Abstract
Introduction
Click chemistry is a synthetic concept introduced by Nobel Laureate KB Sharpless in which modular synthesis is used to rapidly find new molecules with desirable properties. 1 The representative reactions of click chemistry are copper (I)-catalyzed azide–alkyne cycloaddition (CuAAC) triazole annulation and sulfur (VI) fluoride exchange (SuFEx) catalysis.2,3 These reactions have a few widely recognized advantages, such as yields approaching 100%, rapid access to their products, and being largely orthogonal to other reactions. 4 However, the risk of explosion and potential toxicity have restricted their application. 5 In 2019, a new click-chemistry synthesis method was reported to add to the click reaction family. 6 This method uses just 1 equivalent of fluorosulfuryl azide (FSO2N3),7–9 a simple diazotized substance, and can enable the generation of more than 1200 compounds on 96-well plates in a safe and practical manner. Using this method, starting with a large No. of available molecular building blocks of primary amine functional groups, 10 a large No. of differential derivatives with a purity of approximately 70% are directly synthesized on 96-well plates, and their biological activities can be directly screened.
Malignant tumors are common and frequently occurring diseases that seriously threaten human health. 11 Molecular targeted therapy is a tumor treatment method that is different from conventional treatment methods for malignant tumors, such as surgery, chemotherapy, and radiotherapy. Its advantages are high efficiency, low toxicity, and high specificity, which has led to molecular targeted therapy gradually becoming a hot research topic. 12 The research and development cycle of new antitumor molecular targeted drugs is time-consuming, and the cost is high. Structural modification or transformation of the existing small-molecule targeted drugs may speed up the development of antitumor drugs and reduce research costs. 13 Using innovative click-chemistry methods to modify the structure of existing compounds can quickly obtain a large No. of new compounds with diversified structures, which is in sharp contrast to the cumbersome and inefficient methods of traditional medicinal chemistry.
Epidermal growth factor receptor (EGFR) is a signal transduction receptor for epithelial growth factor (EGF) with tyrosine kinase activity on the cell membrane surface. Research has shown that EGFR is highly expressed in a variety of epithelial tumors, and small-molecule targeted drugs have achieved significant efficacy in tumor treatment. 14 Erlotinib is a first-generation small molecule inhibitor targeting EGFR that is suitable for patients with locally advanced or metastatic non-small-cell lung cancer (NSCLC). 15 Erlotinib effectively controls the progression of tumors, with definite clinical efficacy, mild side effects, and good patient tolerance. Erlotinib can be used for the third-line treatment of NSCLC when 2 or more chemotherapy regimens have failed, based on the results of a foreign phase III clinical study. Erlotinib may have clinically significant drug interactions. Therefore, it is emphasized that this product must be used under the guidance of doctors with experience in the use of such drugs and only used in national clinical trial pharmacology bases or tertiary A hospitals. 16
In our present study, the small molecule targeted drug Erlotinib was modified using click-chemistry methods, and 840 new compounds were quickly obtained in one day. Through screening these newly synthesized compounds, we explored the application of click chemistry in the field of new antitumor drug discovery. First, the thiazolyl blue (MTT) method was used to conduct high-throughput screening of the antitumor activity of these novel compounds on A549 lung adenocarcinoma cells at 10 μM. After obtaining 22 compounds with better activity, we synthesized their pure products and measured their IC50 values to inhibit A549 cells. Among them, the compound P19G1 showed the best inhibitory activity, and it inhibited many kinds of tumor cells, such as H1299 lung cancer cells, HeLa cervical cancer cells, RKO colorectal cancer cells, and MCF-7 breast cancer cells, with IC50 values in the range of 1 to 5 μM. Second, further studies found that P19G1 effectively inhibited tumor cell proliferation and migration and promoted tumor cell apoptosis in A549 cells and RKO cells. Finally, we studied the antitumor activity, toxicity, and pharmacokinetic (PK) properties of compound P19G1 in vivo. The research results showed that P19G1 has better absorption, lower toxicity, and significant antitumor activity. This study reveals the potential of new click-chemistry reactions in drug discovery and provides a new paradigm for the application of such reactions in the discovery of antitumor drugs.
Materials and Methods
Preparation of 840 new 1,2,3-Triazole Compounds Using a Modular Click-Chemistry Method
The preparation of the azide library was adapted from a previously reported procedure. 6 A total of 840 azides were stored in a 96-well microplate (50 mM dimethyl sulfoxide [DMSO] solution), Erlotinib was dissolved into its solution in DMSO (25 mM), sodium ascorbate was dissolved into its solution in water (250 mM), and CuSO4 and Tris(3-hydroxypropyltriazolylmethyl) amine (THPTA) were premixed and dissolved into its solution in water (2.5 mM each). The transfer and addition of solutions in this method were carried out with a 12-channel pipette.
Ninety-six azide solutions from the azide library were transferred (10 µL each well, containing 0.5 µmol of an azide) to a 96-well microplate (each empty well with 300 µL volume). To each well, the Erlotinib solution (20 µL, 0.5 µmol) was added, and then CuSO4/THPTA aqueous solution (10 µL, 0.025 µmol each) and sodium ascorbate solution (10 µL, 2.5 µmol) were added. Additional DMSO (37 µL) and water (13 µL) were added; therefore, the bulk volume of the mixture was approximately 100 µL in each well. The microplate was then sealed and incubated in a microplate shaker at 800 rpm and 25 °C for 12 h. The synthetic route is shown in Figure 1A.

Synthesis of 840 new 1,2,3-triazole-containing Erlotinib derivatives using a modular click-chemistry approach and evaluation of their antitumor activity in A549 cells. (A) A total of 840 1,2,3-triazole-containing Erlotinib derivatives were synthesized using the new modular click-chemistry method. (B) Antitumor activity screening was performed using the Erlotinib series of library in A549 cells. A549 cells were treated with 10 μM of compounds for 48 h and the antitumor activity was evaluated using thiazolyl blue (MTT) assay. (C-D) Concentration-response curves and IC50 of 22 pure compounds and Erlotinib in A549 cells. A549 cells were treated with 22 pure compounds at different concentrations for 48 h and the antitumor activity was evaluated using MTT assay. Graphed data are presented as means ± SD. (n = 3).
Synthesis and Isolation of P19G1
Chemicals were acquired from Alfa Aesar, Sigma-Aldrich, and TCI. All chemicals were used without further purification. The preparation of 4-aminophenyl fluorosulfonate 3 and FSO2N3 was adapted from a previously reported procedure. 6 4-(4-(3-((6,7-bis(2-methoxyethoxy)quinazolin-4-yl)amino)phenyl)-1H-1,2,3-triazol-1-yl)phenyl sulfurofluoridate (P19G1) was synthesized, as shown in Figure 2A. 4-aminophenyl fluorosulfonate (25 mmol, 4.78 g) was dissolved in 200 mL N,N-dimethylformamide (DMF), followed by 33 mL KHCO3 solution (3 M aqueous solution, 100 mmol) and 60 mL FSO2N3 solution (0.42 M methyl tert-butyl ether solution, 25 mmol). The reaction mixture was stirred for 4 h at room temperature.

Synthesis and structural characterization of 4-(4-(3-((6,7-bis(2-methoxyethoxy)quinazolin-4-yl)amino)phenyl)-1H-1,2,3-triazol-1-yl)phenyl sulfurofluoridate (P19G1) and evaluation of its antitumor activity in different tumour cell lines. (A) Synthesis of compound P19G1. (B) Concentration-response curves of P19G1 in 20 different tumor cell lines. (C) The IC50 value of P19G1 and Erlotinib in 20 different tumor cell lines. A549 cells were treated with P19G1 and Erlotinib at different concentrations for 48h and the antitumor activity was evaluated using thiazolyl blue (MTT) assay. The data are means ± SD. (n = 3).
After completion, 12.5 mL sodium ascorbate solution (2 M aqueous solution, 25 mmol) was added to quench the reaction. Erlotinib (25 mmol, 10.74 g) and 0.63 mL CuSO4 solution (1 M aqueous solution, 0.63 mmol) were then added, and the reaction mixture was stirred at 60 °C for 4 h and monitored by high-performance liquid chromatography–mass spectrometry (HPLC-MS). The hot solution was then poured into 1 L distilled water, and the yellow solid was precipitated out and collated by filtration. The crude product was then dissolved in 100 mL methanol (CH3OH)/dichloromethane (CH2Cl2) (1:1, v/v), dried over anhydrous Na2SO4, and purified by column chromatography (CH2Cl2/CH3OH = 10:1, v/v), yielding the pure product as a light yellow solid (8.4 g, 55% yield). We further used nuclear magnetic resonance (NMR) and high-resolution mass spectrometry (HRMS) to confirm the structure of P19G1. P19G1 was finally treated with 1 equiv. CH3SO3H in CH3OH quantitatively yields its mesylate for cellular and animal experiments.
Tumor Cell Lines and Cell Culture
Twenty tumor cell lines were used in our research. The human lung cancer cell lines A549 (TCHu150), NCI-H1299 (TCHu160), NCI-H358 (TCHu151), and NCI-H1703 (SCSP-593); the human renal carcinoma cell line 786-O (TCHu186); and the human colorectal cancer cell lines RKO (TCHu116) and HCT116 (TCHu99) were maintained in Roswell Park Memorial Institute 1640 (RPMI1640). The human hepatocarcinoma cell line HepG2 (TCHu72), human glioma cell line U251 (TCHu58), human pancreatic cancer cell line Panc-1 (TCHu98), human cervical carcinoma cell line HeLa (TCHu187), and human breast cancer cell line MCF-7 (TCHu74) were cultured in Dulbecco's modified Eagle's medium (DMEM). The human colorectal cancer cell line LOVO (TCHu82), human gastric cancer cell line AGS (TCHu232), human breast cancer cell lines MDA-MB-468 (TCHu136), MDA-MB-231 (TCHu227), human prostate cancer cell line PC-3 (TCHu158), and human colorectal cancer cell line HT-29 (TCHu103) were cultured in minimum essential medium/F-12 (MEM/F-12). Human chronic myelogenous leukemia cells K562 (TCHu191) and human myeloid leukemia cell line HL60 (TCHu23) were cultured in Iscove's modified Dulbecco's medium (IMDM). These media were supplemented with 10% fetal bovine serum (FBS) and antibiotics (1% penicillin-streptomycin). The method of culturing cells was performed as previously described. 17 All cell lines were purchased from the Shanghai Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences (SIBS, CAS).
MTT and Cell Proliferation Assays
For the high-throughput screening of a library of click-chemistry compounds and antitumor activity testing, the MTT assay was used according to the previous description. 17 The tumor cells were treated with the compounds for 48 h, and then 5 mg/mL MTT aqueous solution was added to the 96-well plates and incubated for 4 h at 37 °C. Dissolution solution (0.012 mol/L HCl, 5% isobutanol, 10% sodium dodecyl sulfate [SDS], dissolved in distilled water) was added to dissolve formazan crystals in these viable cells. 18 The absorbance was measured at 570 nm for each well using a spectrophotometer (Thermos Fisher). The concentration for the high-throughput screening of an Erlotinib compound library was 10 μM. IC50 calculations were performed in GraphPad Prism Version 7.0.
Colony Formation Assay
A549 and RKO cells (500-1000) were seeded into 12-well plates in triplicate. The operation of the cell monoclonal formation experiment was as described before. 19 Cells rested overnight before treatment with the compound. For compound treatment, Erlotinib or P19G1 was supplied to the liquid media layer, and the cell culture medium was changed every 2 days. Two weeks later, cell colonies were stained with 0.05% crystal violet, imaged using a camera (Nikon) and quantified using colony counting software (NIH ImageJ, particle analyser).
Immunofluorescence Staining
A Ki67 immunofluorescence (IF) staining experiment was performed as described before. 20 A549 and RKO cells were incubated with compound P19G1 and Erlotinib for 48 h. After washing with Dulbecco's phosphate-buffered saline (DPBS), A549 and RKO cells were fixed with 4% paraformaldehyde, permeabilized in DPBS with 0.1% Triton X-100, sealed with 1 × goat serum at 37 °C for 1 h, and then incubated overnight with Ki67 fluorescent primary antibody (1:300) at 4 °C. After incubation, the cells were washed with DPBS and then stained with 4′,6-diamidino-2-phenylindole (DAPI, 1:100, Sigma). Images were captured and quantified using a confocal laser scanning fluorescence microscope (Nikon, Japan) and Adobe Photoshop 2019 (Adobe, USA).
Scratch Assay
A549 and RKO cell migration assays were performed under different conditions according to a previous protocol. 21 The cells were inoculated into a 6-well plate, grown to a confluence of 90% to 95%, and then a pipette tip was used to mark out the cell monolayer to form a straight line “scratch”. The cells were treated with a medium containing 1% FBS with different concentrations of Erlotinib and P19G1 for 48 h. At 0, 24, and 48 h after wound scratching, photographs were captured with an inverted microscope (N-SIM, Japan), and the migration ratios of the cells were determined according to ImageJ software (NIH, USA).
Cell Cycle Analysis
A total of 5 × 105 A549 and RKO cells per well were seeded in a 6-well plate and then incubated with different concentrations of P19G1 or Erlotinib. After 48 h, all cells were collected in the well, centrifuged at 1000 rap/min for 10 min, and washed with DPBS, and 1 mL of precooled 70% ethanol was added for fixation. After washing with DPBS, staining operations were performed in accordance with the requirements of the cell cycle kit (Dalian Meilun Biotechnology). A FACSCalibur flow cytometer (BD Bioscience) was used to analyze the samples, and MODFIT software (Verity Software House) was used to determine the cell cycle distribution.
Cell Apoptosis Analysis
Tumor cells were seeded into a 6-well plate (5 × 105 cells per well) overnight and then incubated with different concentrations of P19G1 or Erlotinib. After 48 h, the cells floating in the culture medium were collected, washed with cold DPBS, and removed from the plate by trypsinization. The floating and adherent cells were mixed, and the resuspended cells were washed with cold DPBS and stained with the Annexin V-FITC/PI apoptosis detection kit according to the protocol instructions (Roche Applied Science). After incubating at room temperature for 15 min in the dark, we immediately analyzed the apoptotic cells with a Beckman Coulter flow cytometer (BD Biosciences). The distribution of apoptosis was computed using FlowJoTM software (BD Biosciences).
Transwell Migration Assay
The migration assay was performed according to a previous protocol. 22 Briefly, 8 μm pore size culture inserts (Costar) were placed in 24-well cell culture plates, and each well was divided into an upper chamber and a lower chamber. In the lower chamber, 300 μL of medium with 1% FBS containing different concentrations of compounds was added. In the upper chamber, 300 μL of A549 or RKO cells (5 × 104 cells/mL) were seeded. After 48 h, the cells were fixed with 4% formaldehyde and then stained with 0.05% crystal violet; the nonmigrated cells in the upper chamber were scraped off with a cotton swab. An inverted microscope (N-SIM, Japan) was used to capture images, and the migrated cells were counted with ImageJ software (NIH, USA).
Hoechst 33342 Staining
A Hoechst 33342 staining assay was performed as described previously. 23 A549 and RKO cells (1 × 106 cells per well) were seeded in 6-well plates and incubated overnight. Then, different concentrations of P19G1 or Erlotinib were used to treat the cells for 48 h. After the cells were washed, adherent tumor cells were fixed with 4% formaldehyde. Then, 100 μL of Hoechst 33342 stain was used to treat these cells according to the manufacturer's protocol. 24 Washing was repeated to remove the excess stain, and a fluorescence microscope (Nikon, Japan) was used to capture the images.
Western Blot Assay
P19G1-treated cells were collected and washed with ice-chilled DPBS. Cells were lysed with radioimmunoprecipitation assay (RIPA) reagent supplemented with protease and phosphatase inhibitors (Transgen Biotech, Beijing, China) according to the manufacturer's instructions. Protein concentrations were determined with the Enhanced BCA Protein Kit (Beyotime Biotechnology). The protein samples were then subjected to western blot analysis as described previously. 21 Antibodies against CDK-4 (ab108357, 1:1000), CDK-6 (ab124821, 1:1000), MMP-2 (ab92536, 1:1000), and MMP-9 (ab76003, 1:1000) were purchased from Abcam (Abcam). Antibodies against BCL-2 (15071T, 1:1000), Caspase-3 (9662S, 1:1000), and Cleave-caspase 3 (9661T, 1:1000) were purchased from Cell Signaling Technology (Danvers, MA, USA), and antibodies against β-actin (GB15001, 1:1000) were purchased from Servicebio (Servicebio, Wuhan, China). Detection was analyzed by the Azure Biosystem 25 (Azure c600, Azure Biosystem™).
In Vivo Studies
Four-week-old BALB/c nude mice were used for subcutaneous tumor formation experiments. The operation protocol of the experiment was as described before. 22 After rearing the mice for a period of time, the BALB/C nude mice were injected subcutaneously (6 mice per group) with approximately 1 × 107 A549 cells per mouse. When the tumor volume was greater than 100 mm3, the mice received intratumoral (IT) injections of P19G1 in saline (50 mg/kg body weight), Erlotinib in saline (50 mg/kg body weight), or vehicle control every day. The body weight and tumor volume of these mice were measured every 2 days for 17 days, and the tumor volume was calculated according to the universal method. At the end of the experiment, the nude mice were sacrificed by intraperitoneal (IP) injection of 200 mg/kg of sodium isobarbitone, 26 and the tumor tissues were removed.
For PK studies, 200 g SD female rats were used. The SD rats were administered a single dose orally, intraperitoneally, or via the tail vein, and then anesthesia with 0.5 L/min of isoflurane and blood was taken from the orbital cavity of the rats at 0.25 h, 0.5 h, 0.75 h, 1 h, 2 h, 3 h, 4 h, 6 h, 8 h, 12 h, and 24 h after administration and then centrifuged; the internal standard solution and methanol were added to the mixture, and the supernatant was filtered and measured by HPLC-MS. ImagePro Plus software (NIH, USA) was used to analyze tumor section images.
BALB/C nude mice and SD female rats were acquired from the Shanghai University of Traditional Chinese Medicine Institutional Animal Care and Use Committees (Shanghai, China).
Immunofluorescence and Hematoxylin and Eosin (H&E)
Tumor tissues were fixed in natural buffer formalin. Paraffin embedding, section preparation, and staining were carried out as described previously. 22 Briefly, the paraffin-embedded tissue slides were deparaffinized and treated with PTLink and pretreatment buffer at pH 9.0 (Dako, DK, S2375) and 100 °C for 20 min for antigen retrieval. The sections used for staining were washed in DPBS after pretreatment. Staining was performed using a TUNEL stain kit (ThermoFisher) and specific antibodies for FITC-Ki67 primary antibody (CST, USA) and monoclonal anti-MMP9 antibody (CST, USA). Haematoxylin and eosin imaging and staining were conducted as described previously. These tumor section images were analyzed by ImagePro Plus software (NIH, USA).
Statistical Analysis
All data were statistically analyzed using formal SPSS 16.0 software (SPSS Inc.). The Tukey method was used for multiple comparisons. The significance level was measured using SPSS one-way analysis of variance. A significant difference was considered at a value of P < .05. All data are displayed as the means ± standard deviation (SD).
Ethics Statement
All animal protocols and procedures were approved by the Shanghai University of Traditional Chinese Medicine Institutional. Animal Care and Use Committees and were carried out in accordance with the guidelines outlined in the Guide for the Care and Use of Laboratory Animals published by the National Institute of Health. 27 We make efforts to minimize the No. of animals utilized and to decrease their suffering. All experiments were carried out in compliance with ARRIVE guidelines. 28 Animal ethics number: PZSHUTCM200410003.
Results
Synthesis of 840 new 12,3-Triazole-Containing Erlotinib Derivatives Using a Modular Click-Chemistry Approach and Evaluation of Their Antitumor Activity in A549 Cells
Using the newly prepared azide library to perform diazo transfer-CuAAC steps with Erlotinib in a 96-well microplate, 840 Erlotinib derivatives containing 1,2,3-triazoles were easily obtained (Figure 1A). The derivatives were directly screened for biological activity without purification because it has been reported in the laboratory that 81% 1,2,3-triazoles compounds achieved greater than 70% conversion among the 1,224 CuAAC reactions, and this is a conversion standard that we think is very suitable for direct biological screening.
The MTT method was used to screen the library at a concentration of 10 μM in A549 cells, and the in vitro antitumor activity was judged by the inhibition rate of cell viability. The experimental results showed that 22 compounds out of 840 new Erlotinib series derivatives have more than 60% inhibitory activity on A549 cells, while the inhibitory activity of Erlotinib on A549 cells at this concentration was only 20% (Figure 1B).
Subsequently, we synthesized pure products of these 22 compounds and determined their IC50 values for inhibiting the viability of A549 cells. The results showed that the cell viability of these 22 new Erlotinib series compounds on A549 cells gradually increased with the increase of the compound concentration, and their inhibitory effect was stronger than that of Erlotinib. Among them, P19G1 had the strongest inhibitory effect, and its IC50 value was approximately 1.952 μM (Figure 1C and D).
Synthesis and Structural Characterization of P19G1 and Evaluation of its Antitumor Activity in Different Tumor Cell Lines
4-aminophenyl fluorosulfonate and FSO2N3 were stirred at room temperature for 4 h, and then Erlotinib and CuSO4 solutions were added. The reaction mixture was stirred at 60 °C for 4 h. The crude product was then dissolved, dried over anhydrous Na2SO4, and purified by column chromatography, yielding pure product P19G1 as a light yellow solid (8.4 g) (Figure 2A).
The structure of the obtained P19G1 compound was characterized by 1H NMR (Supplementary Figure S1A), 13C NMR (Supplementary Figure S1B), 19F NMR (Supplementary Figure S1C), and HRMS. The specific map data are as follows: 1H NMR (400 MHz, d6-DMSO) δ 8.50 (s, 1H), 8.42-8.36 (m, 1H), 8.26-8.17 (m, 2H), 8.00-7.87 (m, 4H), 7.65 (d, J = 8.0 Hz, 1H), 7.52 (t, J = 8.0 Hz, 1H), 7.23 (s, 1H), 4.38-4.23 (m, 4H), 3.85-3.71 (m, 4H), 3.38 (s, 3H), 3.36 (s, 3H); 13C NMR (100 MHz, d6-DMSO) δ 156.4, 156.3, 153.6, 152.9, 148.8, 148.1, 147.6, 146.9, 140.2, 140.1, 136.7, 130.3, 129.1, 122.4, 122.2, 120.6, 120.0, 119.1, 118.9, 109.0, 108.1, 103.2, 70.2, 70.1, 68.4, 68.1, 58.4, 58.4; 19F NMR (376 MHz, d6-DMSO) δ 39.48 and HRMS (ESI): Calcd for C28H28FN6O7S+ ([M + H]+) 611.1719, found 611.1717.
Then, P19G1 was given to 20 other tumor cell lines derived from different human organs at different concentrations below 100 μM. The cell viability was determined by the MTT method, and finally, the antitumor activity was judged based on the calculated IC50 value (Figure 2B and C). Experimental results showed that P19G1 has a strong inhibitory effect on most tumor cell lines, among which cell lines with strong inhibitory effects include A549, H1299, MDA-MB-468, and RKO cells.
Inhibition of Tumor Cell Proliferation in A549 and RKO Cells by P19G1
Since P19G1 has a strong inhibitory effect on the two tumor cell Lines A549 and RKO, they were selected to further verify the antitumor mechanism of P19G1. We first evaluated the inhibitory effect of P19G1 on A549 and RKO cell proliferation. Using cell cloning experiments, we demonstrated that P19G1 effectively inhibited the growth of A549 and RKO cells. The area stained by crystal violet steadily decreased with increasing concentrations of P19G1. However, at the same concentration, the inhibitory effect of Erlotinib on both cell lines was not obvious (Figure 3A-D). To further clarify the effect of P19G1 on tumor cell proliferation, a Ki67 IF staining assay was performed. The Ki67 protein has been widely used as a proliferation marker for human tumor cells. 29 As shown in Figure 3E, the experimental results showed that compared with the control group, P19G1 significantly inhibited the expression of Ki67 in A549 cells and RKO cells (Figure 3F, ***P < .001), while the Erlotinib group had no significant inhibitory effect. Finally, a flow cytometry cell cycle assay was performed to evaluate the effect of P19G1 on tumor cell proliferation (Figure 3F and G). The results showed that P19G1 significantly induced A549 and RKO cell cycle arrest at the G1/S phase after 24 h or 48 h of intervention (Figure 3H and I, ***P < .001), while erlotinib showed no significant inhibitory effect.

Inhibition of tumor cell proliferation in A549 and RKO cells by 4-(4-(3-((6,7-bis(2-methoxyethoxy)quinazolin-4-yl)amino)phenyl)-1H-1,2,3-triazol-1-yl)phenyl sulfurofluoridate (P19G1). (A-B) Colony formation assay and quantification of A549 cells treated with P19G1 (1.25, 2.5, 5 μM) for 48 h. (C-D) Colony formation assay and quantification of RKO cells treated with P19G1 (1.25, 2.5, 5 μM) for 48 h. (E) Ki67 immunofluorescence (IF) staining experiments and quantification of A549 and RKO cells treated with P19G1 (5 μM) and Erlotinib (5 μM) for 48 h. Scale bar: 10 μm. (F and H) Quantification of cell cycle distribution and flow cytometry analysis were performed to determine the A549 cell cycle distribution. (G and I) Flow cytometry analysis and quantification of cell cycle distribution were performed to determine the RKO cell cycle distribution. A549 and RKO cells were treated with P19G1 (5 μM) and Erlotinib (5 μM) for 24 h and 48 h and cell cycle distribution were evaluated using Flow cytometry. Data are presented as means ± SD. (n = 3). *P < .05, **P < .01, ***P < .001 versus Control group.
Inhibition of Tumor Cell Migration in A549 and RKO Cells by P19G1
Scratch assays and Transwell migration tests were used to detect the inhibitory effects of P19G1 and Erlotinib on tumor cell migration. For the scratch assay, A549 cells were treated with 2.5 μM and 5 μM P19G1 and Erlotinib for 24 or 48 h before creating a “scratch” in a cell monolayer. Meanwhile, the cell images were captured at the beginning and at regular intervals during cell migration to close the scratch and then compared to quantify the migration rate of the cells (Figure 4A). Our experimental results showed that P19G1 distinctly inhibited the migration of A549 cells compared to the control group (Figure 4B, ***P < .001), but the erlotinib group had no obvious inhibitory effect. In RKO cells, the scratch detection results were similar to those in A549 cells (Figure 4C), revealing that 2.5 μM and 5 μM P19G1 markedly inhibited the migration of RKO cells (Figure 4D, ***P < .001). For the Transwell migration assay, A549 and RKO cells were treated with 5 μM and 10 μM P19G1 and Erlotinib for 48 h, and then the cells were stained with crystal violet and photographed (Figure 4E and F). The results showed that P19G1 had a significant inhibitory effect on the migration of A549 and RKO cells at concentrations of 5 μM and 10 μM, while Erlotinib had no significant effect on Transwell migration (Figure 4G and H, ***P < .001).
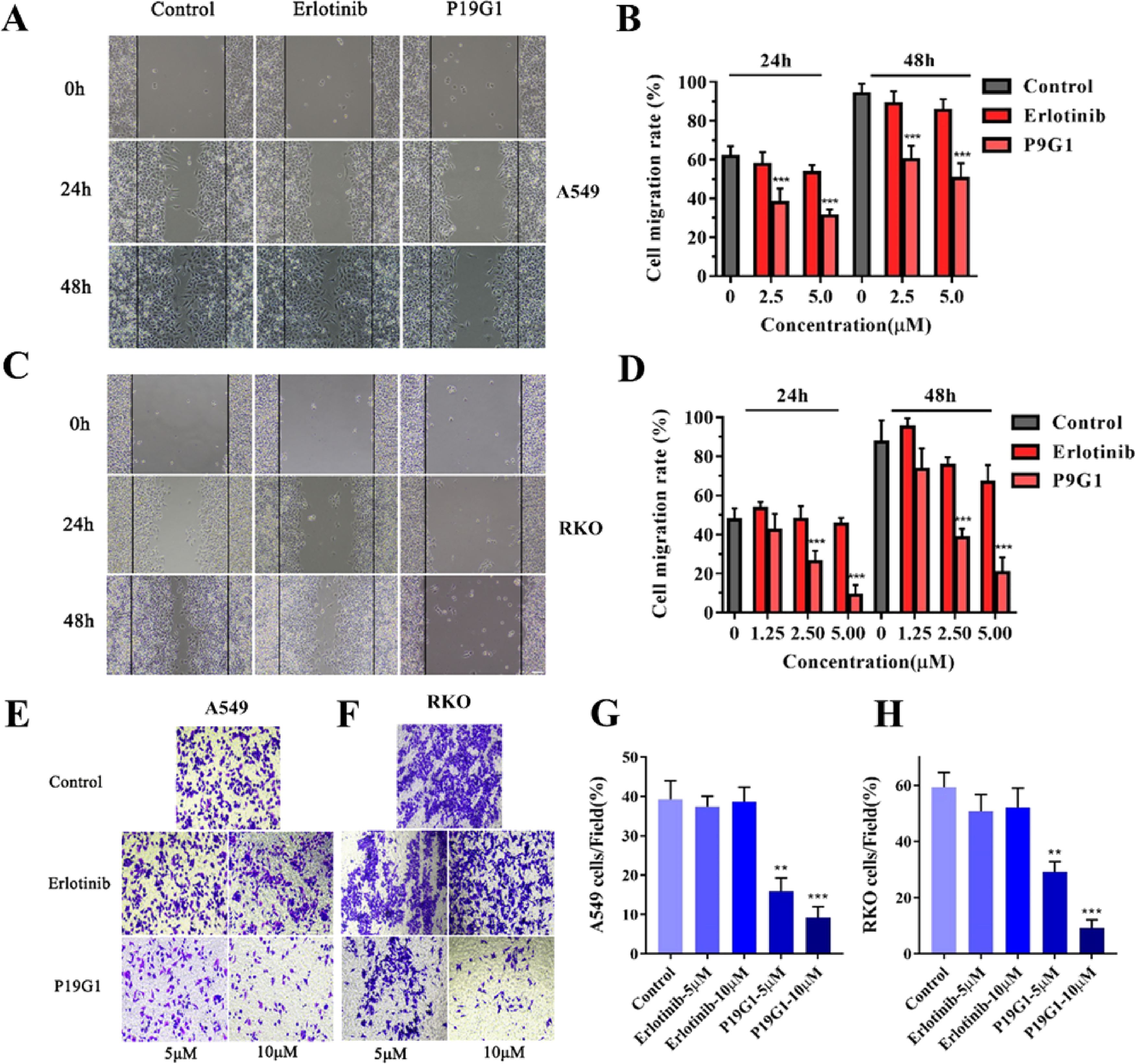
Inhibition of tumor cell migration in A549 and RKO cells by 4-(4-(3-((6,7-bis(2-methoxyethoxy)quinazolin-4-yl)amino)phenyl)-1H-1,2,3-triazol-1-yl)phenyl sulfurofluoridate (P19G1). (A-B) Scratch assay and quantification were conducted to evaluate the A549 cells migration. A549 cells were treated with P19G1 (2.5 and 5 μM) and Erlotinib (2.5 and 5 μM) for 24 h and 48 h (A), and then scratches were photographed and quantified (B). (C-D) Scratch assay and quantification for the evaluation of migration in RKO cells. RKO cells were treated with P19G1 (1.25, 2.5 and 5 μM) and Erlotinib (1.25, 2.5 and 5 μM) for 24 h and 48 h (C), and then scratches were photographed and quantified (D). Scale bar: 50 μm. (E-F) Transwell migration assay for A549 and RKO cells. A549 and RKO cells were treated with P19G1 (5 and 10 μM) and Erlotinib (5 and 10 μM) for 48 h. Scale bar: 50 μm. (G-H) The number of migrated A549 cells and RKO cells was quantified. Data are presented as mean ± SD, n = 3, **P < .01, ***P < .001 versus Control group.
Promotion of Tumor Cell Apoptosis in A549 and RKO Cells by P19G1
Annexin V-FITC/PI apoptosis assays and Hoechst staining assays were used to evaluate the effect of P19G1 on promoting tumor cell apoptosis. For the Annexin V-FITC/PI apoptosis assay, A549 and RKO cells were treated with 5 μM and 10 μM P19G1 and Erlotinib for 48 or 72 h and then double-stained with PI and Annexin V-FITC to analyze cell apoptosis by flow cytometry (Figure 5A and B). Our results indicated that P19G1 significantly promoted A549 and RKO cell apoptosis (Figure 5C and D, ***P < .001), while Erlotinib did not notably promote apoptosis. For the Hoechst staining assay, 2.5 μM and 5 μM P19G1 and Erlotinib were incubated with A549 and RKO cells for 48 h and then stained with Hoechst fluorescent dye. The images were captured by fluorescence microscopy (Figure 5E and F). Our results showed that A549 and RKO cells were deep-colored and undergoing nuclear fragmentation in the P19G1 group.

Promoting tumour cell apoptosis in A549 and RKO cells by 4-(4-(3-((6,7-bis(2-methoxyethoxy)quinazolin-4-yl)amino)phenyl)-1H-1,2,3-triazol-1-yl)phenyl sulfurofluoridate (P19G1). (A and B) Flow cytometry analysis was performed to determine the A549 and RKO cell apoptosis. A549 (a) and RKO (b) cells were treated with P19G1 (5 and 10 μM) and Erlotinib (5 and 10 μM) for 48 h or 72 h. (C) Quantification of apoptosis rate in A549. A549 cells were incubated with P19G1 and Erlotinib in 5 μM (left) and 10 μM (right). (D) Quantification of apoptosis rate in RKO. RKO cells were incubated with P19G1 and Erlotinib in 5 μM (left) and 10 μM (right). (E) Hoechst staining to detect A549 cells apoptosis. A549 cells were treated with P19G1 (2.5 and 5 μM) and Erlotinib (2.5 and 5 μM) for 48 h. (F) Hoechst staining to detect RKO cells apoptosis. RKO cells were treated with P19G1 (2.5 and 5 μM) and Erlotinib (2.5 and 5 μM) for 48 h. Scale bar: 50 μm. Data are presented as mean ± SD, n = 3, ***P < .001 versus Control group.
Alterations in the Expression of Proteins Associated with Proliferation, Migration, and Apoptosis in A549 and RKO Cells Induced by P19G1
Western blotting was used to measure the expression changes of key proteins related to cell proliferation, migration, and cell cycle changes, including CDK-4, CDK-6, BCL-2, Caspase-3, MMP-2, and MMP-9 (Figure 6A and B). The experimental results showed that P19G1 (2.5 and 5 μM) significantly inhibited the expression of CDK-4, CDK-6, MMP-9 and MMP-2 proteins, indicating that it has a significant inhibitory effect on the cell cycle and migration. In addition, P19G1 (2.5 and 5 μM) inhibited the expression of BCL-2 and promoted the lysis of Caspase-3 protein, indicating that it promoted tumor cell apoptosis. Our results demonstrated that the antitumor mechanism of P19G1 inhibited the expression of cell proliferation- and migration-related proteins and increased the expression of apoptosis-related proteins.
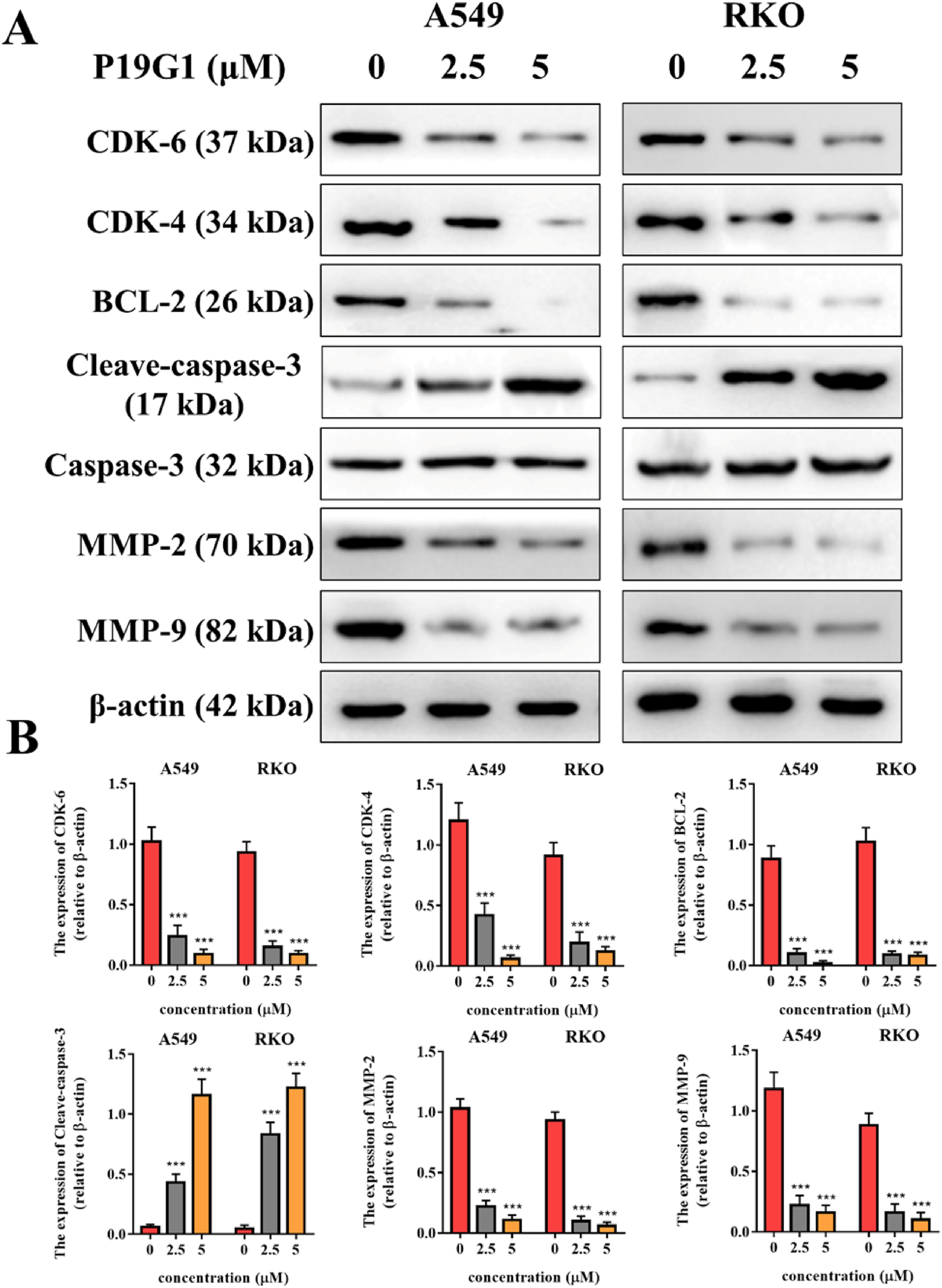
Alterations in the expression of proteins associated with proliferation, migration, and apoptosis in A549 and RKO cells by 4-(4-(3-((6,7-bis(2-methoxyethoxy)quinazolin-4-yl)amino)phenyl)-1H-1,2,3-triazol-1-yl)phenyl sulfurofluoridate (P19G1). A549 and RKO cells were incubated with P19G1 (2.5 and 5 μM) for 48 h. (A) Western blot of CDK-6, CDK-4, BCL-2, Cleaved-caspase 3, MMP-2, and MMP-9 proteins. (B) Quantification of Western blot results. **P < .001, ***P < .001 versus Control group. Data are presented as mean ± SD, n = 3.
Inhibition of Tumor Growth and Metastasis in a Xenograft Mouse Model by P19G1
We further examined the anticancer growth effect of P19G1 in vivo. The PK of the indicated P19G1 doses in plasma after single oral, IP, and intravenous (IV) administrations in SD rats were first evaluated. As shown in Figure 7A, P19G1 at 100 mg/kg was administered as a single oral dose in SD rats, and we found that the plasma concentration of P19G1 was less than 50 ng/mL. After IP injection of 5 mg/kg P19G1 in SD rats, the highest plasma concentration in rats was lower than 500 ng/mL, which was not enough to inhibit tumor growth. After tail vein injection of 5 mg/kg P19G1 in SD rats, the maximum plasma drug concentration was near 1700 ng/mL, while its metabolism was very fast. After 3 h of administration, the compound was completely undetectable in the plasma (Figure 7A). Based on these PK data of P19G1, IT injection was used to evaluate the antitumor activity of P19G1 in vivo.
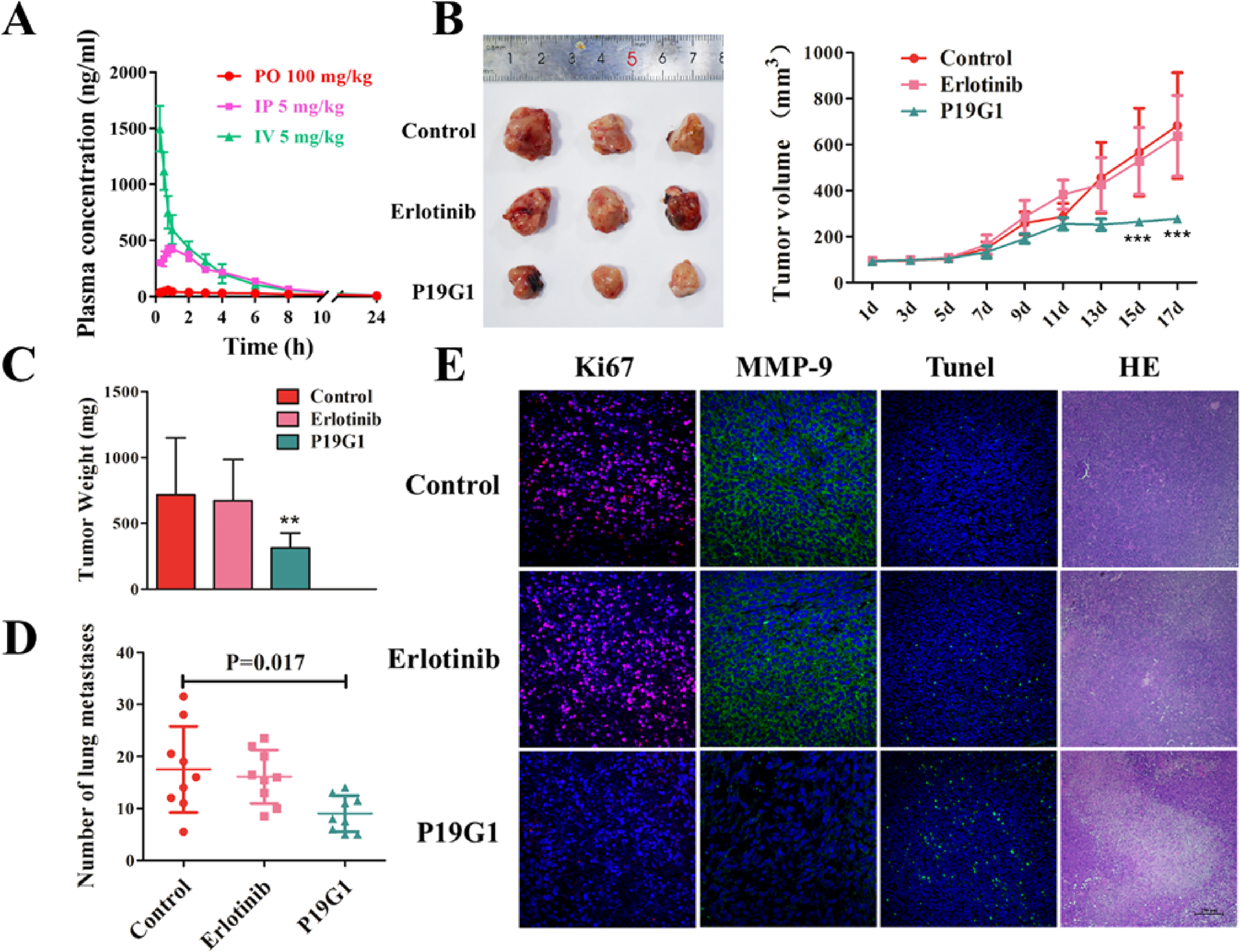
Inhibition of tumor growth and metastasis in a xenograft mouse model by 4-(4-(3-((6,7-bis(2-methoxyethoxy)quinazolin-4-yl)amino)phenyl)-1H-1,2,3-triazol-1-yl)phenyl sulfurofluoridate (P19G1). (A) Pharmacokinetics (PK) of indicated P19G1 doses in plasma after single oral (PO; red circles), intraperitoneal (IP; purple squares), and intravenous (IV; green triangle) administrations in SD rats (n = 3). (B) Representative tumors were excised at day 17 and tumor volume of A549 cell implants in nude mice treated with P19G1 (IT, 50 mg/kg, once per day) and Erlotinib (IT, 50 mg/kg, once per day) or vehicle (IT, once per day). Control versus P19G1 ***P < .001. (C) P19G1 significantly reduced tumor weight. Tumors were excised and weighed at the end of the experiment (day 17). Control versus P19G1 **P < .01. (D) P19G1 reduced the number of pulmonary nodules in nude mice xenograft model. Control versus P19G1 *P < .05. (E) Immunofluorescence (IF) analysis and hemotoxylin and eosin (H&E) staining analysis demonstrated that P19G1 significantly inhibits the proliferation (Ki67 staining) and migration (MMP9 staining) and promotes apoptosis (Tunel staining and H&E staining) in A549 tumor bearing BALB/C nude mice. Scale bar: 250 μm.
Using A549 cell lines to establish a xenograft mouse model, our experimental results showed that the IT administration of 50 mg/kg P19G1 significantly suppressed the tumor size at 15 and 17 days after administration (P < .001, Figure 7B) and weight at 17 days after administration (P < .01, Figure 7C) compared with the control group. Meanwhile, the lung nodules, which represent lung metastases of tumors in the xenograft mouse model (P = .017, Figure 7D), also decreased compared with the control group. While under the same conditions, the Erlotinib group had no obvious inhibitory effect on the xenograft mouse model.
To further clarify the antitumor mechanism of P19G1 in vivo, we performed section staining analysis of xenograft mouse tumor tissues (Figure 7E). Ki67 IF staining results showed that P19G1 significantly inhibited the expression of Ki67 in tumor tissue compared with the control group, indicating that the growth of tumor tissue was inhibited. Through an MMP9 IF staining assay, we found that P19G1 significantly inhibited the expression of MMP9 protein in tumor tissues, indicating that tumor cell migration in tumor tissue was suppressed. Furthermore, P19G1-induced apoptosis in tumor tissues was supported by increased TUNEL-positive staining and changed tumor cell morphology in H&E staining. Together, these results indicate that P19G1 has potent antitumor activity in vivo by inhibiting tumor cell proliferation and migration and promoting tumor cell apoptosis.
Discussion
The azide–alkyne cycloaddition (AAC) is the most widely used reaction type in cycloaddition reactions. 30 It is often used for the rapid construction of 1,2,3-triazole. 31 However, this type of addition reaction did not receive the attention of scientists in the first 100 years after it was discovered owing to the lack of regioselectivity, higher reaction temperature, and longer reaction time. 32 In 2001, Sharpless et al discovered that CuAAC could occur at room temperature, and the reaction once again attracted the attention of scientists. 4 Through development in recent years, CuAAC has become a favorable tool for discovering lead compounds due to its complete specificity, high reliability, and biocompatibility of reactants and is widely used in the anticancer, antiviral, and antibacterial fields. 33
Erlotinib is an inhibitor of the protein tyrosine kinase EGFR containing a quinazoline backbone. Through reversible binding to EGFR, it competitively inhibits the binding of EGFR to ATP and subsequent phosphorylation, thereby blocking the proliferation and anti-apoptosis signaling pathways of a variety of tumor cells and inhibiting tumor growth. However, EGFR mutations and drug resistance limit the application of erlotinib. 34 There are currently 4 generations of inhibitors developed for EGFR. Clicking on the chemical modification of the Erlotinib quinazoline skeleton to find a new generation of EGFR-targeting lead compounds is the focus of this research.
The reaction of Erlotinib with azides was conveniently carried out due to the terminal alkynes in its structure. As a result, 840 new 1,2,3-triazole-containing Erlotinib derivatives were obtained by Click chemistry and our azide library. According to our research on click compounds, the purity of each of these 840 compounds should be greater than 80%. 6 Therefore, these compounds were directly subjected to cell experiments without separation and purification. In A549 cells, 840 Erlotinib derivatives at a concentration of 10 μM were preliminarily screened for cell viability by MTT assay, and 22 compounds were found to inhibit cell viability by more than 60% relative to Erlotinib. Subsequently, these 22 compounds were synthesized to obtain pure products, and their IC50 values were measured in A549 cells. We found that the IC50 value of Compound P19G1 was 2.13 μM, and the IC50 value of Erlotinib was 40.20 μM, indicating that P19G1 has the best inhibitory activity. In 1H NMR of P19G1, chemical shifts in the range of 7 to 9 belong to the hydrogens in the aromatic region, and those in the range of 3 to 4 belong to the aliphatic hydrogens, where the peak at chemical shift 7.65 is split into a doublet and the peak at 7.52 is split into a triplet. An evaluation of the inhibitory effect of P19G1 on many other tumor cells showed that P19G1 exhibited broad-spectrum inhibitory activity in H1299, HeLa, RKO, and MCF-7 cells. The IC50 values of these cells were between 1-5 μM, and the detailed experimental results are shown in Figure 2.
Changes in tumor cell activity are regulated by multiple mechanisms. Furthermore, to investigate the antitumor mechanism of P19G1, we conducted colony formation experiments, Ki67 IF staining experiments, scratch experiments, Transwell migration experiments, Hoechst fluorescent staining experiments, and flow cytometry experiments in A549 and RKO cells. We proved that P19G1 has antitumor effects by inhibiting tumor cell proliferation and migration and promoting tumor cell apoptosis. Moreover, the antitumor mechanism of P19G1 was further demonstrated by Western blot experiments, which revealed that P19G1 altered the expression of proliferation-, migration- and apoptosis-related proteins in A549 and RKO cells. In vivo studies, we noticed the poor PK properties of P19G1, possibly due to the fact that the addition of some groups to the Erlotinib by click chemistry has increased the molecular weight and changed its lipid/water partition coefficient, thus reducing the solubility of P19G1. Despite the enhanced activity in vitro compared to Erlotinib, oral and IP administrations in vivo is still less effective against tumors, therefore, IT injection was used in this study to observe its effect in animal models. Our data revealed that 50 mg/kg P19G1 effectively inhibited the growth and metastasis of A549 tumor cells compared with the control group. However, under the same experimental conditions, the inhibitory effect of Erlotinib was not obvious. The staining results further elucidate the antitumor mechanism of P19G1 in vivo; the compound significantly inhibits the proliferation and migration of tumor cells in tumour masses and promotes tumor cell apoptosis in tumor masses.
It is well known that the antitumor target of Erlotinib is EGFR, but it is still unclear whether the antitumor target of P19G1 after structural modification is changed. In the following research, we will further investigate the antitumor target of P19G1 to find out the reason for its enhanced antitumor activity, which will lay the foundation for further studies.
Conclusions
In conclusion, we synthesized and identified a novel Erlotinib derivative, P19G1, by screening a library of Erlotinib derivatives synthesized by a modular click-chemistry approach, which showed stronger antitumor activity than Erlotinib in tumor cells and xenograft mouse models. Our data revealed that the rapid structural modification of lead compounds using novel modular click-chemistry reactions holds great potential for use in obtaining diverse derivatives for tumor drug screening and development. The acquisition of P19G1 is also an antitumor candidate molecule worthy of development.
Footnotes
Acknowledgments
This work was supported in part by the Shanghai Municipal Health Commission/Shanghai Municipal Administration of Traditional Chinese Medicine (ZY(2021-2023)-0501); Shanghai Frontiers Science Center for Traditional Chinese Medicine Chemical Biology.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the State Administration of Traditional Chinese Medicine of People's Republic of China, the Gansu Province Science Foundation for Distinguished Young Scholars, and National Natural Science Foundation of China (grant numbers GZYYGJ2019059, 20JR10RA348, and 81803779).
Ethical Approval
All animal protocols and procedures were approved by the Shanghai University of Traditional Chinese Medicine Institutional. Animal Care and Use Committees and were carried out in accordance with the guidelines outlined in the Guide for the Care and Use of Laboratory Animals published by the National Institute of Health. We make efforts to minimize the number of animals utilized and to decrease their suffering. All experiments were carried out in compliance with ARRIVE guidelines. Animal ethics number: PZSHUTCM200410003.
Supplemental Material
Supplementary material for this article is available online.
Abbreviations
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
