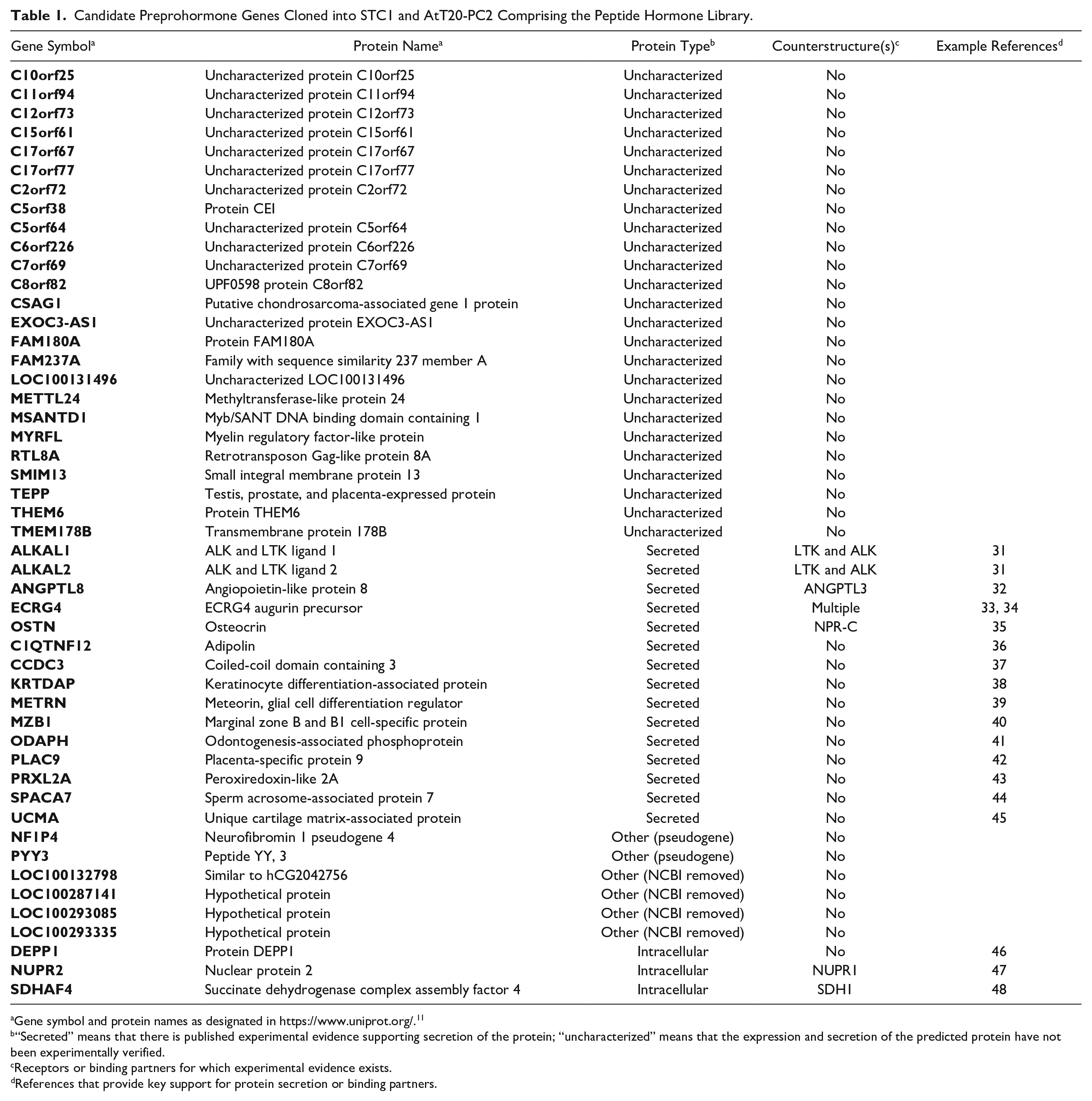
Gene symbol and protein names as designated in https://www.uniprot.org/. 11
“Secreted” means that there is published experimental evidence supporting secretion of the protein; “uncharacterized” means that the expression and secretion of the predicted protein have not been experimentally verified.
Receptors or binding partners for which experimental evidence exists.
References that provide key support for protein secretion or binding partners.
Endocrine Cell Line Sourcing and Culture
AtT20 mouse pituitary cells stably transfected with prohormone convertase 2 (AtT20-PC2) were generously provided by Dr. Richard Mains (University of Connecticut). STC-1 mouse intestinal cells were purchased from ATCC (CRL-3254). HEK293-6E (293-6E) cells (licensed from National Research Council Canada), 12 which lack expression of the prohormone convertases PC1/3 and PC2, were used for the expression of proteins through the constitutive secretion pathway as controls. AtT20-PC2, STC-1, and 293-6E cells were propagated in Dulbecco’s modified Eagle’s medium (DMEM) containing 10% fetal bovine serum (FBS; Corning, Corning, NY) and 100 U/mL penicillin and 100 μg/mL streptomycin (Thermo Fisher, Waltham, MA).
Generation of Preprohormone-Expressing Cell Library
DNA sequences encoding each of the 56 full-length candidate human preprohormones and the positive controls preproglucagon, preproinsulin, and preprochemerin in the PiggyBac expression vector (Lonza, Basel, Switzerland) were electroporated with PBase transposase vector at a 1:1 ratio into AtT20-PC2 and STC-1 cells using the appropriate Lonza Nucleofector kit, following the manufacturer’s protocol. After 48 h, puromycin selection was added (2 µg/mL for AtT20-PC2 and 10 µg/mL for STC-1). Stable cell lines were successfully generated for 49 of the 56 candidate preprohormone genes, but the remaining 7 genes could not be stably transfected into either cell line (
Generation of Conditioned Media from Candidate Preprohormone-Expressing Cells for Screening
AtT20-PC2 and STC-1 cells expressing each of the candidate preprohormones and the controls were thawed, seeded in tissue culture-treated 96-well plates, and passaged every 3–4 days. Cell densities were determined by measurement of supernatant fluorescence at 560 nm excitation and 590 nm emission after a 3 h incubation with Alamar Blue dye (Thermo Fisher DAL1025) and were normalized across cell lines by adjusting culture seeding volumes accordingly during passaging. Normalization for at least three passages was necessary to maximize regulated secretion of hormones for assays, as assessed using positive controls. To induce hormone secretion, the cells were grown to near confluence, and on the day of the assay the cell medium was changed to Opti-MEM (Thermo Fisher 31985070) or DMEM containing 0.5% bovine serum albumin (BSA; Thermo Fisher 15260037), 100 U/mL penicillin, 100 μg/mL streptomycin, and 5 mM N 6 ,2′-O-dibutyryladenosine 3′,5′-cyclic monophosphate (dibutyryl cAMP, Sigma, St. Louis, MO, D0260) for 3 h at 37 °C. The supernatants were then collected, centrifuged, transferred into a 96-well deep well microplate (VWR, Radnor, PA, 75870-796), and immediately assayed for GPCR activity as described in “GPCR Activity Assays and Screening.” The candidate preproproteins were also transiently expressed in 293-6E cells and 48 h conditioned media collected for assays as described previously. 13
GPCR Activity Assays and Screening
GPCRs that have a higher probability of binding to peptide or protein ligands were identified by constructing phylogenetic trees using multiple sequence alignments of selected regions of 816 GPCR encoding genes from UniProt
11
and identifying the GPCRs that lack a confirmed ligand and cluster together with GPCRs that have known peptide ligands. The trees were constructed based on the sequences of domains that may influence ligand specificity, including transmembrane domains, exposed loops, cytoplasmic loops, N-terminal sequences prior to the first transmembrane domain, and C-terminal sequences that follow the seventh transmembrane domain. Seventy-three GPCRs that clustered together with GPCRs known to have protein ligands within one or more of these analyses were prioritized for screening. Eurofins (Luxembourg) DiscoveRx PathHunter eXpress cell lines for 31 of these receptors were available for screening. The complete set of PathHunter eXpress cell lines screened for activity or used as positive controls is listed in
The DiscoveRx PathHunter eXpress assay protocol was modified to adjust cell seeding density, media, reagent, and treatment volumes for screening. The modified reagent and treatment volumes tested were based in part on recommendations from the DiscoveRx technical support team. Optimization was carried out using the GCGR assay (DiscoveRx 93-0241E2), purified glucagon (Sigma-Aldrich G3157-2MG), and glucagon secreted from AtT20-PC2 and STC-1 cell lines or 293-6E as described in the “Generation of Conditioned Media” method section above. The extra cell volume required for automated liquid handling necessitated seeding a reduced cell density compared with the vendor protocol (1740 cells per well vs recommended 2000 cells per well in a 384-well microplate). In a side-by-side test, the reduced cell density had a dose-dependent response with similar EC50, fold induction, and noise compared with the recommended cell density, albeit with a reduced absolute signal and background (data not shown). The conditioned media from the three cell types (AtT20-PC2, STC-1, and 293-6E) was confirmed to be compatible with the PathHunter eXpress assay by testing a glucagon dose–response in each and showing acceptable EC50, fold induction, and noise compared with the recommended assay media (data not shown).
For each GPCR a single-use vial of the corresponding DiscoveRx PathHunter eXpress cell line was thawed and diluted in the DiscoveRx PathHunter Cell Plating Reagent (93-0563R1B) recommended for each cell line to 8.7 × 104 cells/mL. The resuspended cells were pipetted into a sterile 384-individual-well reservoir with pyramid bottom (Thomas Scientific, Swedesboro, NJ, 1145M87), and 20 µL aliquots were seeded in white opaque 384-well plates (Corning 3570) by a Bravo Automated Liquid Handling Platform (Agilent, Santa Clara, CA). The plates were manually sealed with Breathe-Easy adhesive (Sigma-Aldrich Z380059), and after an overnight incubation at 37 °C in 5% CO2, 20 µL of each hormone-conditioned media was applied to duplicate wells using the automated liquid handler. After a 90 min incubation at 37 °C, 5% CO2, 20 µL of freshly prepared detection reagent (DiscoveRx 93-0001L) was added per well. The plates were incubated at room temperature in the dark for a further 90 min, and luminescence was read on an Envision 2103 with 0.2 s integration. For a hormone-expressing cell line to be considered a hit, its corresponding conditioned supernatant had to increase the assay signal ≥3 sigma above background in both duplicate wells. The median background signal and standard deviation were calculated for each cell type (293-6E, AtT20, or STC-1) separately using all wells derived from the conditioned media plate. Up to 12 orphan GPCR DiscoveRx PathHunter eXpress cell lines were tested in a single round of screening, along with one or more of the following as positive controls: the glucagon receptor GCGR (DiscoveRx 93-0241E2) and the chemerin receptors GPR1 (DiscoveRx 93-0335E2) and CMKLR1 (DiscoveRx 93-0313E2) (
Insulin and Glucagon Assays
Mature insulin was measured by enzyme-linked immunosorbent assay (ELISA; Mercodia, Uppsala, Sweden, 10-1113-01), following the manufacturer’s protocol. Glucagon and proglucagon protein levels were measured by ELISA (Bio-Techne R&D Systems, Minneapolis, MN, DY1249), following the manufacturer’s protocol. Insulin activity was measured in the rat hepatoma cell line H4IIE (ATCC CRL1548) grown to passage 13 in Eagle’s minimum essential medium (EMEM) with 10% fetal bovine serum. H4IIE cells were starved overnight in EMEM/0.1% BSA and then treated for 10 min at 37 °C, 5% CO2, with supernatants from AtT20-PC2, STC-1, and 293T cells expressing preproinsulin. The supernatants were diluted in a 1:1 ratio with starvation media prior to treatment, and supernatants from cells expressing vector only with and without exogenous recombinant insulin were used as controls. After treatment, cells were washed with phosphate-buffered saline (PBS), lysed, and phospho-Akt measured according to the R&D Systems ELISA protocol (cat. DYC887). Glucagon activity was measured using the DiscoveRx PathHunter eXpress cells expressing glucagon receptor, as described in “GPCR Activity Assays and Screening” and seeding 2000 cells per well.
Purification of Active FAM237A and FAM237B from AtT20-PC2 Cells
FAM237A-expressing and PiggyBac empty vector-transfected (control) AtT20-PC2 cells were grown in 15 cm plates, and secretion was stimulated by changing the medium to DMEM containing 0.05% BSA, penicillin/streptomycin, and 2 mM barium chloride (Sigma 529591) and incubating at 37 °C for 3 h. FAM237A-expressing and control supernatants were subjected to parallel, identical purification protocols, monitored at each step with the DiscoveRx PathHunter eXpress GPR83 activity assay using the protocol described in “GPCR Activity Assays and Screening” (DiscoveRx 93-0441E2A) and by sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) on a 4%–12% Bis-Tris gel (Bio-Rad, Hercules, CA) in MES running buffer, stained with Sypro Ruby (Thermo Fisher S12000). The supernatants were collected, centrifuged, diluted with 2 volumes of 50 mM Tris (pH 9) and 1 M urea, run over Q HP anion-exchange columns (GE) in 20 mM Tris (pH 9.0) and 1 M urea, and eluted with a linear gradient from 50 mM to 1 M NaCl. This and all subsequent purification steps were performed on an AKTA Purifier (GE, Boston, MA). Aliquots of fractions were dialyzed against 20 mM Tris (pH 8.0) and 150 mM NaCl, and active fractions were pooled and acidified with 1% trifluoroacetic acid (TFA; Sigma T6508). The corresponding fractions from the control supernatants were also pooled and acidified. The pooled samples were run over a 10 × 250 mm C18 column (Higgins Analytical Proto300) in 0.1% TFA, eluting with a linear gradient from 10% to 80% acetonitrile. Aliquots of the fractions were lyophilized and resuspended in 20 mM Tris (pH 8.0) and 150 mM NaCl. Active fractions were pooled and diluted with 0.1% TFA to approximately 10% acetonitrile and then run over a 4.6 × 100 mm C4 column (Vydac 214TP104) and eluted over a linear gradient from 10% to 80% acetonitrile. Active fractions were pooled and lyophilized. FAM237B was purified by the same protocol.
Characterization of AtT20-PC2-Derived FAM237A and FAM237B
For DTT reduction, purified FAM237A protein or control fractions from AtT20-PC2 cells were incubated for 30 min at room temperature with or without 5 mM dithiothreitol (DTT; Sigma). A 25 mM concentration of iodoacetamide (IAA; Sigma) was added to both samples and incubated at room temperature for another 30 min. All of the samples were dialyzed into 20 mM Tris (pH 8.0) and 150 mM NaCl prior to testing in the DiscoveRx PathHunter eXpress GPR83 activity assay. The intact mass of purified FAM237A protein from AtT20-PC2 cells was determined by electrospray ionization mass spectrometry at the University of California, San Francisco Sandler-Moore Mass Spectrometry Core Facility. For N-terminal sequencing, the FAM237A and FAM237B proteins were reduced, run on an SDS-PAGE gel, and transferred to Sequi-Blot PVDF (Bio-Rad 1620184). The relevant bands were cut out and processed by the Tufts University Core Facility for sequencing.
Expression and Purification of PAM
Peptidylglycine α-amidating monooxygenase (PAM) amino acids 1–864 were expressed with a C-terminal human IgG1 Fc tag (PAM-Fc) from the pTT5 vector. CHO-3E714 cells were grown in CD DG44 medium (Thermo Fisher) supplemented with 8 mM
Generation of Active FAM237A and FAM237B from E. coli
The nucleotide sequence of FAM237A corresponding to amino acid residues 34–114, or FAM237B amino acid residues 25–113, with a start codon added, was cloned into the pET-24a vector and then transformed into Rosetta E. coli cells (Sigma 70954) following the manufacturer’s protocol. A colony was grown in Terrific Broth with shaking at 37 °C to an OD600 of approximately 0.5, and then 1 mM isopropyl β-
IP1 Accumulation Assay
293-6E cells were transiently transfected with the nucleotide sequence corresponding to the full-length open reading frame for GPR83 (amino acid residues 1–423) in the PiggyBac vector (or empty vector as a control) by mixing 1:1 with the PBase transposase vector using a 2:1 PEI:DNA ratio. The transfected 293-6E cells were cultured shaking in FreeStyle 293 medium (Thermo Fisher) for 24 h, fed with 0.5% tryptone N1 (Organotechnie), cultured for another 24 h, and then seeded in suspension at 40,000 cells in 7 µL of IP1 stimulation buffer (IP-One HTRF assay, Cisbio, Codolet, France, 62IPAPEB) per well in an opaque low-volume, 384-well plate (Greiner, Monroe, NC, 784075). FAM237A, FAM237B, and control fractions generated from E. coli not expressing the ligands were diluted in IP1 stimulation buffer at twice the final concentration in the assay, and 7 µL was added to the cells immediately after seeding. The plate was sealed and incubated for 60 min at 37 °C, 5% CO2, and IP1 levels were measured versus an IP1 standard curve according to the manufacturer’s instructions.
Results
Identification of Candidate Novel Preprohormone Genes
We developed a method based on the Random Forest of Decision Trees algorithm
10
to identify genes that may represent novel preprohormone convertase substrates. Random Forests is a powerful method for distinguishing classes of individuals based on knowledge of a finite number of individual attributes. The method involves supervised learning with test, training, and unknown datasets containing entries corresponding to the same list of attributes. A significant advantage of this method is that Random Forests are robust to noise and errors. The method as described in “Identification of Candidate Novel Preprohormone Genes,” Materials and Methods, correctly identified all 88 known convertase substrates. This analysis was combined with requirements that the amino acid sequence of the candidate preprohormones be conserved across species and the corresponding genes be co-expressed with the prohormone convertases. Fifty-six candidate preprohormone genes meeting all of these criteria were identified, and 49 were successfully used to construct the screening library as described below (
Table 1
). Vectors encoding the predicted open reading frames for the remaining seven candidate genes failed to generate stable transfectants (
Since the original list of 56 candidate novel preprohormone genes was generated, the majority have limited or no publications supporting function to date, and only 8 have five or more gene-specific references in PubMed as of December 2019. For 40 of the 49 genes within the screening library, 25 of the encoded proteins are functionally uncharacterized, and 15 have been shown to encode secreted proteins. Of the 15 with experimental evidence of secretion, only 5 have reported receptors or binding partners. For the residual nine genes, the National Center for Biotechnology Information (NCBI) has removed four genes and designated two as pseudogenes, although this does not rule out that one or more of these six genes encode functional protein(s). Transcripts from both pseudogenes are detected in the Genotype-Tissue Expression (GTEx) project database (www.gtexportal.org). The remaining three library genes have been reported to encode proteins localized to intracellular organelles. The gene annotations for the screening library are summarized in Table 1 .
Construction and Validation of Novel Hormone-Expressing Endocrine Host Cells
To ensure that the candidate preprohormones in our library were properly processed and secreted in active form, we stably transfected them into cell lines that are known to contain the regulated secretion pathway and to express active hormones upon stimulation with secretagogues, such as cAMP (
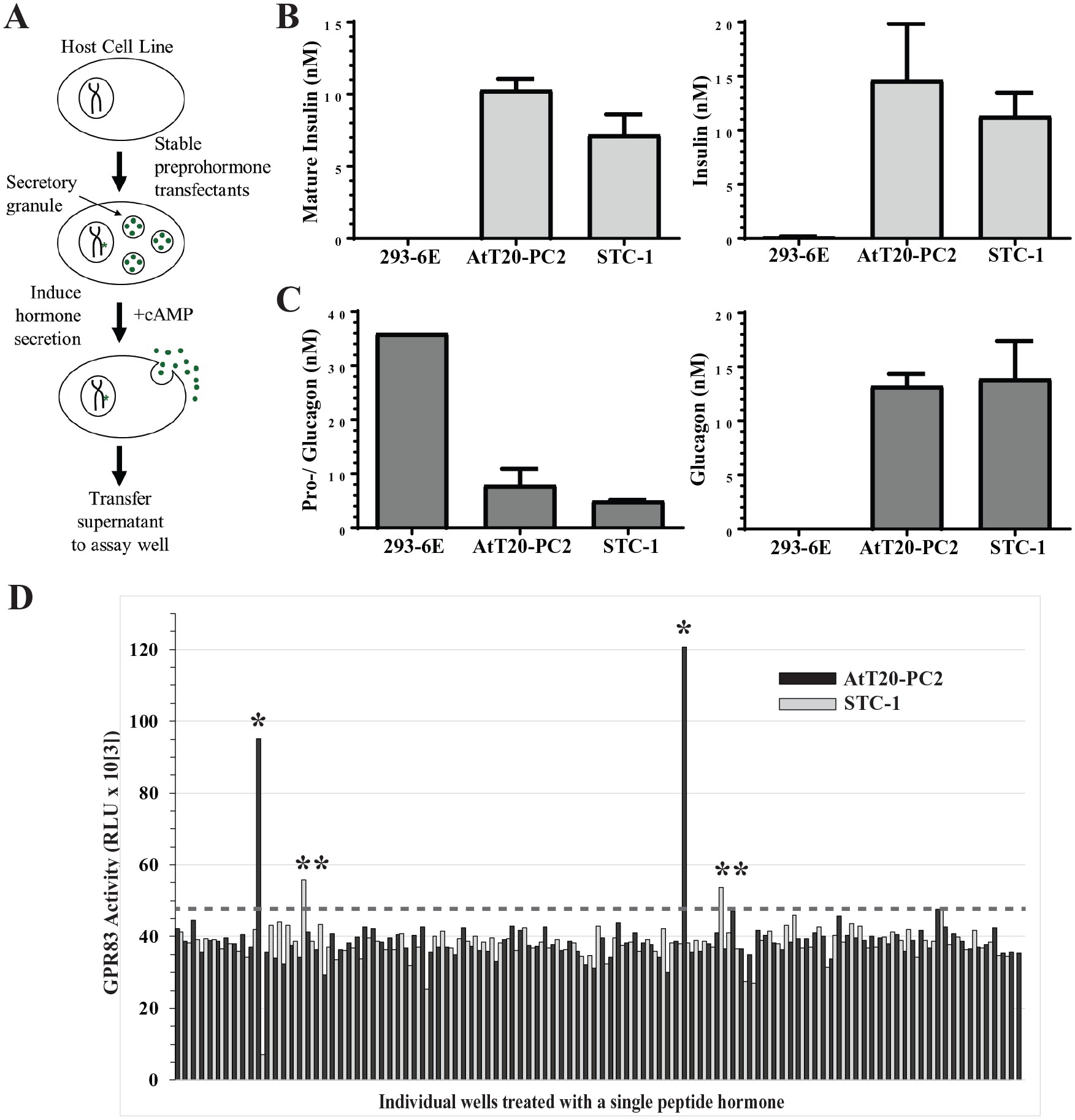
The peptide hormone library produces processed, active hormones and was used for GPCR functional screening. (
To validate that this approach produced properly processed and active hormones, we stably transfected preproinsulin and preproglucagon into AtT20-PC2 and STC-1 cells, collected cAMP-stimulated conditioned media supernatants, and assayed them for expressed protein and for bioactivity as described in “Insulin and Glucagon Assays,” Materials and Methods. Conditioned media supernatants from 293-6E cells expressing preproinsulin and preproglucagon were included in the assay as a negative control, because 293-6E cells lack regulated secretion pathway machinery. 293-6E produced proinsulin protein (data not shown) but no detectable mature insulin protein (
Screening of the Candidate Novel Preprohormone Library for Ligands of Orphan GPCRs
Because the receptors for many hormones are GPCRs, we screened the candidate novel preprohormone library for the ability to activate a set of 31 GPCRs (
Activation of the GPCRs was measured using the PathHunter eXpress cell-based reporter system, which relies on enzyme fragment complementation. CHO or HEK-293 cells were engineered to express (1) the GPCR of interest tagged at the C-terminus with an inactive fragment of the enzyme β-galactosidase, and (2) β-arrestin tagged with the remaining (also inactive) fragment of β-galactosidase. Upon activation, GPCR binding to β-arrestin allows enzyme complementation to occur between the β-galactosidase fragments, resulting in a chemiluminescent signal in the presence of the detection reagents. Each line is confirmed by the manufacturer to express tagged GPCR and β-arrestin by lysing the cells to force enzyme complementation and detecting significant chemiluminescent signal above parental controls. The 31 GPCRs with DiscoveRx PathHunter eXpress cell lines were assayed for ligand-induced signaling after treatment with duplicate conditioned supernatants from the 49 candidate preprohormone and control cell lines expressed by AtT20-PC2, STC-1, and 293-6E cells. Assay optimization and screening is described in “GPCR Activity Assays and Screening,” Materials and Methods.
Supernatants from AtT20-PC2 and STC-1 cells expressing the protein FAM237A induced GPR83 activity 2.8- and 1.4-fold above background, respectively, in the screen (
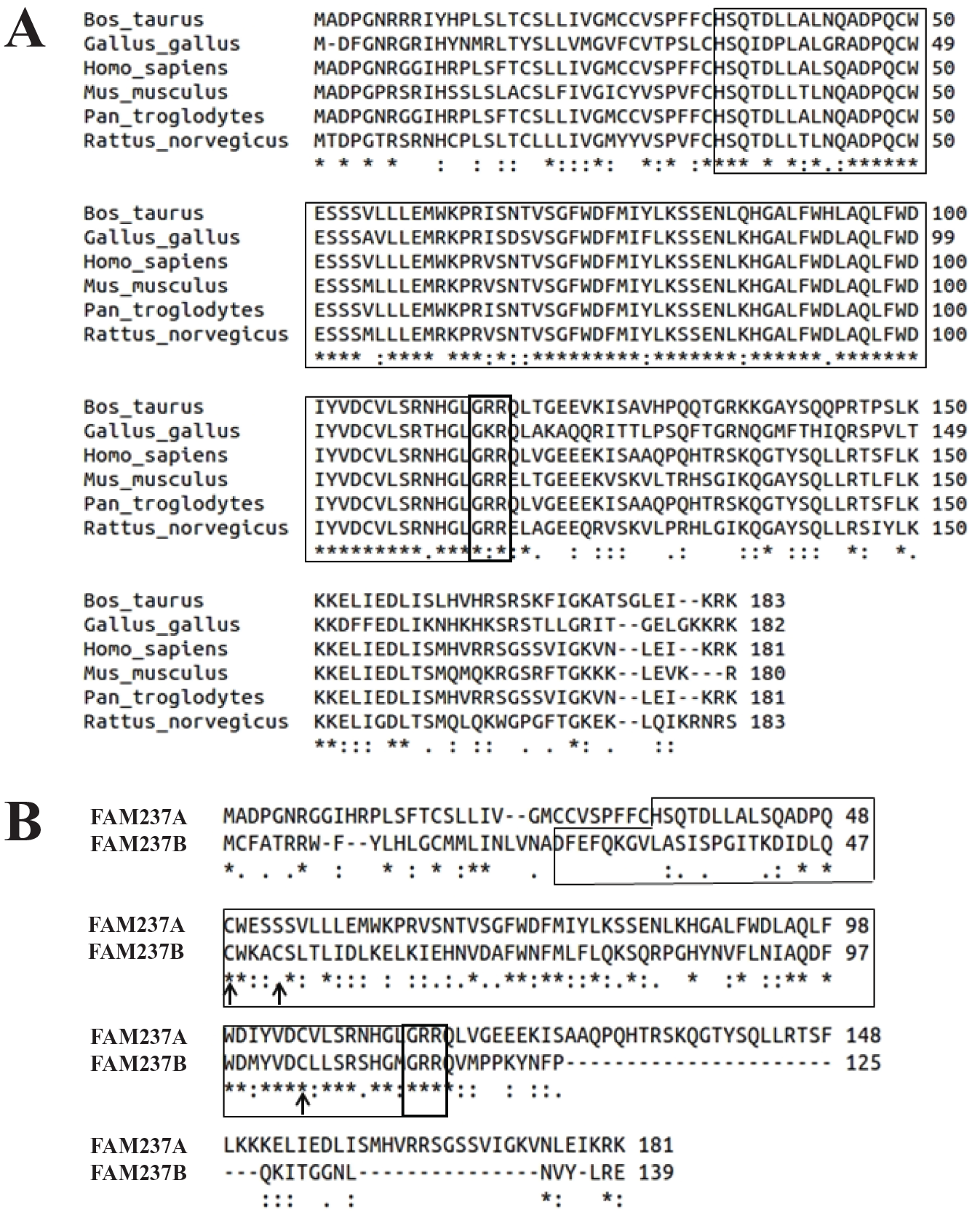

Sequence conservation of FAM237A and FAM237B. (
Purification and Characterization of the Active Species of FAM237A and FAM237B
We selected FAM237A-expressing AtT20-PC2s to purify the active species of FAM237A, because dose-dependent GPR83 activity was consistently greater in AtT20-PC2 cell supernatants compared with STC-1 (
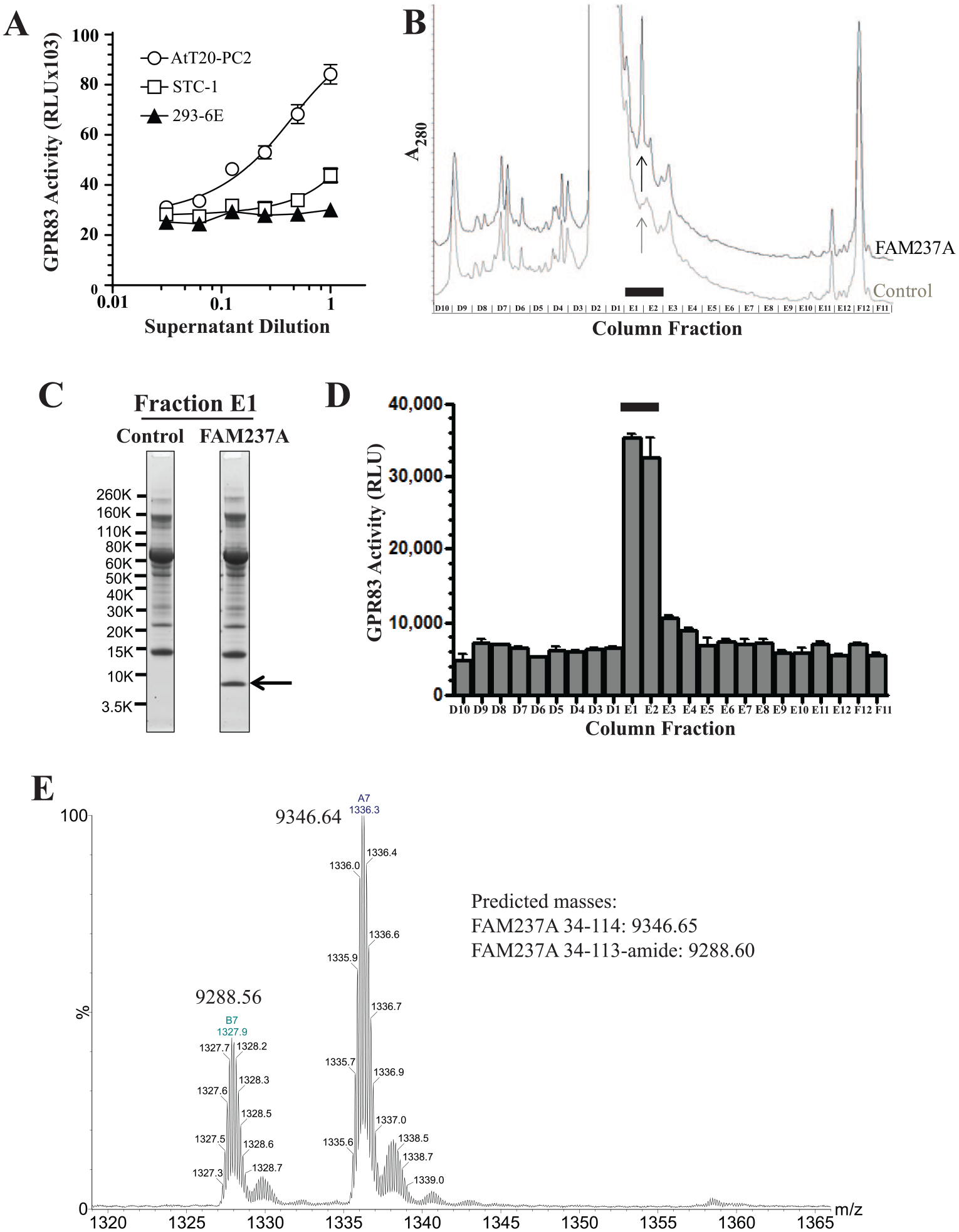
Partial purification and characterization of the active form of FAM237A protein from AtT20-PC2 cells. (
Conditioned supernatants of FAM237A-expressing AtT20-PC2 cells were fractionated by serial chromatography as described in “Purification of Active FAM237A and FAM237B from AtT20-PC2 Cells,” Materials and Methods, using the PathHunter eXpress GPR83 activity assay to identify active fractions throughout the purification. FAM237A-expressing cells were stimulated with barium chloride to enhance release of active hormone, and the conditioned supernatants were subjected to sequential anion-exchange, C18, and C4 reversed-phase chromatography. Conditioned supernatants from AtT20-PC2 cells transfected with control (empty) vector were harvested and subjected to the same purification steps as a control. The A280 profile of fractions from the C18 reversed-phase step for FAM237A-expressing supernatants had an A280 absorbance peak in fractions E1 and E2 that was absent from the control (
Generation of Purified, Recombinant, Bioactive FAM237A and FAM237B from E. coli
To confirm that we had accurately defined the structural features critical for induction of GPR83 activity, FAM237A (residues 34–114) and FAM237B (residues 25–113) were expressed in E. coli and processed in vitro. The proteins were harvested from E. coli inclusion bodies, solubilized, partially purified by anion-exchange chromatography, refolded, and further purified on a Source 15RPC column. Lysates from E. coli transformed with empty vector were used as controls and subjected to the identical purification and refolding protocol. Because mass spectrometry suggested that C-terminally amidated protein was present in the protein purified from AtT20-PC2 cells, the purified, E. coli-produced material was amidated with varying amounts of a PAM-Fc fusion protein generated as described in Materials and Methods and then repurified on the Source 15RPC column. The final E. coli-purified FAM237A material was a single, approximately 9 kDa band by SDS-PAGE (
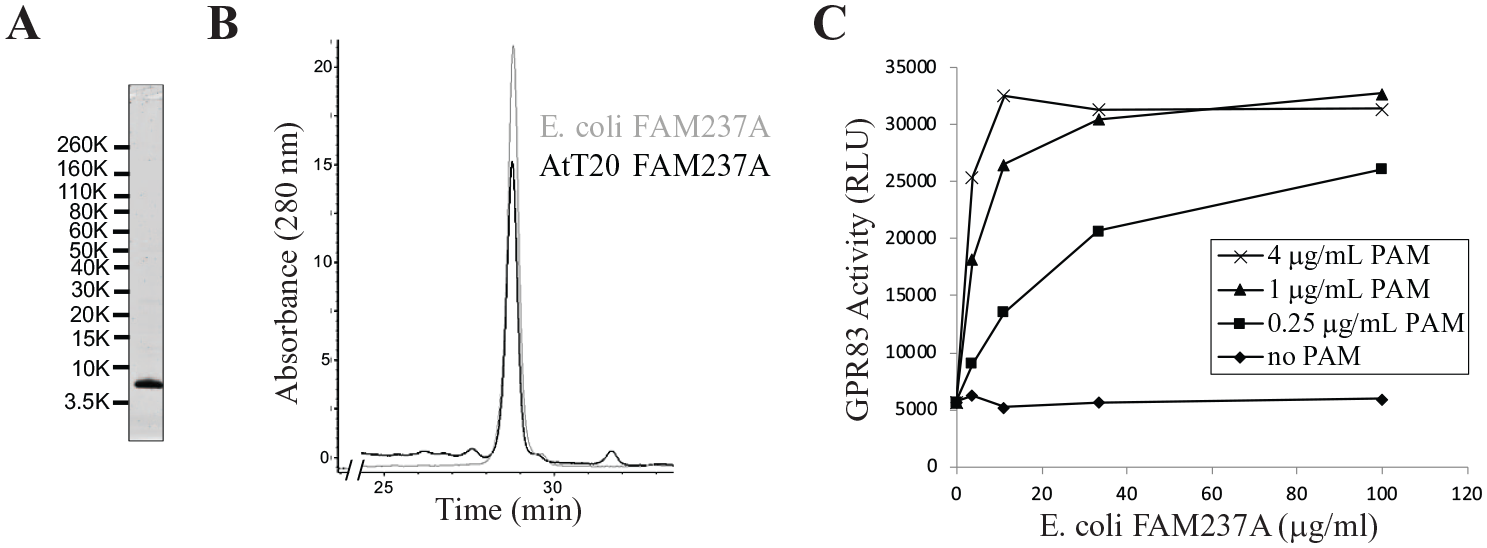
Generation of purified, active recombinant FAM237A expressed in E. coli requires amidation. FAM237A was expressed in E. coli, refolded and purified as described in “Generation of Active FAM237A and FAM237B from E. coli,” Materials and Methods. (
Identification of GPR83 Signaling Pathways Activated by FAM237A/B
Native sequence GPR83 was transfected into 293-6E host cells, which lack endogenous GPR83, and the cells were stimulated with a dose titration of E. coli-purified, bioactive FAM237A, FAM237B, or negative control. The cells were then assayed by using the IP-One HTRF assay as described in “IP1 Accumulation Assay,” Materials and Methods, for production of IP1, an indirect measure of the generation of the second messenger IP3, which is produced downstream of GPR83 coupling to Gαq subunits. Both FAM237A (
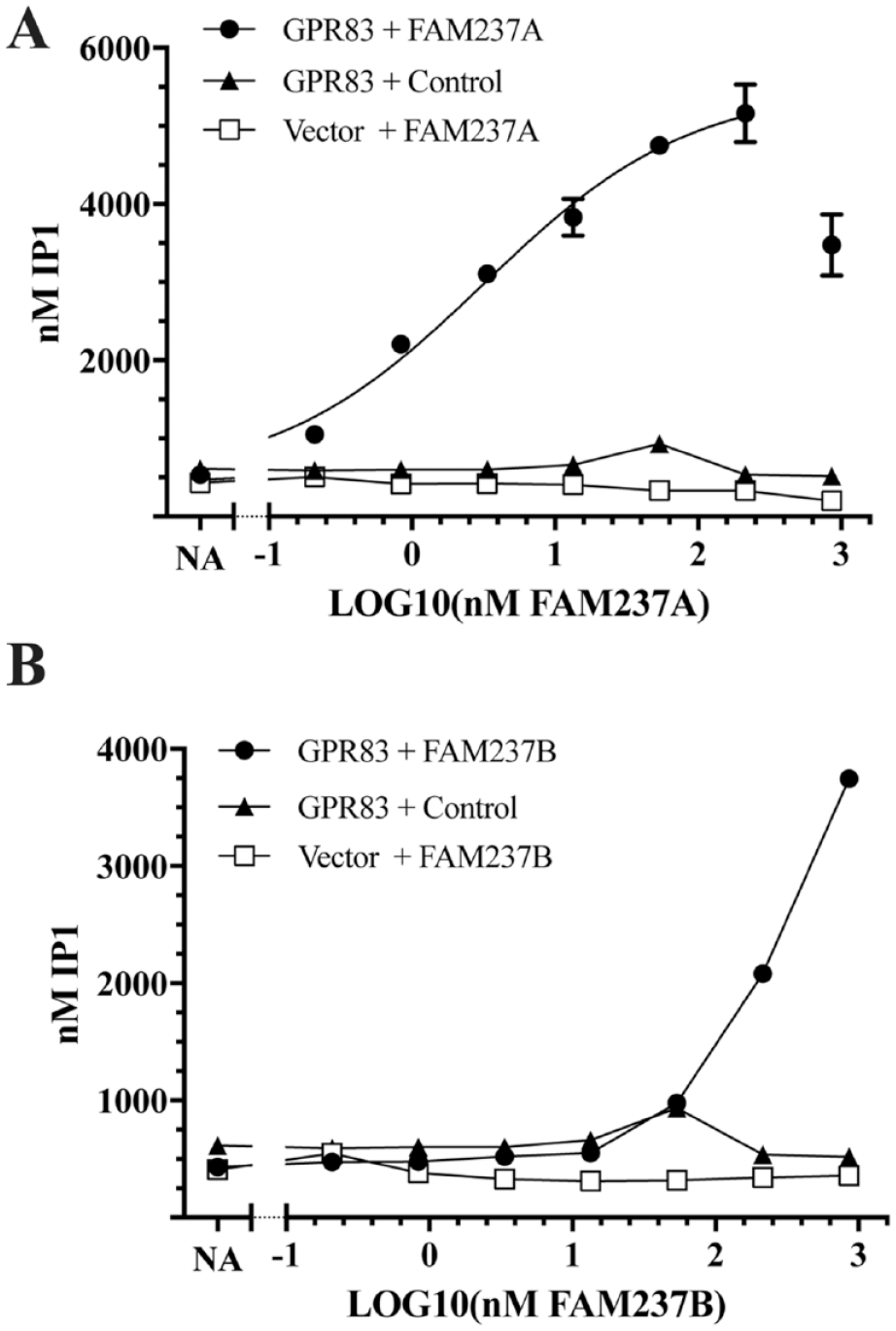
FAM237A and FAM237B stimulate GPR83-dependent IP1 accumulation. 293-6E cells were transiently transfected with native GPR83 or empty vector, treated in duplicate with the indicated concentrations of (
FAM237A Expression in Normal Human Tissues
FAM237A and GPR83 mRNA expression was assessed in normal tissues by using the GTEx project database (www.gtexportal.org). Consistent with published reports, GPR83 is expressed broadly in the human brain,
17
particularly in the cerebellar regions, as well as in the testis and thyroid gland (
Discussion
We describe here a process that was developed to identify novel peptide hormones. Our approach was to generate a set of novel preprohormone candidate genes based on known peptide hormone properties, express candidate proteins in host cell lines with a regulated secretory pathway capable of processing preprohormones into mature, active peptides, and functionally screen for hormone activity using GPCR cell-based reporter assays. Once a novel hormone activity is identified, the host cell lines can be scaled up to purify and characterize the mature active form, which is a key advantage given that standard expression hosts lack the posttranslational processing machinery required for many hormones, hormones often have restricted or very low expression endogenously, and synthesis is often not feasible depending on the posttranslational modifications. Expressing each candidate gene in two cell lines increased the likelihood that the hormone was properly processed and minimized the effect of endogenous expression of known hormones by the host cells, which may confound assay results. It is still possible that some candidate genes are not fully processed in these hosts, which can be addressed by incorporating one or more host lines derived from other tissues into the library. For the seven candidate genes that failed to generate stable lines in either host, preliminary troubleshooting suggested that expression of these genes interfered with host cell expansion in some way. For these clones, in addition to a new host cell, an inducible or weaker promoter might mitigate this issue and enable stable pool production, and cloning into more than one vector should be considered for these and future additions to the library.
We demonstrated the utility of our library by identifying a posttranslationally modified form of FAM237A protein, as well as its homolog FAM237B, that is capable of activating GPR83 via the Gαq signaling pathway. This is the first report of a biological function for FAM237A or FAM237B, showing the ability of our approach to identify novel biology. GPR83 has been shown to signal via the Gαq pathway,19,20 which is consistent with the activation by FAM237A. The ability of FAM237A and FAM237B proteins to activate GPR83 was dependent on specific modifications that are characteristic of peptide hormones and only made by host cells containing the regulated secretion pathway. This supports their identities as prohormones and further demonstrates the utility of the system for identifying novel hormone activities, as standard expression hosts such as 293-6E did not produce active protein. Compared with the STC-1 cell line, AtT20-PC2 cells produced more active FAM237A protein, which could be because these cells originate from the pituitary, where FAM237A expression has been detected. We do not know if the active form of FAM237A or FAM237B described here corresponds to the fully mature endogenous form in humans, since we are using a cell line derived from a murine host. However, the EC50 for FAM237A activation of transiently expressed GPR83 was estimated as 3.2 nM, which is within the range typically observed for other hormone–receptor pairs. 8 Further confirmatory studies that FAM237A (and FAM237B) is a physiologically relevant GPR83 ligand must also be done, including generation of GPR83 binding curves, specific activity in cells endogenously expressing GPR83, and characterization of the mature peptide form in a cell that endogenously expresses FAM237A. GPR83 has been reported to form hetero-oligomers with other GPCRs, including GPR171, 20 GHSR, MC3R, and MC4R; 21 thus, the ability of FAM237A and FAM237B to activate GPR83 hetero-oligomers should also be assessed. The E. coli-produced FAM237B had significantly reduced potency compared with FAM237A in activating GPR83, and comparison of their sequences revealed an unpaired cysteine at position 52 that is not conserved in FAM237A. Unpaired cysteines can reduce the activity of E. coli-produced proteins; however, a C52S substitution had no effect on FAM237B activity. The reduced potency of FAM237B may instead reflect a physiological role distinct from activation of GPR83 homo-oligomers.
The identification of an additional candidate peptide ligand for GPR83 should further facilitate characterization of GPR83 biology. GPR83 has been reported to be activated by multiple modalities, including basal activity regulated by the N-terminal extracellular domain,21,22 Zinc(II), 19 and most prominently the ligand neuroendocrine peptide PEN. 20 PEN is a peptide hormone with no significant homology to FAM237A (2% identity of mature forms by Clustal Omega 11 ) that is derived from the precursor proSAAS encoded by the PCSK1N gene, which is processed into at least five other peptides: SAAS, GAV, PEN, bigLEN, and littleLEN. PEN binding to GPR83 induces a dose-dependent reduction in cAMP and an increase in PLC activity, indicating PEN induces GPR83 to signal through Gαq and the inhibitory Gαi G-proteins. 20 Zinc(II) induces activation of GPR83 signaling via Gαq but not Gαi or Gαs. 19 The dose-dependent increase in IP1 induced by FAM237A is consistent with inducing GPR83 signaling through Gαq. In contrast, we were unable to detect a consistent, dose-dependent effect of FAM237A on GPR83 resulting in an increase or decrease in cAMP levels. While the signaling pathways downstream of GPR83 appear to be context or ligand dependent, GPR83 activation in response to FAM237A is consistent with the published modalities.
The expression pattern of FAM237A in public transcript datasets is also consistent with a role in GPR83 biology. In normal human tissues, GPR83 and FAM237A expression overlap within the brain, as does the published ligand PEN, which is reported to be robustly expressed in the hypothalamus. 20 GPR83 is largely conserved in human and mouse, with the strongest levels within the brain, including cerebellar, hypothalamic, hippocampal, and amygdaloid regions. 17 The expression of FAM237A in the pituitary and pancreatic islet alpha cells is potentially consistent with a role in modulating GPR83 function in the thyroid or in metabolism, respectively.
GPR83 expressed in the central nervous system may play a role in neurological functions, including emotion, learning, reward processing, and metabolism.17,23 Knockdown of GPR83 in the hypothalamic preoptic area reduces core body temperature and elevates circulating levels of adiponectin. 24 Analysis of GPR83 knockout mice has suggested that the receptor may be involved in stress, reward and learning, 25 and the regulation of systemic energy metabolism. 26 GPR83 is also expressed in mouse FoxP3+ regulatory T cells (Tregs) 17 and has been implicated as a mediator of immunosuppressive Treg formation or function during inflammation. Transfer of GPR83-overexpressing CD4+ T cells into mouse models of inflammation is associated with reduction in inflammation and expression of Treg markers in the adoptively transferred cells. 27 This suggests the possibility that FAM237A and/or PEN may be upregulated by inflammation in vivo to activate GPR83 and induce peripheral generation of Tregs, either directly or indirectly.
The fact that GPR83 is the only receptor for which we identified a novel candidate ligand as part of our initial screen may result from a combination of biological and technical causes. The GPCRs in the screen were selected by sequence-based clustering with GPCRs that have confirmed peptide ligands. Despite this enrichment, it is likely that some of these GPCRs do not have peptide ligands and that the candidate hormones in our collection have receptors that are not GPCRs, as is the case for insulin. The GPCR PathHunter eXpress cell lines were not validated to have plasma membrane expression of active GPCRs, and so may lack sufficient surface receptors for a detectable response. The reporter cell line limitations could be mitigated by using different reporter lines and validating surface expression.
It has been noted that the rate of novel peptide discovery and GPCR de-orphanization has slowed, which has been attributed to a number of causes, including the complexity of signaling pathways and posttranslational processing. 9 Our success rate in identifying novel ligand–receptor interactions is consistent with other GPCR de-orphanization attempts using β-arrestin recruitment as a readout, 28 which may be caused by distinct signaling mechanisms of the remaining orphan GPCRs or the pleiotropic nature of GPCR signaling. 29 This strongly argues that orthogonal assays are required to further interrogate the library. Techniques such as label-free dynamic mass redistribution that are agnostic to downstream signaling pathway have been successfully used to identify GPCR ligands in a variety of cellular formats in a high-throughput manner. 30 This would enable GPCRs to be screened in a pooled fashion or in primary cells, which would also allow for the identification of ligands for hetero-oligomers. Given our success in identifying a novel activator for GPR83, future screens should incorporate a broader array of GPCRs, particularly those of therapeutic relevance, to increase the likelihood of identifying novel ligand–receptor interactions.
Beyond direct identification of novel ligand–receptor interactions, we believe that our peptide hormone library is compatible with most in vitro assays for the identification of novel candidate hormone functions, particularly for activities that can only be detected with endogenous receptors under more physiological conditions. To identify hormones that induce complex physiological phenomena, such as behavior or metabolism, the peptide hormone cell line collection is adaptable for in vivo approaches by implantation of host cell lines via alginate beads into appropriate murine model systems to assess the effects on behavior or metabolism. We believe that our approach can contribute to the future characterization of novel hormones, and that the novel hormone we identified, FAM237A, may play an important role in GPR83 function in the central nervous, endocrine, metabolic, or immune system.
Supplemental Material
Supplemental_Material_for_Peptide_Hormone_Platform_Identified_Candidate_GPR83_Ligands_by_Sallee_et_al – Supplemental material for A Pilot Screen of a Novel Peptide Hormone Library Identified Candidate GPR83 Ligands
Supplemental material, Supplemental_Material_for_Peptide_Hormone_Platform_Identified_Candidate_GPR83_Ligands_by_Sallee_et_al for A Pilot Screen of a Novel Peptide Hormone Library Identified Candidate GPR83 Ligands by Nathan A. Sallee, Ernestine Lee, Atossa Leffert, Silvia Ramirez, Arthur D. Brace, Robert Halenbeck, W. Michael Kavanaugh and Kathleen M. C. Sullivan in SLAS Discovery
Footnotes
Acknowledgements
The authors thank DiscoveRx, Jackie Chan, Thomas Bray, Shawn Russell, Grayson Kochi, Kaumudi Bhawe, Nallakkan Arvindan, John Cesarek, Elizabeth Bosch, Hongbing Zhang, and Jin Zhou for technical assistance, and Richard Mains and Lewis T. Williams for guidance and advice. The data used for the convertase co-expression analysis and ![]() were obtained from the GTEx portal, which is supported by the Common Fund of the Office of the Director of the National Institutes of Health, and by NCI, NHGRI, NHLBI, NIDA, NIMH, and NINDS. The work described in this manuscript work was funded by Five Prime Therapeutics Inc.
were obtained from the GTEx portal, which is supported by the Common Fund of the Office of the Director of the National Institutes of Health, and by NCI, NHGRI, NHLBI, NIDA, NIMH, and NINDS. The work described in this manuscript work was funded by Five Prime Therapeutics Inc.
Supplemental material is available online with this article.
Authors’ Note
Kathleen M. C. Sullivan is now affiliated with ChemoCentryx.
Declaration of Conflicting Interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: All authors were employed by Five Prime Therapeutics, and their research and authorship of this article was completed within the scope of their employment with Five Prime Therapeutics.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
