Abstract
Coenzyme Q10 (CoQ10) deficiency syndrome is a rare disease included in the family of mitochondrial diseases, which is a heterogeneous group of genetic disorders characterized by defective energy production. CoQ10 biosynthesis in humans requires at least 11 gene products acting in a multiprotein complex within mitochondria. The high-throughput screening (HTS) method based on the stabilization of the CoQ biosynthesis complex (Q-synthome) produced by the COQ8 gene overexpression is proven here to be a successful method for identifying new molecules from natural extracts that are able to bypass the CoQ6 deficiency in yeast mutant cells. The main features of the new approach are the combination of two yeast targets defective in genes with different functions on CoQ6 biosynthesis to secure the versatility of the molecule identified, the use of glycerol as a nonfermentable carbon source providing a wide growth window, and the stringent conditions required to mark an extract as positive. The application of this pilot approach to a representative subset of 1200 samples of the Library of Natural Products of Fundación MEDINA resulted in the finding of nine positive extracts. The fractionation of three of the nine extracts allowed the identification of five molecules; two of them are present in molecule databases of natural extracts and three are nondescribed molecules. The use of this screening method opens the possibility of discovering molecules with CoQ10-bypassing action useful as therapeutic agents to fight against mitochondrial diseases in human patients.
Keywords
Introduction
Mitochondrial diseases are a large family of genetic disorders characterized by defects in multiple biochemical pathways and cellular processes in mitochondria. 1 Mitochondria are recognized as the massive energy producers in the cell but are responsible for other essential functions not related to ATP production. The functional diversity of mitochondria explains the pleiotropic effects found in mitochondrial diseases and the difficulty in identifying definitive therapeutic procedures. 2
Coenzyme Q10 (CoQ10 or Q10) deficiency is one mitochondrial disorder that is considered a rare disease. 3 CoQ10 is an amphipathic lipid formed by an aromatic polar ring of quinone with redox properties condensed to a hydrophobic polyisoprenoid tail responsible to the membrane attachment. 4 The primary function of CoQ10 is electron transfer between complex I (NADH-quinone reductase) or complex II (succinate-quinone reductase) and complex III (ubiquinol-cytochrome c reductase), 5 but it is also involved in antioxidant protection, nucleotide synthesis as the electron acceptor of dihydro-orotate dehydrogenase, the regulation of the mitochondrial permeability transition pore (mPTP), mitochondrial sulfide metabolism, and the regulation of uncoupler proteins of brown adipose tissue, as well as being an electron acceptor for reactions of β-oxidation. 5 CoQ10 deficiency is considered a rare disease because it affects less than 1 in 2000 persons. As a mitochondrial disease, it produces pleiotropic symptoms in a high number of organs and tissues, 6 and it can be classified in primary deficiency when it is generated by alterations in genes directly involved in CoQ10 synthesis, 7 and in secondary deficiency when the affected genes are involved in other mitochondrial functions not related to CoQ10 synthesis. 8 In humans, 11 genes are needed to produce CoQ10, and human CoQ10 deficiency has been reported by mutations in 10 of them. 7 These multiple origins of the disease have not facilitated the discovery of versatile therapies that apply universally to patients.
Most of our knowledge on coenzyme Q (CoQ) biosynthesis in eukaryotic cells has been provided by research with the yeast Saccharomyces cerevisiae, a yeast model that produces coenzyme Q6 (CoQ6). CoQ shows a conserved structure, a quinone ring that is similar in all organisms bound to an isoprenoid tail composed by a different number of monomer isoprene units depending on the species, 6 units in yeast (CoQ6) and 10 units in humans (CoQ10). All yeast COQ genes show significant homology with the human orthologs; each yeast gene has one counterpart in humans with the exception of yeast COQ1, which is homologous to PDSS1 and PDSS2 human genes, and COQ8, homologous to the ADCK3/COQ8A and ADCK4/COQ8B human genes. 9 In yeast, CoQ6 synthesis involves the assembly of a biosynthetic complex (Q-synthome) at the matrix side of the inner mitochondrial membrane. 10 Q-synthome is composed of proteins encoded by the COQ genes, except for COQ1 and COQ2, responsible for the hexa-isoprenoid tail synthesis and its condensation with the aromatic ring of p-hydroxybenzoic acid (p-HB), respectively. Q-synthome works as a single enzyme catalyzing the conversion of the Coq2 product, hydroxy-hexaprenyl benzoate (HHB), to CoQ6, although the exact composition of the Q-synthome is not clearly established and its catalytic activity has certainly never been measured. However, a shared feature for null mutant strains in COQ genes implicated in the Q-synthome assembly has been reported; the absence of one of the CoQ proteins blocks the complex formation, leading to the accumulation of the first precursor, HHB, instead of the expected substrate of the lacking enzyme. 11 In strains harboring missense mutations of COQ genes, the Q-synthome is assembled and the expected precursor is accumulated as has been demonstrated for missense mutations in the COQ7 gene. 12 We have previously demonstrated that COQ8 gene overexpression in a null COQ7 mutant (coq7Δ) restores the synthesis of 5-demethoxy-CoQ6 (5-DMQ6), 13 the Coq7 precursor. 5-DMQ6 is synthesized by the stabilization of components of the Q-synthome. 10
COQ8 overexpression became an effective strategy for dissecting the yeast CoQ6 biosynthetic pathway. The complex stabilization in the absence of specific CoQ proteins makes it possible to detect and identify diagnostic CoQ6 precursors. 14 The second application of COQ8 overexpression was the identification of molecules bypassing the CoQ6 biosynthesis defects of COQ null mutant yeasts. 15 Small molecules analyzed using this approach, such as vanillic acid (VA), have been proven successful in null COQ6 mutants (coq6Δ) expressing human COQ6 patients’ alleles. 16 Moreover, VA is able to restore CoQ10 biosynthesis and ATP production in human HEK cells lacking the COQ6 gene. 17 Other molecules similar to VA, such as 3,4-dihydroxybenzoic acid, can restore CoQ6 synthesis in coq6Δ mutants overexpressing the COQ8 gene. Greater interest has been aroused by β-resorcylic acid (β-RA) or 2,4-dihydroxybenzoic. β-RA restores CoQ6 synthesis in coq7Δ mutants overexpressing the COQ8 gene; 14 partially restores coenzyme Q9 (CoQ9) levels in a murine model of induced CoQ9 deficiency improving mitochondrial respiration and lifespan; 18 rescues the encephalopathic phenotype in Coq9R239X mice, a CoQ9 deficiency model with a defective Coq9 protein; 19 and blocks renal fibrosis in podocyte-specific Coq6 knockout mice. 20 The most promising effect of β-RA is the rescue of mitochondrial activities and CoQ10 levels in fibroblasts from CoQ10-deficient patients caused by mutations in COQ7 and COQ9 genes.21,22 The existence of bypassing compounds opens a therapeutic pathway for CoQ deficiency treatment that is more affordable than other methods. 23 However, the known bypassing molecules are not a general therapeutic solution for all causes of primary CoQ10 deficiency, and in fact, they are specific for each mutated gene.
The goal of this study was to design and configure a high-throughput screening (HTS) method based on the specificity and robustness of the COQ8 overexpression approach in COQ null mutant yeasts to detect new bypassing small molecules showing a general effect in all cases of primary CoQ10 deficiency. Our results demonstrate that this strategy is successful and leads to discovering molecules with potential value for CoQ10 deficiency treatment, based on an HTS method applied to a pilot subset of 1200 natural product extracts.
Materials and Methods
Strains, HTS Medium, and Reagents
The S. cerevisiae strains used in this work are described in
HTS Conditions
HTS was analyzed in sterile 96-well polystyrene microplates. Aqueous extracts (10 µL per well) from the Fundación MEDINA’s collection 24 (http://www.medinadiscovery.com) were deposited in each microplate using the automated Biomek FX workstation (Beckman Coulter, Indianapolis, IN). Then, 90 µL of the yeast suspension in YPG medium was added per well. Plates were incubated at 30 °C with shaking. Control wells were distributed in the left and right columns. Four controls per microplate were used in this study: (1) culture media controls (the two bottom wells of the right column without yeast [YPG]), (2) positive controls (the six top wells of this column [three top YPD and another three YPG + VA 2 mM]), (3) negative controls (the four top wells of the left column without VA), and (4) other positive controls (the four bottom wells of this column [YPG + VA 0.5 mM]). Microplates were analyzed with 80 extracts per plate (bacterial and fungi samples). All the experiments were performed in duplicate with an initial absorbance of 0.08 U/mL. Extracts were considered positive when the absorbance was at least double the initial absorbance or negative control, and the Z′ score of the plate was higher than 0.85.
Fundación MEDINA’s Natural Products Library Subset
For the primary screening (crude extracts), a subset of 1200 microbial extracts from different modules of the MEDINA natural product collection was used. The microbial extracts were obtained from bacterial and fungal strains cultivated in different nutritional conditions and extracted with acetone (1:1) for 1 h in an orbital shaker. Extracts were then centrifuged at 1500g for 15 min and the supernatant concentrated to half of the original volume (2XWBE) in the presence of a final concentration of 20% DMSO. Extracts were stored at −20 °C in 96-well ABgene V-bottom plates until needed.
Measurement of the Respiratory Rescue
For the primary screening (extracts), the yeast culture density in a nonfermentable carbon source (YPG) was monitored by determining the optical density (OD600) using an EnVision multilayer plate reader (PerkinElmer, Boston, MA) before and after 120 h of culture at 30 °C with shaking (1000 rpm) in a BIOSAN PST-60HL-4 thermo-shaker (Riga, Latvia). In the second screening (fractions), a continuous mode of OD600 monitoring was used from the beginning of the culture (TECAN SPARK 10M reader, Männedorf, Switzerland). Yeasts were cultured in 96-well plates at 30 °C with shaking (510 rpm).
Data Analysis
Extract activities were measured by OD600 and the growth of each sample (extract or compound) was determined by the following equation:
where Tf is the OD600 at the end time and To is the OD600 at time zero. An extract was considered to have activity when its growth at least doubled the initial OD. The Z′ factor predicts the robustness of an assay by taking into account the mean and standard deviation of both positive and negative controls. 25 In all experiments performed in this work, the Z′ factor obtained was between 0.92 and 0.99.
Bioassay-Guided Extract Fractionation
Culture broths generated from positive extracts were extracted by addition of a 1:1 volume of acetone. After shaking for 2 h at 220 rpm, centrifugation at 2700 rpm, and filtration, the acetone was removed under a nitrogen stream and the remaining solution was loaded onto an SP207ss cartridge (1.5 × 5 cm) that was washed with water (100 mL) and eluted with methanol (40 mL) and acetone (60 mL). The combined organic eluates were evaporated, the residue was dissolved in DMSO (400 μL), and 200 μL of this volume was subjected to semipreparative high-performance liquid chromatography (HPLC) on a Gilson GX-281 system equipped with a Zorbax SB C8 column (9.8 × 250 mm) at a flow rate of 3.6 mL/min, using a gradient of H2O:CH3CN (5% CH3CN for 1 min, then up to 100% CH3CN for 35 min, held at 100% CH3CN for 7 min, and back to 5% CH3CN in 1 min), and UV detection at 210 and 280 nm was monitored. Eighty fractions were collected every 30 s from minutes 2 to 42. These fractions were evaporated, dissolved to a final concentration of 20% DMSO, and subjected to a bioactivity test in the second screening.
LC-MS and Database Matching of Known Secondary Metabolites
Liquid chromatography–mass spectrometry (LC-MS) analysis in the low-resolution (LR) or high-resolution (HR) mode was performed as previously described.26,27 A database search was performed against the MEDINA proprietary database of microbial metabolites and the commercial Chapman & Hall Dictionary of Natural Products (v25.1).
CoQ6 Extraction and Quantification from Whole Yeast Cells
Yeast cells were lysed in a breaking buffer (10 mM Tris-HCl, pH 7.6, 1 mM EDTA, and 1 mM dithiothreitol) by vortexing with glass beads and centrifuged (10 min at 700g) to remove debris. The supernatant was centrifuged (60 min at 100,000g) to obtain a total membrane pellet. After protein quantification, samples equivalent to 1 mg of protein were extracted with a hexane method and separated, and CoQ6 was identified with an HPLC–electrochemical detection (ECD) method as previously described. 28
COQ8 Cloning in pCM189 Plasmid
Plasmids and primers used in this study are shown in
Results
COQ8 Overexpression Strategy to Detect CoQ Deficiency-Bypassing Molecules
The HTS approach designed to detect molecules that bypass defects of CoQ6 synthesis in yeast takes advantage of the Q-synthome stabilization produced by COQ8 gene overexpression. This strategy is depicted in
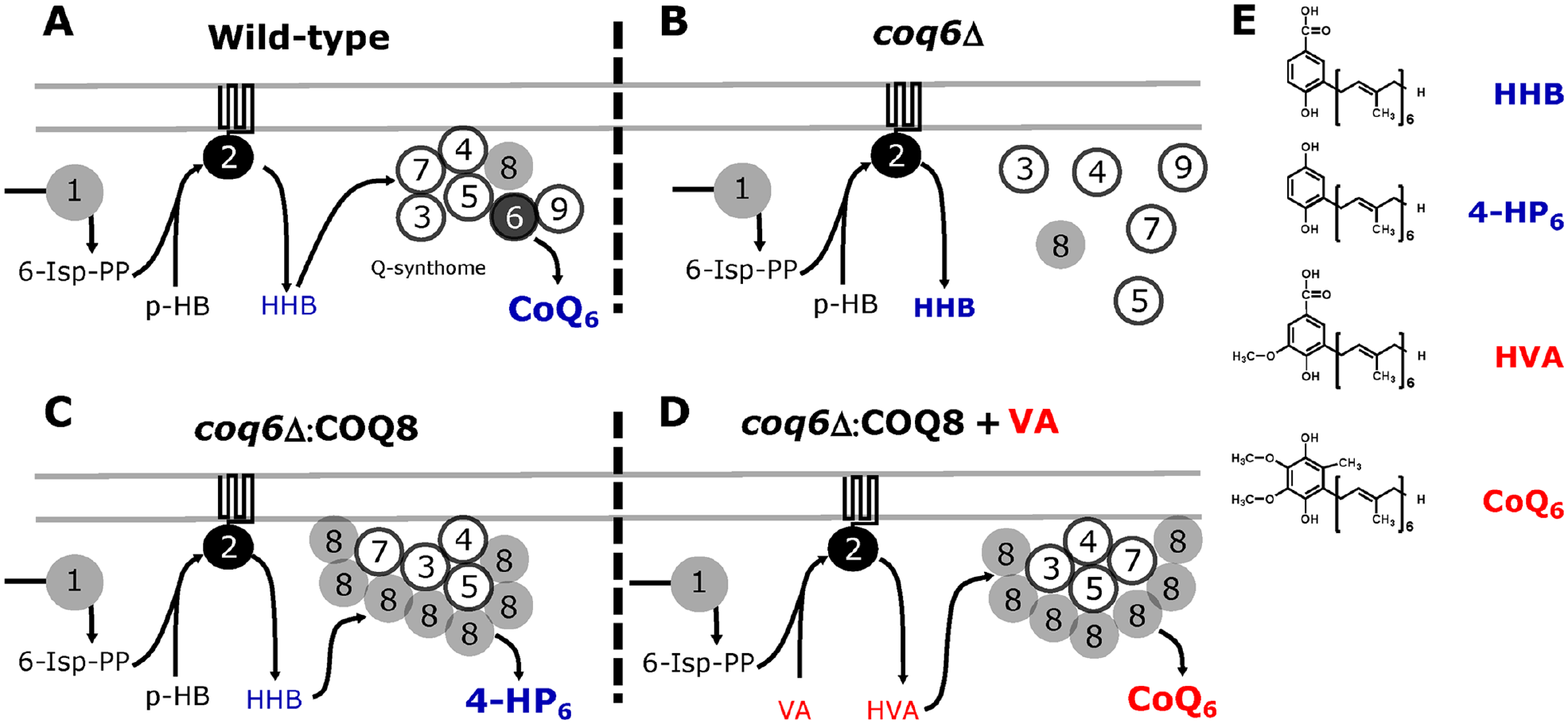
Biological foundations of the CoQ6-bypassing HTS method. (
VA and Glycerol Growth Constitute a Valuable Approach for Detecting CoQ Deficiency-Bypassing Molecules
In order to be successful, the strategy adopted by the HTS requires identifying molecules recovering CoQ6 biosynthesis, taking advantage of Q-synthome stabilization produced by the COQ8 gene overexpression. CoQ6 biosynthesis recovery will be analyzed using the growth in glycerol, a nonfermentable carbon source to activate the respiratory metabolism. In the yeast model, glycerol has been extensively used to detect respiratory-deficient mutants
29
and CoQ6-deficient yeast mutants.
30
In this screening approach, CoQ6 deficiency strains (coq4∆ and coq6∆) are unable to grow in YPG, a rich medium containing glycerol as a carbon source (
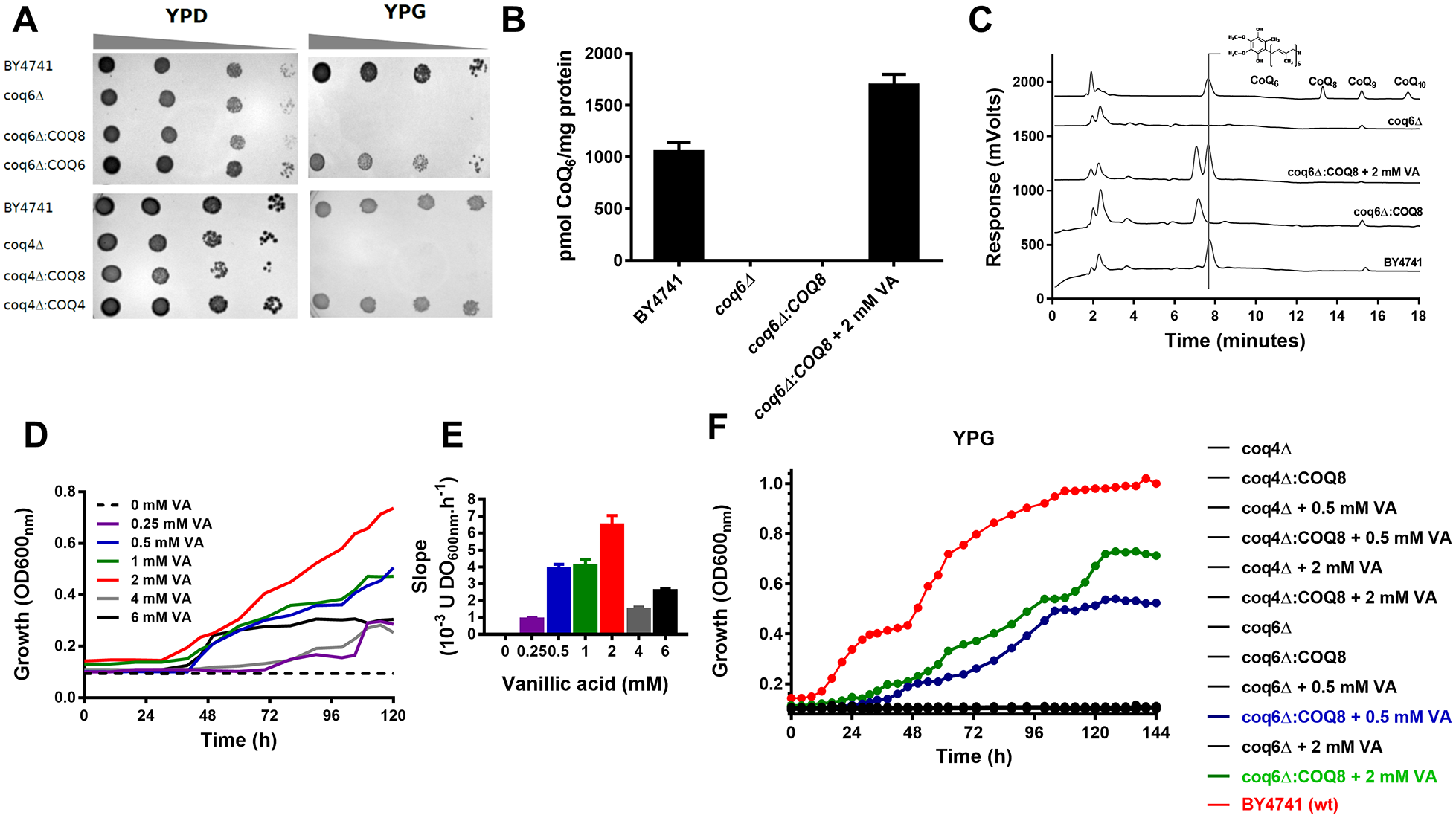
Glycerol growth and VA supplementation are basic components of the CoQ6-bypassing HTS method. (
First Screening Detection of Positive Extracts
The screening was divided into two parts; the first was designed to detect positive extracts recovering the growth in YPG, and the second was performed to detect positive fractions derived from the first screening. A scheme of the full HTS is depicted in
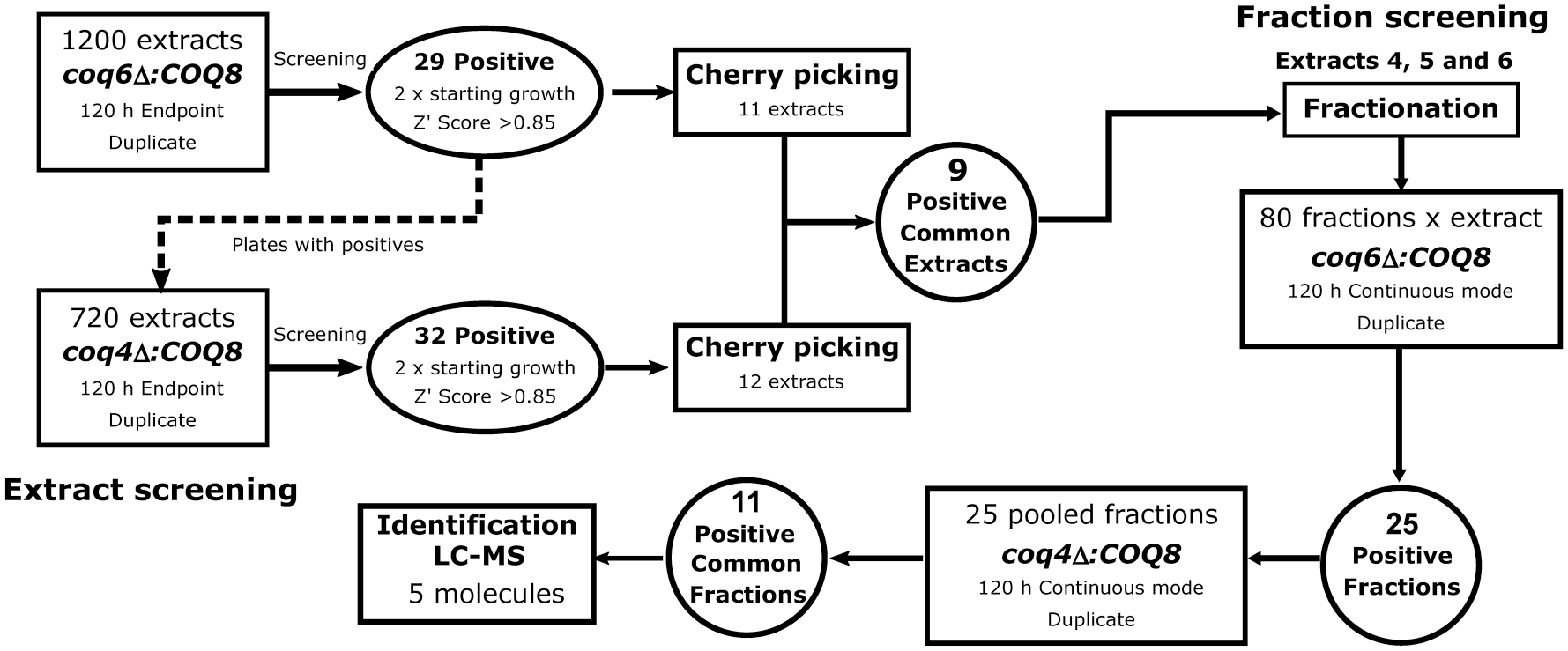
Flowchart of the CoQ6-bypassing HTS method. The first screening performed cell growth measurement in duplicate by an endpoint method using the coq6∆:COQ8 yeast strain. Extract plates with positive results were probed against the coq4∆:COQ8 yeast strain. An extract was considered positive when the final OD of the culture was at least double the initial OD. The Z′ score as a robustness indicator must be above 0.85. Positive extracts were selected and allocated together in a new plate that was probed in duplicate (cherry-picking analysis). Three positive extracts common for both strains (coq6∆:COQ8 and coq4∆:COQ8) were selected for chromatographic fractionation. In the second screening, fractions were probed against coq6∆:COQ8 in duplicate over 120 h and monitored in continuous mode. Positive fractions were collected and probed in the same conditions against the coq4∆:COQ8 strain. Common positive fractions (11) were analyzed by LC-MS to identify new or already known molecules (5).
Summary of Screening Results.
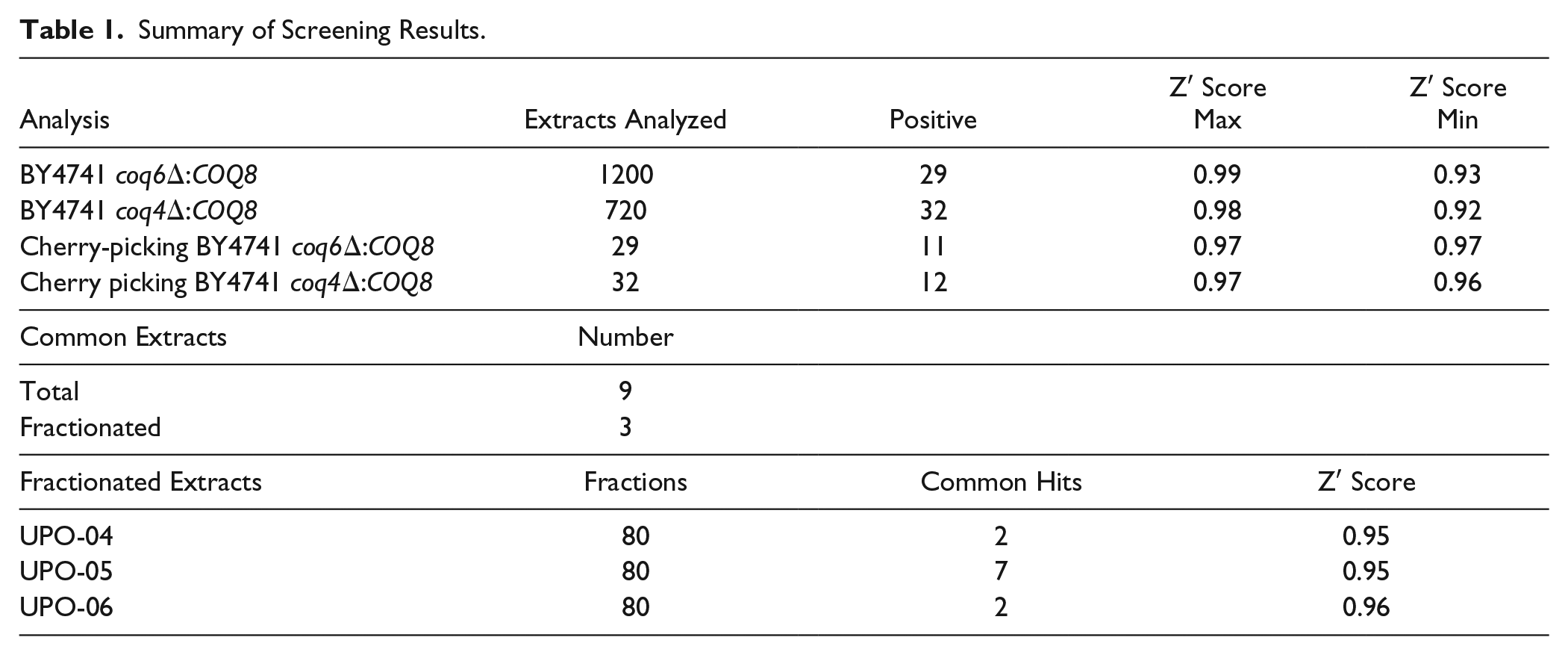
Raw Absorbance of Common Positive Extract.
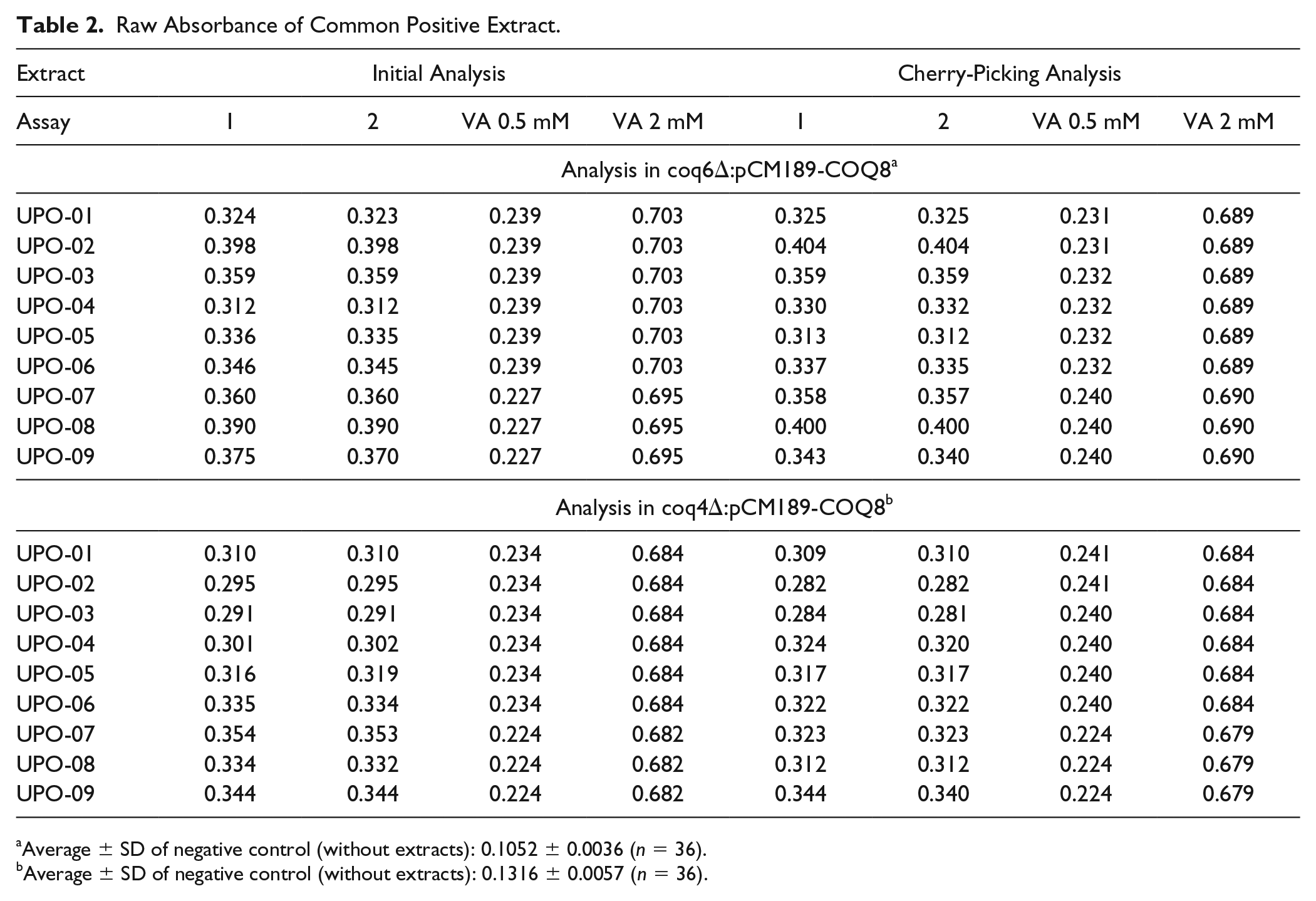
Average ± SD of negative control (without extracts): 0.1052 ± 0.0036 (n = 36).
Average ± SD of negative control (without extracts): 0.1316 ± 0.0057 (n = 36).
Second Screening with Fractionation of Positive Extracts Detected Molecules with CoQ6-Bypassing Activity
The first screening resulted in the finding of nine extracts common to both strains analyzed. Three of them (UPO-04, UPO-05, and UPO-06) were subjected to chromatographic fractionation into 80 fractions arranged in microplates that were used in the second screening (
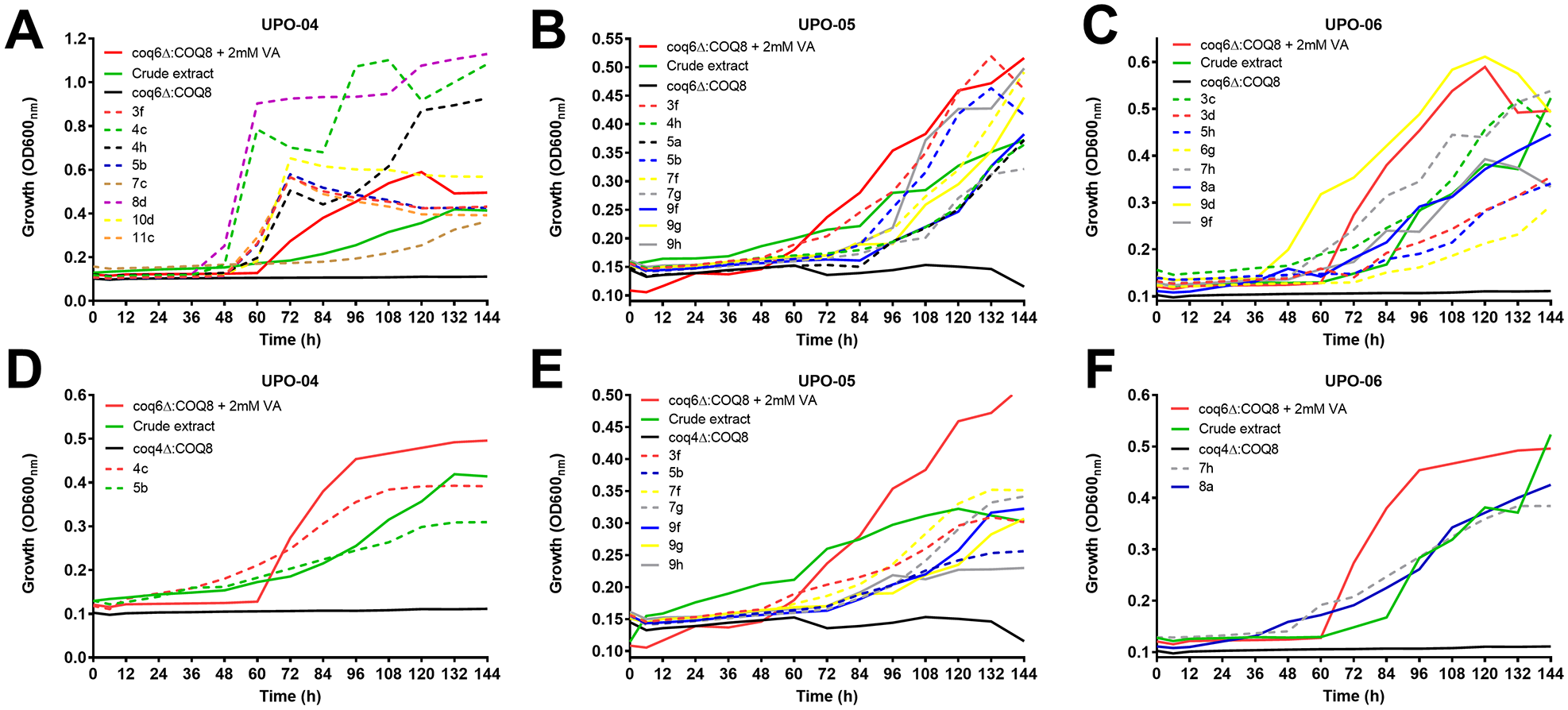
Growth curves corresponding to the second screening of CoQ6-bypassing molecules with fractionated extracts. Plates containing fractionated positive extract (80 samples plus controls) were probed against the coq6∆:COQ8 yeast strain in duplicate for 120 h with shaking at 30 °C. Yeast growth was monitored measuring OD at 600 nm in continuous mode. Data correspond to the average ± SD of both replicas. Only positive wells and controls were included in the plot. Names of fractions correspond to the well position: (
Identification of Potential Bypassing Molecules with Mass Spectrometry
Common positive fractions (11) collected from fractionated extracts were subjected to LC-MS analysis. The identification resulted in the finding of two possible active molecules with a known structure and three molecules (
Discussion
The new strategy described in this study tried to configure an HTS approach to detect molecules produced by bacteria or fungi bypassing CoQ6 deficiency. This activity is the ability to act as an intermediary for the CoQ6 biosynthetic complex or Q-synthome in null mutants when the complex is stabilized by COQ8 gene overexpression. Despite the results obtained corresponding to a pilot study, the strategy followed proved that this HTS method will be useful to identify new and universal molecules bypassing CoQ6 deficiency.
The effectiveness of this HTS approach could be evaluated by previously known CoQ-bypassing molecules such as VA,16,36 which supports the growth of the coq6∆:COQ8 yeast strain in nonfermentable carbon sources at 2 mM. The specific design of this approach makes it possible to detect positive extracts recovering the growth in mutant yeasts that are defective in one specific enzyme, such as Coq6, 34 which are also effective in mutants of a regulatory protein required for the right assembly of Q-synthome, such as Coq4. 37 This property opens the possibility of detecting CoQ-bypassing molecules as therapeutic agents useful for patients with CoQ10 deficiency harboring mutations in all genes involved in CoQ10 synthesis. Several experimental pieces of evidence support this possibility. The crude extracts analyzed (UPO-04, UPO-05, and UPO-06) restore the growth in coq6∆ and coq4∆ mutants only after COQ8 expression, indicating that Q-synthome is required for the extract’s action. The requirement of Q-synthome allows us to discard that the growth in YPG was supported by some type of CoQ molecule present in the extract. Other evidence reinforces the lack of CoQ in the extracts; extracts were obtained with an acetone–water mixture, which is not useful to purify molecules with the polarity of CoQ, and CoQ molecules were not detected in the mass spectrometry analysis performed in the positive fractions of the extracts analyzed. The UPO-06 crude extract not only restores the growth in glycerol media for coq4∆:COQ8 and coq6∆:COQ8 yeast strains but also is able to recover the growth in the coq7∆:COQ8 strain, supporting the idea that the HTS described here is useful for identifying molecules with a universal bypassing action. The definitive evidence of a molecule bypassing CoQ6 deficiency is the ability to synthesize CoQ6 in deficient cells. CoQ6 was detected in the pooled yeasts grown with the positive fractions from the second screening. This means that not only the growth in YPG has been restored, but also CoQ6 biosynthesis is in operation at a wild-type level.
The strength and robustness of this HTS can be deducted by the extrapolation of results obtained from the full collection of natural extracts of the Fundación MEDINA. This Natural Products Library is composed of more than 200,000 extracts, from which only a representative subset of 1200 have been used in this study. Despite the stringency parameter selected in the HTS design, that is, eight replicas per positive extract analyzed, the minimal growth threshold, and the Z′ score measured for all plates, from three positive extracts, 11 fractions with CoQ6-bypassing activity were detected and 5 potential molecules were identified by LC-MS. It is not possible to perform a simple extrapolation, but these results support the potential of the HTS described in this study.
With this limited number of fractionated positive extracts, five molecules have been detected. Even though we have found a high number of common positive fractions (11 in three positive extracts), the number of identified molecules is lower. This apparent low efficiency can be explained by the existence of contiguous positive fractions showing the same molecule, and that the three fractionated extracts belong to phylogenetically related organisms or to the same microorganisms growing under different culture conditions.
The future use of this HTS approach will require increasing the number of natural extracts to be analyzed. However, in the short term, the analysis of the remaining six positive extracts, the identification of the active molecules present in all the extracts, and the test in cellular models derived from patients’ fibroblasts affected by CoQ10 deficiency will allow us to obtain data of the real therapeutic capacity of the identified molecules. Despite the differences between human and yeast, the high similitude in the CoQ biosynthetic machinery 9 and the existence of known molecules working in both models as bypassing agents, such as VA or dihydroxybenzoic acid,14,18,19 support the hypothesis that molecules identified in this study could also be effective in humans.
A larger number of positive extracts detected is not an additional complication, but rather the opposite since it will enable the introduction of new yeast mutants in the first screening, in order to decrease the number of common positive extracts while improving the versatility of the molecules identified to restore the CoQ10 synthesis in patients. A second benefit of extending the analysis to more natural extracts is to eliminate the effect produced by the potent metabolic capacity of the yeast compared with the human model. Compounds with bypassing capacity in yeast may not work properly in humans and may even produce harmful effects since human cells do not have the appropriate enzymatic machinery to redirect the target molecule to a metabolite involved in the CoQ10 synthesis pathway.
The primary CoQ10 deficiency is a rare disease. Recent studies have estimated that the number of potential cases worldwide is 1665 taking into account the known pathogenic variants, or even 123,789 when predicted pathogenic variants are also included in the prediction study, 38 supporting the idea that primary CoQ10 deficiency is an underdiagnosed disease. It is also necessary to take into account patients with secondary CoQ10 deficiency, for which the number of potential cases is still impossible to estimate. Both primary and secondary CoQ deficiency produce common clinical symptoms, and patients can be treated with CoQ10 supplementation. This therapy has showed some cases of success in the past 39 but in general is ineffective; CoQ10 shows a poor uptake by most affected organs (skeletal muscle and nervous system), and only crosses the blood–brain barrier with larger doses. 40 The use of CoQ-bypassing molecules, such as the ones that have been identified in this study, circumvents most CoQ10 supplementation defects. They are small and polar molecules that are able to cross the blood–brain barrier and usually are nontoxic and cheaper. The requirement of the Q-synthome assembly is not an obstacle, since patients maintain the ability to produce CoQ10. The molecular similarities of the CoQ synthesis processes between yeast and humans open the possibility that molecules identified in a more ambitious HTS than that based in this pilot study could be potential therapeutic agents to combat CoQ10 deficiency in humans, a mitochondrial rare disease that does not have a definitive and universal therapeutic strategy.
Supplemental Material
Supplementary_Material – Supplemental material for Design of High-Throughput Screening of Natural Extracts to Identify Molecules Bypassing Primary Coenzyme Q Deficiency in Saccharomyces cerevisiae
Supplemental material, Supplementary_Material for Design of High-Throughput Screening of Natural Extracts to Identify Molecules Bypassing Primary Coenzyme Q Deficiency in Saccharomyces cerevisiae by Aida M. Berenguel Hernández, Mercedes de la Cruz, María Alcázar-Fabra, Andrés Prieto-Rodríguez, Ana Sánchez-Cuesta, Jesús Martin, José R. Tormo, Juan Carlos Rodríguez-Aguilera, Ana Belén Cortés-Rodríguez, Plácido Navas, Fernando Reyes, Francisca Vicente, Olga Genilloud and Carlos Santos-Ocaña in SLAS Discovery
Footnotes
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The research group is funded by the Andalusian Government as the BIO177 group, through FEDER funds (European Commission), from the Junta de Andalucía Proyectos de Excelencia P12 CTS943. The MEDINA authors disclosed receipt of financial support from Fundación MEDINA, a public–private partnership of Merck Sharp & Dohme de España S.A./Universidad de Granada/Junta de Andalucía.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.

