Abstract
During autophagy, autophagosomes are formed to engulf cytoplasmic contents. p62/SQSTM-1 is an autophagic adaptor protein that forms p62 bodies. A unique feature of p62 bodies is that they seem to directly associate with membranous structures. We first investigated the co-localization of mKate2-p62 bodies with phospholipids using click chemistry with propargyl-choline. Propargyl-choline-labeled phospholipids were detected inside the mKate2-p62 bodies, suggesting that phospholipids were present inside the bodies. To clarify whether or not p62 bodies come in contact with membranous structures directly, we investigated the ultrastructures of p62 bodies using in-resin correlative light and electron microscopy of the Epon-embedded cells expressing mKate2-p62. Fluorescent-positive p62 bodies were detected as uniformly lightly osmificated structures by electron microscopy. Membranous structures were detected on and inside the p62 bodies. In addition, multimembranous structures with rough endoplasmic reticulum–like structures that resembled autophagosomes directly came in contact with amorphous-shaped p62 bodies. These results suggested that p62 bodies are unique structures that can come in contact with membranous structures directly:
Introduction
The autophagy–lysosome system is a bulk degradation system of intracellular components including organelles. It has a significant relationship with many diseases such as neurodegenerative diseases, cardiomyopathy, hepatitis, carcinogenesis, type II diabetes, immune diseases, and infectious diseases.1,2 During autophagy, an isolation membrane is formed to engulf cytoplasmic components, and finally enclosed to form the autophagosome. The autophagosome then fuses to a lysosome, and intra-autophagosomal contents are degraded by lysosomal acid hydrolases. During autophagosome formation, microtubule-associated protein light chain 3 (LC3) is conjugated to phosphatidylethanolamine via Atg7 and Atg3, and localized to isolation/autophagosomal membranes.3–8
p62/SQSTM1 is identified as an LC3-interacting protein that is essential for selective autophagy.9,10 Its structure features four distinct domains: PB1, LIR, KIR, and UBA.11–13 The LIR domain is essential for the interaction of p62 with LC3. 13 The KIR domain interacts with Keap1 essential for oxidative stress sensor, the Keap1-Nrf2 pathway. p62 recognizes ubiquitylated proteins via the UBA domain and recruits to the autophagosome via LC3–p62 interaction in the LIR domain.9,10 p62 self-oligomerizes and forms liquid droplets via the PB1 domain in the cells. Recently, it was reported that overexpressed GFP-p62 in the cells form gels with low liquidity. 14 Impairment of the autophagy–lysosome system results in the accumulation of p62 aggregates.9,15 We have reported that rough-surfaced endoplasmic reticulum partially surrounds p62-positive fibrous inclusions in Atg9-deficient brain. 15
Considering the function of p62 in selective autophagy, it is possible that p62 bodies can directly contact membranous structures. In this study, we investigated the ultrastructures of p62 bodies in the cells by in-resin correlative light and electron microscopy (CLEM) of Epon-embedded cells 16 and found that p62 bodies directly come in contact with membranous structures.
Materials and Methods
Cells, Media, and Materials
HeLa cells were obtained from the American Type Culture Collection and were cultured in Dulbecco’s Modified Eagle’s Medium (045-30285; Fujifilm Wako Chemicals, Tokyo, Japan) containing 10% fetal calf serum (12603C; JRH Biosciences/Sigma-Aldrich, Lenexa, KS). Cells were labeled with a propargyl-choline (AOBT7378; AOBIOUS, Gloucester, MA), and copper-catalyzed azide/alkyne cycloaddition was performed according to the manufacturer’s protocol using Alexa Fluor 647 azide (A10277 and C10269; Thermo Fisher Scientific, Eugene, OR). FuGENE HD (E2311; Promega, Madison, WI) transfection reagent was used to introduce the plasmid into cells. For the expression of p62 (GenBank Ac. No. NP_001135771) tagged with mKate2-GGGGSGL at the amino terminus under the control of the CAG promoter, the DNA fragment encoding p62 protein was introduced into an expression plasmid pCAG-mKate2-G (pCAG-CLEM-Red-p62). 16
Sample Preparation, Fluorescent Microscopy, and Electron Microscopy for In-resin CLEM
Sample preparation for in-resin CLEM was performed as described previously. 16 Briefly, cells expressing mKate2-p62 proteins were prefixed with 2.5% glutaraldehyde buffered in 0.1 N phosphate buffer, pH 7.2, at 4C for 1 hr. The fixed cells were washed twice with 0.1 N phosphate buffer, pH 7.2, and postfixed in 1% osmium tetroxide at 4C for 10 min. Fixed cells were incubated in TUK solution (209-20851; Fujifilm Wako Chemicals) at 4C for 10 min. Cells were then dehydrated with a graded series of ethanol and embedded in Epon812 (Oken Shoji, Chiyoda-ku, Tokyo, Japan) at 60C for 72 hr. Thin sections (100 nm) were cut with an ultramicrotome UC6 (Leica Biosystems, Nussloch, Germany) and placed on glass cover slips that were coated with Pt/Au. Sections were observed in a TUK solution using either a BZ-X710 fluorescence microscope (FM; Keyence, Osaka-shi, Osaka, Japan) with a structured illumination microscopy unit or a Nikon A1 confocal FM (Nikon, Minato-ku, Tokyo, Japan). Thereafter, sections were stained with uranyl acetate and lead citrate. Scanning electron microscope (SEM) images were obtained via a JSM-7900F SEM (JEOL, Akishima, Tokyo, Japan) or a Helios NanoLab 660 (FEI, Hillsboro, OR). Transmission electron microscope (TEM) images were obtained via a JEM-1400 flash TEM (JEOL). Line profiles of fluorescent images were analyzed with a BZ-X image analyzer software (Keyence).
Results
Propargyl-choline-labeled Phospholipids Were Detected Inside mKate2-p62 Bodies
To investigate whether or not p62 bodies come in contact with phospholipids, we metabolically labeled phospholipids with propargyl-choline because it incorporates strongly into phospholipids in whole animals. HeLa cells expressing mKate2-p62 fusion proteins were incubated with propargyl-choline for 24 hr, fixed with 4% paraformaldehyde, and propargyl-choline-labeled phospholipids in the cells were reacted with Alexa Fluor 647 azide via copper-catalyzed azide/alkyne cycloaddition (Fig. 1). Red fluorescence of mKate2-p62 was detected in a round shape. In some mKate2-p62 bodies, fluorescent signals of Alexa Fluor 647 propargyl-choline-labeled phospholipids were detected (Fig. 1B). Line profiling of these labeled phospholipids and the p62 bodies in fluorescent images revealed that Alexa Fluor 647 signals were detected in mKate2-p62 bodies (Fig. 1B). These results suggested that phospholipid-based structures, that is, membranous structures, are able to localize inside the mKate2-p62 bodies.
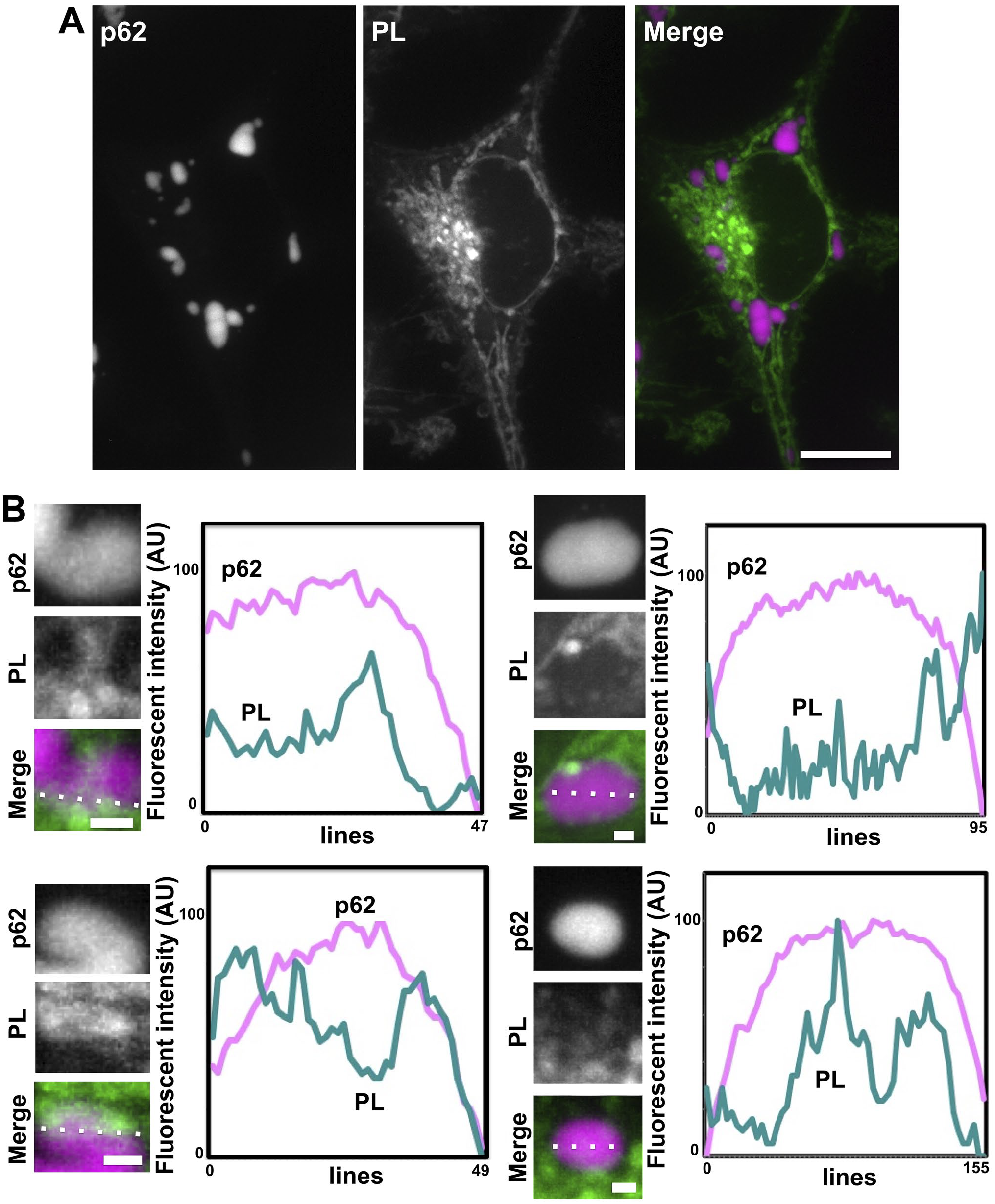
Propargyl-choline-labeled phospholipids were detected in mKate2-p62 bodies. (A) Phospholipids in HeLa cells expressing mKate2-p62 were detected using a click chemistry. The cells were metabolically labeled with propargyl-choline for 24 hr and fixed with a 4% paraformaldehyde at 4C for 1 hr. Cells were reacted with Alexa Fluor 647 azide via a copper-catalyzed click reaction. Fluorescent images of mKate2-p62 and Alexa Fluor 647–labeled phospholipids (PL) were obtained with a BZ-X810 fluorescence microscope with Structured Illumination Microscopy unit (CCD monochrome camera, NIKON Apochromat 100× lens, gain +8 dB, haze reduction). The “Merge” image is a merged image of p62 (magenta pseudo color) and PL (green pseudo color). Scale bars, 10 µm. (B) Line profiles of four independent p62 bodies and Alexa Fluor 647–positive signals in fluorescent images were analyzed. Relative fluorescent intensity of p62 (magenta-colored lines) and phospholipid (green-colored lines) on dotted lines in “Merge” images were analyzed with a BZ-X810 analyzer software. Scale bars, 0.5 µm.
Fluorescent-positive mKate2-p62 bodies Were Detected as Lightly Osmificated Structures by In-resin CLEM of Epon-embedded Samples
To clarify whether the p62 bodies come in contact with membranous structures, we performed in-resin CLEM of Epon-embedded cells expressing mKate2-p62. Fluorescent signals of mKate2-p62 in the 100-nm-thin section were detected as round-shaped fluorescent signals by an FM (Figs. 2A and 3A and F). The ultrastructures of mKate2-p62 in the same section were observed by an electron microscope (EM; Figs. 2A and 3A and F). The ultrastructures of mKate2-p62 corresponding to red fluorescent-positive signals were uniformly lightly osmificated structures (Figs. 2B and 3B).
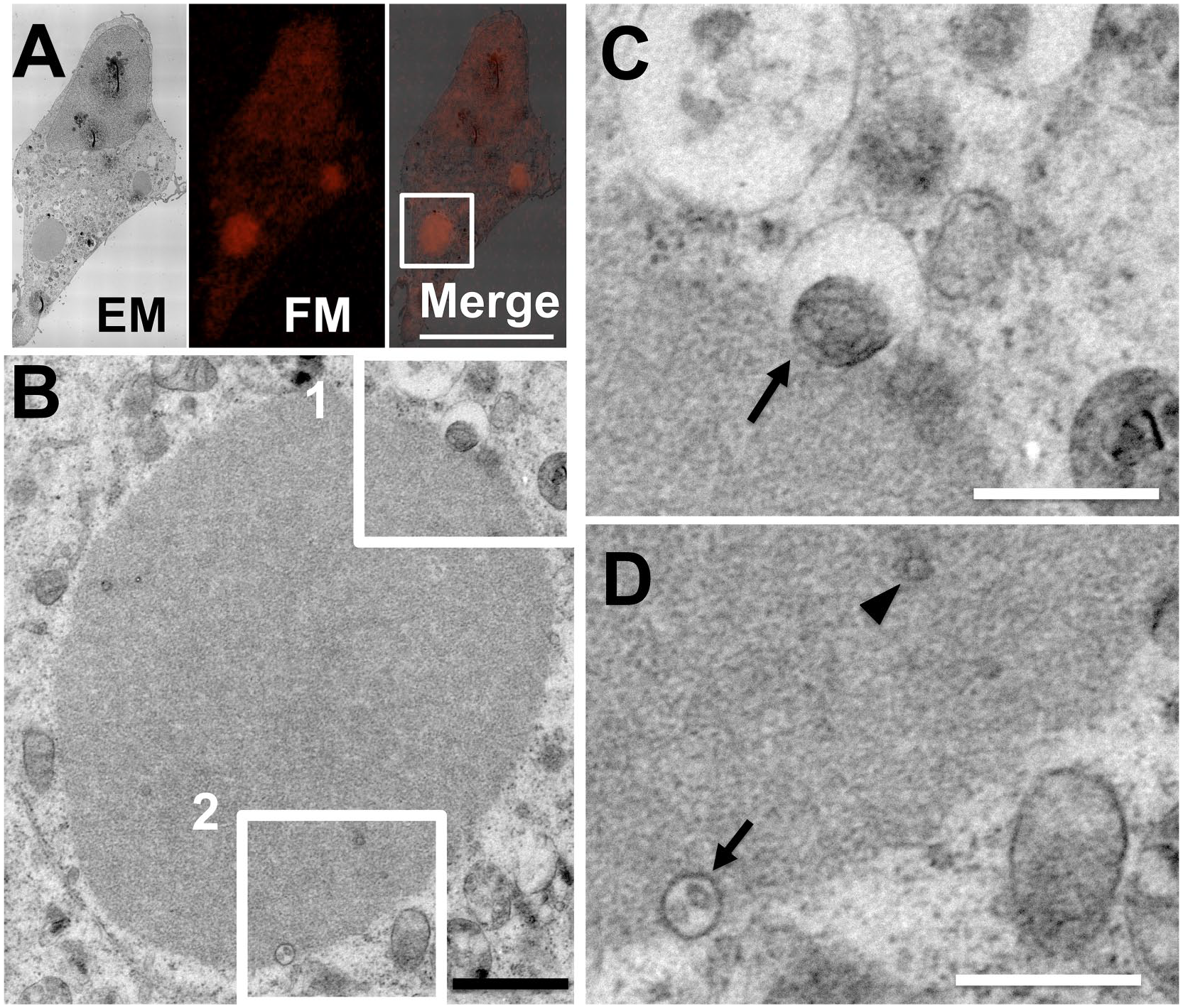
Fluorescent-positive mKate2-p62 bodies were detected as lightly osmificated structures. HeLa cells expressing mKate2-p62 were fixed with a mixture of paraformaldehyde and glutaraldehyde at 4C for 1 hr. After osmium staining and dehydration, cells were embedded in Epon812 resins. A thin section (100 nm) of Epon-embedded cells was prepared. A fluorescent image of the thin section was obtained with a Nikon A1 confocal fluorescence microscope (FM; A). Electron microscopic (EM) image of the same sample was obtained with JEM-1400 Flash transmission electron microscope (JEOL). The “Merge” is a merged image of the FM image with an EM image. Scale bar, 25 µm. The image in (B) indicates magnification of an image corresponding to the boxed area in the Merge image in (A). Scale bar, 1 µm. The images in (C) and (D) indicate higher magnification of images corresponding to the boxed areas 1 and 2, respectively, in (B). Scale bars, 0.5 µm. Arrows indicate membranous structures associated with p62 bodies. Arrowhead indicates a membranous structure in p62 bodies.
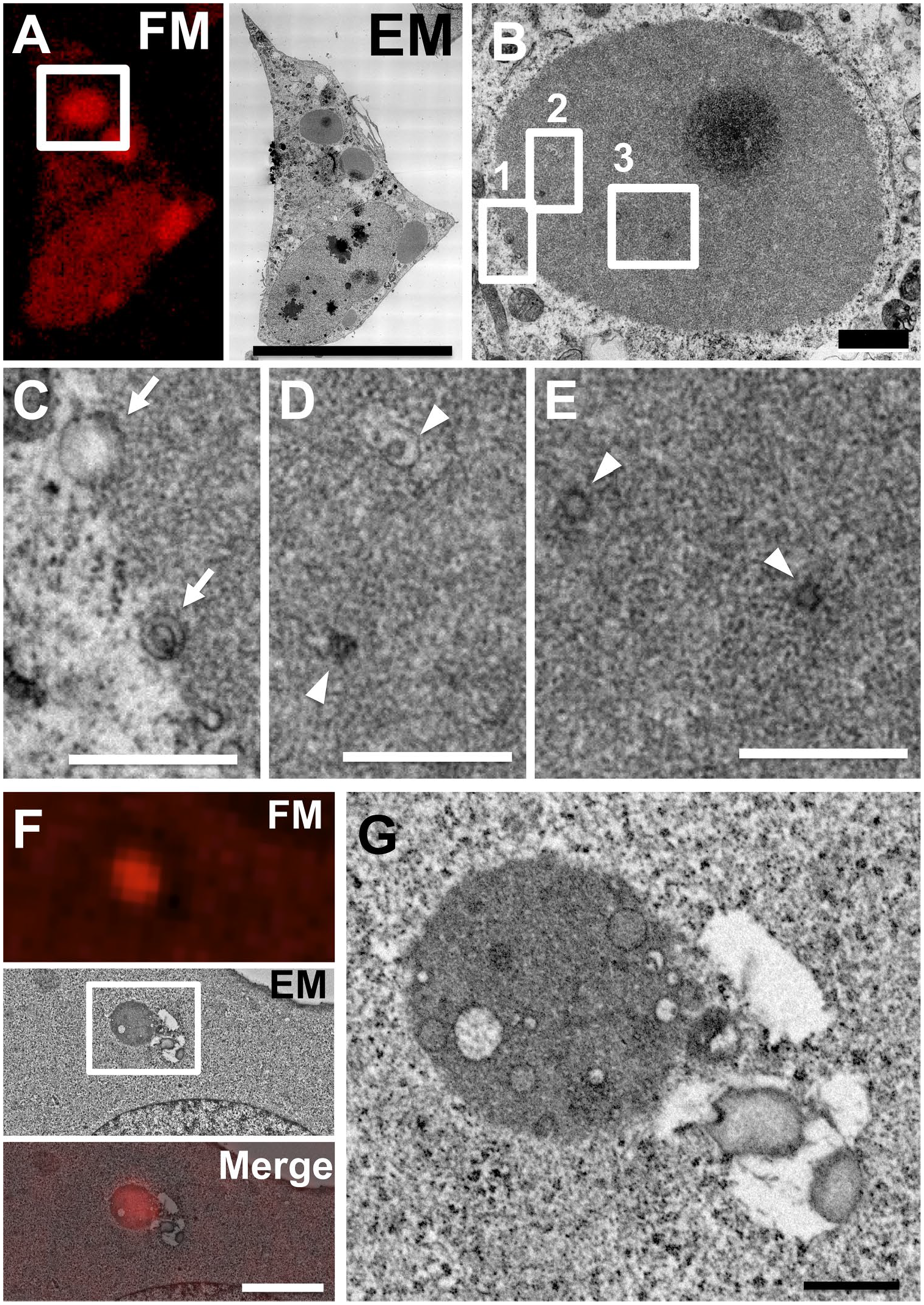
Membranous structures were observed in p62 bodies. Fluorescent images of the thin section were obtained with a Nikon A1 confocal fluorescence microscope (FM). Electron microscopic (EM) images corresponding to the same section are shown in (A) and (F), respectively (A–E, JEM-1400 Flash transmission electron microscope, JEOL; F and G, a Helios NanoLab 660, FEI). Scale bar in (A) and (F), 25 and 4 µm respectively. The images in (B) and (G) indicate magnification of images corresponding to the boxed areas in (A) and (F), respectively. Scale bars, 1 µm. The images in (C), (D), and (E) indicate higher magnification of images corresponding to the boxed areas 1, 2, and 3, respectively, in (B). Scale bars, 0.5 µm. Arrows indicate membranous structures associated with p62 bodies. Arrowheads indicate membranous structures in p62 bodies.
The p62 Bodies Come in Contact With Membranous Structures
We further focused on the ultrastructures on the surface of p62 bodies by in-resin CLEM of Epon-embedded cells. Osmificated membranous structures were detected on p62 bodies (Figs. 2C and D and 3C). The diameter of membranous structures on the surface of p62 bodies is about 611 ± 150 nm (mean ± SE). In addition to their surface, small vesicles and tubulovesicular structures were detected inside mKate2-p62 bodies (Figs. 2D and 3D, E, and G). The diameter of membranous structures in p62 bodies is about 108 ± 13 nm (mean ± SE). The membranous structures in p62 bodies are smaller than those on their surface (p value of Welch’s t-test <0.01).
In the case of p62 bodies, an amorphous-shaped structure was observed (Fig. 4). In-resin CLEM revealed that the amorphous p62 bodies directly contacted with an invasive multilamellar structure, resembling an autophagosome (Fig. 4C). In addition, rough endoplasmic reticulum–like structure was associated with multilamellar structures (Fig. 4C). These results indicated that p62 bodies have unique properties that come in contact with membranous structures.
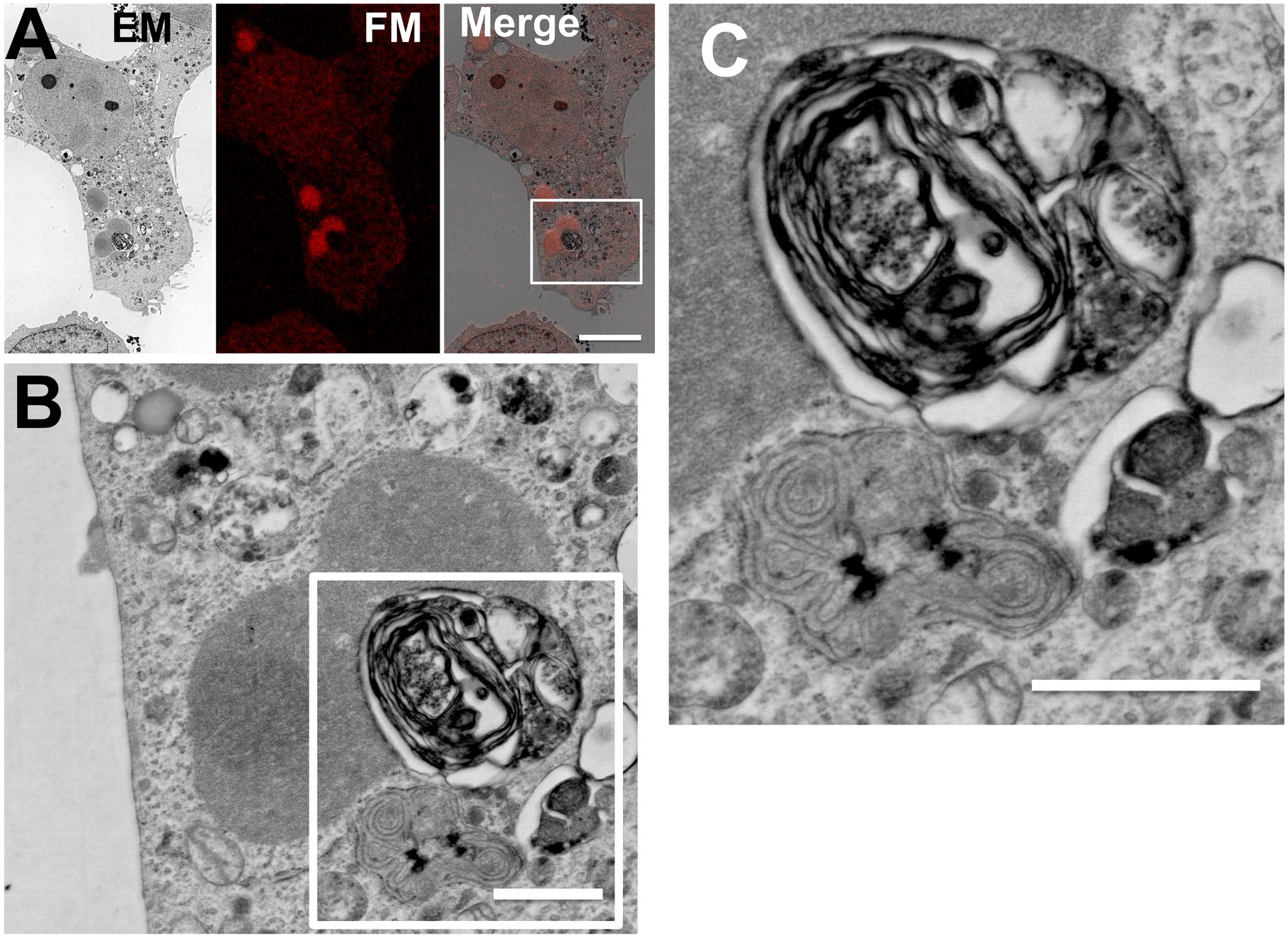
Multimembranous structures with rough endoplasmic reticulum–like structures that resembled autophagosomes make contact with the p62 bodies directly. (A) A fluorescent image of 100-nm-thin section of Epon-embedded cell was obtained by a Nikon A1 confocal fluorescence microscope (FM), and the electron microscopic (EM) image of the same section was obtained by a JSM-7900F scanning electron microscope (JEOL). The “Merge” is a merged image of the FM image with an EM image. Scale bar, 25 µm. The image in (B) indicates magnification of an image corresponding to the boxed area in the image in (A). Scale bar, 1 µm. The image in (C) indicates higher magnification of an image corresponding to the boxed areas in (B). Scale bars, 1 µm.
Discussion
We show evidence that p62 bodies come in contact with membranous structures. When cells expressing mKate2-p62 were incubated with propargyl-choline, propargyl-choline mainly incorporated into phosphatidylcholine. Phosphatidylcholine is one of the major phospholipids and is distributed in the lipid of the membranes to form phospholipid-based organelles and membranous structures. Incorporated propargyl-choline in the phospholipids was reacted with Alexa Fluor 647 azide via a copper-catalyzed click reaction. Therefore, Alexa Fluor 647–positive signals suggested the presence of membranous structures in p62 bodies (Fig. 1). Furthermore, we identified for the first time the ultrastructures of p62 bodies by in-resin CLEM of Epon-embedded cells. Compared with traditional CLEM and immunoelectron microscopy, in-resin CLEM of Epon-embedded cells has an advantage to preserve the intracellular structures and the accuracy of its correlation in the thin sections.17,18 We also found that multimembranous structures with rough endoplasmic reticulum–like structures that resembled autophagosomes were able to contact the p62 bodies.
p62 plays important roles in selective autophagy. During autophagy, the dynamic membrane formation occurs. Membranous structures in p62 bodies were observed under the fed conditions. Therefore, this ability of p62 bodies to come in contact with membranous structures in addition to interacting with ubiquitylated proteins could be essential for the formation of the isolation/autophagosomal membrane during selective autophagy. Inside p62 bodies, the content of some vesicles has a similar density as the p62 bodies themselves. However, other vesicles have content with a different density when observed in EM images, especially in Fig. 3G. p62 interacts with ubiquitylated proteins, Keap1 and LC3. The difference in the density of the contents of the vesicles would derive from the difference in p62-interacting proteins. Considering that p62 is essential for selective autophagy, we will further investigate these membranous structures in and on p62 bodies during selective autophagy by morphological and biochemical approaches.
Footnotes
Acknowledgements
We thank Dr. Tomoko Takahashi for construction of plasmids.
Competing Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
IT and YU conceived of and designed the study. IT, TH, MS, ST, AT, YF, JY, JAOT, and SK carried out the experiments. IT and YU wrote the manuscript with input from all other authors. All authors read and approved the final manuscript.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work is partly supported by the Project for Elucidating and Controlling Mechanisms of Aging and Longevity from the Japan Agency for Medical Research and Development (AMED JP19gm5010003 and 20gm5010003 to Y.U) and by the MEXT-supported Program for the Strategic Research Foundation at Private Universities (to Y.U). This work was also supported by grants from the Japan Society for the Promotion of Science to I.T. (JSPS KAKENHI 15K12749 and 20H05342) and J.Y. (JSPS KAKENHI 20K22744) and the Research Institute for Diseases of Old Age, Juntendo University School of Medicine to I.T., S.K., and J.A.O.T.
