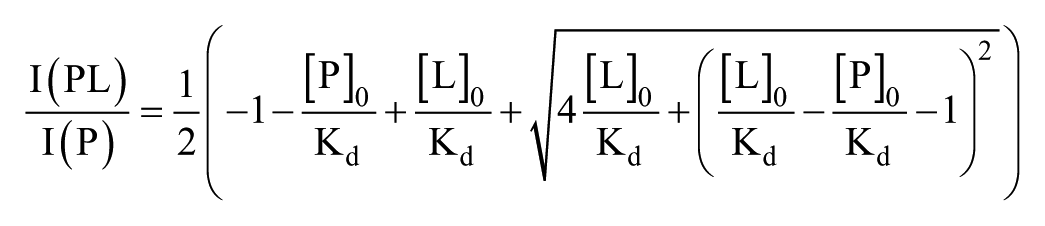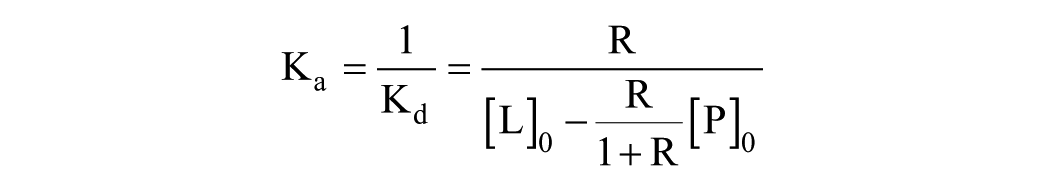Abstract
Native electrospray ionization mass spectrometry (ESI-MS) was applied to analyze the binding of compounds generated during fragment-based drug discovery (FBDD) campaigns against two functionally distinct proteins, the X-linked inhibitor of apoptosis protein (XIAP) and cyclin-dependent kinase 2 (CDK2). Compounds of different molecular weights and a wide range of binding affinities obtained from the hits to leads and lead optimization stages of FBDD campaigns were studied, and their dissociation constants (Kd) were measured by native ESI-MS. We demonstrate that native ESI-MS has the potential to be applied to the stages of an FBDD campaign downstream of primary screening for the detection and quantification of protein–ligand binding. Native ESI-MS was used to derive Kd values for compounds binding to XIAP, and the dissociation of the complex between XIAP and a peptide derived from the second mitochondria-derived activator of caspases (SMAC) protein induced by one of the test compounds was also investigated. Affinities of compounds binding to CDK2 gave Kd values in the low nanomolar to low millimolar range, and Kd values generated by MS and isothermal titration calorimetry (ITC) followed the same trend for both proteins. Practical considerations for the application of native ESI-MS are discussed in detail.
Introduction
Fragment-based drug discovery (FBDD)1–3 is a pharmaceutical approach for generating small-molecule inhibitors. It involves the identification of small (< ~350 Da), low-affinity molecular fragments that are subsequently evolved into lead compounds by iterative cycles of medicinal chemistry.
A range of biophysical methods are used in FBDD, including nuclear magnetic resonance (NMR), protein x-ray crystallography, surface plasmon resonance (SPR), and thermal shift (Tm) measurement to study weaker interactions. The affinity of protein–ligand interactions during the first stage of an FBDD campaign would typically be in the micromolar to low tens of millimolar range and would decrease to nanomolar or subnanomolar for a lead compound after the hit-to-lead phase (H2L) and lead optimization (LO) stages. In the H2L phase, initial fragment hits are optimized into “lead” compounds that have good potency and other promising properties, and in the LO phase, a lead compound is further optimized so that the resulting “candidate drug” has all the in vitro and in vivo properties required to make it a suitable candidate for clinical trials. The various techniques to analyze ligand binding often require large amounts of purified protein and compounds, and may also be restricted in their use by factors such as compound solubility 3 and crystallizability. There may also be a requirement for immobilization of the protein on a surface (SPR) or the use of reporter molecules such as dyes (Tm) for fluorescence-based methods, and these can impose their own limitations on the analysis of protein–ligand binding events. Sensitive complementary techniques are therefore required to address these challenges and to support FBDD campaigns by detecting weak noncovalent binding and generating estimations of dissociation constants (Kd) values. Weak-affinity chromatography (WAC) combined with mass spectrometry (MS) is an emerging technology for fragment screening based on selective retention of fragments by a drug,4,5 and it is demonstrated here that native MS can also be applied during an FBDD campaign.
The use of native MS as a primary screening tool has previously been reported, but despite the development of native MS methods for studying noncovalent interactions between proteins and small molecules,6,7 as described in a 2013 review by Poulsen, 8 application of native MS to the LO and H2L phases of an FBDD campaign, downstream from the primary fragment screening, has not been widely reported.
Maple et al. 9 described a primary fragment-based screen of 157 compounds by native MS using a TriVersa NanoMate (Advion)10,11 for automated nanospray infusion of protein–ligand complexes, which was completed and analyzed in 6 h. A significant amount of time was required to achieve the optimal instrument conditions, but it was demonstrated that the throughput of native MS is comparable to that of NMR or isothermal titration calorimetry (ITC), and required less protein and compound. Native MS could therefore be integrated into an FBDD program for both screening and postscreen characterization of ligand binding. Woods et al. 12 and Drinkwater et al. 13 used native MS as a complementary method to X-ray crystallography, SPR, and ITC to demonstrate the applicability of native MS as a complementary technique in FBDD.
Native MS can preserve weakly bound protein–ligand complexes in the gas phase, is rapid and sensitive, uses comparatively low amounts of protein and compounds, and requires neither crystallization, derivatization, nor immobilization. It is able to measure dissociation constants over a wide dynamic range, and Kd values in the range of low nanomolar to high millimolar have previously been reported.14,15
In this study, the applicability of native MS to analyze the binding of early-stage, low-affinity fragments, as well as higher-affinity compounds, generated in the H2L and LO stages was investigated, and the key experimental parameters were determined. Compounds discovered by Astex’s FBDD platform16,17 against the X-linked inhibitor of apoptosis protein (XIAP), which regulates the apoptotic response, and cyclin-dependent kinase 2 (CDK2), a regulatory element for natural cell progression, were analyzed. The study was composed of a small set of compounds, which were hits from primary screens and still cover a wide affinity range and were used to demonstrate the utility of native MS.
Native MS yielded information about the stoichiometry of ligand binding to XIAP, and absolute binding affinities for four noncovalent protein–ligand complexes were determined. The relative binding strengths of the second mitochondrial activator of caspase (SMAC) consensus peptide and one of the test compounds targeting XIAP was also measured. Moreover, the ability of one of the test compounds to disrupt the XIAP-SMAC peptide interaction was analyzed in a competition format. For the purpose of validation, three of the four compounds were also analyzed by ITC for their binding to XIAP, and the results were in good agreement with MS results. Eight noncovalent CDK2–compound complexes were also studied, and their binding affinities were quantified by native MS. Binding affinities of five of the eight compounds analyzed by MS were also determined by ITC, and the Kd values were found to be in the low-nanomolar to low-millimolar range. Practical considerations and the limitations for the application of native MS during the H2L and LO stages of an FBDD campaign are discussed.
Experimental
All solvents were purchased from Sigma Aldrich (Buchs, Switzerland). Recombinant CDK2 and XIAP protein, SMAC peptide composed of nine amino acids from the C-terminus, and all compounds ( Suppl. Table S1 ) were obtained from Astex Pharmaceuticals. The protocol for the expression and purification of XIAP is published in Chessari et al., 16 and that of CDK2 can be found in the Supplemental Information .
ITC measurements were performed on a Microcal VP-ITC instrument (MicroCal, Northampton, MA) at 25 °C. XIAP measurements were done in two solutions: 20 mM ammonium acetate (NH4Ac), pH 7.5, and 50 mM HEPES, 100 mM NaCl, 1 mM tris(2-carboxyethyl)phosphine (TCEP), pH 7.5. Both solutions gave similar results, but HEPES was used for the ITC measurements because of its higher buffering capacity. For CDK2, a buffer of 50 mM NH4Ac and 5% DMSO, pH 7.5, was used for all measurements. All ITC experiments were configured with protein in the sample cell and compound in the injection syringe, and data were fitted to a single-site binding model using Origin 7.0 software.
Solubility measurements were carried out by 1H quantitative NMR (qNMR) at 500 MHz using the QUANTAS approach 18 with dual suppression (WGATE and PRESAT for the water resonance and PRESAT for the acetate resonance). Aliquots of small molecules (100 mM in DMSO-d6) were diluted 20-fold with buffer (50 mM NH4Ac, 15% D2O, pH = 7.5) to generate samples with nominal concentrations of 5 mM. A 2.5 mM sample of p-hydroxybenzoic acid, prepared by dilution of a 50 mM stock in DMSO-d6 × 20 with buffer, was used as the qNMR standard. Data were processed using MNova (qNMR plug-in) reprocessing software.
MS experiments were carried out under native-like conditions using 20–50 mM NH4Ac adjusted to pH 7.5. In the case of XIAP, 10 mM imidazole was added. Stock protein solutions were desalted and buffer exchanged against NH4Ac, and stock solutions of the SMAC peptide and compounds were prepared either in NH4Ac or in DMSO. The final concentration of DMSO in protein–ligand samples for MS analysis was 1% v/v. The protein concentration was kept constant at 5 μM, and the compound concentration was varied from 1 to 5000 μM. Electrospray ionization (ESI) spectra were acquired with a hybrid quadrupole time-of-flight mass spectrometer (Q-TOF ULTIMA, Waters/Micromass, Manchester, UK) in positive ion mode. Sample solutions were directly infused with gold/palladium-coated borosilicate glass nano-ESI emitters (Thermo Fisher Scientific, Reinach, Switzerland) using a commercial nano-ESI ion source (Waters/Micromass). The operating parameters of the MS were adjusted as follows: capillary voltage = 1.5 kV; cone and first ion tunnel RF1 voltages = 40 and 30–50 V, respectively; backing pressure = 0.3–0.5 bar; and the ion source was maintained at ambient temperature. The hexapole collision cell was filled with argon (purity 5.0, PanGas, Zurich, Switzerland) and the collision energy offset was 2–5 V. The collision energy was kept low to minimize dissociation of the noncovalent complexes in the gas phase.
To determine Kd values, the experimentally derived relative peak areas were used. It was assumed that the ionization efficiencies for the apoprotein and the complex were equal, which allowed the use of the peak area ratios of the free protein and the complex from the mass spectrum instead of their concentrations. In cases where the compound is small compared with the protein, such that the size and surface properties of the protein and the complex are similar, uniform ionization efficiencies are expected. 19 These conditions apply to the studied complexes, since the mass difference between the protein and the complex with its highest-molecular-weight compound is < 3.5%. Two methods based on the same binding model were used: the titration approach where the data are fitted to the equation derived by Daniel et al.: 20
and the direct ESI-MS approach, 21 which uses the following equation:
where R = I(PL)/I(P) as defined above. In both cases, Kd was measured for each charge state independently as well as for the deconvoluted spectrum. The Kd calculations and the fitting of the titration curves were performed using MATLAB software (2013a, MathWorks, Natick, MA).
Results and Discussion
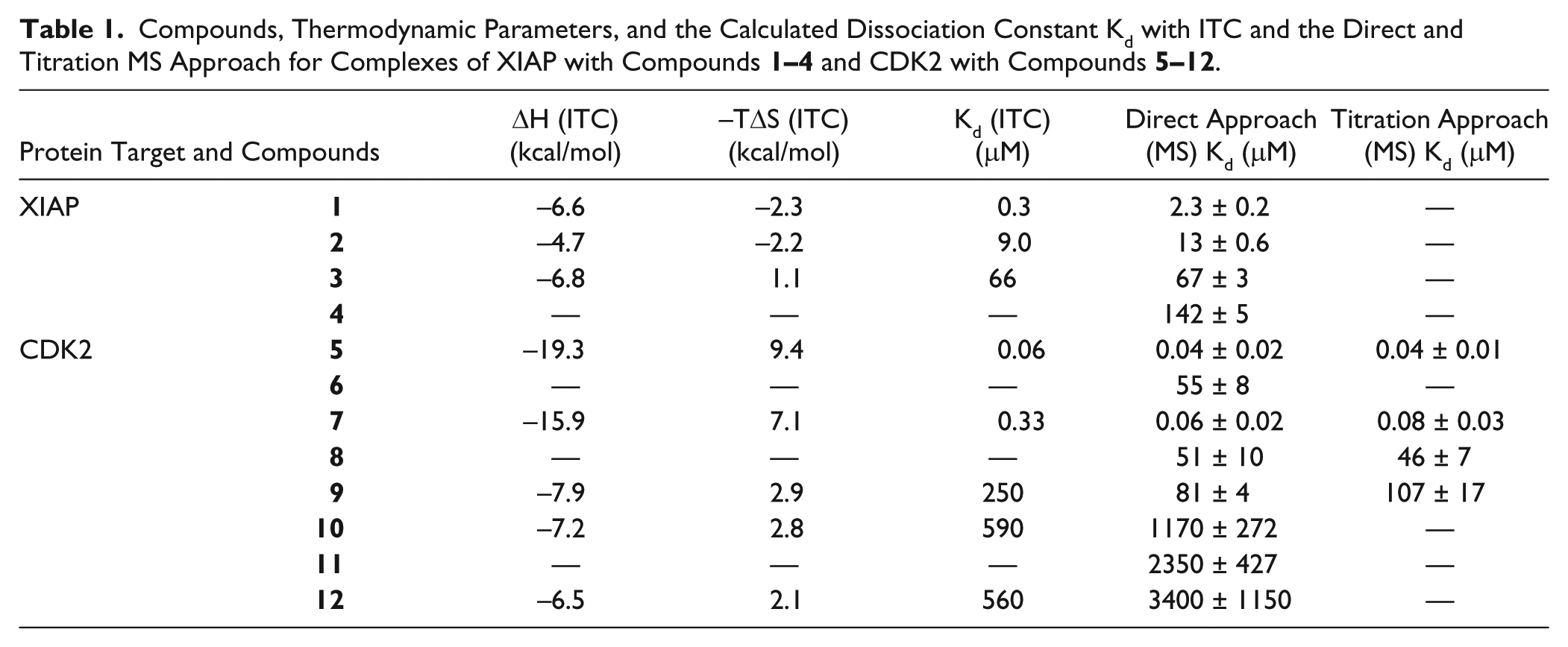
The chemical structures and physicochemical properties of the compounds used in this study are summarized in Table 1 and Supplemental Table S1 . In Supplemental Table S2 , the calculated mass difference between the protein–ligand complex and the apoprotein is given.
Compounds, Thermodynamic Parameters, and the Calculated Dissociation Constant Kd with ITC and the Direct and Titration MS Approach for Complexes of XIAP with Compounds
Mass spectra were obtained for 5 μM XIAP incubated with a 10-fold molar excess of the four compounds
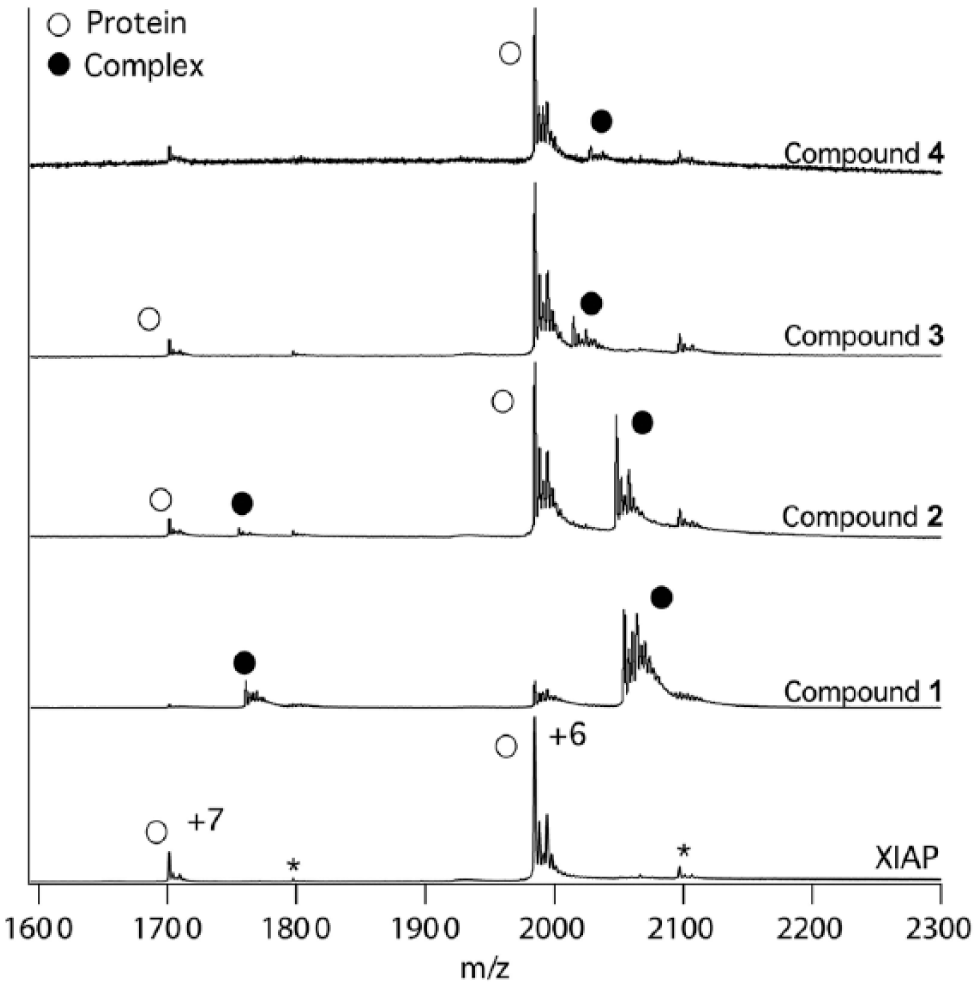
Representative nano-ESI-MS spectra of 5 μM XIAP in complex with 50 μM compounds
Zinc is required for a native conformation of XIAP, and the calculated mass (11,904 ± 2 Da) shows that XIAP was fully zinc bound, which proves that during the native MS experiments, it remained folded. For each charge state in
Figure 1
, a second peak was observed at m/z = 1800 (7+) and m/z = 2100 (6+) that had the same intensity in the presence of each of the compounds and was independent of the compound concentration. The mass difference between these peaks and those of the non-ligand-bound protein was 673 Da, which was subsequently shown by high-performance liquid chromatography (HPLC) fractionation and MS/MS peptide sequencing to be a hexapeptide with the sequence AVPYPQ, which is very similar to the high-affinity N-terminal consensus sequence of the SMAC protein, AVPI. The peptide was a component of the bacterial growth medium, which co-purified with the (inhibitor of apoptosis) IAP proteins and proved recalcitrant to complete removal despite multiple purification steps. Interestingly, this hexapeptide was also not competed away by high-affinity compounds or the SMAC peptide (
Suppl. Fig. S1
). This is not uncommon in protein–ligand binding experiments, and our data underscore that MS is capable of distinguishing the species that are present and that bind (or do not bind) to the protein investigated. The same approach was taken to compare binding of compounds
The Kd of compound
For Kd values < 100 μM, accurate fits for the titration curves were generated. However, at weaker affinities the R values will not change dramatically,
23
and in a titration experiment the ligand will not saturate the protein and the binding curve will tend to a straight line, as observed for the titration of compound
Of the compounds for which Kd values were determined by MS, five of the eight CDK2 compounds and three of the four XIAP compounds were also benchmarked against Kd values determined by ITC (see
Fig. 2
and Suppl. Fig. S6
). ITC experiments could not be done for all the compounds due to limited protein supply and/or compound insolubility, highlighting the key benefit of native MS, that it requires comparatively low quantities of protein. ITC data for the lowest and highest Kd values that could be confidently measured before compound solubility issues compromised these measurements (e.g., compound
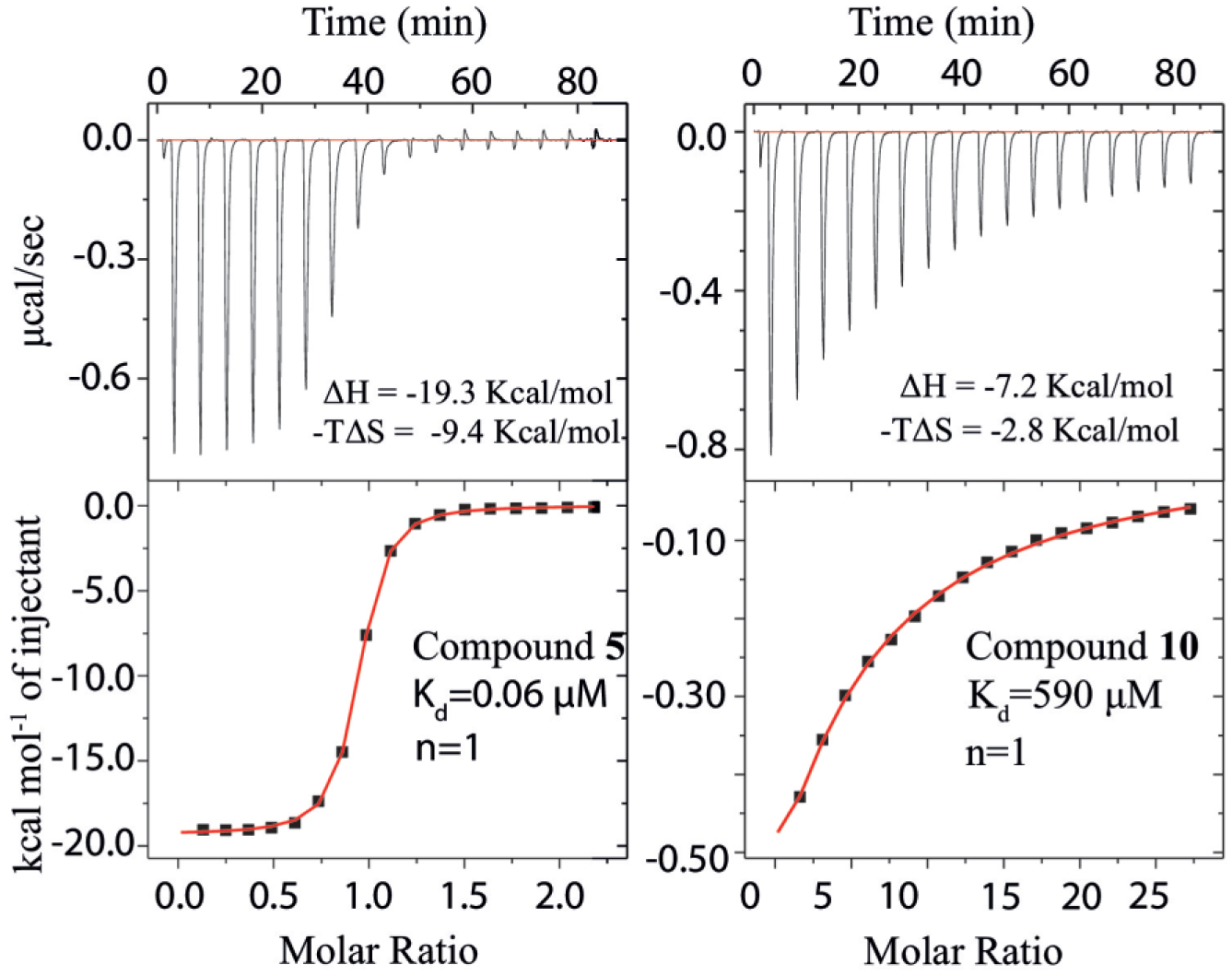
Measurement of thermodynamic parameters, including dissociation constant (Kd) and stoichiometry (n), by ITC for interactions between CDK2 with compounds
When designing native MS experiments for determination of accurate binding affinity values, it is essential to optimize the following factors: (1) compound concentration—the exact concentration of the protein and the compound, as well as the solubility of the compound, must be known to reliably measure affinities; (2) solvent—a volatile buffer suitable for ESI-MS must be used to ensure protein stability and compound solubility; (3) MS parameters—they require fine control in order to minimize gas-phase dissociation of weak protein–ligand complexes and simultaneously desolvate and desorb adducts; and (4) protein–ligand ratio—this must be controlled to minimize nonspecific binding of the ligand to the protein.
Compound Concentration
DMSO is used routinely for fragment solubilization and storage prior to use in a screening experiment. The final sample will therefore contain a small percentage of DMSO, typically 1%–5%, so the effect of DMSO on CDK2 was investigated ( Suppl. Fig. S7 ). DMSO concentrations up to 5% were tested for the effect on charge state distribution, and a decrease in the number of charge states was observed at higher DMSO concentration, suggesting compaction of the protein, which is in agreement with previously published observations. 26 DMSO concentrations of < 10% result in narrow charge distributions; however, more charging is observed at higher DMSO concentrations, indicating that supercharging may be causing the protein to unfold, and this may alter the binding affinity of noncovalent complexes. 27 It is therefore critical that the DMSO concentration be kept as low as possible.
Poor solubility of some fragments may render accurate concentration determination difficult even in the presence of a high percentage of DMSO; for instance, compound
ESI-Compatible Buffer
In order to detect weakly bound compounds, the signal-to-noise (S/N) ratio of the protein–ligand complex was optimized ( Suppl. Fig. S8 ). ESI is sensitive to nonvolatile solvent additives, such as buffer salts; therefore, protein solutions are electrosprayed from a solvent that differs from the original purification or storage buffer. Consequently, an effective desalting step is critical to ensure the minimum level of adduction of the protein by nonvolatile buffer components. 28 In this study, small protein sample volumes were desalted by centrifugal gel filtration using Micro Bio-Spin P-6 Gel Columns (BIORAD, Cressier, Switzerland) in NH4Ac, or by centrifugal filtration (Amicon Ultra-0.5 10K, Merck Millipore, Darmstadt, Germany) with up to three cycles of dilution and concentration to exchange the nonvolatile buffer for NH4Ac.
It has been demonstrated previously 29 that addition of imidazole at a high concentration to the nano-ESI solution can increase the stability of gas-phase complexes, therefore representing the solution-phase species more accurately. Imidazole acts as a nonspecific, sacrificial ligand, which prevents dissociation of the specifically bound ligand by enhanced evaporative cooling of the complex. A final concentration of 10 mM imidazole was added to XIAP–ligand complexes, which resulted in an increase in the relative abundance of the complex ions and a higher S/N ( Suppl. Fig. S9 ). No difference was observed in the intensities of the CDK2–ligand ion peaks in the presence of imidazole, which shows that each protein has to be individually optimized for native MS experiments.
MS Parameters: Collisional Dissociation
The collision energy was optimized to minimize dissociation of weak complexes in the gas phase, as this would lead to artificially high Kd values and false-negative results, which would not be representative of the solution phase. An example of a false-negative result in which complete dissociation of the protein–ligand complex was observed can be seen in Figure 3 , top panel. Conversely, insufficient collision energy led to incomplete desolvation and a broad adduct peak distribution, which rendered the data uninterpretable ( Fig. 3 , bottom panel).
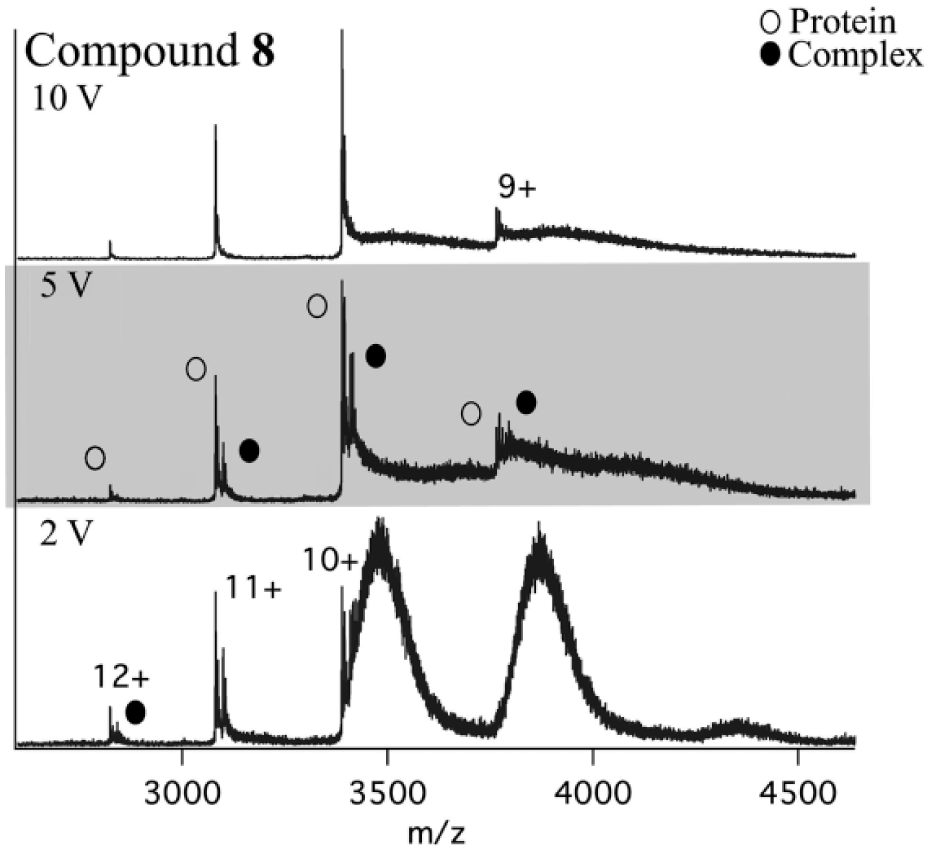
Native ESI-MS spectra of noncovalent complex of 60 μM compound
Collision-induced dissociation (CID) experiments were performed with XIAP to investigate the stability of the noncovalent XIAP–compound complexes in the gas phase. The collision energy offset was varied until the selected parent ions of the noncovalent complexes were completely dissociated, and the optimum collision energy to study XIAP complexes was determined from a plot of the normalized percentage of the intact complex against the collision energy offset ( Suppl. Fig. S10 ).
Nonspecific Protein–Ligand Interactions
High concentrations of compounds may lead to nonspecific interactions with the target protein ( Fig. 4 ). According to the electrospray model, 30 the initial ESI droplets undergo solvent evaporation until the Rayleigh limit is reached, at which point they undergo fission, releasing several multiply charged droplets containing none, one, or multiple molecules of analyte. The probability of a droplet containing more than one analyte molecule is increased at higher analyte concentration; therefore, the occurrence of nonspecific binding would also be expected to increase in that instance. For weak binders, a high concentration of compounds was used; therefore, the probability of nonspecific interactions occurring is increased. In this experiment, nonspecific binding was clearly observed when the compound was at a 30-fold molar excess with regard to protein, but was reduced at a 20-fold molar excess of compound and was eliminated at a 10-fold molar excess. A mathematical approach has been developed 31 to correct the calculation of Kd values in cases where nonspecific binding occurs, which yielded Kd values similar to those derived in solution-phase experiments. Additionally, solution-phase competition experiments with a higher-affinity tool compound monitored by ESI-MS can discriminate specific from nonspecific interactions. 22
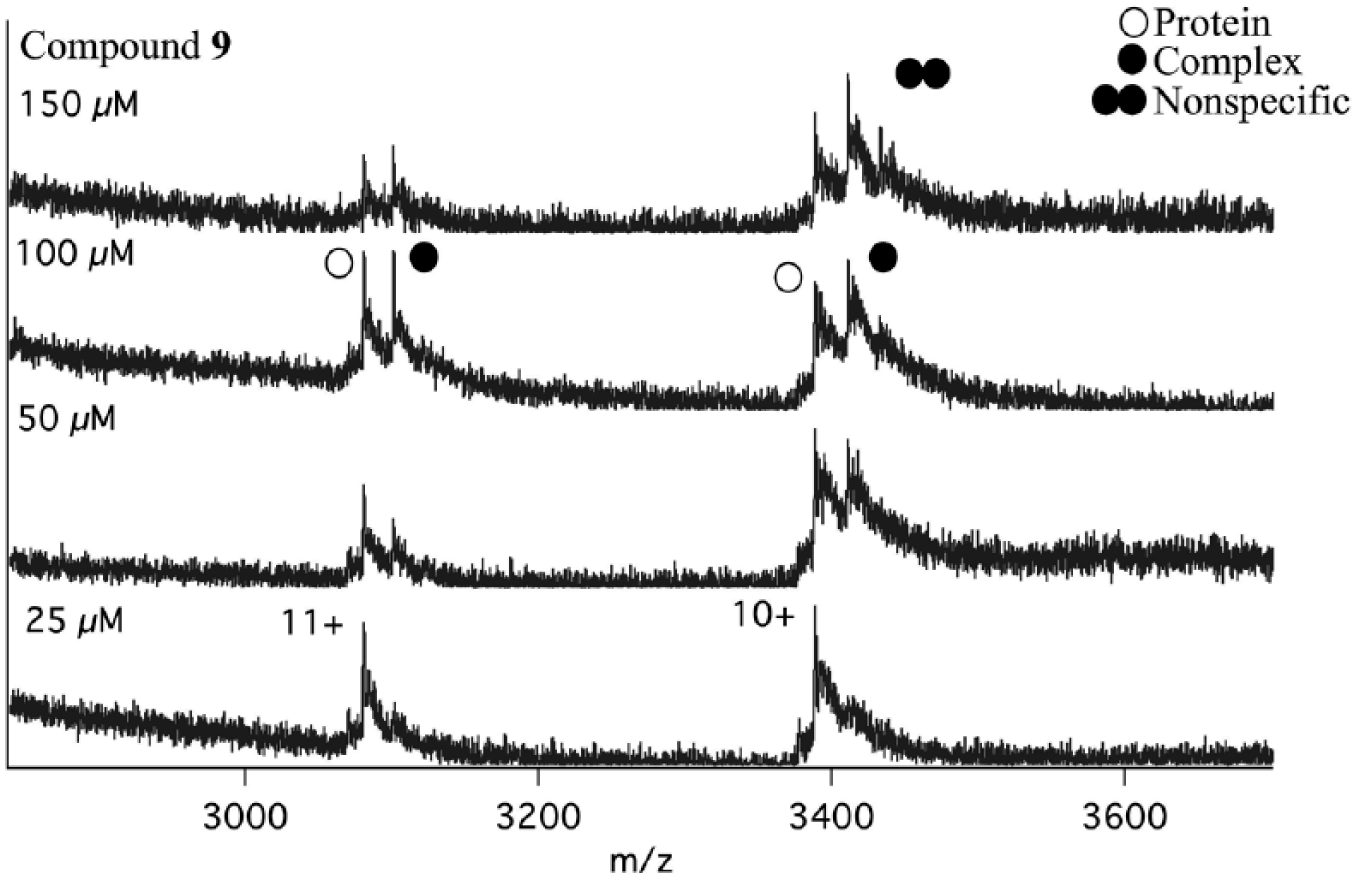
Native ESI-MS spectra of titration of compound
Native MS experiments can also be configured in a competition format, which has the advantage that it can directly visualize the replacement of one ligand with a competitive ligand via the observed mass difference between the various protein–ligand species. The influence of test compounds
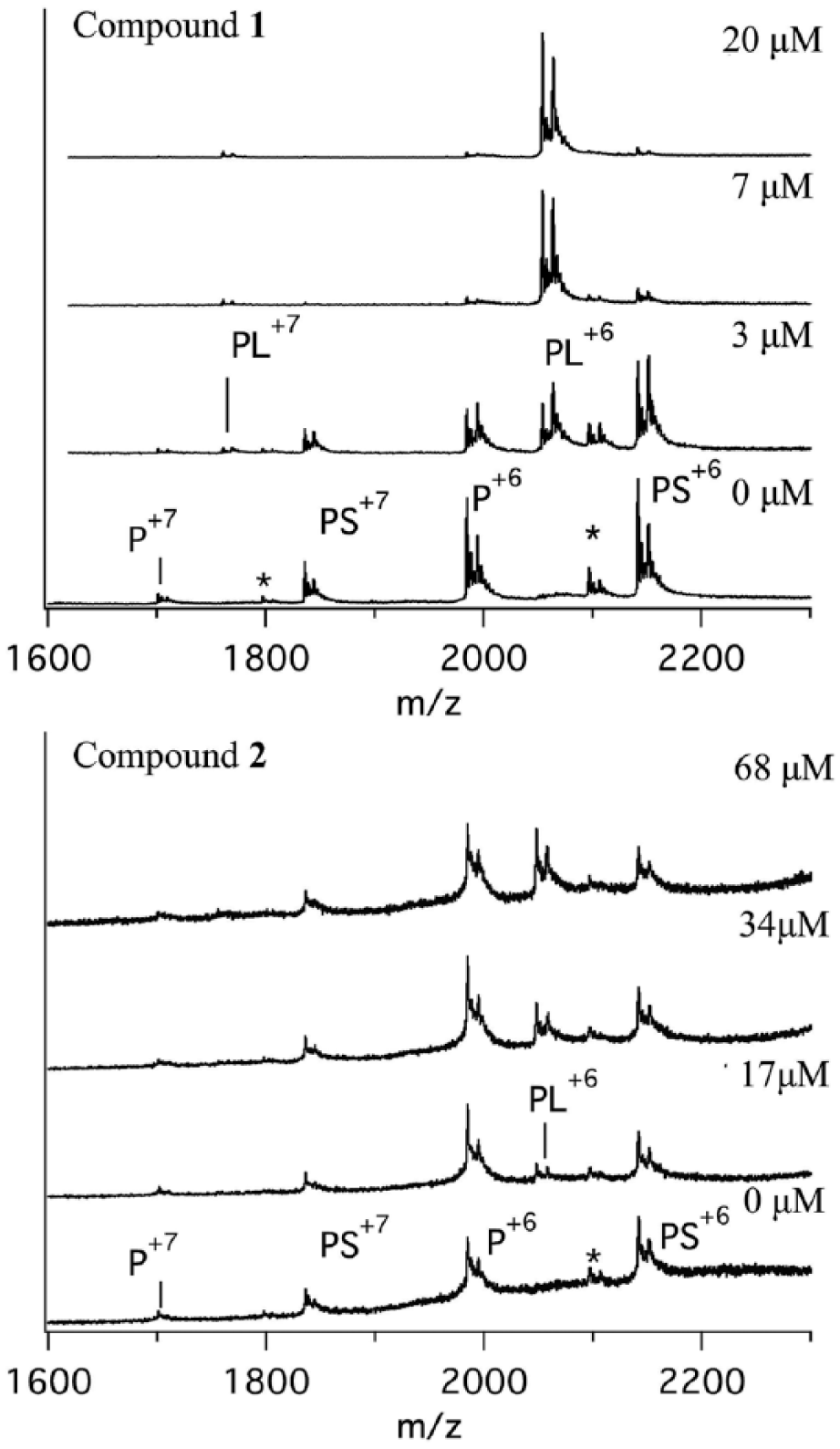
Nano-ESI mass spectra of XIAP-SMAC peptide complexes (PS) in the presence of different concentrations of the compounds (L). The PS signal clearly decreased with increasing concentration of compound
Various techniques are applied in the course of an FBDD campaign to analyze a large number of fragments and related molecules derived from successive cycles of medicinal chemistry. Each of these techniques is subject to its own limitations, such as high protein consumption due to low compound binding affinity and/or solubility, or a requirement for labeling or immobilization. Native MS is a rapid and sensitive technique and can provide a direct measure of ligand binding stoichiometry and affinity for ranking compounds. The use of native MS as an orthogonal screen for hit generation by the detection of weak noncovalent binding fragments has already been demonstrated previously, 9 but it can also serve as a powerful tool to complement the other stages of the FBDD process. 7 In this study, the aim was to investigate the extended applicability of the technique to not only fragments but also higher-affinity compounds generated in the H2L and LO stages of two Astex FBDD campaigns against CDK2 and XIAP.
The conditions required to generate reliable binding data for complexes of XIAP and CDK2 with FBDD-derived compounds, which possess a broad range of affinities, were tested. These included compound solubility, ESI buffer compatibility, MS parameters, and the protein–ligand ratio.
The data indicated that with careful experimental design, native MS can reliably detect noncovalent protein–ligand binding, enabling determination of Kd values and ranking of relative binding affinities over a wide dynamic range. In an FBDD environment, multiple factors influence the design of the experimental approach,2,3 and an important consideration is the availability of the relevant protein, which may be limited and thereby restrict application of techniques such as X-ray crystallography, NMR, and ITC, which require large amounts of protein. Native MS can help to address this issue, but each protein target must be individually evaluated for its suitability for MS-based ligand binding experiments. The protein must, however, be stable in MS solvent over the time frame of the experiment, and the effect of DMSO-containing buffers on the protein and its complexes must be determined. Experimental conditions should be configured to ensure that the native conformation is retained, which is competent to form a complex with low-affinity ligands that do not dissociate in the gas phase. The experimental procedure must remove adducts in order that weak ligand binding with low abundance can be discerned from spectral noise, and ligand binding to low-molecular-weight compounds can be sufficiently resolved from the apoprotein peak. Sample preparation and analysis is comparatively rapid, and thus all the proposed optimization steps may be achieved within an appropriate time frame in the context of a drug discovery project.
Current MS technology is capable of analyzing protein–protein interactions, including those involving membrane proteins,32,33 and while analysis of fragments binding to these proteins may not yet be possible, it is conceivable that fragment binding may be detectable using native MS methods in the future. To further augment the analytical platform available for compounds derived from the H2L and LO phases of FBDD campaigns, we show here that native nano-ESI-MS is a technique that can be applied to generate a better understanding of protein–ligand interactions, especially when protein availability, compound solubility, and other factors may be limiting for other techniques.
Supplemental Material
SI_for_Application_of_Native_ESI-MS_to_FBDD_by_Gavriilidou_et_al – Supplemental material for Application of Native ESI-MS to Characterize Interactions between Compounds Derived from Fragment-Based Discovery Campaigns and Two Pharmaceutically Relevant Proteins
Supplemental material, SI_for_Application_of_Native_ESI-MS_to_FBDD_by_Gavriilidou_et_al for Application of Native ESI-MS to Characterize Interactions between Compounds Derived from Fragment-Based Discovery Campaigns and Two Pharmaceutically Relevant Proteins by Agni F. M. Gavriilidou, Finn P. Holding, Joseph E. Coyle and Renato Zenobi in SLAS Discovery
Footnotes
Acknowledgements
We are grateful to Dr. Jing Dong for his assistance in protein production and Dr. Torren Peakman for performing the NMR experiments to determine the compound solubility. We thank the Swiss National Science Foundation (grant nos. 200020_140398 and 200020_159929) for financial support of this research. All original data, evaluations, plots, and so forth, will be stored in a curated data archive at ETH Zurich (![]() ) under the DOI 10.3929/ethz-b-000264348.
) under the DOI 10.3929/ethz-b-000264348.
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.