Abstract
Protein acylation, exemplified by lysine acetylation, is a type of indispensable and widespread protein posttranslational modification in eukaryotes. Functional annotation of various lysine acetyltransferases (KATs) is critical to understanding their regulatory roles in abundant biological processes. Traditional radiometric and immunosorbent assays have found broad use in KAT study but have intrinsic limitations. Designing acyl–coenzyme A (CoA) reporter molecules bearing chemoselective chemical warhead groups as surrogates of the native cofactor acetyl-CoA for bioorthogonal labeling of KAT substrates has come into a technical innovation in recent years. This chemical biology platform equips molecular biologists with empowering tools in acyltransferase activity detection and substrate profiling. In the bioorthogonal labeling, protein substrates are first enzymatically modified with a functionalized acyl group. Subsequently, the chemical warhead on the acyl chain conjugates with either an imaging chromophore or an affinity handle or any other appropriate probes through an orthogonal chemical ligation. This bioorganic strategy reformats the chemically inert acetylation and acylation marks into a chemically maneuverable functionality and generates measurable signals without recourse to radioisotopes or antibodies. It offers ample opportunities for facile sensitive detection of KAT activity with temporal and spatial resolutions as well as allows for chemoproteomic profiling of protein acetylation pertaining to specific KATs of interest on the global scale. We reviewed here the past and current advances in bioorthogonal protein acylations and highlighted their wide-spectrum applications. We also discussed the design of other related acyl-CoA and CoA-based chemical probes and their deployment in illuminating protein acetylation and acylation biology.
Introduction to Protein Acetylation
Lysine acetylation of proteins is an essential posttranslational modification (PTM) that regulates plethoric biological processes, including gene transcription, the cell cycle, apoptosis, metabolism, signal transduction, and cell differentiation.1–8 Although nonenzymatic processes are present, the site-specific acetylation reaction is dominantly catalyzed by protein lysine acetyltransferases (KATs) that transfer the acetyl group from acetyl–coenzyme A (acetyl-CoA, Ac-CoA) to the ε-amino group of specific lysine residues in proteins ( Fig. 1A ). A dozen KAT members in mammalian cells have been identified and characterized both genetically and biochemically, which include the GNAT representative members GCN5, PCAF, and HAT1; five MYST family members (MOF, TIP60, MORF, MOZ, HBO1); p300 and CBP.9–16 A few other proteins, such as CLOCK, NAT10, and NCOAT, though mentioned as KAT members, remain poorly characterized and are sometimes considered as orphan or noncanonical KAT members.16,17 Importantly, many KATs were found to associate with cancer incidence, progression, and metastasis.15,18,19 Some KATs were found to efficiently catalyze other short-chain fatty acylation as well, especially protein propionylation and butyrylation.20–23 On the other end of the acylation spectrum, long-chain fatty acylation has also gained increasing interest in recent years, of which the most extensively studied modifications are N-myristoylation on the N-terminal glycine residues catalyzed by N-myristoyltransferases (NMTs) and S-palmitoylation on the cysteine residues via a thioester bond catalyzed by palmitoyltransferases (PATs).24–26 Side chain myristoylation of lysine residues is also identified, but no corresponding acyltransferase has been reported so far.
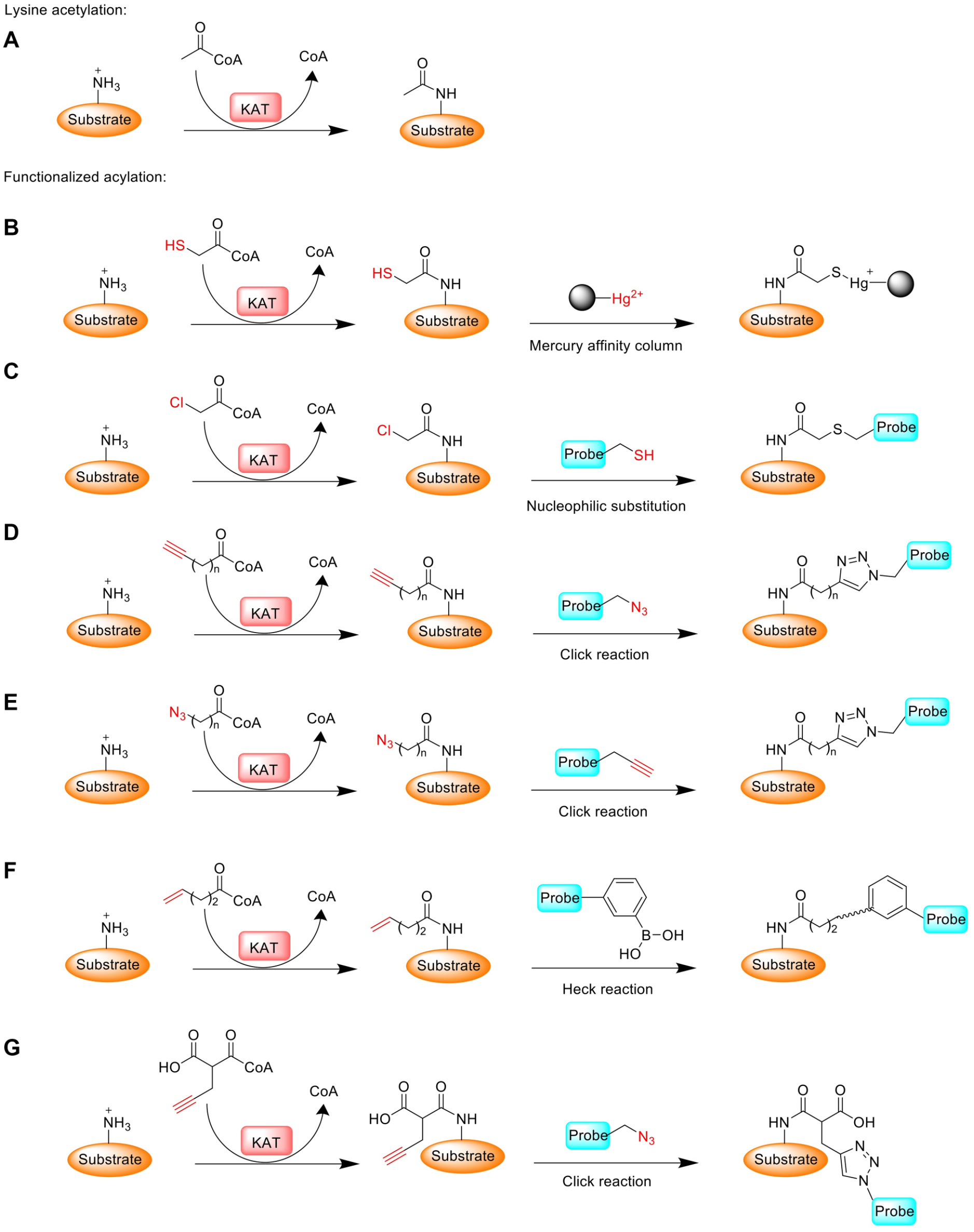
Schematic of protein lysine acetylation and its reformulated bioorthogonal acylation using functionalized acyl-CoA surrogates. (
Proteomic studies have revealed the presence of hundreds to thousands of acetylated proteins and acetylation sites throughout the cell, which suggests the prevalence of lysine acetylation in nearly every facet of cell physiology.27–32 Since KAT enzymes are key regulators for cancer biology, high-throughput assays have been designed to screen lead inhibitors for specific KATs with the goal of developing new anticancer drugs.16,33–35 While the importance of KATs in physiology and disease is widely recognized, the functional annotation of KAT enzymes in regulating key biological pathways is far from completely understood. In particular, how the acetylomes of individual KAT enzymes are distinguished from one another and how the substrate distribution of KATs is affected by various intracellular or environmental stimuli demand further investigation. A clear biochemical, structural, and proteomic understanding of KAT substrate specificity and the impact of individual KATs in (patho)physiology regulation is greatly needed. Elucidation of molecular targets of KATs represents a leading step toward fully dissecting the roles of KATs in gene regulation and their functions beyond the chromatin biology realm. In this regard, mass spectrometry (MS)-based profiling has provided a plethora of information about acetylated proteins,5,36,37 albeit with limited information on enzyme–substrate correlations. Protein microarray is also an effective method in KAT substrate identification on a large scale. 38 Using yeast proteome microarrays, Lin et al. identified many nonchromatin substrates of the nucleosome acetyltransferase of H4 (NuA4) complex. 39
A technical challenge in the biochemical study of lysine acetylation is that the transferred acetyl group is chemically inert, which renders the direct chemical detection of KAT-medicated protein acetylation practically difficult. Standard methods of acetylation detection typically rely on radioisotope-labeled acetyl-CoA or antibody recognition of acetylated lysine residues.16,35,40,41 The application of high-throughput MS techniques for the label-free detection and quantitation of lysine acetylation reaction products is also used to characterize KAT enzyme activities and inhibitors. 42 All the methods have satisfying sensitivity and reliability in acetylation detection, but they suffer from such issues as lacking substrate specificity, loss of tempospatial information, operational discontinuity, and high costs. In order to gain amenable signals for detection and capture, a number of research efforts have been made to explore acetyl or fatty acyl derivatives that contain small-size bioorthogonal functional reporter groups to make acetyl-CoA and acyl-CoA surrogates that can be used by KATs for substrate labeling.16,40,43–45 These bioorthogonal approaches provide a new dimension of technical accessibility to explore the landscape of lysine acetylation and acylation. The evident benefit is that the chemical reporter moiety on fatty acyl groups can be utilized for conjugation with fluorescent probes or affinity tags via a chemoselective chemical ligation to detect acylated proteins ( Fig. 1 ). For in vitro studies, functionalized short-chain fatty acyl-CoAs are directly used to replace acetyl-CoA for substrate labeling. For cellular studies, functionalized short-chain fatty acids (SCFAs) are typically used to circumvent the cell permeability issue, which is based on the principle that inside cells SCFAs are converted by endogenous acyl-CoA synthetases into functionalized fatty acyl-CoAs that are used by respective KATs to label their acetylated substrates. The major focus of the following sections is to review and discuss the development of various acyl-CoA reporters and their fatty acyl precursors for use in bioorthogonal probing of protein acetylation and acylation. We also discuss other related CoA-based chemical probes and their employment in acetyltransferase mapping, inhibition, and drug discovery.
Thiol-Containing Acetate Analog for KAT Substrate Labeling
With 2-mercaptoacetate as an acetate surrogate, Sterner and Allfrey first reported a cell-permeable chemical reporter for monitoring postsynthetic acetylation to target acetylated proteins in duck erythrocytes. 46 They proposed that the incorporated acetate analog would be metabolically converted to mercaptoacetyl-CoA that subsequently underwent histone mercaptoacetylation ( Table 1 ). The study indeed showed that the incorporated sulfhydryl groups can be utilized by endogenous KATs to be introduced into chromosomal proteins such as histones and HMG proteins. Furthermore, mercaptoacetylated proteins were selectively recovered from the biological mixture by mercury-affinity column chromatographic techniques and then subjected to MS analysis ( Fig. 1B ). Where this method falls short is that it is limited for the labeling and isolation of KAT substrates lacking sulfhydryl amino acids. For most proteins, the sulfhydryl group present on the side chain of cysteine residues will interfere with and complicate labeling efficiency and specificity.
Acetyl-CoA Surrogates Combined with Wild-Type or Mutant KATs to Label Their Substrates.
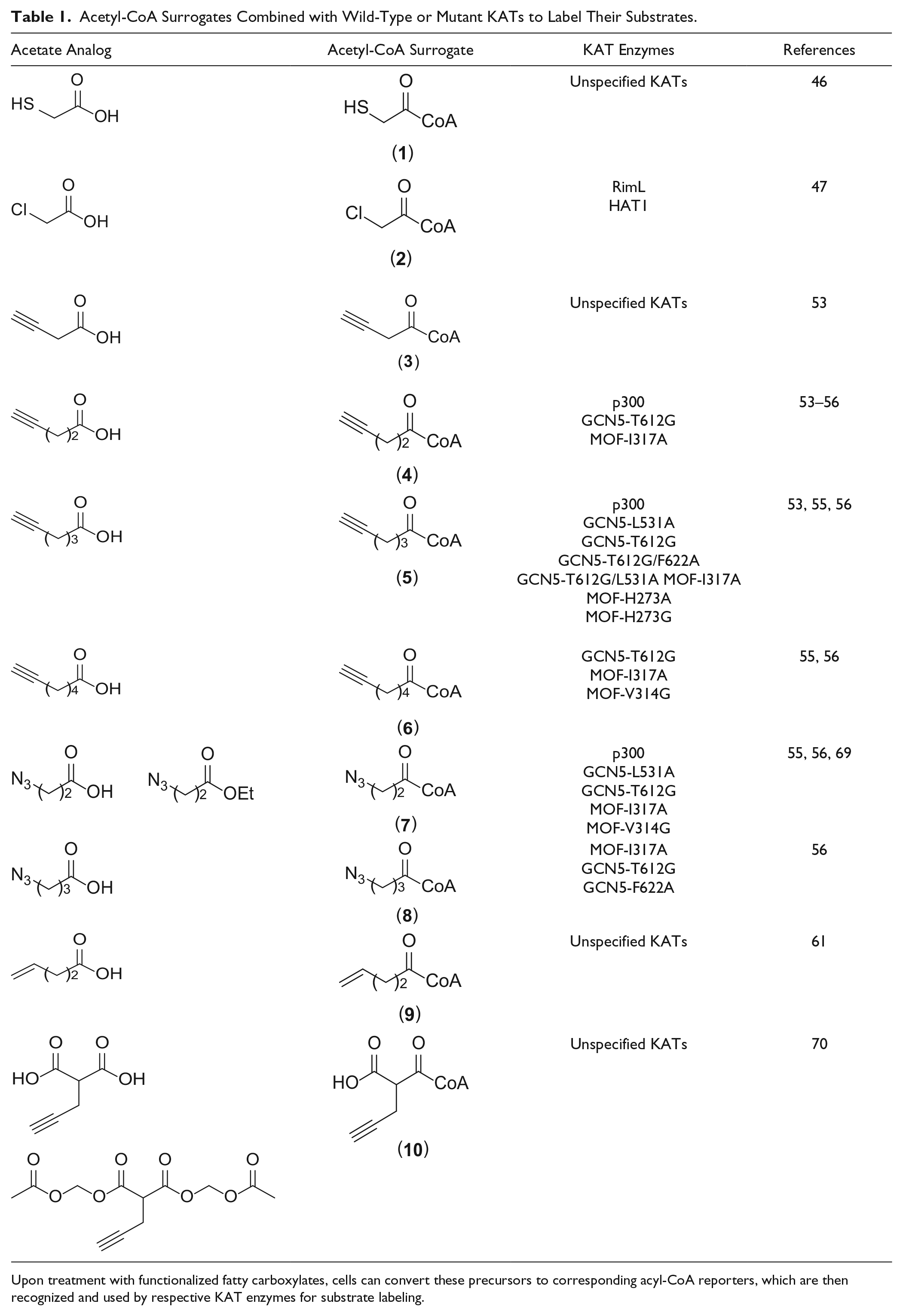
Upon treatment with functionalized fatty carboxylates, cells can convert these precursors to corresponding acyl-CoA reporters, which are then recognized and used by respective KAT enzymes for substrate labeling.
Halide-Labeled Acetyl-CoA for KAT Substrate Labeling
The use of chloroacetyl–coenzyme A (ClAcCoA) ( Table 1 ) to identify protein substrates for some members of the GNAT superfamily was reported by Blanchard’s group. 47 One such GNAT member is histone acetyltransferase 1 (HAT1), the first identified HAT enzyme. 48 Using ClAcCoA as an acetyl-CoA surrogate in junction with sulfydryl-functionalized fluorophores, Blanchard et al. developed this orthogonal method to label yeast HAT1 substrates ( Fig. 1C ). 47 Histone H4, a known substrate of HAT1, can be chloroacetylated by HAT1 rapidly and selectively in vitro. Moreover, RimL, which is an acetyltransferase in E. coli, also showed a good activity with the reporter. The chloroacetylated products are stable enough with a slow rate of hydrolyzation, whereas they are reactive enough to be conjugated with thiol nucleophiles such as TAMRA-cysteamine for fluorescent visualization. Potentially, thiol-containing reagents can be designed with affinity handles for liquid chromatography (LC)-MS/MS analysis to make the proteome-wide identification of substrates for the GNAT members. A unique property of ClAcCoA is that, during KAT catalysis, the chloroacetyl group transferred to the substrate can react with the other product, CoASH, to generate an CoA-acetyl–substrate conjugate. When [3′- 32 P]-ClAcCoA is used, the product of the reaction becomes radiolabeled, and for protein substrates, it can be observed after sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) and autoradiography. 47
Alkyne or Azide-Functionalized Acyl-CoA Reporters for KAT Substrate Labeling
Biomolecular labeling with azide or alkyne-functionalized chemical reporters and subsequent bioorthogonal ligation have gained prominent recognition in the chemical biology field.45,49 The incorporated alkyne or azide reporter group serves effectively as a chemical warhead to selectively react with azide/alkyne-containing probes via copper-catalyzed azide−alkyne cycloaddition (CuAAC) chemistry.50–52 Owing to the much stronger chemoselectivity and bioorthogonality of CuAAC reaction compared with thiol-halide nucleophilic substitution reactions, the utilization of clickable reporter groups for biomolecular labeling is a prime choice in bioorganic chemistry. The design of clickable acyl-CoA surrogates bearing an alkyne functional group ( Table 1 ) to identify KAT substrates was first explored by Hang’s group to provide a chemoproteomic strategy to profile the acetylome of KAT members p300/CBP.53,54 p300 showed a good activity toward 4-pentynoyl CoA (4PY-CoA) ( Table 1 ) as the cofactor for substrate labeling, which suggests that 4PY-CoA serves as a bioorthogonal chemical reporter to study p300 activity. 54 In kinetic characterization, 4PY-CoA exhibited a 10-fold lower kcat/KM value compared with acetyl-CoA. 55 Consistent with the data reported by Hang’s group, we found that p300 has great cofactor promiscuity toward the alkyne-functionalized acetyl-CoA analogs 4PY-CoA, 5-hexynoyl CoA (5HY-CoA), and 6-heptynoyl CoA (6HY-CoA) ( Fig. 1D ). 56 Additionally, we found that the azido-containing analog 3-azidopropanoyl CoA (3AZ-CoA) ( Table 1 ) is strongly recognized by the p300 HAT domain with comparable or stronger activity than Ac-CoA based on kinetic characterization. Also, there is around an eightfold higher specificity for 3AZ-CoA toward p300 than that of 4PY-CoA, suggesting that 3AZ-CoA is another excellent acetyl-CoA surrogate to identify protein targets of the p300/CPB KATs ( Fig. 1E ). 55 Importantly, the labeling activities of 3AZ-CoA for the other wild-type KATs, such as MOF, Tip60, MOZ, MORF, GCN5, and PCAF, were quite weak, which demonstrates the unique pairing property between 3AZ-CoA and p300.55,56
Although several short-chain clickable acyl-CoAs are active for wild-type p300/CBP to label its substrates, most KATs cannot efficiently take clickable acyl-CoAs for substrate labeling. The major obstacle is that the active pocket of most KATs is too small to accommodate the acyl head group of acyl-CoA. To overcome this problem, we performed experiments to engineer the active site of the KATs GCN5 and MOF to expand their cofactor binding capability to accommodate the bulkier synthetic cofactors. 56 As such, the engineered KATs will be able to be used in junction with synthetic acetyl-CoA surrogates to establish bioorthogonal probes to investigate cellular substrates of these KAT enzymes. In our study, we tested five functionalized acetyl-CoA analogs containing alkynyl functional groups, including 4PY-CoA, 5HY-CoA, 6HY-CoA, and two azide-functionalized acyl-CoA molecules, 3AZ-CoA and 4-azidobutanoyl CoA (4AZ-CoA) ( Table 1 ). The installed alkynyl and azido functional group is expected to facilitate downstream detection and characterization of labeled KAT substrates.
For GCN5 KAT, we analyzed its active site structure and identified several conserved bulky residues surrounding the acetyl warhead group of acetyl-CoA: L531, M534, I576, F578, T612, F622, and Y645. 56 To expand the cofactor binding pocket for it to accommodate bigger size acyl groups, we replaced each of these residues with smaller ones, that is, alanine or glycine. To identify the engineered enzyme forms that are active to the Ac-CoA substitutes, the entire panel of acetyl-CoA analogs was screened in histone modification reactions catalyzed by both the wild-type and engineered GCN5 proteins. As expected, wt-GCN5 exhibited a strong activity toward acetyl-CoA, the cognate cofactor of KATs, but was inert toward all the acetyl-CoA substitutes. On the other hand, several engineered GCN5 proteins, for example, GCN5-L531A/G, -I576A, -T612A/G, and -F622A, exhibited appreciable activities to the synthetic analogs at varied degrees. In particular, the single mutant GCN5-T612G was active toward all the tested cofactors, with 4AZ-CoA being the weakest. 55 Further, GCN5-T612G/F622A and -T612G/L531A mutants exhibited excellent activity toward 5HY-CoA.
We also engineered the active site of the MYST member MOF. 56 The crystal structure of the MOF–Acetyl-CoA complex 57 shows that the acetyl moiety of the cofactor is surrounded by several bulky residues: V314, I317, I333, P349, P352, and L353. H273 is another potential residue that may affect the enzyme activity.58–60 We carried out mutation of each residue to Ala or Gly to expand the enzyme active site for acyl-CoA binding. Activities of each MOF mutant toward the synthetic acetyl-CoA analogs were screened through enzymatic modification of the histone H4 tail peptide. Among the tested MOF mutants, H273A and H273G recognized 5HY-CoA, and V314G recognized 6HY-CoA and 3AZ-CoA. Strikingly, MOF-I317A was active toward all the acetyl-CoA substitutes. The double mutant MOF-I317A/H273A retained a reasonably good activity to 5HY-CoA. More details of the work can be found in Yang et al. 56
Except alkyne/azide-functionalized acyl-CoA as chemical reporters for probing protein acylation, Dekker and coworkers introduced sodium 4-pentenoate as a bioorthogonal chemical reporter to label endogenous acylated proteins followed by Heck coupling. 61 In their studies, RAW 264.7 cells were incubated with sodium 4-pentenoate to form intracellular 4-pentenoyl-CoA ( Table 1 ) to establish 4-pentenoyl-labeled proteins. The labeled histones were then coupled with fluorescein or biotin-labeled phenylboronic acid via oxidative Heck reaction using EDTA-PdII as a catalyst ( Fig. 1F ). 62 Fluorescence-based imaging showed that the 4-pentenoyl labeling of histones was dose dependent and a strong fluorescent signal was seen for histone H3. Compared with alkynic reporters, the authors suggested that the lack of histone H4 labeling was due to the poor cell permeability of the alkenic reporter. This study provides an alternative orthogonal reporter method for metabolic labeling of KAT substrates.
Application of Bioorthogonal Acyl-CoA Reporters to Detect and Profile KAT Substrates
Bioorthogonal acyl-CoA reporters provide unprecedented opportunities to establish a powerful chemical biology protocol to detect and study protein lysine acetylation in ways that cannot be accomplished by classic molecular biology methods. The added chemoselective functionality on the acyl group in substrates can be utilized to achieve various applications. First, fluorescent and other optical probes can be chemoselectively linked to the clickable reporter group to image acylated products and their subcellular location without recourse to the reliance on an antiacetyl antibody or radioisotope. In principle, the labeling and imaging can be performed in live cells, which will provide spatially resolved images of KAT substrate distribution in different organelles. Second, the acylated substrates can be further labeled with affinity handles for their pulldown enrichment from complex biological mixtures. This methodology coupling with modern MS/MS allows for proteomic profiling of KAT substrates. Since the chemical-modifying group is drastically different from the native acetyl group, it is technically more facile to identify acetylated substrates caused by overexpression or knockdown of a particular KAT of study. Third, the chemoreactive group in acetylated substrates can potentially be linked to any other biomolecules of interest that bear complementary clickable moieties to investigate KAT-mediated protein–protein interactions and other biological processes. Last but not least, the currently available high-throughput assays for KAT drug discovery are mostly based on antibody-based detection of the acetyl product or spectroscopic detection of the side product CoA.16,35 Bioorthogonal linking of acylated KAT substrates with fluorescent or bioluminescent probes could generate new protocols for KAT drug screening.63,64
The methodology of using bioorthogonal acyl-CoA reporters for KAT substrate identification and profiling typically involves two steps: the first step is protein substrate labeling by acyl reporters that are enzymatically acted on by respective KATs; then the labeled targets are detected with an imaging or affinity probe via a chemoselective bioorthogonal chemical conjugation. 65 In practice, the reporter labeling can be carried out in two different protocols that depend on the biological contexts to be studied. In one protocol, functionalized acyl-CoA and respective recombinant KATs are used directly to install acyl reporters onto proteins in purified forms or complex cell lysates. This is technically desirable because there is no requirement of plasma membrane penetration. The limitation, however, is that KATs are not present in their native forms and cell lysate proteomes do not retain their intact cellular milieu. Therefore, there is a higher chance of introducing artificial interference. The other protocol is through metabolic labeling in living cells. Acyl-CoA is widely believed to have no or little membrane permeability; therefore, it cannot be directly applied to interrogate protein acetylation and acylation in cells. To circumvent the membrane passage issue, functionalized SCFAs are used as precursors of respective acyl-CoA and added in tissue culture media. A number of studies have shown that functionalized SCFAs can cross the plasma membrane to enter cells. Once inside the cells, these molecules are converted to respective acyl-CoA derivatives by the cell’s own biosynthetic machinery. Acyl-CoA is eventually used as a cofactor by KATs to acylate their substrates.
The KAT-mediated protein labeling followed by site-selective chemical conjugation provides ample opportunities for detection, visualization, and further characterization of KAT targets. Most commonly, labeled proteins are either visualized by coupling to fluorescent dyes such as fluorescein or tetramethylrhodamine (TAMRA) for in-gel fluorescence imaging, or isolated and enriched by specific affinity pulldown experiments ( Fig. 2 ). 66 MS/MS proteomic profiling of the bioorthogonally labeled KAT substrates is of great significance to provide insights into the function of KATs. To facilitate affinity enrichment of labeled protein substrates for proteomic study, Hang’s group developed highly efficient Na2S2O4-cleavable azobenzene–biotin tags (azido-azo-biotin) that allow pulldown of labeled proteins on streptavidin beads and selective elution of affinity-attached proteins for MS-based protein identification ( Fig. 2 ). 67
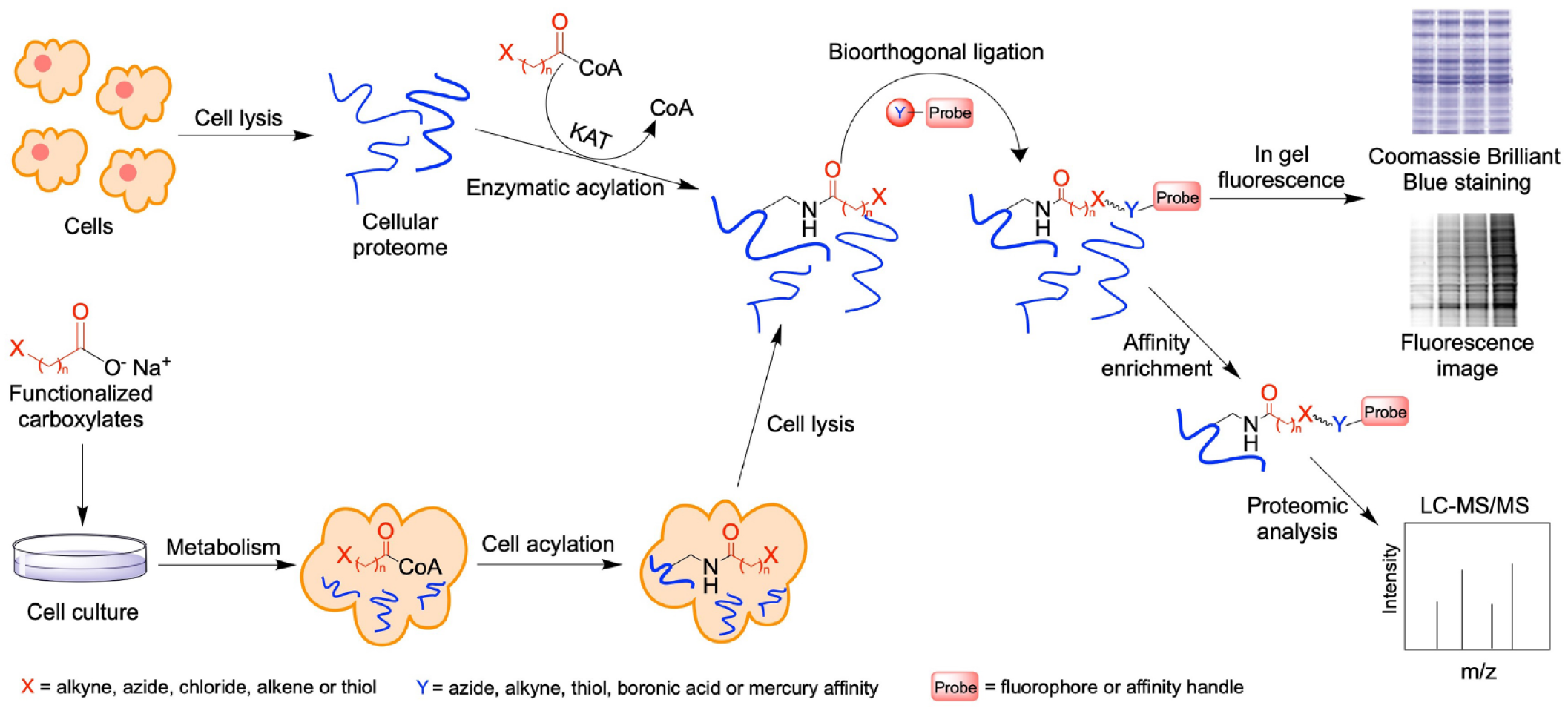
General procedures to profile KAT substrates using bioorthogonal reporters. The reporter labeling can be carried out in two different ways: (1) Acyl-CoA and respective recombinant KATs are used directly to install acyl reporters onto proteins in purified forms or in complex cell lysates. (2) Functionalized SCFAs are used as precursors of respective acyl-CoAs to treat cells in tissue culture. Once entering the cells, these precursor molecules are converted to respective acyl-CoA derivatives by the cell’s own biosynthetic machinery. Following reporter labeling, modified substrates are conjugated via bioorthogonal ligation to install either fluorophores for fluorescent imaging or affinity handles for pulldown enrichment and proteomic MS profiling.
Several research groups have demonstrated substrate labeling with functionalized acyl-CoA, bioorthogonal click conjugation, and downstream protein detection and analysis. Hang et al. detected the feasibility of cell-permeable bioorthogonal chemical reporters to profile protein lysine acetylation. 68 Culturing Jurkat T cells with 3-butynoate, 4-pentynoate, and 5-hexynoate followed by CuAAC fluorescent detection, all three reporters showed that the metabolic labeling was time and dose dependent. The application of sodium 4-pentynoate to Jurkat T cells identified 194 modified proteins as KAT substrate candidates through affinity enrichment by reacting with a cleavable azido-biotin tag followed by LC-MS/MS analysis. 68 Furthermore, the MS analysis also revealed that the modification sites on protein by 4-pentynote moiety were the same as known acetylation sites on histones H2B, H3, and H4 in cells, which provides validating evidence that these short-chain alkyne-acyl carboxylates can monitor protein acetylation in living cells as efficient chemical reporters ( Fig. 2 ). In addition, the same group applied 4PY-CoA together with purified KAT p300 to profile and identify p300 substrates from nuclear extracts of HeLa cells. 54 This chemical proteomics experiment identified several known protein substrates of p300, several new candidate p300 substrates, and the sites of modification. These results are an excellent demonstration that bioorthogonal chemical proteomics allows the rapid substrate identification of individual protein acetyltransferases in vitro.
We applied 3AZ-CoA as a bioorthogonal surrogate of acetyl-CoA for understanding the substrate profile difference between KAT p300 and GCN5, two representative KATs in mammalian cells. 55 In this study, protein extracts of human embryonic kidney 293T cells were subjected to 3AZ-CoA with wild-type p300 and GCN5-T612G mutant. The 3-azidopropionylated substrates were subsequently labeled with a sodium dithionite-cleavable alkyne–azo-biotin through the CuAAC reaction. Following protein enrichment on streptavidin-coated resin, we conducted LC-MS/MS studies from which more than 400 proteins were identified as GCN5 or p300 substrate candidates. The identified proteins either are p300 or GCN5 unique or shared by the two KATs, and are extensively involved in various biological events, including gene expression, the cell cycle, and cellular metabolism. We validated two novel substrates of GCN5 by co-transfection studies, IQGAP1 and SMC1. These results reveal extensive engagement of GCN5 and p300 in cellular pathways and provide insights in understanding their functional redundancy and distinction in biological pathway regulation.
To boost the cell permeability of functionalized SCFAs, Meier and coworkers developed a set of ester-modified SCFA pro-metabolites inspired by masking the polar groups in analogy to pro-drugs. 69 After treating the cell with 3-azidopropionyl-ester ( Table 1 ), LC-MS metabolomics analysis confirmed the formation of corresponding 3AZ-CoA. The modified proteins can be conjugated with fluorescent probes with CuAAC reaction visualized through the fluorescent gel-based assay. Again, the pro-metabolic labeling was also time and dose dependent. Interestingly, inhibition of p300 activity by a small-molecule inhibitor decreased the labeling efficiency of some but not all proteins, which suggests that p300 works with other KATs to establish protein acylation. After conjugation to alkyne biotin via CuAAC and enrichment by streptavidin-agarose, the labeled proteins were subjected to LC-MS/MS analysis; the results showed that 41% of the enriched proteins have been identified by Hang and coworkers, and some new acetylated marks were discovered.
Relatedly, Li and coworkers developed an alkyne-functionalized malonyl ester reporter to study lysine malonylation. 70 This alkynyl diacetoxy-methyl pro-drug (MalAM-yne) ( Table 1 ) can get into cells, release the corresponding functionalized malonic acid, and be converted into alkyl malonyl-CoA, which is subsequently utilized by hypothetic malonyltransferases (note that no bona fide malonyltransferases have been authenticated) to label their substrates ( Fig. 1G ). 70
Design of Other CoA Conjugates as Chemical Probes to Explore KAT Biology
In addition to the above-described functionalized acyl-CoAs as chemical reporters for the detection of KAT substrates, other CoA-based molecules have been designed to study KATs, most of which are affinity-binding ligands. 16 Cole’s group was among the first to design and use peptide–CoA conjugates as bisubstrate ligands for selective inhibition of KATs. In their original work, Lys-CoA and H3K14CoA were found to be submicromolar inhibitors for p300 (IC50 = 0.5 μM) and PCAF (IC50 = 0.3 μM), 71 respectively. We later designed H4K16-CoA conjugates for inhibition of the MYST family KAT enzymes Esa1 and TIP60.72,73 We recently found that the H4K12-CoA compound is a nanomolar potent inhibitor for HAT1. 74 These bivalent inhibitors are particularly useful for substrate–enzyme co-crystalization for x-ray structural elucidation of KAT catalytic mechanisms.75,76 A caveat is that althougth these bisubstrate probes are found to be selective for their KAT targets in biochemical counterscreen assays, target promiscuity could arise in complex proteome contexts and depends on the concentrations used. 77
CoA-based ligands are also designed to incorporate affinity tags for the pulldown of KATs of interest in functional proteomics study. Cole et al. designed a reactive affinity probe, the sulfoxycarbamate derivative of CoA (
Fig. 3
,
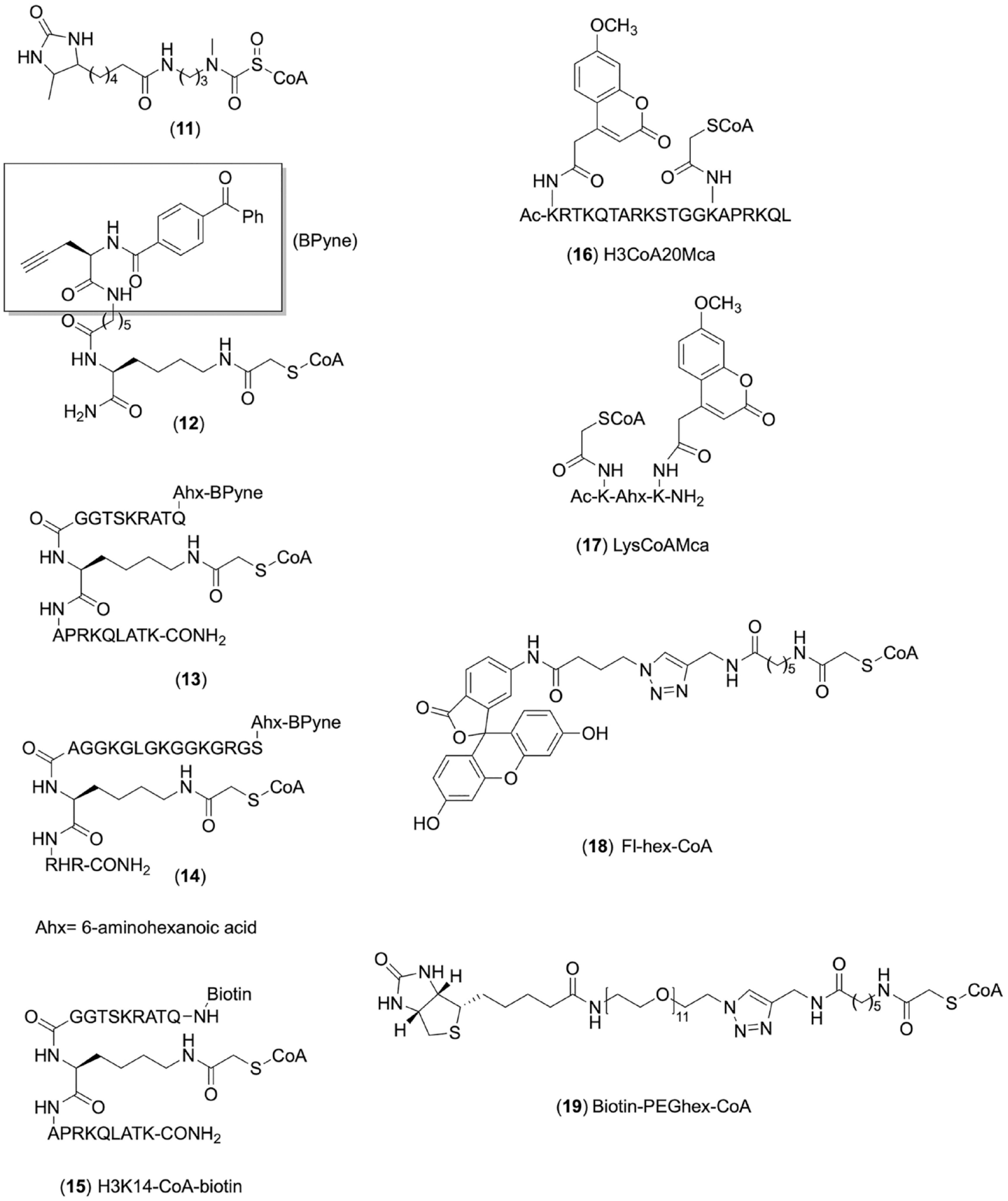
Chemical structures of other CoA-based conjugate probes for KAT activity studies.
To study KAT activities in complex settings and further investigate corresponding mechanisms and develop specific KAT inhibitors, Meier’s group made clickable photoaffinity CoA-based proteomic probes inspired by structures of bisubstrate inhibitors of KATs.
79
Different affinity probes (
Fig. 3
,
Meier and coworkers applied H3K14-CoA–biotin (
Fig. 3
,
The development of chemical probes targeting the cofactor binding sites is also of value for setting up biochemical assays to detect inhibitor-caused KAT activity changes. We created methoxycoumarin-labeled peptide–CoA conjugates (
Fig. 3
,
Long-Chain Fatty Acid Reporters to Study N-Myristoylation and S-Palmitoylation
Except for studying KAT-catalyzed acetylation and short-chain fatty acylation with specific chemical reporters, the development of bioorthogonal long-chain fatty acid chemical reporters has also gained increased interest over the past few years. Ploegh’s group first synthesized a series of varied-chain-length ω-azido-fatty acids containing 12, 14, 15, and 16 carbons (
Fig. 4
,
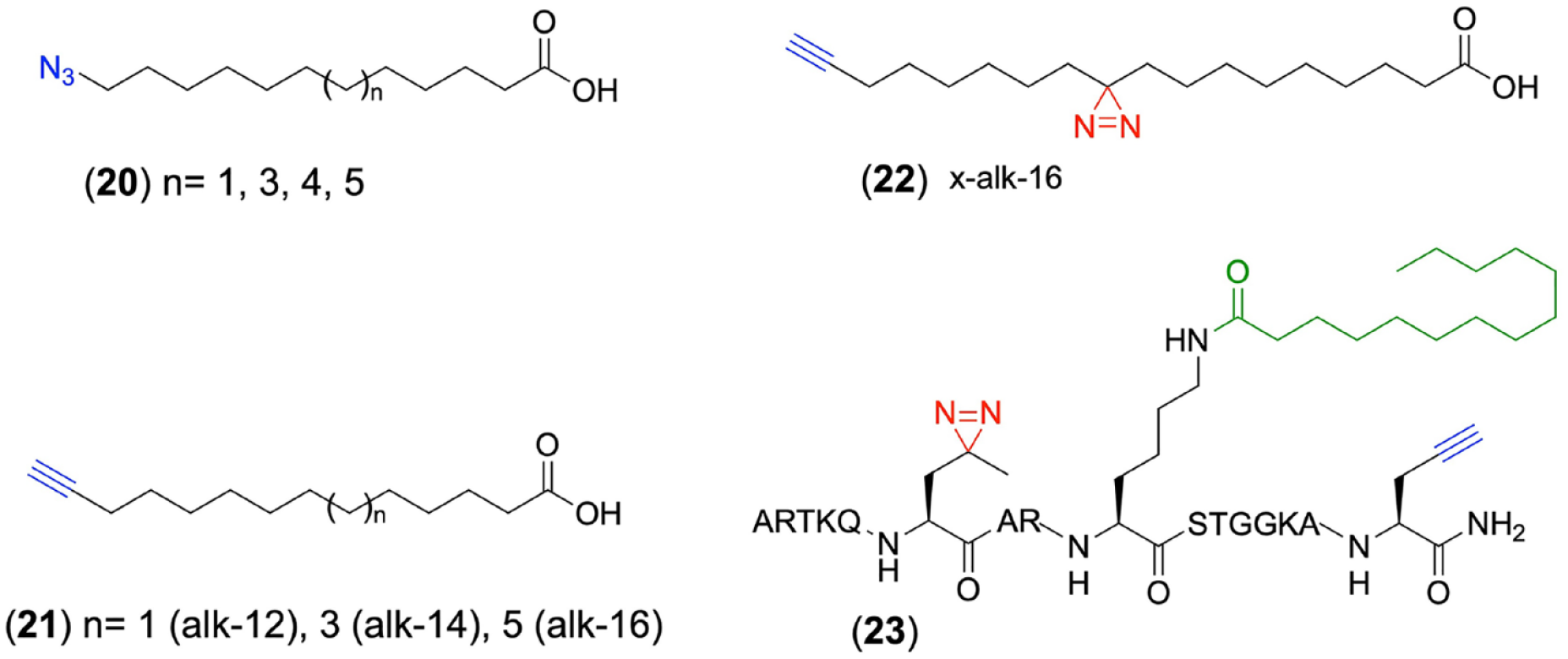
Chemical structures of long-chain fatty acid reporters for studying protein myristoylation and palmitoylation.
Later on, to improve the labeling sensitivity of fatty-acylated proteins, Hang’s group prepared a panel of alkynyl–fatty acids with different lengths (alk-12, alk-14, and alk-16) as chemical reporters (
Fig. 4
,
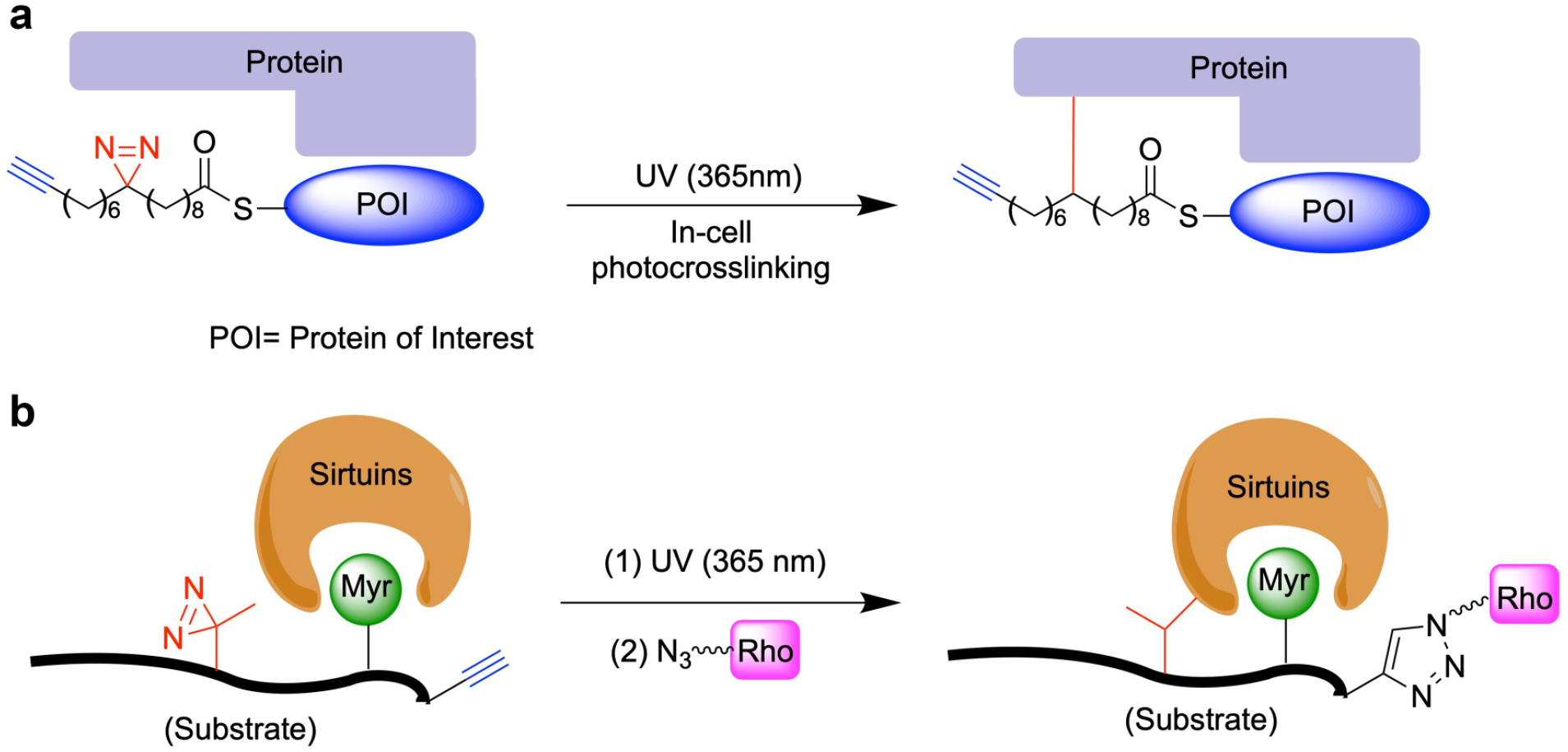
Schematic of using click chemistry and photoaffinity probes for the capture of (
To study the de-fatty-acid acylase activities of mammalian sirtuins, the Li group synthesized a chemical reporter based on a Lys9-myristoylated histone H3 peptide (H3K9Myr) with the replacement of Thr6 by a diazirine-containing photoreactive amino acid (photo-Leu) (
Fig. 4
,
Summary and Outlook
Defining the scope and properties of the acetylomes is a vigorous tour de force of research for biologists in the postgenomic era. Identifying and profiling cellular substrates of KATs is of immediate significance for understanding their roles in both normal physiology and disease conditions. The effort of acetylome profiling can be facilitated by innovative chemical biology reporters. Data have proven that wild-type KATs or rationally engineered enzyme forms are efficient for labeling KAT substrates when used in conjunction with appropriately designed chemoreactive acyl-CoA molecules.53,56 For instance, p300/CBP can take 4PY-CoA and 3AZ-CoA as efficient reporters for chemical labeling of their substrates. For the GCN5, we found that both GCN5-T612G/F622A–5HY-CoA and GCN5-T612G/L531A–5HY-CoA are superior matching pairs for selective labeling of its substrates. GCN5-T612G was active with several synthetic cofactors: 4PY-CoA, 5HY-CoA, 6HY-CoA, and 3AZ-CoA. 55 For MOF, we identified MOF-I317A and MOF-I317A/H273A, which showed marked activity with 5HY-CoA. MOF-I317A was also very active toward 3AZ-CoA. Although in this review we have emphasized chemical reporter design for KAT-mediated lysine acetylation, conceptually equivalent, long-chain acylation of protein (e.g., palmitoylation and myristoylation) with bioorthogonal chemical warheads has been well developed.45,65,85,89,90
Currently, direct usage of acyl-CoA chemical reporters is limited to labeling KAT substrates in purified proteins/peptides or in cell lysates that do not pose membrane barriers. The lack of cell permeability of CoA-based reporters technically hinders their further utilization in detecting activities of KAT enzymes in intracellular contexts. Applying functionalized fatty carboxylates as precursors of acyl-CoA has proven to be a practical approach, but metabolic conversion of fatty acid to fatty acyl-CoA could be limited by the cellular level of acyl-CoA synthetases. 91 For efficient substrate labeling, it would be ideal to use cell-permeable acyl-CoAs for direct interrogation of intracellular acetylation. Interestingly, a recent study revealed a CoA transport mechanism in which CoA was hydrolyzed extracellularly by ectonucleotide pyrophosphatases to 4′-phosphopantetheine, a biologically stable molecule able to translocate through membranes via passive diffusion. 92 Inside the cell, 4′-phosphopantetheine was enzymatically converted back to CoA by the bifunctional enzyme CoA synthase. Phenotypes induced by intracellular CoA deprivation were reversed when exogenous CoA was provided. This finding offers a new insight into cellular delivery of bioorthogonal acetyl-CoA surrogates.
The bioorthogonal acyl-CoA cofactors in conjunction with their matching KATs form versatile reporters to identify KAT targets, thereby expanding the chemical biology tool repertoires for functional annotation of the lysine acetylome in different biological systems. Due to substrate promiscuity of KATs and possible unknown acyltransferases, bioorthogonal labeling is implemented largely in a way to identify KAT substrates globally and robustly. Although hundreds and even thousands of acetylated substrates and sites have been profiled by proteomic screening, our knowledge about how individual KAT members make contributions to construct the map of the acylome sea remains limited. Given that functions of individual KATs are distinct from each other and directly associated with the subacetylomes that they introduce to biological systems, in our view the ultimate goal is that the bioorthogonal chemical reporters should be applied for the discovery of subacetylomes of individual KAT members. In this regard, besides the identification of substrate profiles for the members of the three major canonical KAT families, we also expect that the bioorthogonal reporters can be used to define enzymatic activities of poorly characterized KATs, such as NAT10 and NAA50. 79 Moreover, enzymatic acyl labeling combined with bioorthogonal ligation with fluorescent probes can be used to detect and characterize potential KAT inhibitors by competing labeling through easily accessible in-gel fluorescence imaging. This will be highly valuable for the drug discovery of KAT inhibitors. Except for their use in profiling cellular substrates of KATs, functionalized acylate and acyl-CoA reporters have potential to be valuable to study the activities and functions of lysine deacylases or acylation reader proteins (e.g., bromodomains). In particular, lysine residues modified with fatty acyl groups that contain photoaffinity labels could be utilized to identify downstream protein erasers or readers ( Fig. 5 ). Furthermore, the biological systems to which orthogonal acylation can be applied should be further expanded. Thus far, the bioorthogonal labeling of protein acetylation and acylation is mainly carried out only in cell line models. We anticipate seeing these chemical proteomics advances to be utilized to identify acetylated substrates associated with different disease phenotypes, such as inflammation and malignancies in tissues and animal models. Overall, the methodological advances in bioorthogonal acylation labeling provide a powerful chemical biology tool to better understand enzymatic functions of KATs, illuminate disease mechanisms mediated by KATs, and develop corresponding therapeutic drugs for disease treatment.
Footnotes
Acknowledgements
We thank the National Science Foundation for its support (grants 1507741 and 1808087 to Y.G.Z.).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: We acknowledge the support of the National Science Foundation (grants 1507741 and 1808087 to Y.G.Z.).
