Abstract
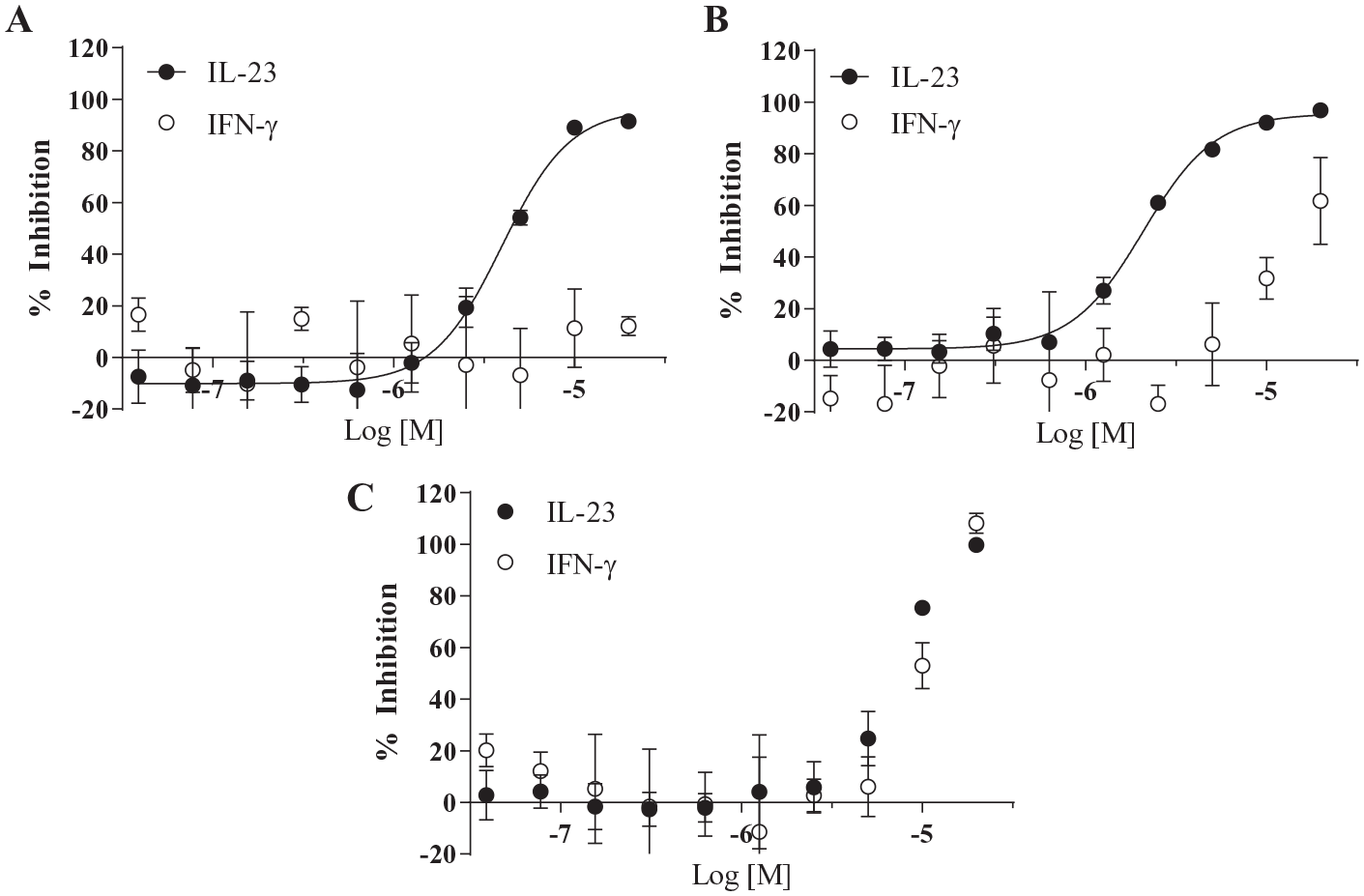
Interleukin-23 (IL-23) is a key cytokine implicated in the pathogenesis of autoimmune disorders, including psoriasis and ulcerative colitis. Although targeted IL-23 antibody therapeutics are used clinically, there are no small-molecule therapeutics that selectively inhibit IL-23 signaling. To address this gap, we developed a high-throughput screening strategy employing an IL-23-responsive cell-based luciferase reporter gene assay as the primary screen, with cellular cytotoxicity and off-target counter screening assays to identify IL-23 pathway-specific inhibitors. The primary screening assay utilized avian DT40 cells, genetically engineered to overexpress IL-23R, IL-12Rβ1, STAT5, and firefly luciferase, in a 1536-well format. Treatment of these cells with IL-23 resulted in the phosphorylation and activation of STAT5, which was completely inhibited by the pan-JAK inhibitor tofacitinib. Assay performance was robust, with signal-to-background >7-fold and Z′ > 0.5 over 40 screening plates (approximately 24,000 compounds), with a hit rate of 5% (>66.9% activity cutoff). Of these 1288 hits, 66% were identified as cytotoxic by incubating the IL-23 reporter cells with compound overnight and measuring cell viability. Further assessment of specificity via examination of impact on off-target IFN-γ signaling eliminated an additional 230 compounds, leaving 209 that were evaluated for dose–response activity. Of these compounds, 24 exhibited IC50 values of <7 µM and ≥80% inhibition of IL-23 activity, with >3-fold selectivity over IFN-γ inhibition, thus representing promising starting points for prospective IL-23 pathway small-molecule inhibitors.
Introduction
Interleukin-23 (IL-23) is a pleiotropic cytokine implicated as a prominent driver of autoimmune inflammatory pathology. In disorders including psoriasis and inflammatory bowel disease, IL-23 is aberrantly produced by dendritic cells and monocytes/macrophages and elicits a host of downstream effector activities. These include IL-23 modulation of adaptive and innate responses through its role in the maintenance and expansion of T helper 17 (Th17) and γδ T cells and activation of innate lymphoid, natural killer (NK), and natural killer-like (NKT) cells.1–4
IL-23 is a heterodimeric protein consisting of the p40 and p19 subunits, 5 with the p40 subunit also shared by IL-12 (p40-p35). The p40 cytokine subunit interacts with the IL-12Rβ1 receptor subunit, 6 making this receptor subunit common to both IL-23 and IL-12 receptor complexes, while p19 binds IL23-R.7,8 Dimerization of the receptor subunits upon cytokine binding results in the recruitment of JAK and subsequent phosphorylation of STAT (pSTAT), which then translocates to the nucleus and initiates gene transcription.7,9 Although initial antibody therapies were developed to target IL-12 via binding p40 and inhibiting receptor interaction, the discovery of IL-23 and elucidation of the common p40 subunit revealed that these therapies inhibited both cytokines. With the subsequent characterization of IL-23 function and recognition of the benefits of its inhibition independent of IL-12 inhibition, IL-23-selective antibodies targeting the p19 subunit were developed.10,11
The clinical success of antibodies that target IL-23 has clearly demonstrated that modulation of the IL-23 pathway can result in safe and effective therapies for patients with some immune-mediated diseases. An orally administered, small-molecule inhibitor of IL-23 signaling may result in benefit to additional populations of patients. However, the identification of small molecules that are capable of disrupting large, high-affinity protein–protein interfaces, such as those between cytokines and receptors, has historically been a significant challenge. 12 Larger, more diverse screening libraries, combined with the development of cell-based systems to employ as screening tools, may provide better approaches to enable the identification of IL-23 inhibitors than traditional biochemical screens. Advantages of cell systems include the physiologically relevant structure of prospective target proteins and the presence of signaling pathway components and other requisite accessory proteins. In the case of IL-23, this would translate to an IL-23-responsive cell system containing the full-length IL-23R/IL-12Rβ1 receptor complex that is capable of binding cytokine and engaging intracellular effectors such as JAK/STAT. Herein we describe the use of the IL-23 iLite reporter gene cell system13–15 in 1536-well miniaturized format to screen a pilot set of approximately 20,000 small-molecule compounds for IL-23 pathway inhibitors.
Materials and Methods
Steady-Glo Assay
iLite IL-23 cells (Svar Life Science, Malmö, Sweden) were thawed rapidly in a water bath at 37 °C for 1 min with agitation. Following removal of 570 µL of 600 µL, cells were resuspended in growth media (RPMI [Thermo Fisher, Waltham, MA] + 9% heat-inactivated fetal bovine serum [HI-FBS; Corning Life Sciences, Pittson, PA] + 1% penicillin-streptomycin [Thermo Fisher] + 25 mM HEPES [Corning Life Sciences]) at 6.67 × 105 cells/mL and plated at 3 µL/well (2000 cells/well) in a 1536-well assay plate. Plates were then centrifuged at 800 rpm, 1 min, and incubated at 20 °C for 20–30 min or incubated at 37 °C in 5% CO2 overnight before initiating treatment with compounds and/or cytokines.
For validation experiments, anti-IL-12p40 (ustekinumab) or anti-IL-23p19 (guselkumab) monoclonal antibodies were sourced from Janssen Pharmaceuticals (Spring House, PA). Cytokines for development studies (IL-12, IL-1β, and TNFα) were sourced from R&D Systems (Minneapolis, MN) (cat. 219-IL/CF, 201-LB/CF, and 210-TA/CF, respectively). During development, several 1536-well plates sourced from Corning (Corning, NY) were tested, including nonbinding polystyrene (NB PS), tissue culture-treated polystyrene (TC PS), tissue culture-treated cyclic olefin copolymer (TC COC), and nontreated cyclic olefin copolymer (NT COC) (cat. 3729, 3727, 4571, and 9118-BC, respectively).
For high-throughput screening (HTS), compounds solubilized in 100% DMSO (Millipore, Burlington, MA) were acoustically dispensed at 10 nL into each well of the 1536-well NT COC assay plates using an Echo 555 (Labcyte, San Jose, CA), leading to a final DMSO concentration of 0.2%. Human IL-23 (53 kDa; eBioscience cat. 14-8239, San Diego, CA, or Janssen Pharmaceuticals) was added at 75 pM (EC80) in 2 µL/well to all test wells using a Tempest (Formulatrix, Bedford, MA). An IC100 control column was created using 6.7 nM anti-IL-23p19 (MW =150 kDa; Janssen Pharmaceuticals), and an EC0 control column was included with buffer and 0.2% DMSO. Assay plates were incubated at 37 °C for 4.5 h before adding 5 µL/well of Steady-Glo (Promega cat. E253B, Madison, WI) reagent with the Multidrop Combi (Thermo Fisher). Plates were incubated at 20 °C for 15 min and luminescence was read on a Pherastar FS (BMG LabTech, Cary, NC) for off-line validation studies (
(
(
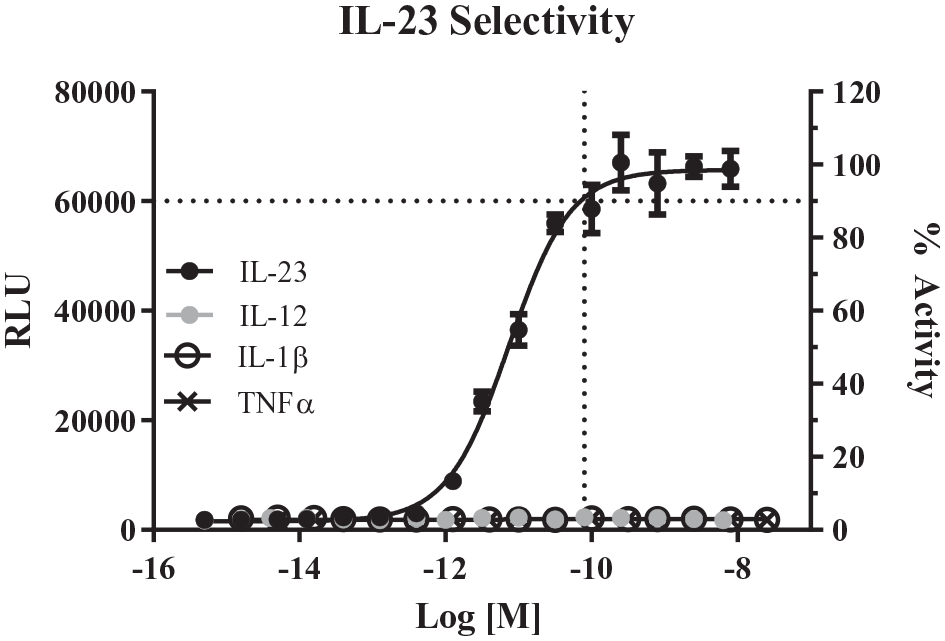
iLite IL-23 cells plated at 2000 cells/well respond to human IL-23 (EC50 = 6.5 pM) but not IL-12, IL-1β, and TNFα cytokines (n = 6 per dose) in a 1536-well plate. Horizontal line indicates the EC90 obtained with 75 pM IL-23, which was used for subsequent screening efforts. Error bars represent the standard deviation for n = 6, and data were fitted to a four-parameter logarithmic fit indicated by the solid black line.
(
(
(
CellTiter-Glo Assay
iLite IL-23 cells were plated as described above and incubated overnight at 37 °C. For HTS, compounds solubilized in 100% DMSO (Millipore, Burlington, MA) were acoustically dispensed at 10 nL into each well of the assay plates using an Echo 555 (Labcyte), leading to a final DMSO concentration of 0.2%. Human IL-23 (53 kDa; eBioscience cat. 14-8239, or Janssen Pharmaceuticals) was added at 75 pM (EC80) in 2 µL/well to all test wells using a Tempest (Formulatrix). Assay plates were incubated at 37 °C for 24 h before adding 5 µL/well of CellTiter-Glo (CTG; Promega cat. G7573) reagent with the Multidrop Combi (Thermo Fisher). Plates were incubated at 20 °C for 15 min, and luminescence was read on a Pherastar FS (BMG LabTech) for off-line validation studies and on a Pherastar FSX on the robotic platform (
IFN-γ Assay
iLite IFN-γ cells (Svar Life Science, Malmö, Sweden) were plated in the same manner as the iLite IL-23 reporter assay but with 4000 cells/well and incubated overnight at 37 °C. For HTS, compounds solubilized in 100% DMSO (Millipore, Burlington MA) were acoustically dispensed at 10 nL into each well of the assay plates using Echo 555 (Labcyte), leading to a final DMSO concentration of 0.2%. Human IFN-γ (17 kDa; R&D Systems, Minneapolis, MN) was added at 24 pM final concentration with a Tempest (Formulatrix) and 10 nL of DMSO solubilized compound was added with an Echo 555 (Labcyte) acoustic dispense. Assay plates were incubated for 4.5 h at 37 °C before adding 5 µL/well of Steady-Glo (Promega cat. E253B) with the Multidrop Combi (Thermo Fisher). Plates were incubated at 20 °C for 15 min, and luminescence was read on a Pherastar FS (BMG LabTech) for off-line validation studies and on a Pherastar FSX on the robotic platform ( Fig. 5B–D ). The automated assay protocol was executed on a HighRes system, driven by the Cellario software. Data were normalized to percent inhibition, where 0% inhibition was defined by control wells treated with 24 pM IFN-γ and 100% inhibition was defined by control wells treated with 24 pM IFN-γ + 1 µM of an internal JAK inhibitor.
Detection of Phosphorylated STAT5 Using TR-FRET
iLite IL-23 cells were thawed as above but resuspended to 2.5 × 106 cells/mL in growth medium. This suspension was dispensed at 8 µL/well into a 384-well plate (Greiner cat. 781080, Greiner-BioOne, Monroe, NC). IL-23 diluted in growth medium with either DMSO (1%) or tofacitinib (Sigma, St. Louis, MO; 3 µM final) was then dispensed at 4 µL/well to cells and incubated at 37 °C for 90 min. Next, 4× lysis buffer was prepared per kit instructions (cat. 64AT5PEG, Cisbio, Codolet, France), added to the plate at 4 µL/well, and shaken for 10 min at 20 °C. Lastly, 3.5 µL/well detection buffer was added to the plate, shaken for 10 min, and incubated overnight at 20 °C prior to reading the TR-FRET signal on the Pherastar FS (BMG Labtech).
Data Analysis
Figure 5 and Supplement Figure S3 were created with Spotfire version 10. All remaining plots, including dose–response curves, were generated with GraphPad Prism version 7 (La Jolla, CA).
Results and Discussion
Assay Validation
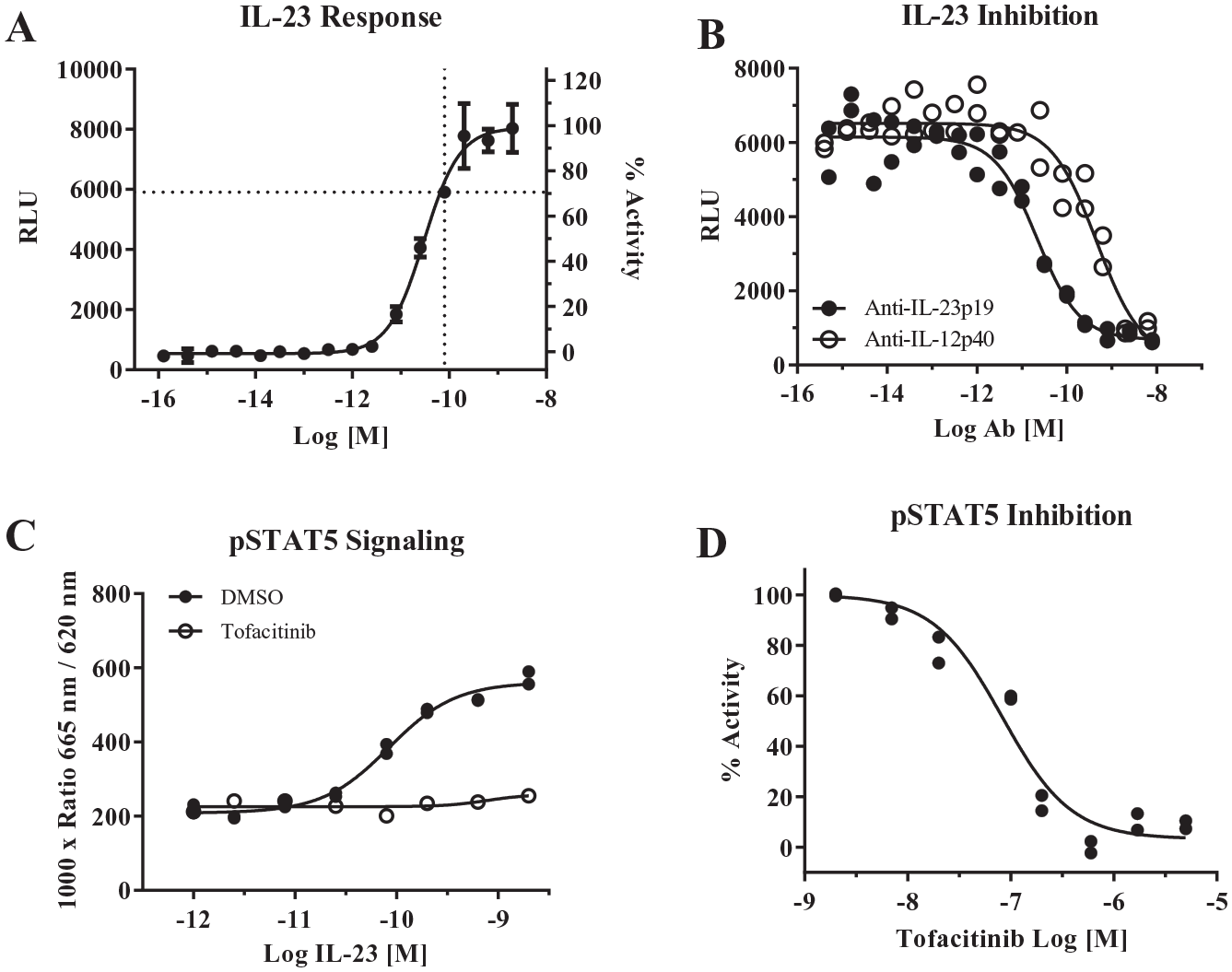
Initial experiments were performed to determine the responsiveness of IL-23 iLite cells to IL-23. Although IL-23 iLite cells were engineered to express Renilla luciferase under a constitutively active promotor that could be used to normalize IL-23-induced firefly activity, 15 we observed that the use of Dual-Glo reagent to ratiometrically measure both firefly and Renilla luciferase resulted in more variable data than measuring firefly luciferase activity alone. All subsequent assay development utilized Steady-Glo reagent, measuring only firefly luciferase activity. First, a concentration-dependent increase in luciferase activity was observed with a 5 h incubation of IL-23 iLite cells with IL-23 (EC50 = 0.028 nM) ( Fig. 1A ). Next, using an approximate EC80 concentration of IL-23 (0.075 nM) to activate the cells, antibodies specific to either the p19 or p40 subunits of IL-23 inhibited the luciferase activity in the IL-23 iLite cells ( Fig. 1B ; anti-p19 IC50 = 0.022 nM; anti-p40 IC50 = 0.46 nM), indicating that induction of the luciferase response is dependent on binding to both IL-23R and IL12Rβ1, respectively. Although IL-23 signaling is driven primarily by pSTAT3, IL-23 iLite cells were engineered to overexpress pSTAT5 to generate a robust response. 15 IL-23 treatment also resulted in STAT5 phosphorylation in a concentration-dependent manner (EC50 = 0.079 nM) that was blocked by 3 µM of the JAK inhibitor tofacitinib ( Fig. 1C ), verifying that the JAK/STAT pathway is engaged in this cellular model upstream of luciferase activity. A titration of tofacitinib against 0.075 nM IL-23 resulted in an IC50 of 83 nM in the pSTAT5 assay, consistent with published potencies in primary cell assays 16 ( Fig. 1D ). Together, these data indicated that the IL-23 iLite cell line was sufficiently sensitive and specific to IL-23 to continue additional assay development.
Assay Optimization
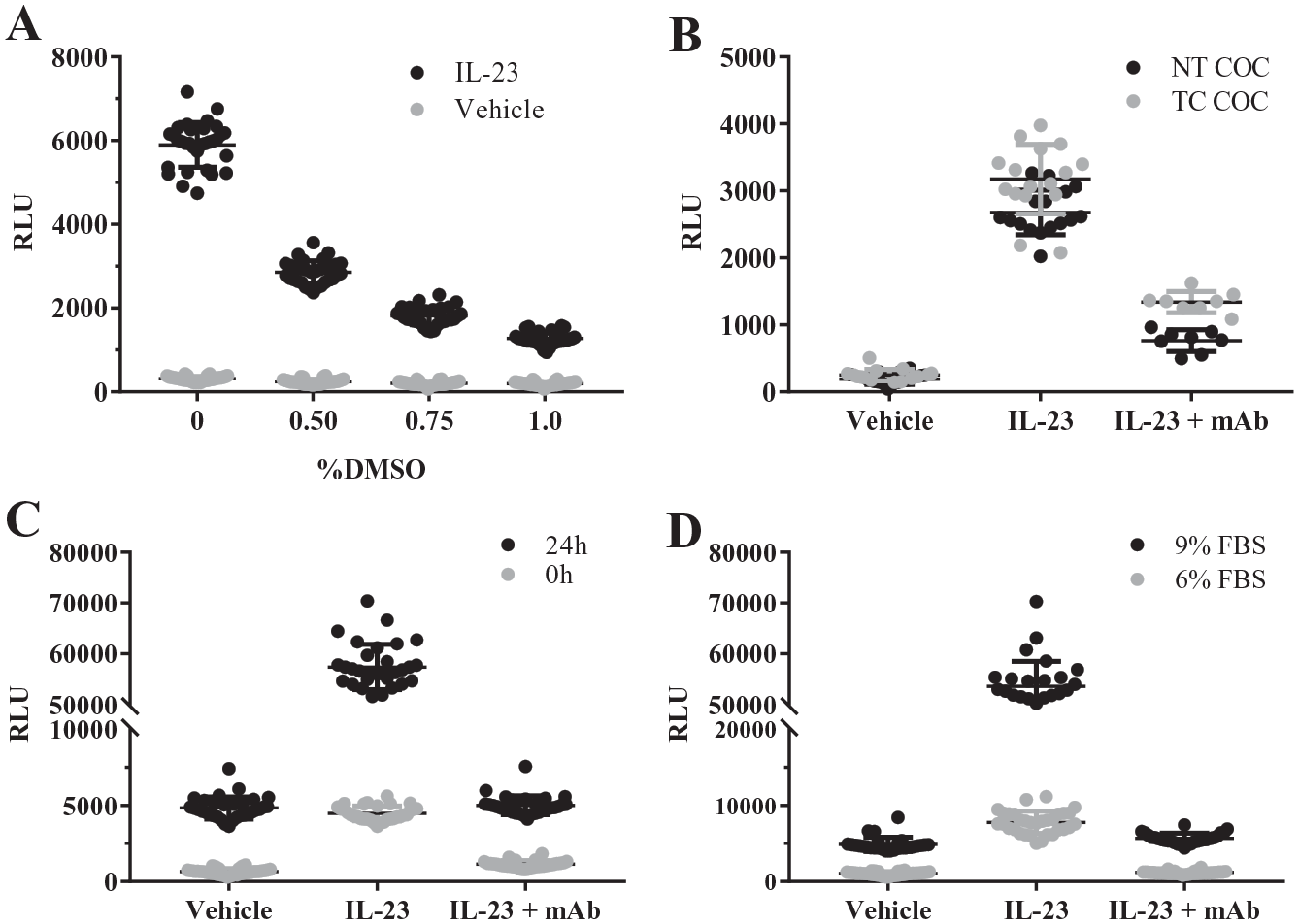
To enable a 1536-well, small-molecule screen for inhibitors of IL-23 signaling, several parameters of IL-23 iLite cell culture were examined in more detail, including DMSO tolerance, plate coating, cell incubation time, and serum content ( Fig. 2 , Suppl. Table S1 ). Incubation of IL-23-stimulated cells with 0.5% DMSO for 4–6 h resulted in a 2.1-fold reduction of signal compared with 0% DMSO. This DMSO effect was concentration dependent, as higher DMSO concentrations lead to a more severe loss of signal ( Fig. 2A ). This observation indicated a low DMSO tolerance for iLite cells, and further assay conditions limited DMSO exposure to 0.25%. In a parallel experiment to examine cell viability, DMSO treatment of up to 0.9% had no effect on cell viability ( Suppl. Fig. S1 ), suggesting that DMSO was not toxic to the iLite cells. For robust miniaturization, plate-type selection can be critical to successful assay development. Both PS and COC plates containing TC or NT treatments were compared for assay performance. Although PS is commonly used for miniaturized assays, COC is a rigid plastic resistant to potential warping by the robotic grippers, making it more attractive for automation. 17 Comparison of TC- and NT-treated assay plates revealed similar signal-to-background (S/B) for IL-23-induced response (S/B = 13 and 14, respectively; Suppl. Table S1 ); however, the TC treatment reduced the effect of 1.3 nM of anti-IL-23p19 antibody from 79% to 67% ( Fig. 2B ). The reduction in efficacy of the anti-IL-23p19 antibody may be due to interaction with the TC treatment; therefore, NT plates were utilized in further experiments. Following evaluation of these 1536-well plate types with IL-23 iLite cells tested at a range of densities, a condition utilizing 2000 cells/well in 1536-well black, NT, COC plates was selected for further assay optimization based on the strong signal window (S/B = 12) and Z′ (0.6) ( Suppl. Table S1 ). Despite the sensitivity of the luciferase response to DMSO, significant signal improvement was made when cells were cultured for 24 h prior to IL-23 treatment, increasing the S/B from 3.9 to 11.5 ( Fig. 2C , Suppl. Table S1 ). As serum may be a source of nonspecific binding for small-molecule compounds, 18 the effect of serum percentage on assay signal was examined ( Fig. 2D ). Lowering FBS from the vendor-recommended 9% to 6% FBS in the presence of 75 pM IL-23 led to significantly diminished luminescence and poor Z′ (<0.1), indicating that reduced serum levels cannot be tolerated. A cell preincubation time of 24 h made the most significant improvement in the assay. Under these optimized conditions, we verified that the IL-23 cells remained specific for IL-23 relative to other cytokines ( Fig. 3 ).
Pilot Screen Results
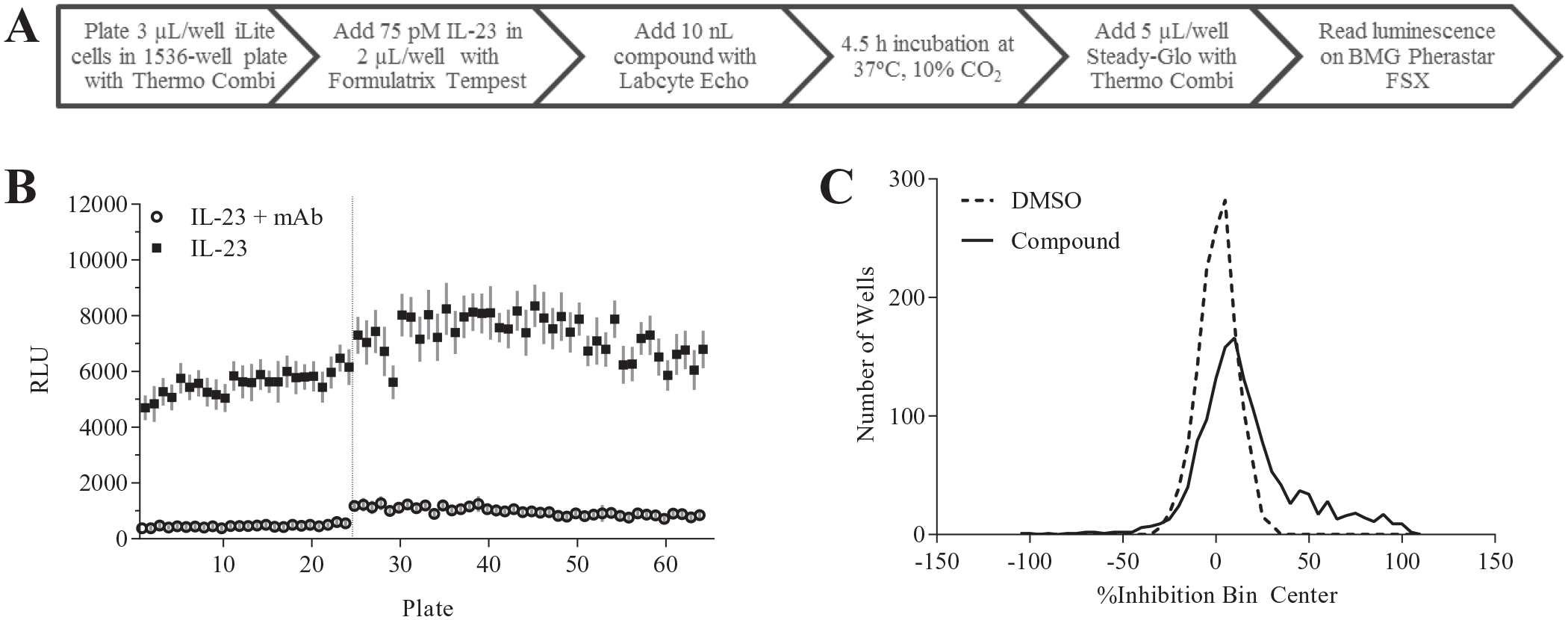
With the assay optimization steps outlined above, the final automated protocol was established for initiation of the pilot screening (
Fig. 4A
). Targeting a manageable hit rate of approximately 1%, a small set of 1408 compounds from the Janssen internal screening collection were tested at 2.5, 5, and 10 µM to determine the hit rate at each concentration. Using a mean (µ) + 3 times standard deviation (σ) cutoff, a 15% hit rate (>47% inhibition) was observed at the 10 µM dose and a 10% hit rate (>32% inhibition) was observed at the 5 µM dose. Even the lowest dose of 2.5 µM with a cutoff of 26% inhibition gave a 4% hit rate. The underlying cause of the relatively high hit rate was not readily apparent. Therefore, instead of launching a full-library screen, a set of approximately 24,000 compounds representing a diversity of chemotypes within our library were tested at 10 μM in the iLite IL-23 assay run across two separate days (
Fig. 4B
,
Suppl. Fig. S2
). The S/B and Z′ values across the 56 plates were 10 ± 3 and 0.6 ± 0.1, respectively. Interspersed throughout the screening days, we also tested plates treated only with DMSO and controls. These DMSO plates had very low percentage coefficients of variation (10 ± 1%, n = 7 plates) with the DMSO wells exhibiting a symmetrical distribution around 0% inhibition (
Fig. 4C
), indicating that the assay itself did not have inherently high variability. Furthermore, intra-assay reproducibility was high, as indicated by a 60% confirmation rate in all three replicates and low standard deviation across replicates above the hit cutoff (
Suppl. Fig. S3A,B
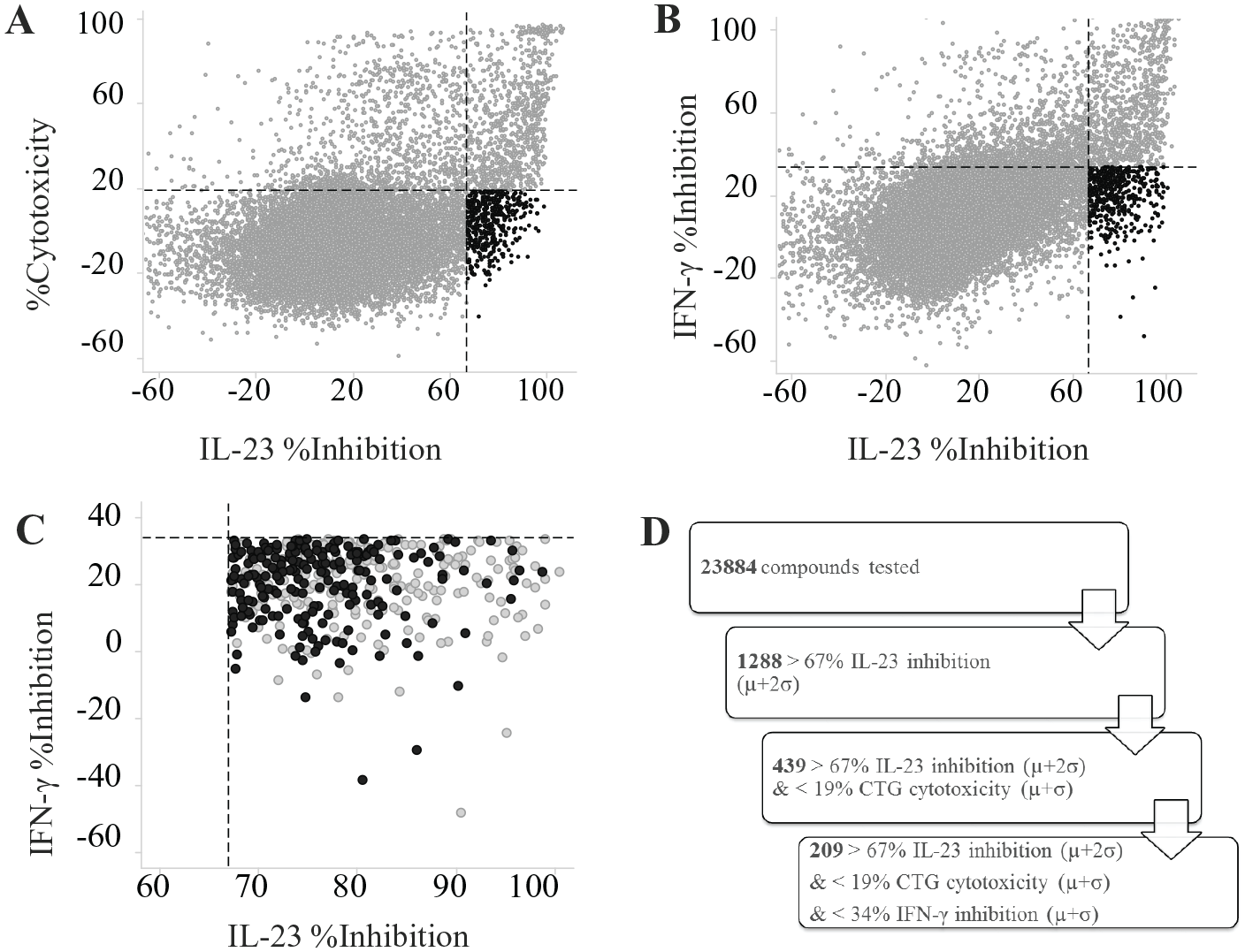
). Taken together, these data suggest that the assay is sensitive to a wide array of small molecules; however, the reduction in IL-23 response in the assay may not be due to direct inhibition of IL-23 binding or IL-23-specific signaling. Therefore, we examined all 24,000 compounds for two potential sources of off-target inhibition, namely toxicity and nonspecific inhibition of downstream signaling. The CTG counter assay (S/B = 42 ± 5, Z′ = 0.6 ± 0.2, n = 61) examined direct compound toxicity to the IL-23 iLite cells with an overnight compound treatment. An iLite IFN-γ counter assay (S/B = 6 ± 1, Z′ = 0.4 ± 0.1, n = 58) with the same reporter system as the iLite IL-23 cell assay was also run to identify nonspecific inhibitors of both the JAK/STAT pathway and the luciferase reporter. A high percentage of the 1288 compounds that were active in the iLite IL-23 assay (>67% inhibition) were active in each of the counter screens (66% in CTG and 67% in iLite IFN-γ) (
The above data illustrate the strengths and weaknesses of cell-based assay development and miniaturization even when starting with strong assay statistics (e.g., S/B and Z′; Fig. 4 , Suppl. Fig. S2 ) and selectivity for the target(s) of interest ( Fig. 3 ). In the case of the iLite IL-23 cell-based assay, one of the surprising challenges was the sensitivity to DMSO. Although this observation did not prevent further success in miniaturizing the assay or in testing compounds per se, the high sensitivity of IL-23 signaling in these cells to DMSO stood out relative to more “standard” screening cell lines such as HEK or CHO cells. 19 It is worth noting that DT40 cells have been used extensively to study environmental toxins, 20 often employing DMSO as a vehicle present up to 1% in assay solutions 21 and using similar readouts such as CTG, 22 suggesting that DMSO sensitivity is not necessarily a feature of DT40 cells themselves. However, the significant degree of overlap between iLite IL-23 signal inhibition and inhibition in the CTG assay, resulting in removal of ~2/3 of the original hits ( Fig. 5D ), indicates a level of sensitivity to small-molecule toxicity that may be unusual. In the present study, the enhanced sensitivity to DMSO did not contribute directly to effects on cell viability ( Suppl. Fig. S1 ); however, whether this sensitivity could synergize with compounds that have a propensity toward cellular toxicity leading to a relatively high hit rate in the CTG assay would require further exploration. The additional removal of hits due to off-target inhibition of IFN-γ is less surprising, particularly given the potential overlap in downstream kinases. Even with these challenges, compounds with selective inhibition of IL-23 signaling were identified and subsequently prioritized for further evaluation in additional biochemical and cell-based assays, such as IL-23-treated purified NK or NKT cell preparations,23,24 which is beyond the scope of the present study.
Supplemental Material
Supplemental_Material_for_Optimization_of_HTS_for_IL-23_by_Varghese,_et_al_revise_SLAS_1 – Supplemental material for Optimization of a High-Throughput Cell-Based Screening Strategy to Identify Small-Molecule Inhibitors of IL-23 Signaling
Supplemental material, Supplemental_Material_for_Optimization_of_HTS_for_IL-23_by_Varghese,_et_al_revise_SLAS_1 for Optimization of a High-Throughput Cell-Based Screening Strategy to Identify Small-Molecule Inhibitors of IL-23 Signaling by Teena M. Varghese, Paul L. Dudas, Samantha J. Allen, Jonathan E. Schneeweis and Michael F. A. Finley in SLAS Discovery
Footnotes
Acknowledgements
We thank Glenda Castro for technical assistance.
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: All authors were employees of Janssen Pharmaceutical R&D at the time of submission.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
