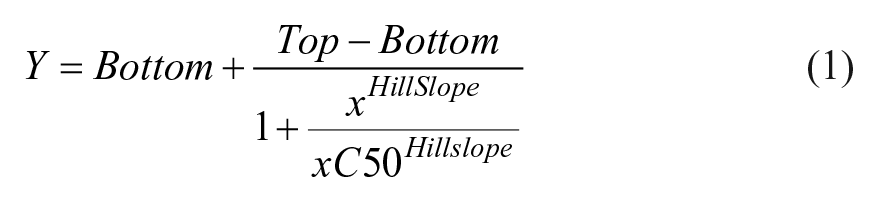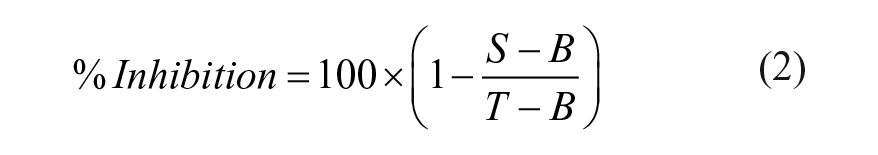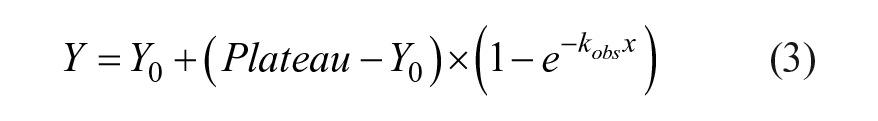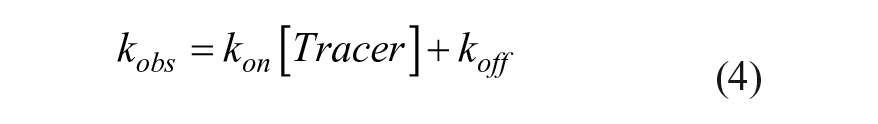Abstract
Protein kinases are intensely studied mediators of cellular signaling. While traditional biochemical screens are capable of identifying compounds that modulate kinase activity, these assays are limited in their capability of predicting compound behavior in a cellular environment. Here, we aim to bridge target engagement and compound-cellular phenotypic behavior by utilizing a bioluminescence resonance energy transfer (BRET) assay to characterize target occupancy within living cells for Bruton’s tyrosine kinase (BTK). Using a diverse chemical set of BTK inhibitors, we determine intracellular engagement affinity profiles and successfully correlate these measurements with BTK cellular functional readouts. In addition, we leveraged the kinetic capability of this technology to gain insight into in-cell target residence time and the duration of target engagement, and to explore a structural hypothesis.
Keywords
Introduction
Kinases are among the most successful target classes in drug discovery, being targeted with small-molecule inhibitors in diverse therapeutic areas.
1
Bruton’s tyrosine kinase (BTK) is a member of the Tec family of nonreceptor tyrosine kinases and is validated as a key pathological driver in autoimmune diseases such as rheumatoid arthritis (RA) and multiple sclerosis.
2
Drug discovery efforts have resulted in at least one marketed kinase drug (ibrutinib), which binds irreversibly to BTK via cysteine 481 in the kinase domain (
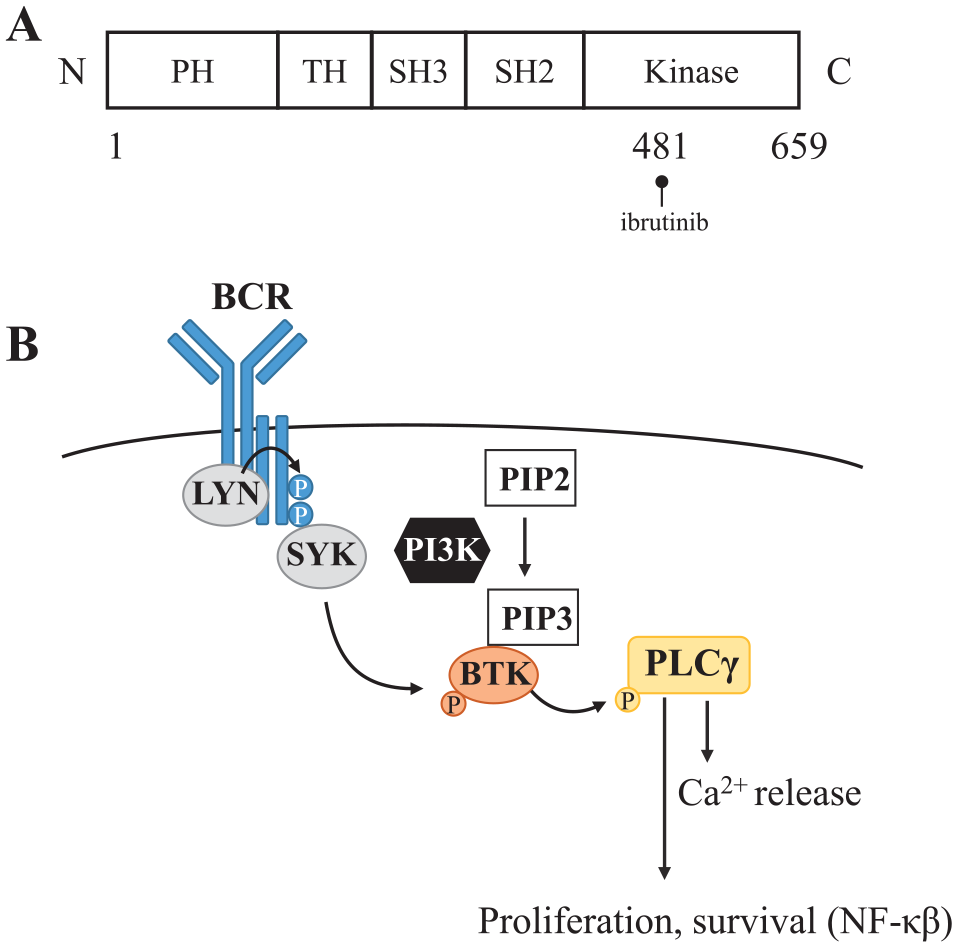
BTK structure and signaling pathway. (
In the absence of cellular target occupancy readouts, cellular BTK activity is traditionally measured via pathway analysis methods. The BTK-mediated signaling events that link B-cell receptor (BCR) ligation to downstream signaling have been well characterized (
Here, we have applied a quantitative method to measure BTK engagement in cells, and to evaluate the relationship between cellular target occupancy and modulation of second messenger responses downstream of BTK. Toward these goals, we applied the NanoBRET14,15 cellular target engagement technology to assess direct BTK target occupancy in real time in a cellular context. Accordingly, we successfully miniaturized the BTK NanoBRET assay to 384-well plate format and determined the optimal conditions for the real-time analysis of BTK residence time. Using our adapted NanoBRET assay system, we were able to triage BTK inhibitors based on intracellular binding affinity. We observed a quantitative correlation between target occupancy and cellular functional activity across a diverse chemical set of BTK inhibitors. We further extended the analysis to a non-steady-state system, where the duration of BTK occupancy can be quantified for select inhibitors. Among the tested compounds, we characterized a reversible inhibitor with durable and potent occupancy of BTK in living cells, providing proof of concept that an extended intracellular engagement of BTK can be achieved via noncovalent and covalent mechanisms of action.
Materials and Methods
Cell Transfection and Cryopreservation
Cryo-banked cells were used in the bioluminescence resonance energy transfer (BRET) target engagement experiment. HEK293 cells (ATCC, Manassas, VA) were transfected with BTK-NanoLuc fusion plasmids (pF31K [CMV/Neo], Promega, Madison, WI) using FuGENE HD transfection reagent (Promega) according to the manufacturer’s protocol. First, BTK-NanoLuc DNA was mixed with transfection carrier DNA (pGEM3ZF vector, Promega) in a 1:10 mass ratio for a final total DNA concentration of 10 μg/mL in phenol red free Opti-MEM (Gibco, ThermoFisher Scientific, Gaithersburg, MD). FuGENE was added into the DNA solution for a final 3% (v/v) concentration, after which the resultant solution was incubated at room temperature for 15 min to form the transfection complex. HEK293 cells were resuspended at 200,000 cells/mL in DMEM (Gibco) supplemented with 10% fetal bovine serum (HyClone, GE Healthcare, Chicago, IL) and 1% HEPES (Gibco). The cells were transfected with the transfection complex solution at a volume ratio of 1:20 (complex–cell suspension), after which cells were plated into a flask and incubated overnight at 37 °C with 5% CO2 supplementation.
Following overnight transfection, cells were harvested with 0.05% trypsin/EDTA (Life Technologies, ThermoFisher Scientific, Carlsbad, CA) and resuspended to a density of 1 million cells/mL in Recovery cell freezing media (Gibco). Cells were aliquoted to cryo-vials (Corning, Corning, NY) and were frozen first at −80 °C in a CoolCell storage unit (Corning) before being transferred to long-term storage in liquid nitrogen.
Steady-State (Endpoint) BRET Measurements
The NanoBRET assay was miniaturized from a 96-well format 16 by employing automated liquid dispensing. BRET measurements were made in white 384-well nonbinding surface plates (Corning). Cells were thawed and resuspended at 200,000 cells/mL in Opti-MEM without phenol red (Gibco) before seeding in a 384-well plate for a final count of 10,000 cells/well. Chemical compounds were resuspended in DMSO (Sigma-Aldrich, St. Louis, MO) and transferred to assay plates via acoustic dispensing (EDC Biosystems, Fremont, CA).
For the live-cell assay conditions, the plates were incubated for 2 h at 37 °C with 5% CO2 supplementation. NanoBRET NanoGlo Substrate and Extracellular NanoLuc Inhibitor (Promega) were added according to the manufacturer’s guidelines. Assay plates were incubated at room temperature for 2 h before adding NanoBRET NanoGlo Substrate (Promega).
For all BRET experiments, the Envision 2015 (PerkinElmer, Waltham, MA) equipped with a BRET2 Enh mirror and BRET 647 nM/75 nM Bandwidth and BC703 460 nM/80 nM Bandwidth filters was used for readouts. Each read was integrated for 0.5 s.
Probe Kd Determination
Affinity for probe 4 or 5 16 for BTK-NanoLuc in the HEK293 cells was determined by titration. Briefly, a twofold dilution series was prepared in DMSO, after which the entire dilution series was diluted with four parts probe dilution buffer (Promega) to prepare a 20× working probe reagent. This dilution series was titrated onto BTK-NanoLuc-expressing cells and equilibrated in the presence and absence of a saturating dose (10–20 µM) of corresponding unlabeled probe parent compound for each probe. BRET was measured as described above. BRET ratios were converted to milliBRET (mBRET) values by multiplying by 1000, after which the mBRET values were plotted as a function of probe concentration. The apparent affinity of the probe (EC50) was determined by fitting to the four-parameter dose–response equation (eq 1) using GraphPad Prism software (GraphPad, La Jolla, CA).
Compound Target Engagement Measurements at Equilibrium
Chemical inhibitors were synthesized in-house as previously described.4,5,17–20 Compounds were diluted threefold in DMSO for a total of 11 points. Final assay conditions used a high concentration of 20−50 μM for most compounds. Probe 4 was used for equilibrium measurements, employing a final concentration of approximately EC50–80. For live-cell measurements, 100 μM probe 4 was added. Plates were incubated and BRET was measured as indicated above.
To determine compound potency (IC50), BRET ratios for each dilution series were first normalized to a percent inhibition value via eq 2, where S represents sample BRET, B represents background BRET, and T represents total BRET. In this equation, the background control (B) consists of BTK-NanoLuc-expressing cells and no probe 4. The total BRET ratio (T) consists of BTK-NanoLuc-expressing cells, probe 4, and an amount of DMSO at a concentration equivalent to that added when testing compounds.
Following normalization, the percent inhibition values are plotted as a function of compound concentration in GraphPad Prism and fit to eq 1.
Kinetic Probe Characterization and Cellular Residence Time Assay
All kinetic parameters were assessed using adherent HEK293 cells transfected as described above. Probe 5 was used for kinetic experiments due to its fast association kinetics in cells. HEK293 cells in Opti-MEM medium containing 1% FBS and transfected with BTK-NanoLuc were plated in 96-well tissue culture-treated plates (Corning) and incubated overnight.
The association kinetics of probes were characterized in kinetic dose–response experiments. A probe dilution series was added to the cells and BRET was measured kinetically on a Glomax Discover instrument (Promega) at 3 min intervals using detection reagents as described above. BRET data were plotted as a function of time and fit to the one-phase association equation in GraphPad Prism (eq 3) to obtain an observed rate constant (kobs) for each compound.
The kobs values were replotted as a function of probe concentration, after which the kon and koff values were determined by linear regression using the observed rate constant equation (eq 4), where koff is apparent from the y intercept and kon is apparent from the slope.
Cellular residence time was assessed via cellular washout. After transfection, adherent HEK293 cells expressing BTK-NanoLuc were treated with a saturating dose of test compound, 20- to 100-fold above the IC50 value measured via live-cell equilibrium titration. Cells were incubated with compounds for 2 h, after which the medium was removed and replaced with fresh Opti-MEM to induce an equilibrium shift. NanoGlo Substrate and extracellular NanoLuc Inhibitor were added to each well, after which probe 5 was added to a final concentration of 0.25 µM and BRET was measured kinetically at 3-min intervals using a Glomax Discover Instrument. BRET data were plotted as a function of time.
Biochemical Enzymatic Activity Assay
The biochemical activity of compounds was measured at KM,ATP using Caliper technology as previously described.4,20 Compounds were diluted threefold in DMSO for a total of 11 points and were tested for BTK activity at the kinase’s ATP Km in a reaction buffer (20 mM HEPES, pH 7.4, 10 mM MgCl2, 0.015% Brij 35 surfactant, and 4 mM DTT in 1.6% DMSO). Final assay conditions included compound, 1 nM human recombinant BTK (Invitrogen, Carlsbad, CA), 1.5 μM fluorescein-labeled peptide, and 20 μM ATP. Reactions were incubated for 1 h before quenching with 35 mM EDTA. The reaction mixture was read using capillary electrophoresis on the Caliper LabChip 3000 (Caliper, Hopkinton, MA) at an excitation wavelength of 488 nm and an emission wavelength of 530 nm. Percent inhibition was calculated according to eq 2 and curves were fitted to eq 1 to determine the IC50 for each inhibitor.
Calcium-Flux Ramos Cell Assay
Human Ramos B-cell assays were performed as previously described.4,20 Human Ramos (RA1) B cells (ATCC CRL-1596) were suspended to 2 × 106 cells/mL in phenol red free RPMI (Invitrogen 11835-030) supplemented with 50 mM HEPES (Invitrogen 15630-130) and 0.1% BSA (Sigma A8577). The cells were added 1:1 with calcium loading buffer (BD PBX calcium assay kit no. 640177; BD, Franklin Lakes, NJ) and incubated at room temperature in the dark for 1 h. Afterward, cells were exchanged to a phenol red free RPMI media with 50 mM HEPES and 10% FBS for a final concentration of 1 × 106 cells/mL. A total of 150,000 cells/well were plated into 96-well poly-
Results
Miniaturization of the BTK NanoBRET Assay and Adaptation for Binding Kinetics Analysis in Living Cells
BRET is highly sensitive to molecular distance and orientation between the donor (luciferase) and acceptor (fluorescent probe) pair.15,21 This property enables a bona fide measurement of direct molecular interactions inside intact cells.
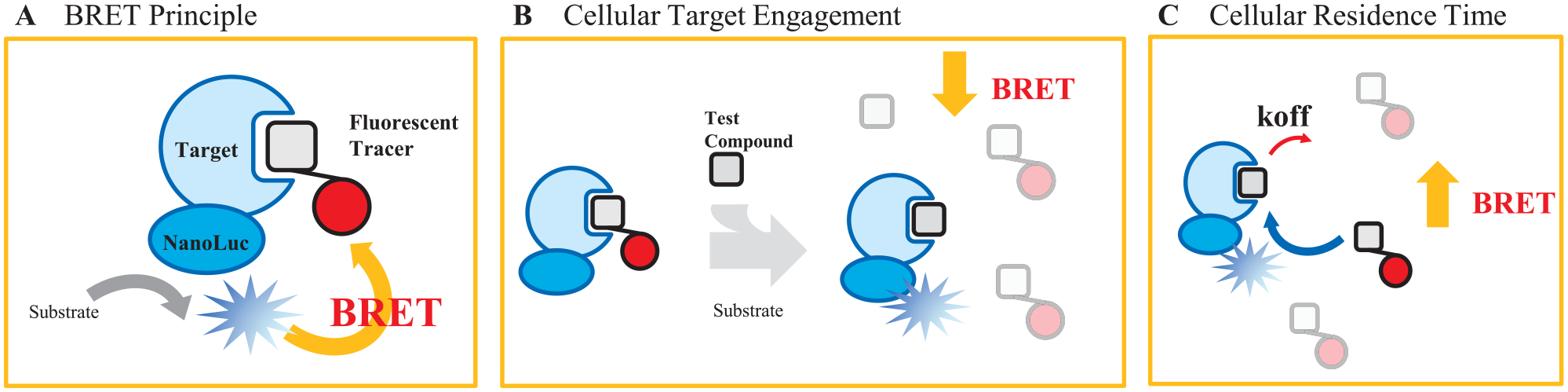
NanoBRET technology enables high-throughput target engagement and residence time measurements in live cells. (
To establish the BTK cellular target engagement assay in-house, the affinity of the competitive probe was first determined by the titration of varying probe concentrations onto live HEK293 cells transfected with the optimal C-terminal genetic fusion of BTK-NanoLuc.
16
Using both fluorescent probes 4 and 5 (from Vasta et al.
16
), strong, dose-dependent BRET signals were observed, shown in
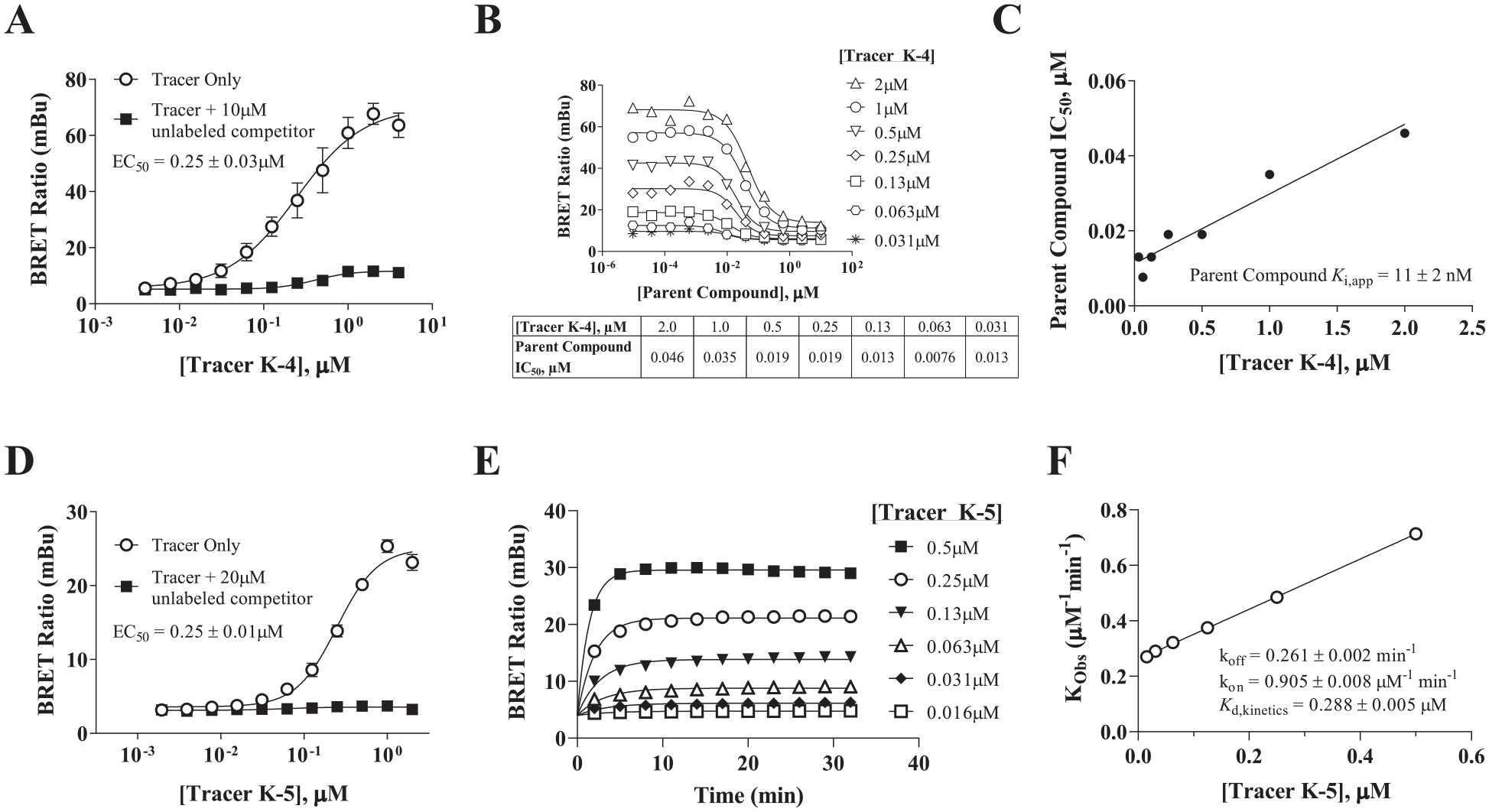
Kd and kinetic characterization of the BTK target engagement assays. Binding affinity and/or kinetic parameters of Bodipy-labeled probes 4 and 5 to BTK-NanoLuc were characterized in live cells. (
We also evaluated the probe binding kinetics by titrating increasing concentrations of probe and monitoring BRET at BTK-NanoLuc kinetically over time (
Characterization of Target Occupancy for Diverse Chemical Series of BTK Inhibitors in Living Cells
To evaluate the utility of the BRET method to profile intracellular BTK target engagement, we tested a panel of structurally diverse inhibitors in a 384-well plate format with a range of in vitro potencies spanning at least three orders of magnitude. The design, synthesis, and optimization of these inhibitors has been reported previously.4,5,17–20,25 At Kd concentrations of the probe, the BTK BRET target engagement assay serves as a measure of the apparent occupancy for BTK-NanoLuc in live cells. Following 2 h of treatment in the presence of probe 4, robust dose–response characteristics were observed for the various inhibitor treatments. Signals were normalized against vehicle control-treated samples such that maximal inhibition corresponded to 100% signal inhibition.
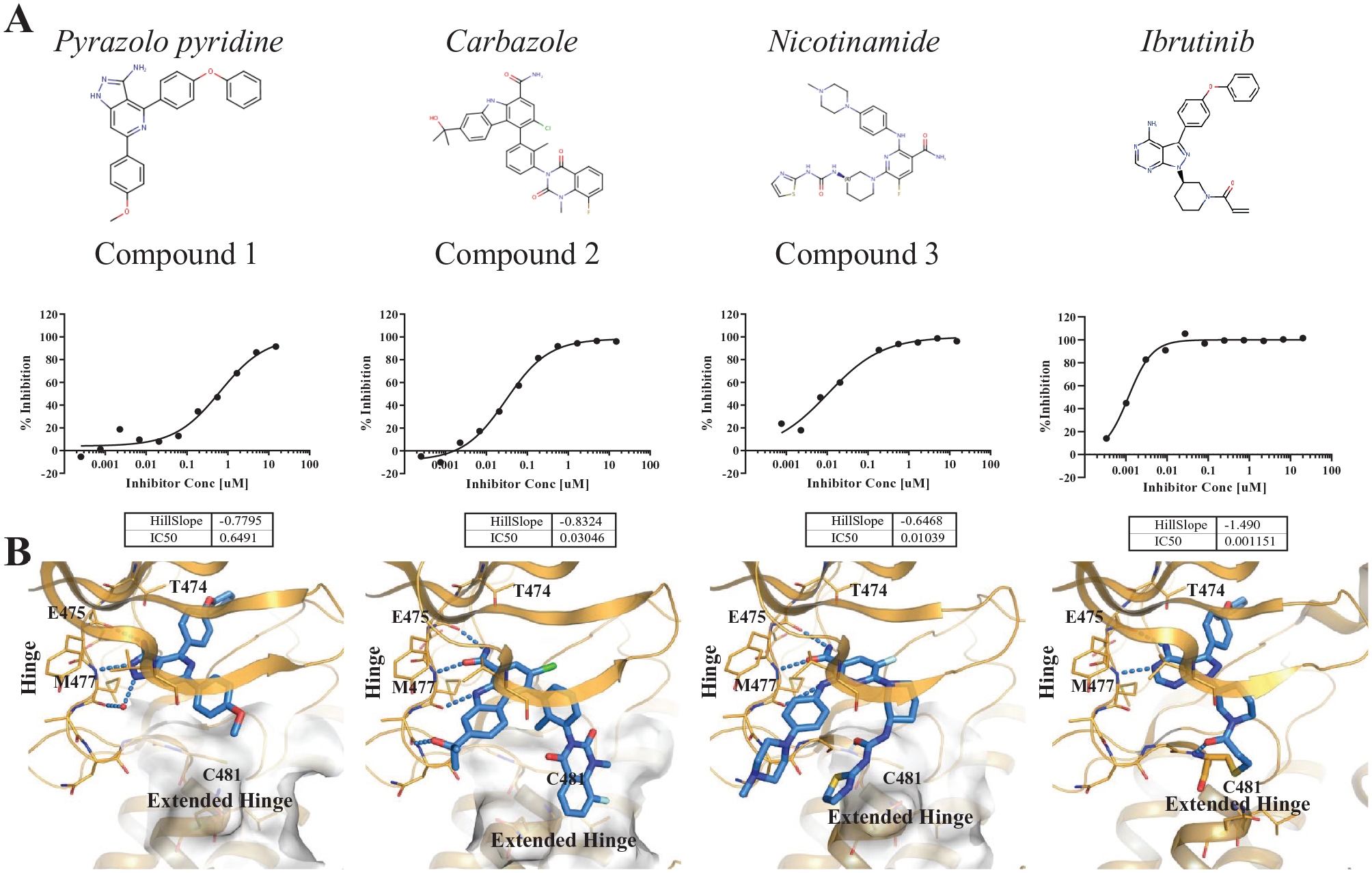
Cellular target engagement IC50 measurement using BRET. (
To better understand the potency differences across these chemotypes, we examined the crystal structures and molecular models of representative compounds in BTK (
In-Cell Target Engagement Correlates Quantitatively with Modulation of BTK Function
The ideal target engagement format provides a direct assessment of compound binding at the selected intracellular target, therefore providing a bridge between reductionist biochemical assays and cellular pathway analysis readouts. To what extent do BTK in vitro measurements actually correlate with cellular target engagement? To address this question, we analyzed in vitro BTK enzymatic activity data and compared it with cellular BRET target engagement data for the test set of compounds.
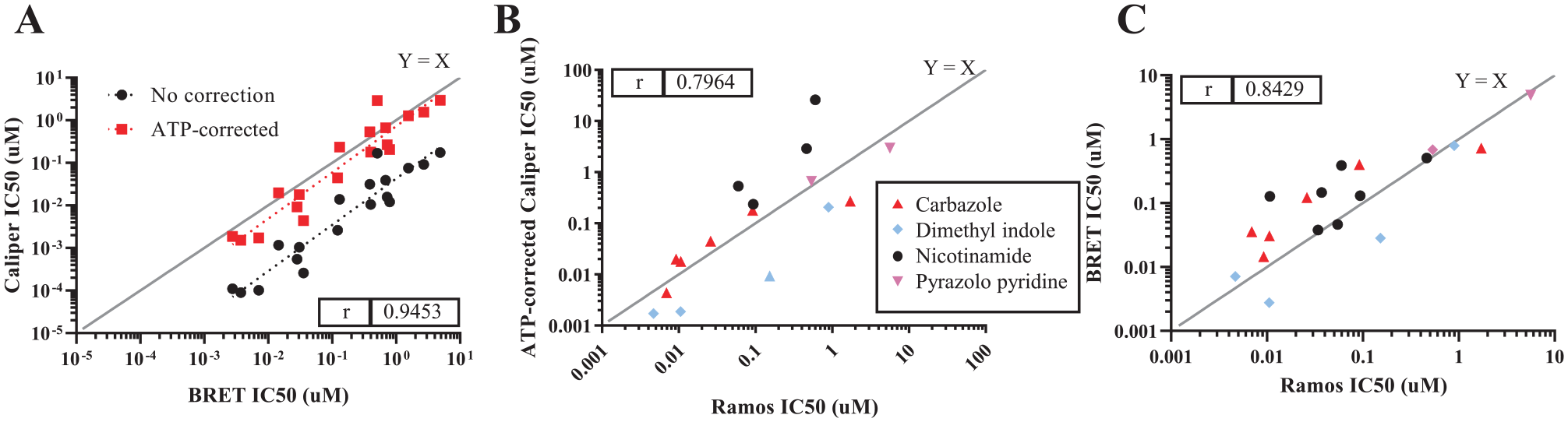
Comparison of BRET with in vitro enzymatic and cellular functional assays. (
Next, we investigated whether the cellular target engagement method could establish a functional link between physical binding and modulation of endogenous cellular kinase activity. Accordingly, we performed a correlation analysis between binding affinity and cellular function. To detect a proximal functional readout of BTK inhibition, we measured Ca2+ flux in Ramos cells.
Extended Target Engagement of a Noncovalent Inhibitor at BTK in Living Cells
Because of the high cellular potency of several of these compounds, we further examined how these compounds behave kinetically and whether any of them offer protracted target binding. We utilized the NanoBRET technology to measure residence time comparing the potent, reversible compound 2 against the irreversible ibrutinib. Cells were pretreated with compound at a saturating concentration (5- to 10-fold IC50) for extended times before washout and the addition of excess competing probe 5. Notably, the off-rate is an intrinsic compound property that follows first-order kinetics and is independent of compound concentration. Remarkably, limited dissociation of compound 2 was observed over the course of 120 min, resembling the kinetic profile of the irreversible ibrutinib inhibitor, thus supporting a cellular half-life (T1/2) of >120 min for this reversible compound. The extended residence times can be explained from the structural model of compound 2, in which the quinazoline moiety is shown to make van der Waals and water-mediated hydrogen bonding interactions with the extended hinge (
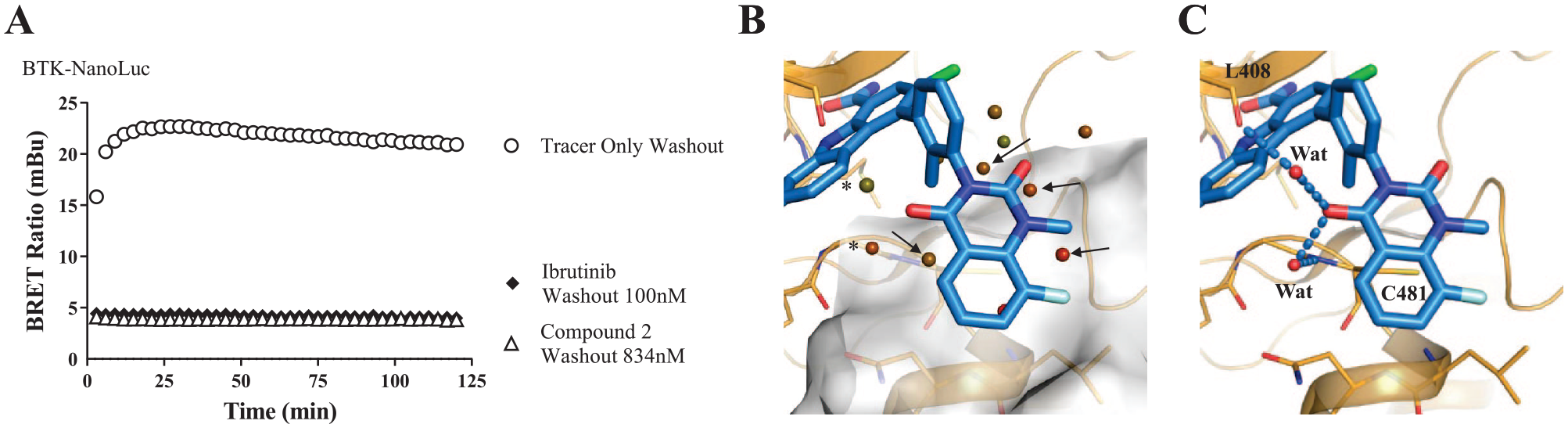
Cellular residence time evaluation for reversible and irreversible BTK inhibitors. (
Discussion
In summary, we have found that NanoBRET methods can provide quantitative analysis of BTK intracellular target engagement and residence time, and serve as a useful tool for the large-scale characterization of cell-active BTK inhibitors, offering a potentially valuable template to evaluate compound structure–activity relationships for kinase inhibitors in cells. We have demonstrated that intracellular binding can be correlated with functional potency for a broad chemical inhibitor set, establishing biochemical-to-cellular connectivity and a framework to explore these relationships. Notably, additional broadly applicable technologies have also emerged that leverage measurements of target stability, mass spectrometry, and chemical proteomics, as well as fluorescence imaging, to measure binding characteristics directly in living cells and correlate them with phenotypic results, utilizing in some cases label-free and proteome-wide assessments. A broader survey of these methodologies and their translational value in the context of drug discovery is nicely presented in Robers et al. 27 and beyond the scope of this discussion. Notably, one of the unique capabilities of this approach versus others is that it can be used to assess in-cell binding kinetics via dynamic monitoring of the BRET signal in live cells. Such kinetic assays proved to be beneficial in further characterizing compounds that may have desirable engagement kinetics, highlighting open system behaviors that may not be evident from steady-state analysis. This opens the potential to study additional pharmacology, such as kinetic selectivity, where it may be possible to achieve on-target selectivity through extended residence time at the primary target and transient engagement to off-targets (such as LCK in the case of BTK). Thus, the cellular BRET assay technology provides a powerful and scalable approach for evaluating key engagement parameters, providing a profiling tool for target-oriented screening and mechanistic studies.16,22 With the emergence of new genome editing and expression techniques, the BRET method will also be expandable to additional physiologically relevant cell types. Our results support that this method has the capacity to translate binding parameters into functional pharmacology in living cells.
Supplemental Material
Supplementary_Material – Supplemental material for A High-Throughput BRET Cellular Target Engagement Assay Links Biochemical to Cellular Activity for Bruton’s Tyrosine Kinase
Supplemental material, Supplementary_Material for A High-Throughput BRET Cellular Target Engagement Assay Links Biochemical to Cellular Activity for Bruton’s Tyrosine Kinase by L. L. Ong, J. D. Vasta, L. Monereau, G. Locke, H. Ribeiro, M. A. Pattoli, S. Skala, J. R. Burke, S. H. Watterson, J. A. Tino, P. L. Meisenheimer, B. Arey, J. Lippy, L. Zhang, M. B. Robers, A. Tebben and C. Chaudhry in SLAS Discovery
Footnotes
Acknowledgements
We would like to thank Amy Hart and Bill Pitts for critical reading of the manuscript. Special thanks to the kinome, cellular technologies, and data management and analytics teams for assay and data support.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Supplemental material is available online with this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.