Abstract
Camelid-derived nanobodies are versatile tools for research, diagnostics, and therapeutics. Certain nanobodies can function as intrabodies and bind antigens within the eukaryotic cytosol. This capability is valuable for the development of intracellular probes and targeted gene therapies. Consequently, many attempts have been made to produce nanobodies that are intracellularly stable and resistant to aggregation. Pursuit of these intrabodies generally focuses on library design or nanobody selection method. Recent variations of library design have yielded diverse libraries capable of producing nanobodies against a wide variety of antigens. Novel screening methods have also been developed, yielding nanobodies with high affinity for intracellular antigens. These screening techniques can have advantages over phage display methods when nanobodies against intracellular antigens must be rapidly produced. Some intracellular screening methods convey the additional advantage of selecting for other desired intrabody characteristics, such as antiviral action or conditional stability. This review summarizes the recent developments in both library design and selection methods aimed at producing intrabodies.
Introduction
The fields of medicine and cellular biology have been revolutionized by the development and production of monoclonal antibodies. Antibodies are essential proteins of the immune system that bind foreign antigens with high affinity and specificity. This binding depends on the sequence of the variable domains of the heavy- and light-chain (VH and VL) portions of the antibody (
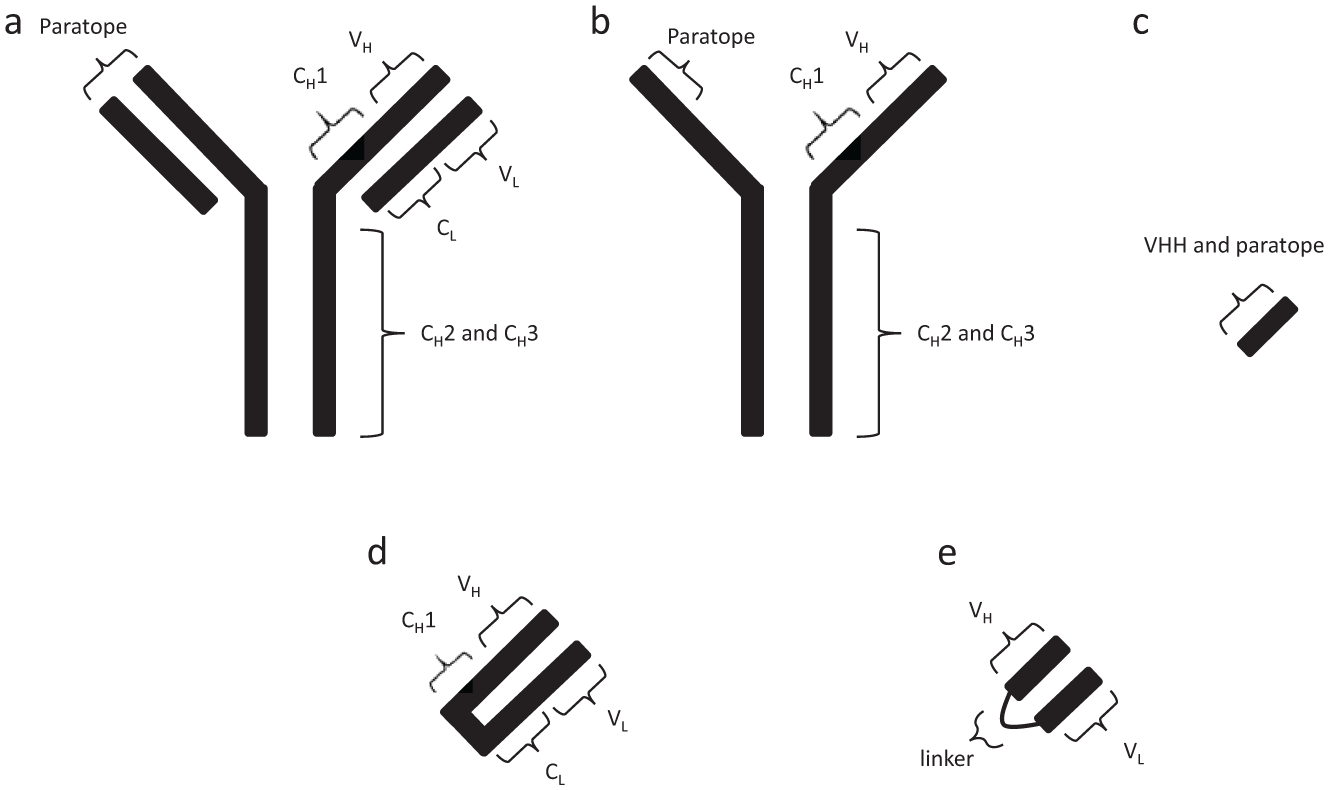
Basic structure of antibodies. (a) Structure of full-sized conventional antibody: consists of three constant heavy-chain regions (CH1, CH2, and CH3), one constant light-chain region (CL), one variable light-chain region (VL), and one variable heavy-chain region (VH). Paratope is the binding site that interacts with the antigen. (b) Heavy-chain-only antibody unique to the Camelid family. Paratope does not rely on light chain for binding. (c) An isolated camelid VH domain is referred to as a VHH. VHHs have fully functional paratopes. (d) Fab format. Includes VH, CH1, VL, and CL but does not include the Fc domain. (e) scFv format. Includes only the VH and VL domains connected by a linker of variable length.
Other antibody formats, including Fabs and scFvs, have also been developed but are limited in their utility when compared to VHH domains. scFv and Fab formats combine parts of heavy chains and light chains with and without linkers, respectively (Fig. 1e and 1d). The multidomain nature of Fabs and scFvs, however, makes their manipulation more complex in comparison to VHH domains. A further limitation is that the reducing environment of the cytoplasm prevents the formation of disulfide bonds in Fabs and scFvs. This hampers the expression and stability of Fabs and scFvs intracellularly,3,4 although there are exceptions when scFvs have been functionally expressed in the cytoplasm. 5 VHHs generally contain a single disulfide bond but are usually soluble within the cytoplasm. even when the disulfide bond is not formed. 6 The presence of a second disulfide bond may improve nanobody stability extracellularly, but seems to have a destabilizing effect on nanobodies when they’re exposed to a reducing environment similar to the cytoplasm. 7
The utility and relative simplicity of nanobodies have made them useful tools in many areas of molecular biology. In the years since their discovery, nanobodies have been engineered to serve a variety of roles, including fluorescent intracellular probes,8–14 molecular tools for protein knockout experiments, 15 in vivo modulators of plant enzymes, 16 aids in protein crystallization, 17 diagnostic tools,18,19 and potential human therapeutics.20–23
The unique advantages of nanobodies stem from their size, shape, and structure, all of which differ significantly from conventional antibodies. Nanobodies are ~15 kDa, around 10 times smaller than most full-sized antibodies. This small size and relative simplicity allow most nanobodies to be produced in standard prokaryotic expression systems. The shape of the nanobody paratope is also advantageous for certain applications. Unlike conventional antibodies, nanobodies have a slightly convex antigen-binding site, allowing them to bind concave or otherwise sterically hindered antigens. 24 This ability allows nanobodies to act as enzyme inhibitors,25,26 presumably by binding the cleft of enzyme active sites, sites that are generally unavailable to standard antibodies. Other advantages, like a reduced tendency to fold improperly or aggregate, make nanobodies especially attractive for use as intrabodies within the cytoplasm. 27
A variety of techniques exist for delivery of intrabodies into the cytoplasm as potential therapeutic agents. Generally, these techniques involve either the delivery of nucleic acids encoding the intrabody or the delivery of already-synthesized protein. Nucleic acid delivery of nanobodies has been accomplished in mice by viral delivery and transplantation, 28 yielding a limited antiviral effect. Due to the safety concerns surrounding viral vectors and gene therapies, however, protein delivery is considered a safer option for clinical development. 29 To deliver intrabodies in protein form to the cytoplasm, several techniques have been developed, including cationic resurfacing,30,31 bioreversible esterification, 32 and fusion with cell-penetrating peptides.33,34 Protein delivery has the added advantage that disulfide bonds are formed before nanobodies are exposed to reducing agents of the cytosol. This may allow nanobodies to function as therapeutics that would otherwise misfold when expressed in the cytoplasm.
Even with effective delivery strategies, intracellular antigens can be difficult to target for a variety of reasons. Among these reasons are complications in expressing and purifying the antigen of interest as well as difficulties in obtaining nanobodies that are functional in reduced form. In vitro selection requires expression and purification of the target antigen, which can be complicated when an antigen has many conformations or posttranslational modifications. An example of this is the HIV Vpr protein that has been difficult to produce at milligram levels. 35 This difficulty may have been related to a failure in selecting an anti-Vpr nanobody by standard phage display, followed by a success in selecting such an antibody by intracellular yeast two-hybrid screening. 36 Besides antigen expression, nanobody expression can also be problematic for reasons that remain unclear. 37 Beyond selection method, library design can also affect intrabody functionality. The main types of nanobody libraries include immune, naïve, and synthetic, each of which possess certain advantages in the development of nanobodies with intracellular functionality. This review highlights the recent developments in nanobody library design in relation to intracellular function, and it discusses the various techniques for intrabody selection, beyond the standard methods of phage display and biopanning.
VHH or Nanobody Libraries
Immune Libraries
The most common VHH or nanobody library is produced by camelid immunization. These immunizations are performed every few weeks until the adaptive immune response climaxes. Then, the camelid’s blood is drawn, the lymphocytes are isolated, and messenger RNA (mRNA) is extracted and reverse transcribed into DNA. The VHH gene is then amplified by PCR to create a library of VHH gene sequences, many of which code for nanobodies that specifically bind the immunization agent. Because this technique is so widely used, details of the process are not included in this review. For reference, a complete protocol on the production of immune libraries has been described by Pardon et al. 38
The main benefit of immune libraries is that they take advantage of camelid physiology to produce nanobodies with the desired affinity. As in other mammals, a foreign antigen in camelids prompts the proliferation of B cells and the production of antibodies. Affinity maturation stimulates the proliferation of B cells that produce antibodies with adequate stability and high affinity for the antigen in question. Thus, affinity maturation acts to enrich nanobodies with affinity for the immunizing antigen. Consequently, immune libraries have much higher concentrations of relevant nanobodies than any other type of nanobody library.
While immune libraries can effectively generate nanobodies against a particular antigen, the production of immune libraries is also costly and time-consuming. Camelid immunization requires access to a camelid and a 3- to 6-week period while the immune response climaxes. Only after this process can blood be drawn, and a library assembled for subsequent rounds of screening. Once created, an immune library can produce nanobodies against only the antigen(s) used in the immunization step. This significantly limits the versatility of immune libraries and necessitates the use of animals in every successive attempt to generate a novel nanobody.
Other limitations of immune libraries come from the process of immunization and affinity maturation. Because the camelid HCAbs generated in an immune response function extracellularly, there is no guarantee that an immune VHH library will yield nanobodies that function intracellularly. 39 In addition, the process of immunization does not allow for control of antigen conformation or chemical environment. This can be especially problematic for antigens that have a variety of conformations, like membrane proteins. 40 Another limitation of immune libraries is the immune tolerance of the camelid. Amino acid sequences that exist naturally in camelid organisms are non-immunogenic, and so cannot be targeted by nanobodies selected from an immune library. This represents a bottleneck that severely limits immune library versatility. To overcome these limitations, two other types of libraries have been developed.
Naïve Libraries
Nanobody libraries can also be assembled from camelids that have not been immunized against any particular antigen. These libraries, referred to as naïve or non-immune, have the primary advantage of containing a much wider variety of VHH sequences. High diversity allows one naïve library to be a source of multiple nanobodies. This versatility eliminates the need to generate a new immune library for every new antigen being targeted. To this effect, Verheesen et al. 41 produced a single naïve library that produced nanobodies against emerin, actin, tropomyosin-1, and nuclear poly(A)-binding protein. Other naïve libraries have produced nanobodies against human fibroblast growth factor-1 42 and human procalcitonin. 43 Because no vaccination step is necessary for naïve library creation or screening, naïve libraries are well suited to produce nanobodies against antigens that are either non-immunogenic or highly toxic.
Despite the advantages of naïve VHH libraries, several limitations hinder the use of naïve libraries as universal sources for antigen-specific nanobodies. Because of the immense diversity found in naïve VHH libraries, the range of possible screening methods is limited to phage display and ribosome display that have higher capacity and sensitivity than other screening methods. 44 While library diversity has clear advantages, it may come with the dead weight of VHH sequences that have not been affinity maturated, and thus lack both thermodynamic stability and antigen affinity. Thus, a trade-off exists in immune and naïve libraries between diversity and functional stability. In light of this challenge, considerable effort has been put forth to understand what factors influence VHH or nanobody fold stability and use this knowledge to rationally design libraries of useful, stable nanobodies.
Synthetic Libraries
In 2005, Bond et al. 45 showed that the antigen-specific portions of nanobodies called complementarity-determining regions (CDRs) could accommodate variations far beyond those seen in natural HCAbs while still allowing the domain fold to remain stable. Since this discovery, many attempts have been made to produce synthetic nanobody libraries by altering residues in the CDR regions. While most attempts to create a synthetic VHH library involve randomizing the CDR sequences to some degree, several limitations prevent full randomization to create a truly universal library. First, stability is negatively affected by full randomization, because the CDR loops play a role both in antigen binding and in the stability of the domain fold. 46 This results in considerable inefficiency when designing a useable synthetic library. A second limitation is the finite capacity and sensitivity of screening methods. Phage display, the most extensively used screening method, can generally manage libraries of 1010 to 1012 clones, which is much less than the >1030 different VHH sequences that would be produced from full CDR randomization. 47
To cope with the limitations involved in synthetic VHH library production Man-Seok Ju et al. 47 analyzed naturally occurring CDR sequences and limited the variation within the CDR loops to a select group of amino acids that most often occur in nature. Stop codons and cysteines were excluded to prevent premature termination or addition of unwanted disulfide bonds. These randomized CDRs were grafted onto a humanized VHH antibody that had previously shown high stability and high yields when produced in E. coli. These limitations on sequence variation significantly reduced library size but still had an estimated diversity of 1.8×1010, which was sufficient to select nanobodies that could specifically bind interleukin-1β (IL1β), amyloid-β, and vascular endothelial growth factor (VEGF).
In 2016, Moutel et al. 48 created a synthetic VHH library that yielded many nanobodies that are stable within the cytoplasm and resistant to aggregation. The scaffold was selected from a library of several hundred clones that were isolated from both immune and naïve libraries in previous studies. 49 These VHHs were filtered by an assay that screened for solubility and resistance to aggregation within the cytosol of E. coli. 50 Once a stable scaffold was selected, it was humanized by changing seven of its framework residues to match the sequence of human VH3. New CDR loops were grafted, from a nanobody designed to bind lamin, into the humanized nanobody scaffold and shown to maintain both lamin affinity and intracellular stability. After validating intracellular expression of this nanobody compared to other known nanobodies, it was used as a scaffold for a partially randomized synthetic library. Randomization was done by holding CDR1 and CDR2 regions mostly constant and similar to naturally occurring sequences, while the CDR3 loop was fully randomized with lengths of 9, 12, 15, and 18 amino acids. Only Cys was excluded from the possible amino acid pool to avoid additional disulfide bonds and major structural alterations. This library proved effective in producing nanobodies against many antigens, including green fluorescent protein (GFP), tubulin, and Her2. In addition, most of the nanobodies selected appeared to function without aggregation within eukaryotic cytosol.
Several other attempts to produce synthetic VHH libraries have yielded successful results. Goldman et al. 51 produced a semisynthetic library by using error-prone PCR on a naïve library, which was then panned by phage display to isolate a variety of effective binders. In 2018, Suzuki et al. 52 created a VHH library by randomizing the CDR sequences and then performing selections using cell-free complementary DNA (cDNA) display technology. While this technique produced high-affinity nanobodies, the authors acknowledged the limitation that cell-free screening yields nanobodies that do not always express well using standard microorganism expression systems.
Producing Intrabodies from VHH Libraries
The methods for creating VHH libraries are varied and increasingly versatile. Most immune, naïve, or synthetic libraries are capable of producing nanobodies that will also function as intrabodies. In cases when a particular nanobody exhibits poor solubility in the cytoplasm or otherwise displays poor functionality, other library types and variations are available that could improve nanobody performance. The above sections are meant as a resource for investigators who are targeting difficult intracellular antigens or who require nanobodies that express especially well within the cytoplasm.
Nanobody Screening Methods
Following library construction, screening methods are used to select for nanobodies with particular desirable properties. Among these characteristics are antigen affinity, stability, ability to inhibit or neutralize an enzyme, resistance to aggregation, and intracellular expression. To efficiently select for these properties in combination with antigen specificity, several methods have been used with varying degrees of success.
Phage Display
Phage display is the most widely used method for VHH library screening and can be used with large VHH libraries, generating an estimated 106–1011 different nanobody-presenting phages in a population of 1012 phage particles. 53 Figure 2 describes a general workflow of this method, although considerable variation exists in various protocols. Phage display is by far the most common screening method. Most phage display does not, however, select for intracellular stability. One possible exception to this limitation, as described by Speck et al., 54 includes using the TAT excretion pathway in otherwise standard phage display screening. The TAT pathway allows the VHH domain to fold within the prokaryotic cytoplasm before display occurs. This adjustment may ensure that selected nanobodies fold efficiently in the cytoplasm and may result in more stable intrabodies. Because most phage display does not select for intracellular stability, however, phage display will not be the focus of this review.
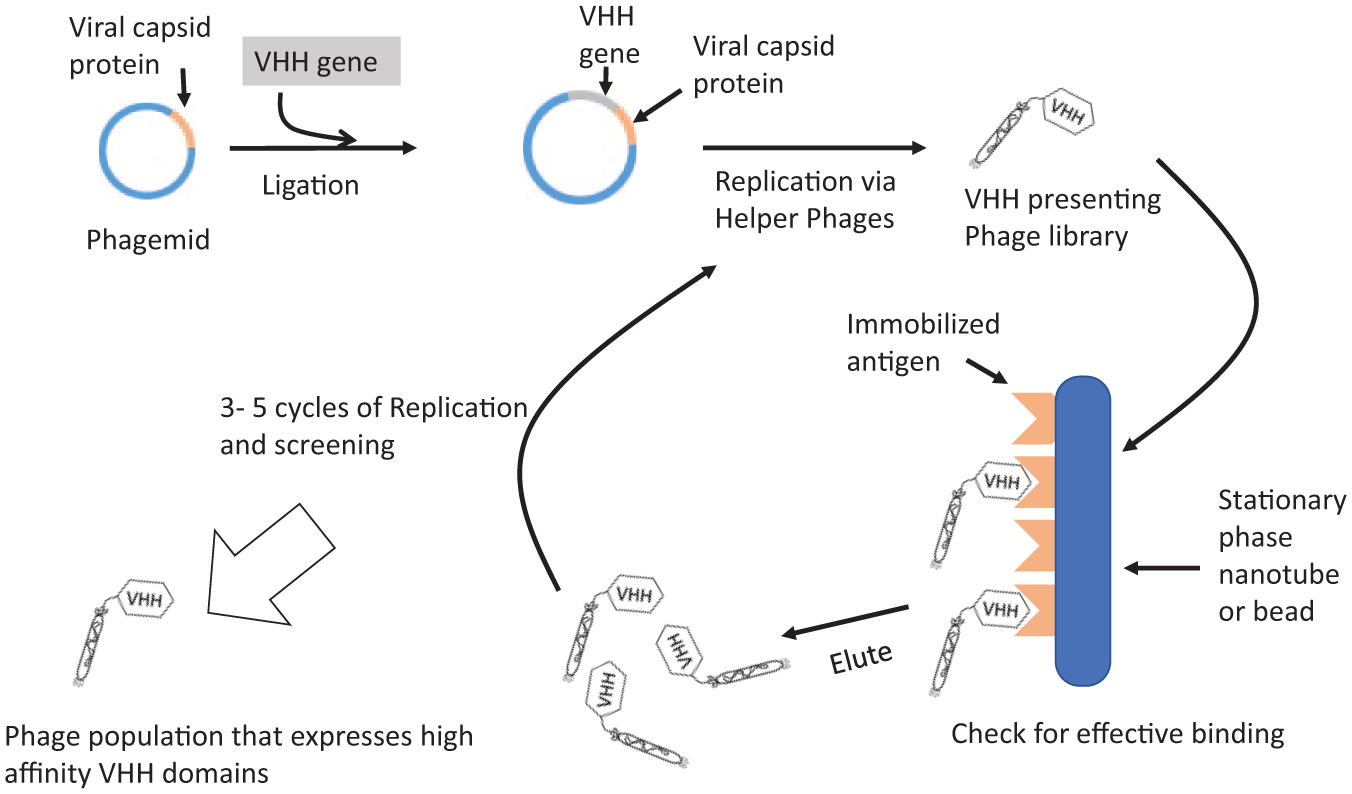
Generalized protocol for phage display selection. VHH library is cloned into a phagemid adjacent to a capsid protein. Expression of VHH plasmid produces a nanobody displaying phage. Immobilized antigen is then used to select for the VHHs with the highest affinity. Bound phages are reamplified, and the selection process is repeated.
Yeast Two-Hybrid
Yeast two-hybrid (Y2H) systems have been used extensively to investigate protein–protein interactions and have been applied to VHH selection. The basis of Y2H is the interaction of a bait protein with a target protein, both of which are fused to proteins that need to interact for the transcription of a reporter gene to occur. The primary advantage of this system is that selection ensures the intracellular solubility and binding affinity of the nanobodies produced. For additional selection specificity, either nuclear or cytosolic Y2H systems 55 can be used depending on the nanobody’s intended location of action. Another advantage of Y2H is the ability of yeast to produce large proteins that require posttranslational modification. In the case of large eukaryotic protein antigens, the Y2H system may be the most appropriate selection system because yeast can presumably synthesize and fold the protein antigen in the same way it would exist in other eukaryotic hosts. In comparison to phage display, Y2H also simplifies the selection process because no purified, immobilized antigen and no helper phage stock are required. To date, Y2H systems have successfully isolated intracellularly stable nanobodies against HIV Vpr protein, 36 PCV2 Cap protein, 56 and Newcastle disease virus HN protein. 57
Despite the successful selection of several nanobodies by Y2H, it remains a much less common technique than phage display. Because of yeast’s relatively low transformation efficiency compared to E. coli, large libraries are more easily handled using a phage display process. If Y2H selection were to be used with synthetic or naïve libraries, additional enrichment steps would likely be necessary, such as the two-step procedure used by Hyland et al. 5
Bacterial Two-Hybrid
A bacterial two-hybrid (B2H) system has also been developed and applied to nanobody selection. In 2012, Pellis et al. 39 published a single-step method for the selection of nanobodies that are functional and stable within the cytoplasm of E. coli. The process works by creating two plasmids. One plasmid codes for the bait protein, which was in this case GFP, HIV integrase, or nucleoside hydrolase. The other plasmid codes for the target protein, which was a variety of VHH domains from an immune library. As in other two-hybrid systems, the target and bait proteins are expressed as fusion proteins that must come in close proximity for the transcription of a specific reporter gene to occur. In this case, the reporter gene endowed the bacteria with the ability to grow on selective media, allowing for isolation of nanobody-carrying colonies in a single selection step.
Nanobodies isolated by Pellis et al. using B2H were shown to have similar antigen affinities and stabilities when compared to nanobodies isolated from the same immune library by phage display. An exception was a B2H nanobody against HIV integrase whose thermodynamic stability as measured by denaturation in guanidium chloride was higher than any other antibody fragment known to the authors as of 2012. This same nanobody also displayed the highest binding affinity of the nanobodies evaluated in the study. This result shows that another potential advantage of a B2H selection system is high affinity and thermodynamic stability, higher in at least one instance than those of nanobodies produced by phage display. Additional advantages of B2H selection reported by Pellis et al. are the relatively low background when compared to phage display and the one-step character of the B2H protocol. These advantages together could shorten the process of nanobody selection considerably.
Although this selection method has produced exceptional results in one study, it has not been used extensively since its invention. As the chemical environment of the E. coli’s cytoplasm differs from that of eukaryotic cytoplasm, it remains unclear whether B2H carries a special advantage in the development of intrabodies whose intended use is in the eukaryotic cytoplasm. Like Y2H selection, B2H does not allow for direct control over antigen conformation. While Y2H and B2H should theoretically produce the antigen of interest in the same form as produced in yeast or bacteria, respectively, it is difficult to assess whether fusion to the bait and target proteins could affect normal folding of the antigen in question. This issue of fusion proteins is, however, common to most selection methods, including phage display, and so is not a prohibitive complication in most circumstances.
Since B2H selection has been carried out on only immune libraries, it is unknown whether it would be useful with larger naïve or synthetic libraries. Assuming transformation efficiency is the major limiting factor in large-library screening, B2H selection should be compatible with large libraries as well. Considering its simplicity and efficiency, B2H stands as one of the most useful and underused selection tools for nanobody development.
Chloramphenicol Acetyltransferase Fusion
Maxwell et al.
50
developed another tool for selecting nanobodies with intracellular functionality by fusing VHH domains to the enzyme chloramphenicol acetyltransferase (CAT) (
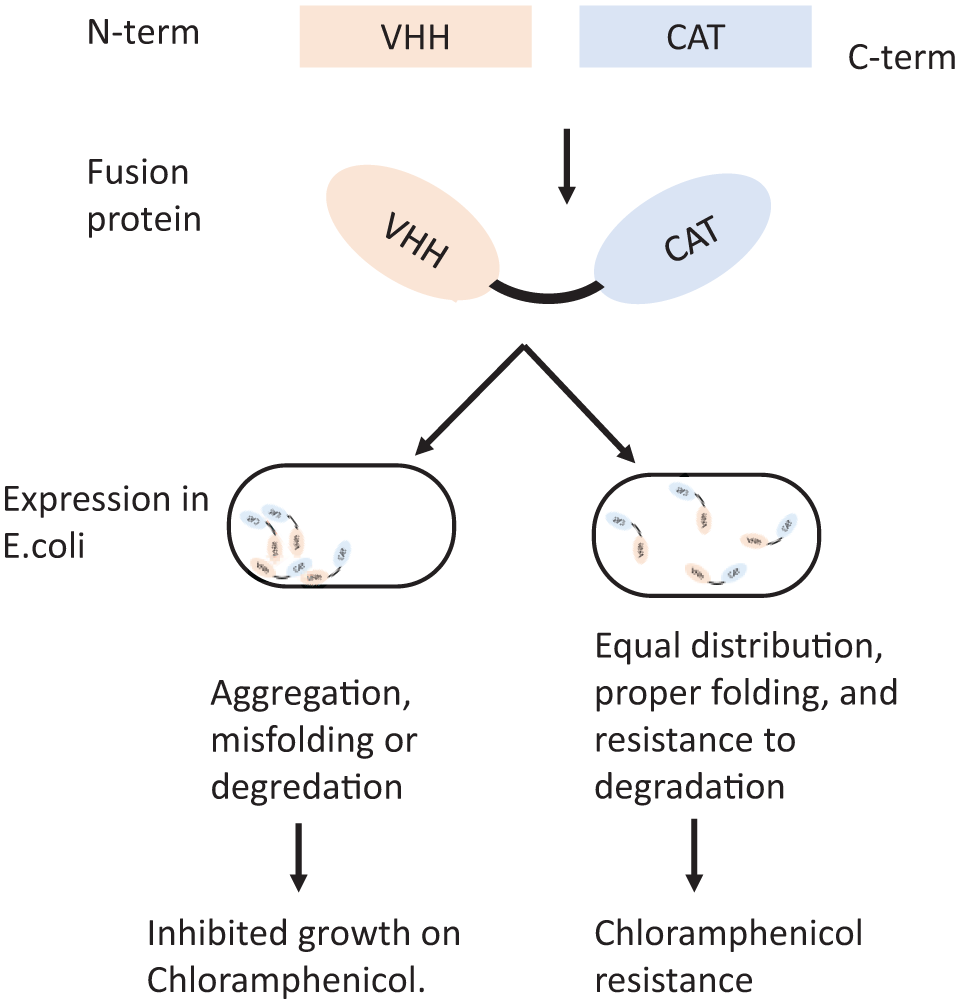
Chloramphenicol acetyltransferase (CAT)–VHH fusion for selection of stable VHHs that aren’t prone to aggregation or degradation. Even distribution within the cytosol of Escherichia coli results in higher chloramphenicol resistance.
Fusion with Disulfide Bond Enzyme
Another method for producing stable intrabodies came from work that was done to improve the solubility of single-chain antibody fragments by the expression of a disulfide bond isomerase.58,59 This technique was demonstrated with camelid-derived nanobodies by Olichon and Surrey 60 while designing functional VHH–fluorescent protein fusions. Such fusion proteins are challenging because nanobodies and fluorescent proteins fold most effectively in opposite redox environments. When produced in the cytoplasm of E. coli, GFP can fold effectively, but VHH domains generally cannot fold because disulfide bonds will not form in this reducing environment. As a solution, Olichon and Surrey tagged VHH–GFP fusions with DsbC, a disulfide bond isomerase that catalyzes the formation of disulfide bonds in the VHH domains, resulting in a stable fluorescent nanobody.
While this method adds an extra level of complexity to nanobody production, it also expands the range of nanobodies that can function and remain folded in the cytoplasm. Even without the fluorescent protein fusion, a nanobody–DsbC fusion format may be a more stable scaffold for useful nanobodies that would otherwise misfold on expression in the cytoplasm.
Lentiviral Screening
In 2016, Schmidt et al. 61 presented a novel technique for efficient selection of nanobodies that are both intracellularly stable and capable of neutralizing viruses. Figure 4 shows the general procedure for this method. First, inactivated influenza or vesicular stomatitis virus (VSV) particles were used to immunize a llama against a multitude of viral antigens. After collecting the llama’s lymphocytes, the camelid VHH sequences were amplified and cloned into a lentiviral plasmid instead of the more common phagemid used for phage display. Using this plasmid library, lentivirus particles were produced and used to transfect human A549 cells, which were subsequently exposed to a lethal dose of influenza or VSV. The surviving cells were those that contained a VHH that was both stable within the cytoplasm and capable of neutralizing the virus.
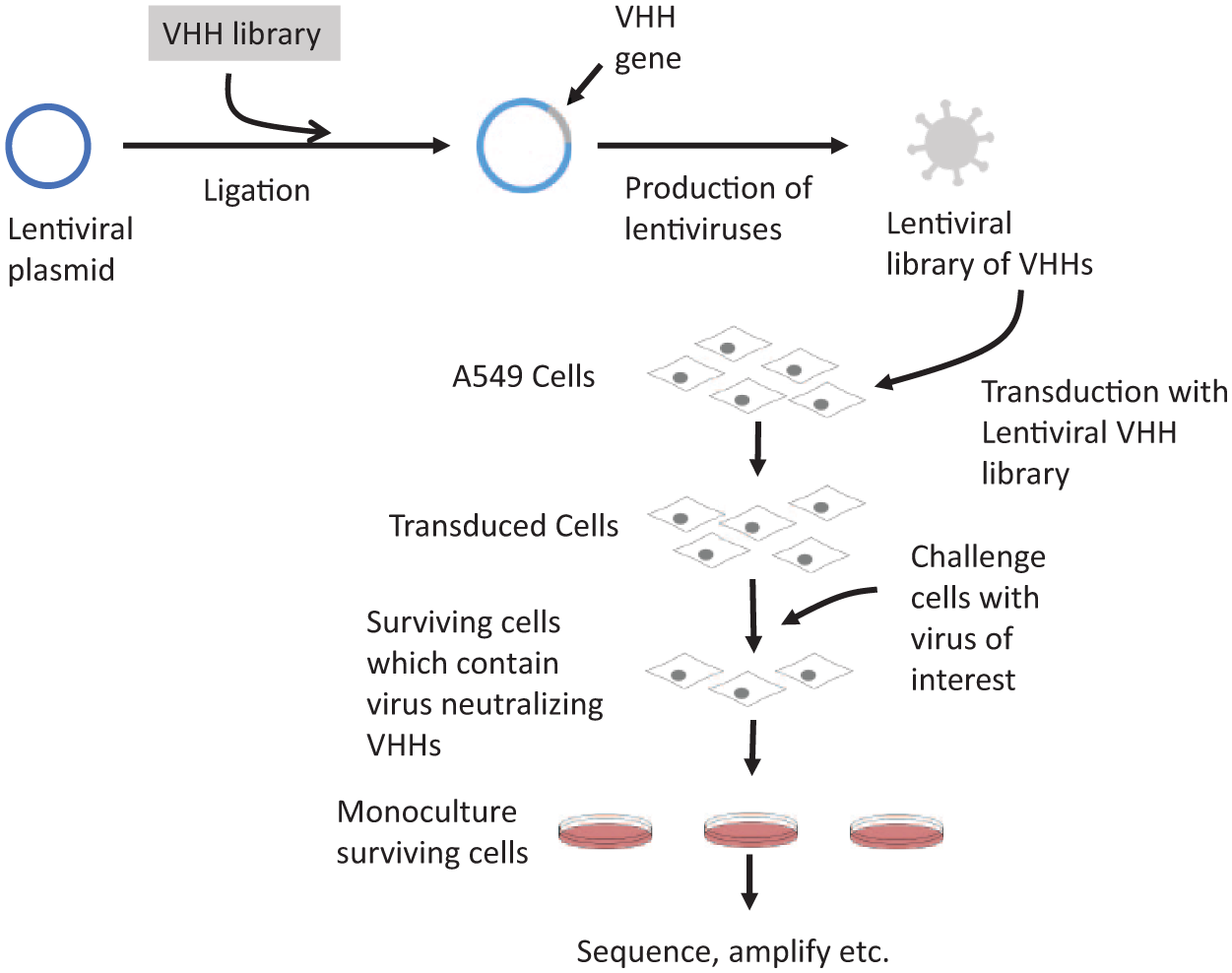
Lentiviral screen for virus-neutralizing nanobodies. Immune camelid VHH library is ligated into a lentiviral plasmid. Lentivirus particles are produced and used to transform A549 cells. Transduced cells are challenged with an active form of the same virus that the camelid was originally challenged with. Surviving cells are monocultured, and their VHH domains are sequenced and amplified.
This relatively recent selection method for antiviral nanobodies has the advantage of narrowing the selected nanobodies to only those that are stable within the cytoplasm of human cells. In addition, this selection method provides information about which epitopes should be targeted for effective neutralization of a particular virus. This has considerable import in the field of gene therapy, in which an intrabody could potentially be used to endow human cells with immunity against viral infection. If applied in other contexts, functional nanobody selection methods like this one could inform future nanobody engineering by providing information about what epitopes and antigens should be targeted to bring about a desired effect.
Yeast Presentation
Another versatile technique for nanobody selection was published in 2018 by McMahon et al. 62 This method uses a 649-amino-acid linker that joins the VHH domains to a glycosylphosphatidylinositol anchor, which secures the VHH to the yeast cell wall. Screening is then performed by incubating the yeast library with the antigen of interest, which has previously been fluorescently labeled. Several rounds of magnetic cell sorting (MACS) using antifluorophore beads are followed by fluorescence-activated cell sorting (FACS) to isolate single cells that bound to the fluorescently labeled antigen. By completing several rounds of selection and alternating the fluorophore, antifluorophore nanobodies were avoided.
As described by McMahon et al., this selection method has several unique advantages over others. First, this method can be used without purified antigens. This is especially useful for proteins that are modified posttranscriptionally, like peptide hormones, and do not express well in prokaryotic systems. McMahon et al. showed that the secreted hormone adiponectin could be expressed as a FLAG-tagged fusion protein and then collected from cell culture media. Without purifying this antigen-containing media, the yeast library was able to bind adiponectin and was screened using a fluorescently labeled anti-FLAG antibody and several rounds of MACS and FACS. Of note, this yeast surface display library has been made publicly available free of charge, contrasting sharply with many selection systems that have been heavily commercialized.
Conditionally Stable Nanobodies
A unique screening method developed by Tang et al. 63 created a completely new class of nanobody-based molecular tools. Using an existing anti-GFP nanobody, a library of random mutants was generated and then inserted into HEK293T cells as a fusion with blue fluorescent protein (BFP). These cells were also infected with adenovirus to stably produce GFP. Screening was then performed to identify cells that had enhanced BFP signal that depended on GFP expression. From these cells, nanobodies were isolated that are stable only when their corresponding antigen, in this case GFP, is present. A sequence analysis of these nanobodies allowed for the identification of three distinct mutations that, when transferred to other nanobodies, yielded the same conditionally stable behavior in eight out of nine cases. Tang et al. subsequently showed a variety of applications made possible by these selectively stable nanobodies. By fusing an enzyme of interest to a conditionally stable nanobody, tight control over enzyme function can be regulated in mice models. One example of this, shown by Tang et al., was tight regulation of recombinase activity, based on the coexpression of GFP in the mice. Another experiment in the same study showed that recombinase activity could be tightly controlled by creating a triple fusion consisting of a recombinase and two different conditionally stable nanobodies. This allowed for recombinase activity to be limited to cells in which the antigens of both of these conditionally stable nanobodies are expressed. This technique, while relatively new, could significantly affect the way tissue-specific genetic experiments are conducted and create more efficient gene knockout models.
Conclusion
Nanobodies have already proven to be invaluable as research tools and therapeutics, but their intracellular applications have yet to be fully exploited. Their potential use as high-affinity binders intracellularly is particularly attractive for the development of intracellular probes, cell-penetrating biologic drugs, and gene therapies. The small size and relatively simple structure of nanobodies make them especially suitable for these applications. Nanobodies are not, however, immune from denaturation, aggregation, and the other structural weaknesses common to all proteins and protein therapeutics.
To engineer nanobodies that express well in eukaryotes and resist aggregation, three general types of VHH libraries can be used. Immune libraries are the most costly and time-consuming, but they have the highest proportion of VHH domains that will bind the antigen of interest. The disadvantages of these immune libraries are their limited versatility and relatively low sequence variation. To address the issue of narrow utility, naïve VHH libraries have been generated by compiling VHH genes from many different camelids. This option provides excellent sequence diversity but lacks the advantages granted by in vivo affinity maturation. Impressive efforts have been put forth more recently to develop synthetic VHH libraries that are diverse in sequence while also maintaining high stability and solubility. These synthetic libraries are useful sources for nanobodies but have also provided valuable information on the relationship between sequence and function in the framework and CDR regions. Understanding these structural factors has informed other fields of protein engineering, allowing for the engineering of noncamelid autonomous antibody fragments. By altering the framework regions within an antibody fragment, autonomous antibody fragment libraries have been created using human heavy-chain 64 and human light-chain 65 fragments. These human VH and VL domains are similar to nanobodies but may have the added benefit of lower immunogenicity when used as human therapeutics.
Another element of intrabody design is the screening method. While phage display continues to be the most common selection method, many other methods have been developed, including bacterial two-hybrid, yeast two-hybrid, lentiviral screening, ribosome display,66,67 bacterial surface display,68,69 yeast surface display, 62 whole-cell biopanning, 70 and others. Each screening method has certain advantages regarding the production of nanobodies. Perhaps more important than these advantages, however, are the biological discoveries that novel screening methods can produce. For example, the lentiviral screening system developed by Schmidt et al. 61 yielded information not just on nanobody design but also on viral protein epitopes that can be targeted for viral neutralization. Another example is the bifluorescent model used by Tang et al. 63 to determine several point mutations that can render nanobodies conditionally stable. Indeed, novel screening methods that have been developed, and perhaps those that will be developed in the future, can contribute more than the nanobodies they select. Screening methods provide powerful tools for basic research and discovery as they simultaneously serve as tools for nanobody engineering.
Nanobodies have a broad range of potential applications and are continually becoming more useful as the methods for nanobody production improve. In the realm of camelid-derived intrabodies, library designs and screening methods are both key to maximizing stability and utility. Improvements in both library design and selection methods can accelerate the development of useful nanobodies and contribute to other discoveries in protein chemistry and target identification.
Footnotes
Declaration of Conflicting Interests
The author declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author received no financial support for the research, authorship, and/or publication of this article.
