Abstract
The generation of cell extensions is critical for matrix remodeling in tissue invasion by cancer cells, but current methods for identifying molecules that regulate cell extension formation and matrix remodeling are not well adapted for screening purposes. We applied a grid-supported, floating collagen gel system (~100 Pa stiffness) to examine cell extension formation, collagen compaction, and collagen degradation in a single assay. With the use of cultured diploid fibroblasts, a fibroblast cell line, and two cancer cell lines, we found that compared with attached collagen gels (~2800 Pa), the mean number and length of cell extensions were respectively greater in the floating gels. In assessing specific processes in cell extension formation, compared with controls, the number of cell extensions was reduced by latrunculin B, β1 integrin blockade, and a formin FH2 domain inhibitor. Screening of a kinase inhibitor library (480 compounds) with the floating gel assay showed that compared with vehicle-treated cells, there were large reductions of collagen compaction, pericellular collagen degradation, and number of cell extensions after treatment with SB431542, SIS3, Fasudil, GSK650394, and PKC-412. These data indicate that the grid-supported floating collagen gel model can be used to screen for inhibitors of cell extension formation and critical matrix remodeling events associated with cancer cell invasion.
Introduction
Metastasis involves the dissemination of cancer cells from the primary tumor to distant sites and accounts for ~90% of cancer-related deaths. For developing new therapies to mitigate metastasis, a large array of resources and screening techniques have been applied, which include high-throughput methods for assessing the therapeutic efficacy of molecules from chemical libraries, 1 alternative splicing programs via binding RNA screening, 2 and somatic copy number screening. 3 A particularly useful approach is exemplified by a high-throughput assay that was developed to screen novel compounds 1 based on identification of drug target molecules in various cancer cell lines. 4
The metastatic process encompasses multiple, well-choreographed interconnected processes, of which local cell invasion is a critical, initiating step. The ability of cancer cells to invade is dependent on local degradation of the extracellular matrix (ECM) and cell migration through dense ECM barriers. These processes rely on tightly coordinated engagement of signaling pathways to alter cytoskeletal dynamics, pathological ECM production and remodeling, and increased cell motility. 5 One of the critical features of metastatic potential and the aggressiveness of individual tumors is their ability to cross tissue boundaries (e.g., basement membranes), which requires ECM degradation. Invasion is facilitated by the formation of actin-rich membrane protrusions (invadopodia), which enable cancer cells to adhere to the ECM and to translocate through matrix pores. 6 The mechanisms by which metastatic cells interact with and traverse the ECM are incompletely defined. This lack of understanding is multifaceted but is due in part to the current shortcomings of two-dimensional (2D) and three-dimensional (3D) models to model faithfully the structure and biochemical properties of extracellular tissue environments in vivo. Many cell culture models are widely used for mechanism-based studies and for drug discovery; these include multicellular tumor spheroids, tumor microenvironment models, co-culture cell assays, stem cell models, and simple model organisms. 7 Several model systems have been used to examine some of the fundamental processes in metastasis, including cell extension formation, cell adhesion, tail retraction, tunnel formation, matrix proteolysis, and tissue boundary crossing. The dimensionality of these systems (i.e., 2D or 3D culture) plays a key role in the fidelity of cell-based assays for modeling of in vivo situations and their applicability to early processes in cancer cell invasion.
Cell invasion into the matrix is often studied using costly, labor- and time-intensive, 2D stiff plastic substrates 8 or by measuring migration of cells into denuded zones in monolayers of cultured cells on or into 3D matrices. 9 None of these culture conditions provides insights into matrix remodeling in the tumor environment, which is critically important for understanding cancer metastasis and for identifying novel chemotherapeutics. 10 Cognizant of these limitations, we assessed the utility of a novel, grid-supported floating collagen gel system 11 that provides acute control of matrix geometry and physical boundary conditions and that measures cell extension formation, collagen compaction, and collagen degradation. With this system, we screened a kinase inhibitor library using several different cell types to identify potential molecules that block early steps in metastasis and collagen remodeling associated with cancer invasion.
Materials and Methods
Reagents
Bovine dermal type I collagen was purchased from Advanced BioMatrix (San Diego, CA; 6.0 mg/mL: ~97% type I collagen). Latrunculin B, NSC23766, ML-141, (±)-Blebbistatin, CK-666, SMIFH2, TGX221, Roscovitine, CGP57380, SB431542, PD0325901, genistein, and SIS3 were purchased from Calbiochem (San Diego, CA). PF573228, VX-680, and Fasudil-HCL were purchased from Selleckchem (Houston, TX). Cell-permeable C3 transferase was purchased from Cytoskeleton (Denver, CO). An antibody (clone 4B4; blocks β1 integrin function) was purchased from Beckman Coulter (Brea, CA). Nilotinib was purchased from Reagents Direct (Encinitas, CA). sunitinib malate, NSC109555, and GSK650394 were purchased from Sigma-Aldrich (Oakville, ON, Canada). NSC693868 was purchased from Tocris (Bristol, UK). Rabbit polyclonal Rac1/2/3 antibody, rabbit polyclonal CDC42 antibody, and rabbit polyclonal phospho-FAK (Tyr397) antibody were purchased from Cell Signaling (Danvers, MA). Mouse monoclonal antibody to Rho A was from Santa Cruz Biotechnology (Dallas, TX). Rabbit polyclonal FAK antibody and mouse monoclonal GAPDH antibody were purchased from Millipore (Darmstadt, Germany). An affinity purified rabbit antibody that recognizes the 3/4 type I collagen cleavage site was purchased from Immunoglobe (Himmelstadt, Germany). Alexa Fluor 568-conjugated goat antirabbit antibody, Alex Fluor 488-conjugated phalloidin, and rhodamine phalloidin were purchased from Invitrogen (Waltham, MA). IRDye 800CW antirabbit goat IgG (H + L) and IRDye 680RD antimouse goat antibody were obtained from Mandel (Guelph, ON).
Cell Culture
Diploid fibroblasts grown from human gingiva (HGFs) were obtained from healthy donors and used at passages 4 to 6. Cells were maintained in alpha-minimum essential medium (α-MEM) supplemented with 10% heat-inactivated fetal bovine serum (10% FBS) and 10% antibiotic solution (10% Ab; 146 U/mL penicillin G, 50 µg/mL gentamycin, and 0.25 µg/mL amphotericin). NIH 3T3 mouse fibroblasts were maintained in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% FBS and 10% antibiotics. Human breast adenocarcinoma cells (MCF7; low-grade metastatic cancer) were maintained in α-MEM supplemented with 10% FBS and penicillin and streptomycin (100 U/mL and 100 µg/mL). Human breast adenocarcinoma cells (MDA-MB-231; a highly metastatic cancer) were maintained in DMEM supplemented with 10% FBS and penicillin and streptomycin. Cells were maintained at 37 °C in a humidified atmosphere with 5% CO2.
Preparation of Floating Gels
Floating gels were prepared as previously described. 11 Briefly, nylon mesh sheets with square openings (200 µm × 200 µm) were purchased from Dynamic Aqua-Supply (Surrey, BC, Canada) and cut into pieces (2 cm × 2 cm or 0.7 cm × 0.7 cm). Collagen solutions (1 mg/mL) were prepared from bovine dermal type I collagen, neutralized with 0.1 M NaOH to pH = 7.4, and poured on to a flat, hydrophobic surface followed by placement of nylon meshes to fill the grids with collagen (~70 µL). Grids were incubated with collagen until polymerization was complete (~90 min) and were gently detached from the hydrophobic surface by addition of 1× phosphate-buffered saline (PBS). Floating gels were inverted and immersed in cell culture medium. Cells were planted at low densities to avoid the inclusion of more than one cell in single grid space and allowed to adhere and spread to the floating gels.
Preparation of Attached Collagen Gels
Collagen solutions (1 mg/mL) were prepared from bovine dermal type I collagen, neutralized with 0.1 M NaOH to pH = 7.4, and then poured on glass-bottom dishes or tissue culture plastic. Collagen gels were incubated for 5 min at room temperature and then air dried.
Collagen Stiffness
A cylindrical probe (flat probe tip = 25 µm in diameter), comparable to the diameters of spread cells, was used for indentation of collagen and estimation of stiffness. 12 Cyclic loading and unloading indentations were conducted on floating collagen gels and attached collagen gels. Loading/unloading indentation was analyzed by calculating irreversible deformation and maximum supported load.
Morphology and Imaging
Cells were fixed with 4% paraformaldehyde (PFA) and permeabilized in 0.2% PBS–Triton X-100. Cell morphology and cell extension formation were visualized by Alex Fluor 488–conjugated phalloidin (1:50) to stain actin filaments. Cell nuclei were stained with 4′,6-diamidino-2-phenylindole (5 µg/mL). Immunofluorescence images were collected with a 20× objective optically interfaced to an Eclipse TE300 inverted fluorescence microscope (Nikon, Melville, NY). Cell morphometry was performed with ImageJ software (National Institutes of Health, Bethesda, MD). Cell extensions were quantified as cell processes that extended from the cell body by >2 µm. The length of cell extensions was measured from the cell centroid to the tip of each extension. The mean length of cell extensions per cell was calculated by summing the length of all cell extensions for each cell and dividing by the number of extensions.
Rac1, CDC42, and Rho A Activation Assays
Pull-down assays were conducted as described. 13 HGFs (2 × 107 cells) were plated on collagen (1 mg/mL) in 15 cm dishes with vehicle or NSC23766 (50 µM, a Rac 1 inhibitor), ML-141 (10 µM, an inhibitor of cdc 42), and cell-permeable C3 transferase (2.0 µg/mL, an inhibitor of Rho) in 10% FBS/α-MEM. After 90 min, cells were lysed in lysis buffer at 4 °C. Cell lysates were incubated with glutathione transferase–fusion protein, which contained the p21-binding domain as expressed in Escherichia coli and bound to glutathione-Sepharose 4B beads (GE Healthcare, Piscataway, NJ) or with Rhotekin beads (Cytoskeleton) for 1 h at 4 °C. Samples were separated by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE; 12% gels) prior to immunoblot with the appropriate antibody to Rho, Rac1, or cdc42.
Immunoblotting
Proteins separated by SDS-PAGE were transferred to nitrocellulose membranes (Bio-Rad, Mississauga, ON, Canada). Membranes were blocked and probed with indicated antibodies at 4 °C overnight, followed by incubation with fluorescent secondary antibodies for 1 h at room temperature. Odyssey (Li-Cor, Lincoln, NE) and Image Studio software (Li-Cor) were used for detection and analysis of the immunoblots.
Compound Library
For testing the efficacy of the new assay for drug screening, we used a kinase inhibitor library, which consisted of 480 drugs, that was compiled by the Ontario Institute for Cancer Research. The library includes biologically active molecules, off-patent drugs, and natural products as described earlier. 14 Compounds for in-depth follow-up studies were obtained from Sigma-Aldrich.
Cell Extension Assay
Drug dilution and injection into culture wells was performed at the Simple Modular Assay and Robotic Technology (SMART facility; Mount Sinai Hospital, Toronto, ON, Canada). 1 For initial screening, NIH 3T3 cells were plated on collagen grids (size: 0.7 cm × 0.7 cm) at 1.4 × 103 cells per well in 200 µL of 10% FBS/DMEM with or without compounds in eight-well chamber slides. Drugs were immediately added to a final concentration of 1, 0.1, or 0.01 µM. After 6 h, 3T3 cells were fixed with 4% PFA, permeabilized in 0.2% PBS–Triton X-100, and stained as described above.
Confocal Microscopy and Local Collagen Fiber Alignment
Collagen fibers were imaged using confocal reflectance imaging with an inverted confocal laser scanning microscope (Leica TCS, Mannheim, Germany) with a 40× oil immersion objective lens. Fast Fourier transform (FFT) was used to extract the orientation of collagen fibers from acquired images using confocal reflectance microscopy as described. 11 To quantify the direction of the fibrils in the original image, the pixel intensities in the resulting FFT image were analyzed over 180° around the peripheries of cell extensions. The summation process was performed using Oval Profile, an ImageJ plug-in. The resulting plot was the sum of pixel intensities between 0° and 180°. A perfect random image would result in constant pixel intensity at different angles. The alignment index was quantified by calculation of the area under the intensity curve within ±10° of the peak intensity value.
Collagen Degradation
MDA-MB-231 cells (1.2 × 104 cells) were plated on collagen grids with vehicle or test compounds (0.01 µM) in 10% FBS/DMEM. Compounds were injected into cultures at 0 or 12 h. After 24 h, cells were fixed with 4% PFA, blocked with 1% bovine serum albumin in PBS, and immunostained with an antibody that recognizes a neoepitope in the collagen type I chain created by interstitial collagenase cleavage (3/4 collagen fragment, affinity purified rabbit antibody; 1:100) at 4 °C overnight. After washing with PBS, the cells were stained with Alexa Fluor 568–conjugated goat antirabbit secondary antibody (1:100) and Alexa 488 phalloidin (1:50). Cells were imaged with a confocal laser scanning microscope with a 40× oil immersion objective lens. Degraded collagen (i.e., 3/4 collagen fragments) was measured using Image J software. Briefly, in Image J, background signals (obtained with images of cells stained with isotype control antibodies) were subtracted using the sliding paraboloid. The local image contrast was enhanced with CLACHE plugin, which was followed by binary processing. The intensity of 3/4 collagen immunostaining was measured using the “Analyze Particles” procedure; 3/4 collagen immunostaining was normalized to nondegraded immunostained collagen.
Statistical Analysis
All continuous variable data are expressed as mean ± SEM. Statistical analyses were performed using one-way or two-way analysis of variance followed by Tukey test for multiple comparisons. A Student unpaired t test was used for comparisons involving two means. Statistical significance was set at p < 0.05. For all conditions and compounds that were evaluated, at least four different experiments with three internal replicates were analyzed. For imaging, at least 10 cells from three replicates from four different experiments were analyzed for each condition. The Z-factor (for estimating assay validity) of number of cell extensions was calculated as described. 15
Results
Cell Extension Formation in Floating and Attached Collagen Gels
Collagen matrices (2D and 3D) are widely used to model cell behavior in connective tissues, but several features of matrix structure and mechanics can strongly affect measurements that are made with these models. Here, we used floating collagen gels supported peripherally by nylon grids, a system that models a relatively soft interstitial matrix constrained by rigid physical boundaries at the border of each gel compartment (200 µm × 200 µm). This system facilitates integrated studies of cell-mediated collagen remodeling and cell extension formation 11 by providing a flat imaging plane (to enhance image quality) and in which matrix mechanics are more closely aligned with those of connective tissues in vivo compared with what is provided by matrix proteins attached to cell culture plastic. In this context, an indentation assay to evaluate the stiffness of the collagen showed that floating collagen gels were 28-fold softer than attached collagen gels (p < 0.001; Fig. 1A ).
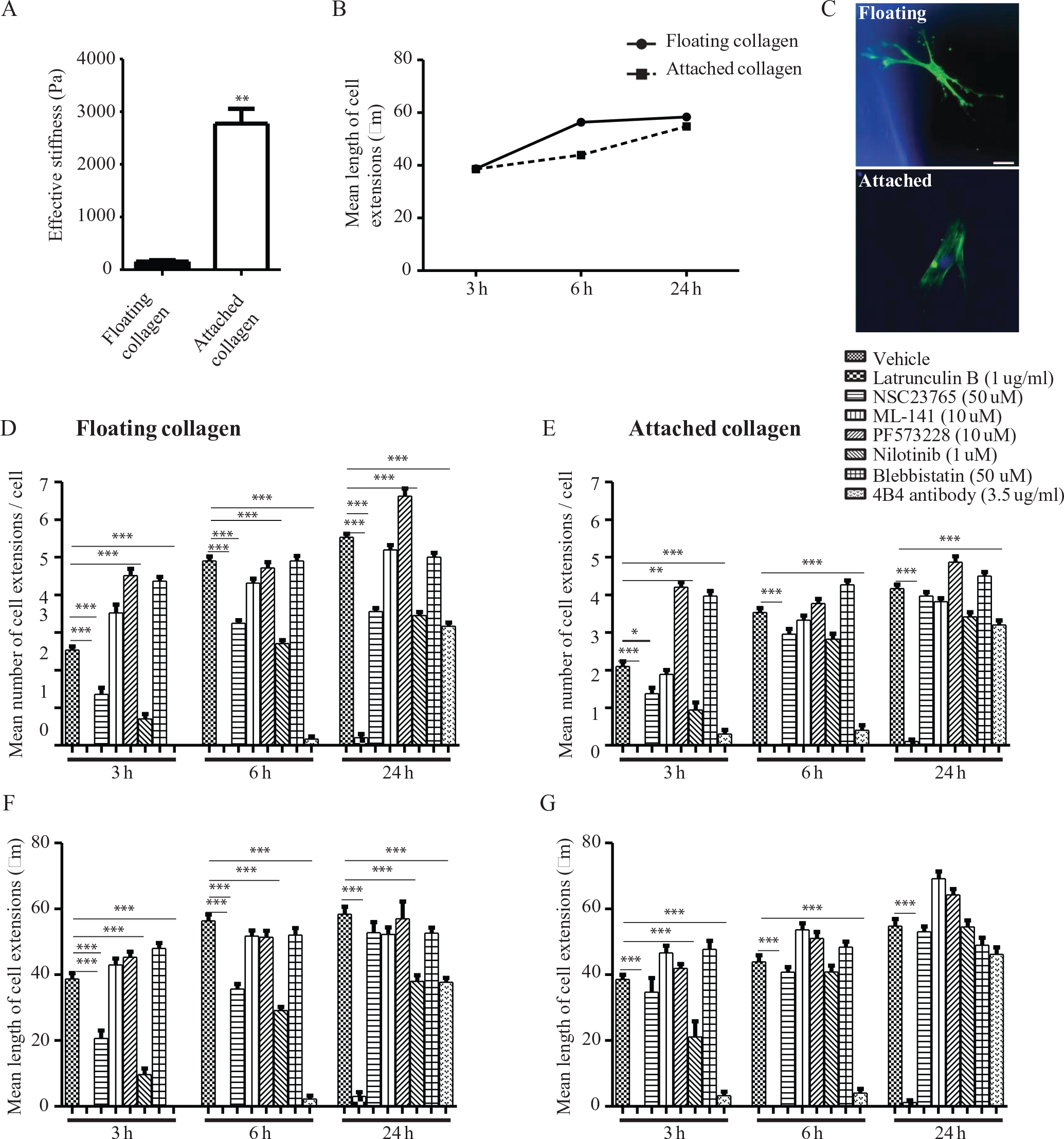
Collagen stiffness and effect of various inhibitors on cell extension formation. (
In studies of HGFs, diploid cells that rapidly remodel collagen matrices by local release of interstitial collagenases,
16
the mean length of cell extensions was ~1.3-fold longer in the floating collagen gel model than in attached collagen gels at 6 h after plating. For both culture models, the mean length of cell extensions increased progressively after time of plating (
Fig. 1B
We examined critical processes required for generation of cell extensions by diploid fibroblasts (HGFs) in the floating gel model with the use of specific inhibitors (
Fig. 1D
The effect of these same inhibitors on extension formation in cells plated on attached collagen gels was examined and compared with vehicle controls (
Fig. 1E
Comparisons of Cell Extension Formation in Different Cell Types
The experiments described above used the floating collagen gel assay to assess cell extension formation by HGFs. These cells can be used to evaluate certain aspects of the supporting peritumor stroma in vitro 18 but are not appropriate for studies of matrix invasion by epithelial cancer cells. Accordingly, we compared the effect of the inhibitors described above on four different cell types including the aforementioned HGFs, along with 3T3 cells (a mouse fibroblastic cell line), MCF7, and MDA-MB-231 cells ( Fig. 2A–D ). For cells treated with latrunculin B or 4B4 antibody, the mean number of cell extensions was reduced compared with untreated cells (p < 0.0001). Treatments of HGFs and 3T3 cells with nilotinib showed that the mean number of cell extensions was reduced compared with untreated cells (p < 0.0001). In contrast, MCF7 and MDA-MB-231 cells did not show inhibition of cell extension formations with these treatments (p < 0.0001 between cell groups), which is consistent with a recent study on the effect of nilotinib on matrix invasion by MDA-MB-231 cells. 19 For 3T3 cells treated with the FAK inhibitor, PF573228 (IC50 = 30–100 nM 20 ), the extent of inhibition of cell extension formation was markedly less than HGFs, while for MCF7 and MDA-MB0231 cells, cell extension formation was not affected by PF573228. These findings may reflect earlier data that inhibition of cell adhesion and extension formation by PF573228 are dependent on the expression of the fibroblast phenotype. 20
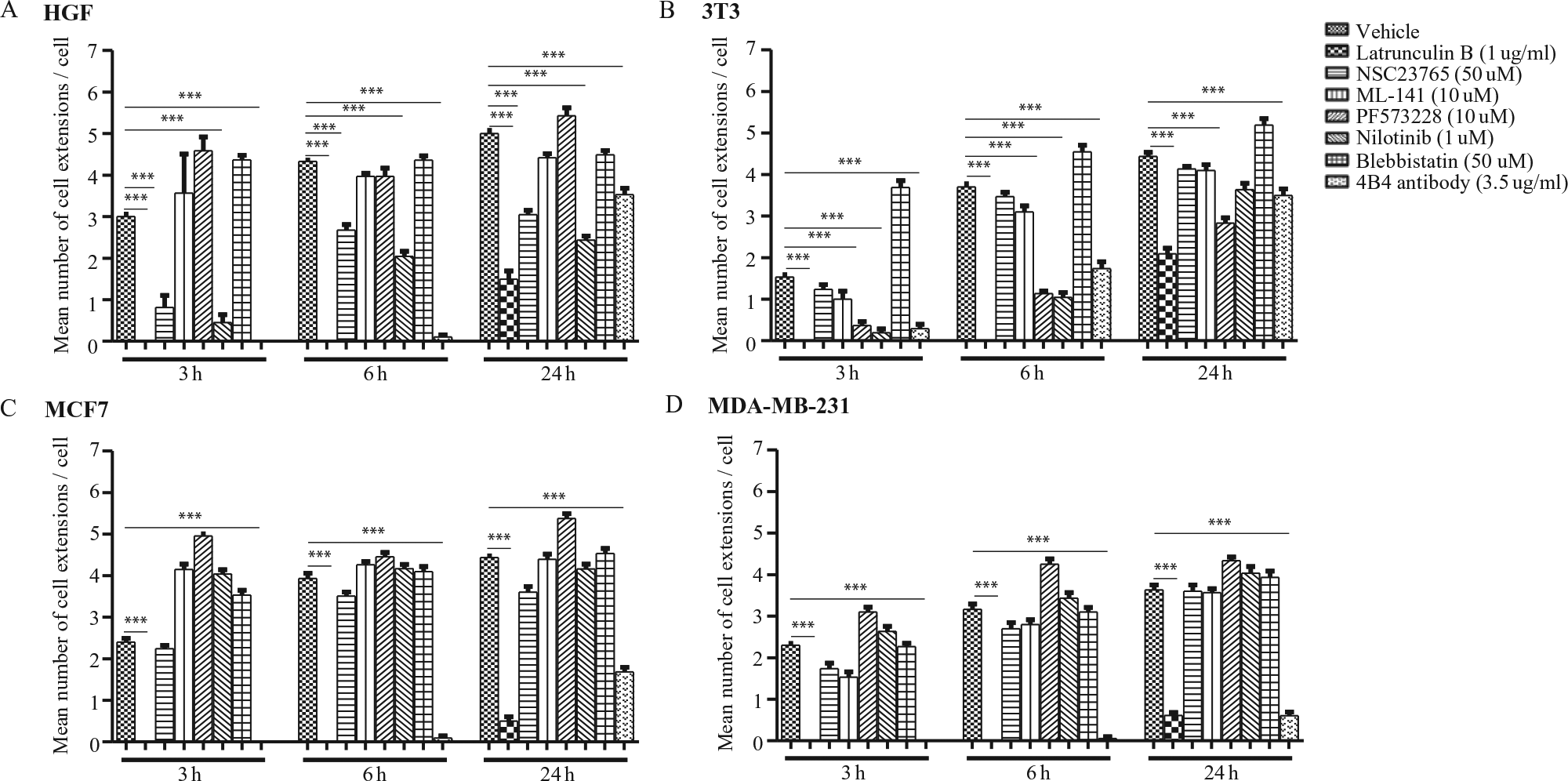
Effect of various inhibitors on cell extension formation in four different cell types. Human gingival fibroblasts (
Formin and Arp2/3 Inhibition of Cell Extension Formation in Diploid Fibroblasts
After attachment to matrix proteins such as collagen, anchorage-dependent cells extend protrusions, which are thought to manifest initially as filopodia or lamellipodia, organelles that are associated with different actin nucleation machineries (i.e., formins and Arp2/3, respectively,). 21 As described above, based on inhibitor studies used with the floating collagen gel system, Rac and cdc42, which are strongly associated with the generation of cell extensions, 22 did not appear to be particularly important for the formation of cell extensions. Notably, the formation of filopodia also involves, in part, actin nucleation by formins, 21 whereas lamellipodia formation is thought to involve Arp23. 21 Accordingly, we assessed whether cell extension formation in the floating collagen gel system was dependent on formins or the Arp2/3 signaling pathways. We treated HGFs for 6 h with SMIFH2 (a formin FH2 domain inhibitor; IC50 = 28 µM; 1–100 µM) 23 or with CK-666 (an Arp2/3 complex inhibitor; IC50 = 17 µM; 1–200 µM) 24 using concentration ranges based on previous studies. 23 Compared with vehicle-treated HGFs, the mean number of cell extensions was dose dependently reduced by SMIFH2 (~89% reduction with 10 µM SMIFH2; Fig. 3A ). However, treatment with CK-666 did not affect cell extension formation compared with vehicle controls ( Fig. 3B ).
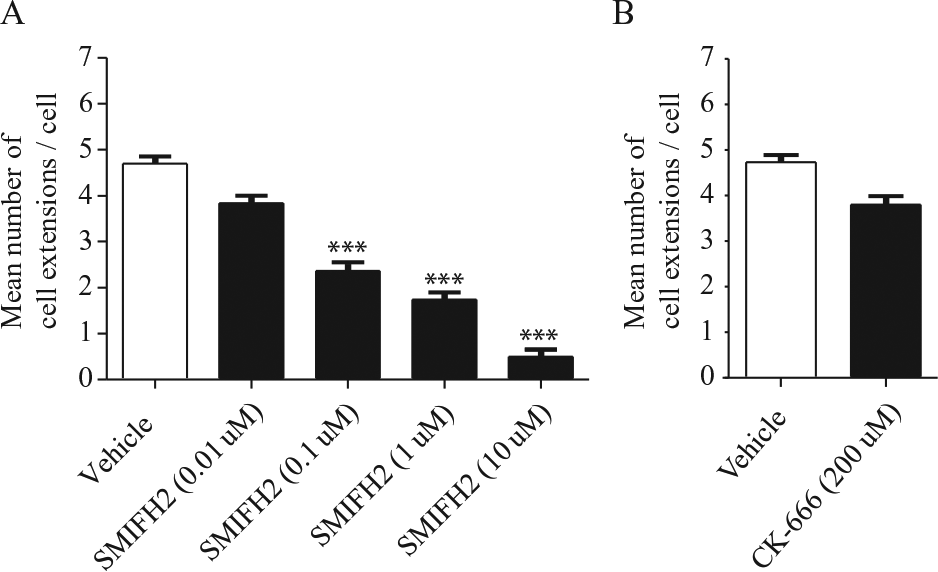
Effect of SMIFH2 and CK-666 inhibitor on cell extension formation in human gingival fibroblasts (HGFs). HGFs were planted on floating collagen gels with or without SMIFH2 (0.01, 0.1, 1, or 10 µM) (
Screening of a Kinase Inhibitor Library
The time required for image analysis described for the data above was ~2 min per well (i.e., after image acquisition). This finding suggested that the floating collagen gel assay may enable rapid and facile measurements of cell extension formation that can be probed with well-described inhibitors that are likely to affect fundamental mechanisms of extension formation. The unexpected finding that a kinase inhibitor (i.e., nilotinib), which selectively blocked cell extension formation in the floating gel model but not the attached collagen gel model, suggested the idea of using the floating gel system to screen a kinase inhibitor library and identify potential new regulators of cell extension formation (kinase inhibitor library: 480 compounds examined at 0.01, 0.1, and 1 µM). Based on inhibition of cell extension formation measured at 6 h, we selected compounds that strongly affected cell extension formation compared with vehicle-treated cells (i.e., >50% reduction in cell extension formation). From screening of these 480 compounds, we found that, in particular, 14 compounds (at 0.01 µM) significantly reduced (p < 0.01) cell extension formation, collagen remodeling, and collagen degradation ( Table 1 ). Based on reduction of the number of cell extensions by >50%, we performed detailed studies of cell extension formation, collagen degradation, and collagen compaction with the floating collagen gel model for these 14 molecules.
Hit Selection of Compounds.
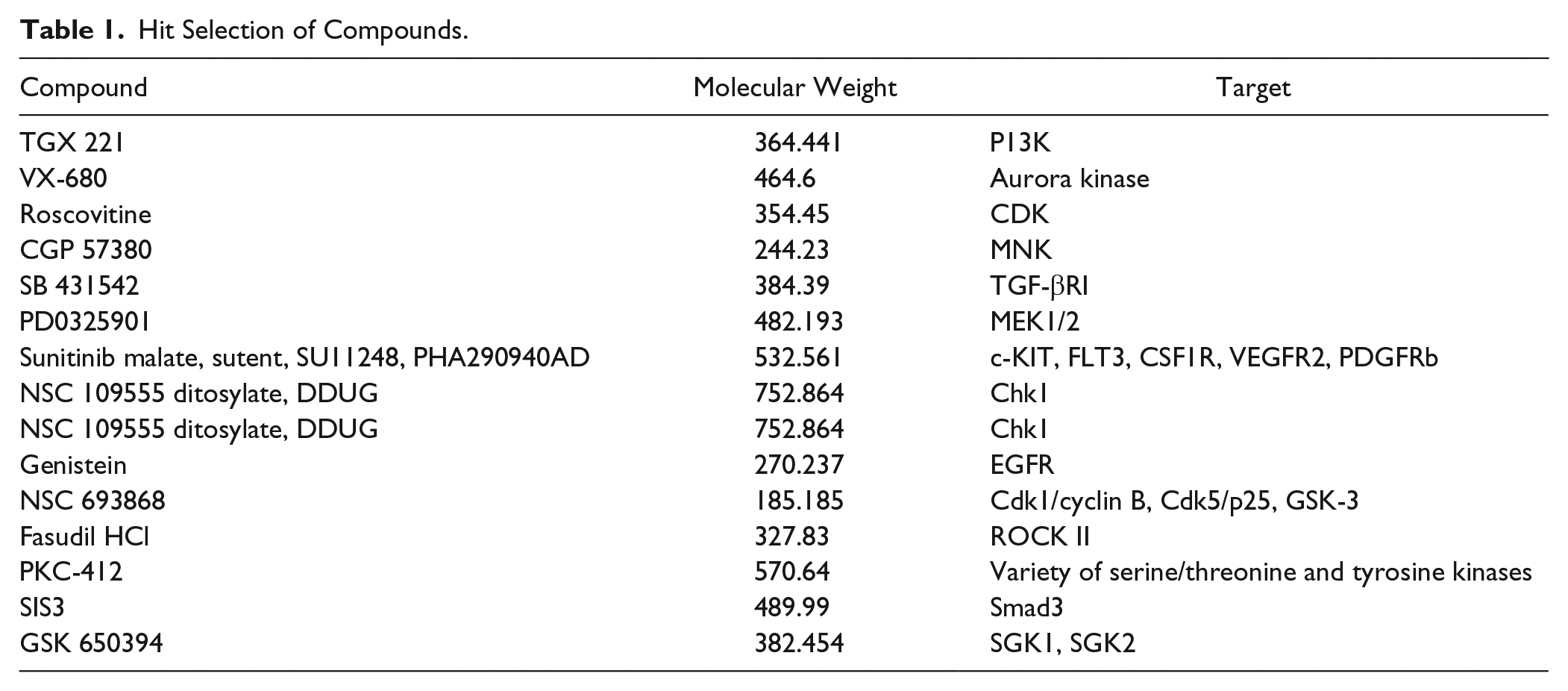
We initially examined HGFs as they express DDR1, 25 rapidly form cell extensions, and express high levels of interstitial collagenases that degrade ECM collagen. 16 HGFs were treated for 24 h with the kinase inhibitors at a low concentration (0.01 µM) to assess which of these inhibitors is most likely to affect cell extension formation and collagen remodeling at relevant pharmacological dosages. Treatment with TGX221, VX680, SB431542, Fasudil, PKC-412, SIS3, and GSK650394 reduced collagen alignment (p < 0.0001) and the number of cell extensions (p < 0.0001) compared with vehicle-treated cells ( Fig. 4A–C ). HGFs treated with PD0325901, sunitinib, genistein, or NSC693868 showed a reduced number of cell extensions compared with vehicle-treated HGFs ( Fig. 4A–C ).
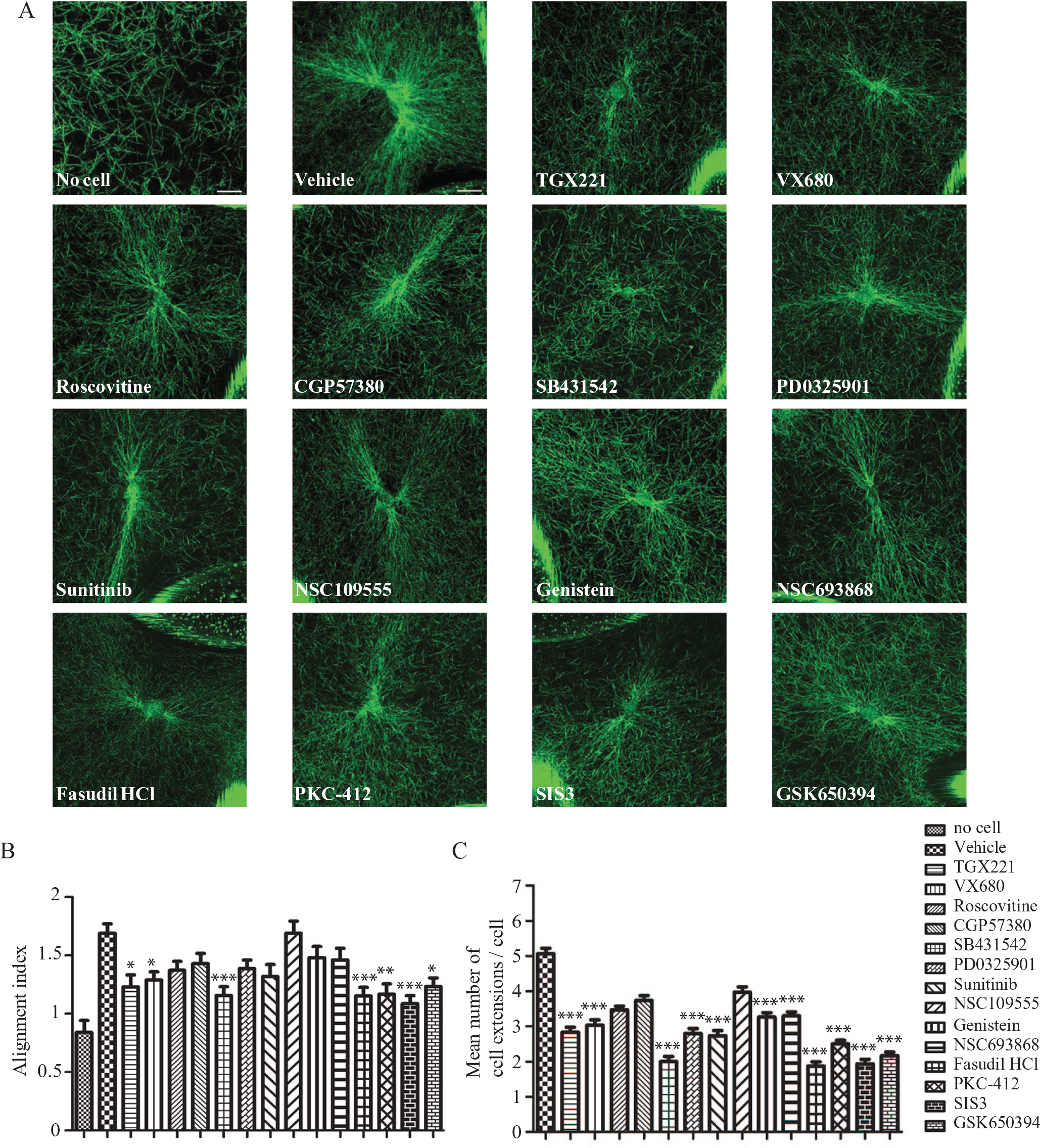
Effect of 14 kinase inhibitors on cell-induced aliment of collagen network in human gingival fibroblasts (HGFs). HGFs were plated on floating collagen gels with or without 0.01 µM TGX221, VX680, roscovitine, CGP57380, SB431542, PD0325901, sunitinib, NSC109555, genistein, NSC693868, Fasudil HCl, PKC-412, SIS3, and GSK650394 for 6 h. Collagen fibers were imaged using confocal reflectance microscopy (
When triple helical type I collagen is cleaved by interstitial collagenases, 3/4 and 1/4 collagen fragments are generated as a result of the hydrolysis of a single Gly-Ile/Leu bond in each of the collagen α chains. 26 We assessed collagen degradation by MDA-MB-231 cells with the use of a neoepitope antibody that recognizes 3/4 collagen fragments. 27 We used this assay because MT1-MMP and MMP2, which contribute to invasion and collagen degradation, are strongly expressed by MDA-MB-231 cells. 27 In cells treated with the 14 kinase inhibitors identified above (0.01 µM for 24 h), cell extension formation was reduced (>50%; p < 0.0001). This time of treatment (24 h) was based on previous in vitro studies that evaluated cancer cell invasion into collagen matrices. 28
MDA-MB-231 cells treated with CGP57380, SB431542, PD0325901, sunitinib, NSC693868, Fasudil, PKC-412, SIS3, and GSK650394 also exhibited greatly reduced pericellular collagen degradation (
Fig. 5A
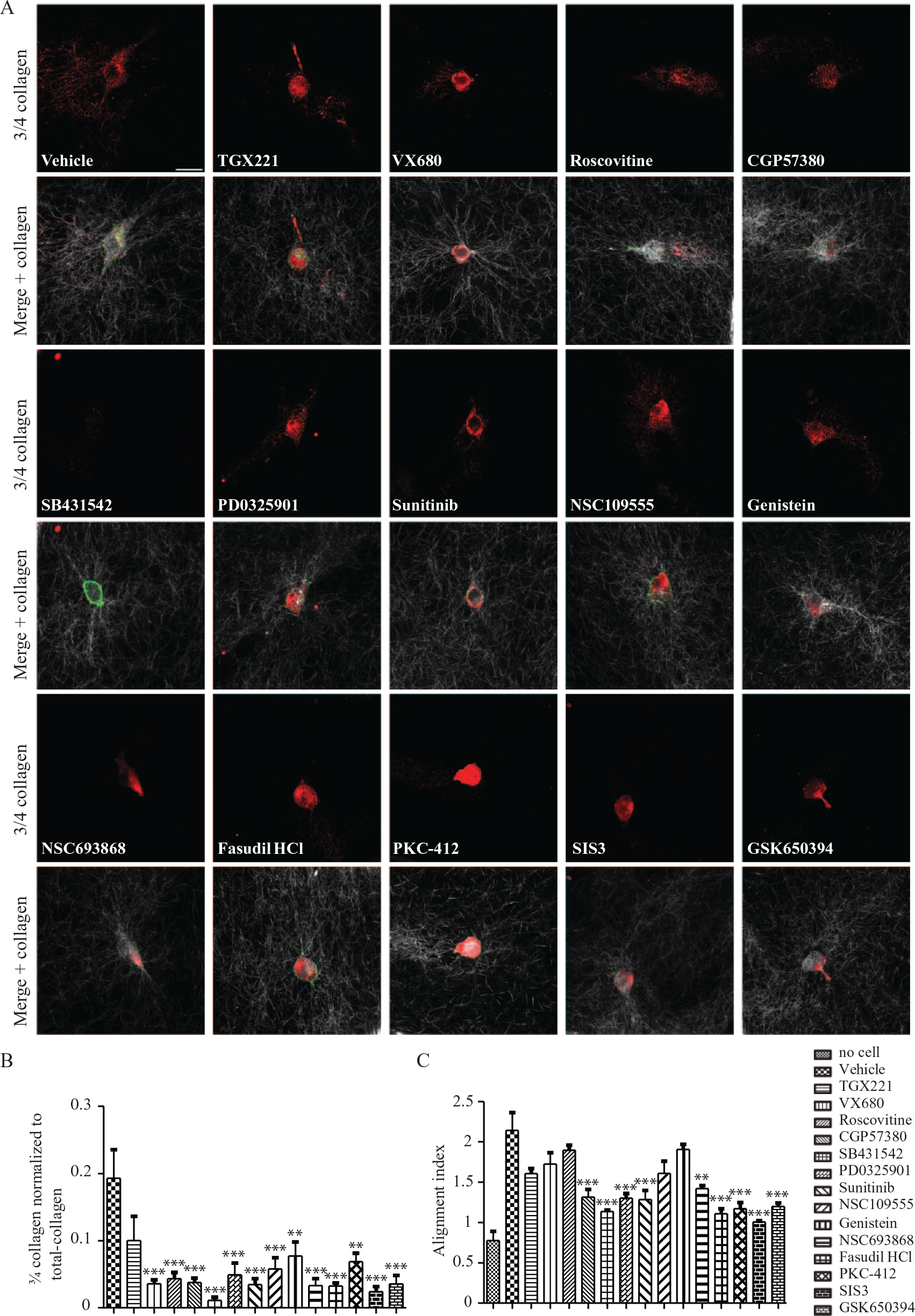
Effect of 14 kinase inhibitors on collagen degradation and collagen alignment in MDA-MB-231 cells. MDA-MB-231 cells were plated on floating collagen gels with or without 0.01 µM TGX221, VX680, roscovitine, CGP57380, SB431542, PD0325901, sunitinib, NSC109555 genistein, NSC693868, Fasudil HCl, PKC-412, SIS3, and GSK650394 for 24 h. Immunofluorescence staining for cleaved collagen (3/4 collagen; red), F-actin (green), and collagen fibers (white/gray) (
Discussion
Our main finding is that a grid-supported, floating collagen gel model system 11 can be used to provide rapid, integrated assessments of cell extension formation, collagen alignment, and collagen degradation by cancer cells using a tyrosine kinase inhibitor library as a test case for efficacy of screening. The data indicate that this system can be used to screen for potential therapeutics that target early events in cancer metastasis. The assay showed strong inhibition by a Rock II inhibitor, which reduces cell spreading and the formation of stress fibers in hepatic stellate cells. 29 We also found that several compounds strongly inhibited cell extension formation, including SB431542, which is a selective inhibitor of Smad3 phosphorylation, and SIS3, which blocks transforming growth factor β–induced phosphorylation of Smad3. Notably, Smad3 activation has been observed in fibrotic areas of cancer cachexia adipose tissue. 30 The assay also showed strong inhibition of cell extension formation by SGK1, which was earlier demonstrated to regulate cell proliferation and differentiation in RKO cells, a human colon carcinoma cell line. 31 Further, our data indicated that Fasudil, SB431542, SIS3, and GSK650394 markedly down-regulated cell extension formation, matrix remodeling, and collagen degradation in fibroblasts and cancer cells, suggesting that the ROCK, Smad, and SGK kinase signaling pathways are key factors in metastatic processes related to cell extension formation, matrix remodeling, and collagen degradation.
Previous studies used 3D collagen matrices to examine cell behavior in connective tissue environments that would hopefully more accurately model the in vivo situation (than 2D systems). These studies indicate that matrix structure and mechanics are important factors that critically modify cell migration and cell morphology. Previous work has shown that 3D models can provide new insights, which superseded information derived from 2D matrix systems and that can clarify the functions of podosomes and invadopodia. 32 Consistent with these general advances, we show here that the measurement of cell extension formation, matrix remodeling, and collagen degradation in a grid-supported floating collagen gel model could provide an enhanced, integrated approach for drug screening in a high-content approach.
The generation of cell extensions involves the generation of cell adhesions, ECM degradation, and the reorganization of existing fibrillar collagen, 33 processes that were examined using four different cell types. In examining the role of cell adhesions, we focused on β1 integrin and DDR1, which are critical proteins for the formation of adhesions to collagen in many different cell types. 34 Integrins cooperate with cell surface proteases to enable ECM degradation that is associated with cell extension formation. Indeed, β1 integrins and MT1-MMP are clustered to enable pericellular collagen degradation in nascent cell extensions. 35 In addition, DDR1 and DDR2 are critical for cell adhesion and migration on collagen, matrix remodeling, and tumor metastasis. 36 We found that cell extension formation absolutely required β1 integrin function. Indeed, our results showed that blockade β1 integrin inhibited cell extension formation in fibroblasts and cancer cell lines, whereas DDR1 inhibition inhibited cell extension formation in cells that expressed DDR1.
We found that in particular, cell extension formation in the grid-supported floating collagen gel model was dependent on formin-mediated actin assembly. Cell extensions are thought to rely on the a priori formation of lamellipodia and filopodia, both of which require actin nucleation. Formins are an important family of actin nucleators 37 and are characterized by formin homology 1 (FH1) and FH2 domains. 37 FH2 dimers as assembly of actin can be stimulated in which the formin stabilizes actin dimers during the nucleation process. 21 One of these formins, mDia2, which is associated with microtubules, is concentrated at the tips of growing filopodia and may play an important role in cell extension formation, tissue remodeling, and sensing of matrix stiffness. 38
Previous studies showed that compared with 2D, plating cells on 3D matrices facilitated studies of cell morphology and tissue organization. 39 Our use of the floating collagen gel model to model cell extension formation, matrix remodeling, and collagen degradation provided a rapid and integrated system to screen a large number of compounds as potential inhibitors of cancer cell invasion. Notably, the alignment of collagen fibers is temporally and spatially correlated with the generation of cell extensions, 40 and our results here generally showed that those compounds that inhibited cell extension formation also affected collagen alignment. In summary, screening compounds with the use of the grid-supported floating collagen gel model may facilitate the linkage of cancer cell biology with the identification of new inhibitor of cancer cell invasion.
Footnotes
Acknowledgements
We are grateful for the assistance provided by Wilson Lee for confocal microscopy. We thank Alessandro Datti of the SMART Facility at the Mount Sinai Hospital (Toronto, ON, Canada) for his support and advice that related to the screening assays.
Supplementary material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The research was funded by the joint NSERC/CIHR-CHRP program (Canada) awarded to C.A.M. C.A.M. is supported by a Canada Research Chair (Tier 1) in Matrix Dynamics.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
