Abstract
Pseudomonas aeruginosa histidyl-tRNA synthetase (HisRS) was selected as a target for antibiotic drug development. The HisRS protein was overexpressed in Escherichia coli and kinetically evaluated. The KM values for interaction of HisRS with its three substrates, histidine, ATP, and tRNAHis, were 37.6, 298.5, and 1.5 μM, while the turnover numbers were 8.32, 16.8, and 0.57 s–1, respectively. A robust screening assay was developed, and 800 natural products and 890 synthetic compounds were screened for inhibition of activity. Fifteen compounds with inhibitory activity were identified, and the minimum inhibitory concentration (MIC) was determined for each against a panel of nine pathogenic bacteria. Each compound exhibited broad-spectrum activity. Based on structural similarity and MIC results, four compounds, BT02C02, BT02D04, BT08E04, and BT09C11, were selected for additional analysis. These compounds inhibited the activity of HisRS with IC50 values of 4.4, 9.7, 14.1, and 11.3 µM, respectively. Time-kill studies indicated a bacteriostatic mode of inhibition for each compound. BT02D04 and BT08E04 were noncompetitive with both histidine and ATP, BT02C02 was competitive with histidine but noncompetitive with ATP, and BT09C11 was uncompetitive with histidine and noncompetitive with ATP. These compounds were not observed to be toxic to human cell cultures.
Introduction
Resistance to multiple antimicrobial agents in pathogenic bacteria is spreading throughout the world and has become a serious problem that significantly threatens public health. 1 These multi-drug-resistant (MDR) bacteria have negatively impacted the clinical setting, with increasing numbers of patients treated in hospitals for infection with drug-resistant bacteria. In recent reports, the Infectious Disease Society of America (IDSA) has summarized the current and future public health burden resulting from drug-resistant bacteria, citing a current annual cost to the U.S. health care system for antibiotic-resistant infections of as much as $34 billion and growing. 2 Pseudomonas aeruginosa is an opportunistic Gram-negative pathogen and a common cause of nosocomial infections, and the leading cause of mortality in cystic fibrosis patients. 3 This bacterium has the remarkable ability to acquire resistance, intrinsic as well as acquired, against multiple groups of antimicrobial agents, including β-lactams, aminoglycosides, and fluoroquinolones, and is referred to as a member of the class of pathogens called “superbugs.” 4 The drug-resistant nature of these bacteria make treatment of infections both difficult and expensive and has resulted in an unmet need for the development of new antimicrobial agents with distinctly different modes of action than the current armament of antibiotics.
Aminoacyl-tRNA synthetases (aaRSs) play a central role in protein biosynthesis by catalyzing the covalent attachment of the amino acids to the cognate tRNA and are crucial for cell growth and viability. Therefore, bacterial aaRS enzymes, which are very different at the amino acid sequence level than their eukaryotic counterparts, are valid targets for development of potential therapeutics. 5 The aaRSs are evenly grouped into two different classes, with each class containing 10 members. Class I aaRSs contain two consensus sequences (HIGH and KMSKS) in their active site and bind ATP through a conserved structural domain (the Rossman fold), while class II aaRSs contain three conserved structural motifs (motifs I, II, and III) composing the ATP binding site. 6 Both classes can be further divided into a number of subclasses based on structural similarities, and within the class II enzymes, there are three distinct subgroups. Subclass IIa enzymes are homodimers and contain a C-terminal insertion that interacts with the tRNA, while in subclass IIb enzymes, this insertion is located within the N-terminal. The subclass IIc enzymes are tetramers, and tRNA binding elements appear to be less conserved than in the other class II aaRSs. 7 Histidyl-tRNA synthetase (HisRS) is a class IIa aaRS composed of three conserved domains: the catalytic domain that resides in the N-terminal, the C-terminal domain that harbors the tRNA anticodon binding site, and a HisRS unique insertion domain containing 50–75 residues forming an α-helix between motifs II and III. 8 HisRS also contains two additional unique motifs that facilitate histidine (His) binding: histidine A (HisA, RGLDYY) and histidine B (HisB, GGRYDG). The HisA and HisB motifs are highly conserved among all known HisRS enzymes.8–12 Upon His binding, the HisA motif is rearranged to increase hydrogen bonding capabilities, and the shape and electrostatic complementarity of the binding pocket. This results in a conformational change optimizing the amino acid binding pocket specificity for His, thus enhancing fidelity. 9
In this work, HisRS from P. aeruginosa was purified and enzymatically characterized by measuring its kinetic parameters (KM, Vmax, and kcat) governing interactions with His, ATP, and tRNAHis. It was then developed into a screening platform using scintillation proximity assay (SPA) technology 13 and used to screen 1690 natural and synthetic compounds. Four compounds were identified as inhibitors of P. aeruginosa HisRS and characterized for inhibitory activity and toxic potential to human cell cultures.
Materials and Methods
Materials
Oligonucleotides were from Integrated DNA Technologies (Coralville, IA). All other chemicals were from Fisher Scientific (Pittsburgh, PA). DNA sequencing was performed by Functional Bioscience (Madison, WI). Radioactive isotopes were from PerkinElmer (Waltham, MA). Escherichia coli tolC mutant, P. aeruginosa PAO200 (efflux pump mutant), and P. aeruginosa hypersensitive strain (ATCC 35151) were a kind gift from Urs Ochsner (Crestone Pharma, Boulder, CO). All other bacteria were from the American Type Culture Collection (ATCC) (Manassas, VA). Penicillin-streptomycin solution was from Mediatech, Inc. (Manassas, VA). The synthetic compound library was from TimTec LLC (Newark, DE), and the natural compound library was from MicroSource Discovery Systems (Gaylordsville, CT). All compounds were provided at or above 95% homogeneity.
Gel Electrophoresis and Protein Analysis
Sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) was performed using 4%–12% polyacrylamide precast gels (Novex NuPAGE; Invitrogen, Waltham, MA) with MOPS running buffer (Invitrogen). EZ-Run Rec Protein Ladder was from Fisher Scientific. Gels were stained with Simply Blue Safe Stain (Invitrogen). Protein concentrations were determined by the method of Bradford 14 using bovine serum albumin (BSA) as the standard.
Cloning and Purification of P. aeruginosa HisRS
The genes encoding P. aeruginosa HisRS were amplified by PCR (Bio-Rad MJ Mini Thermo Cycler, Hercules, CA) from P. aeruginosa PAO1 (ATCC 47085) genomic DNA using the forward primer (5′-ATATGCTAGCAGCAA-GTCCCTGCAAGCCA-3′) designed to add an NheI restriction site to the 5′ end of the gene and the reverse primer (5′-CATTGGATCCTCAAGCCTGTGCCAGACAGG-3′), which was designed to add a BamHI restriction site to the 3′ end of the gene. The PCR product was inserted into a pET-28b(+) plasmid (Novagen, Darmstadt, Germany) digested with NheI/BamHI. The recombinant plasmid was transformed into E. coli Rosetta 2(DE3) Singles Competent Cells (Novagen).
Bacterial cultures were grown in terrific broth containing 25 µg/mL of kanamycin and 50 µg/mL of chloramphenicol at 37 °C, and the target protein was expressed at an optical density (A600) of 0.6–0.8 by the addition of isopropyl β-D-1-thiogalactopyranoside (IPTG) to 0.5 mM. Growth of bacterial cultures was continued for 3 h postinduction, and the bacteria were harvested by centrifugation (10,000g, 30 min, 4 °C). Bacterial cells were lysed and fraction I lysate was prepared as previously described. 15 P. aeruginosa HisRS was precipitated by the addition of ammonium sulfate (AS) to 60% saturation and further purified to greater than 98% homogeneity using nickel-nitrilotriacetic acid (NTA) affinity chromatography (Perfect Pro, 5 Prime), followed by dialysis (two times) against a buffer containing 20 mM Hepes-KOH (pH 7.0), 40 mM KCl, 1 mM MgCl2, 0.1 mM EDTA, and 10% glycerol. Purified proteins were fast frozen by immersion in liquid nitrogen and stored at −80 °C.
ATP:PPi Exchange Reactions
ATP:PPi exchange reactions (100 µL) were carried out for 20 min at 37 °C in a buffer containing 50 mM Tris-HCl (pH 7.5), 10 mM potassium fluoride (KF), 8 mM MgOAc, 1 mM dithiothreitol (DTT), 2 mM [32P]PPi (50 cpm/pmol), and 0.05 µM P. aeruginosa HisRS, as described. 16 In reactions in which the concentration of ATP was varied (100, 200, 300, 400, 500, and 750 μM), the His concentration remained constant at 2 mM; alternatively, when the concentration of His was varied (3.12, 6.25, 12.5, 25, 50, 100, and 200 μM), the ATP concentration was held constant at 2 mM. The exchange of PPi in these reactions was measured between 1 and 5 min, and initial velocities for exchange of PPi were determined. The data were fit to the Michaelis–Menten steady-state model using XLfit (IDBS, Alameda, CA) and the KM, Vmax, and kcat for the interactions of P. aeruginosa HisRS with ATP and His were determined.
Aminoacylation Assays to Determine Kinetics
Aminoacylation was measured using filter binding assays (50 µL) containing 50 mM Tris-HCl (pH 7.5), 7.5 mM MgCl2, 2.5 mM ATP, 1 mM DTT, 75 µM [ 14 C] His, and 0.1 µM P. aeruginosa HisRS, as previously described. 13 The tRNAHis concentrations were 0.03, 0.6, 0.9, 1.2, 1.8, 2.4, 3, and 4.5 μM. Assays were stopped at time intervals between 1 and 5 min. Initial velocities for tRNA aminoacylation were calculated for all concentrations, and the data were fit to the Michaelis–Menten steady-state model using XLfit (IDBS) to determine the KM, Vmax, and kcat values.
Chemical Compound Screening
tRNA aminoacylation was monitored using a SPA, as previously described.
16
The concentration of P. aeruginosa HisRS in the screening assays was 0.2 µM. The screening reactions were carried out in 96-well microtiter plates (Costar). Compounds were dissolved in 100% DMSO, and the final concentration of compounds in the screening assay was 132 µM. Briefly, 2 μL of compound (3.3 mM) was equilibrated by the addition of 33 μL of the protein/substrate mix described above for the aminoacylation assay (minus tRNA). Control reactions contained only DMSO with no compound. This mixture was incubated at ambient temperature for 15 min, and then reactions were started by the addition of 15 μL of E. coli tRNA (80 μM total tRNA or 2.5 μM tRNAHis), followed by incubation for 30 min at 37 °C. Reactions were stopped by the addition of 5 μL of 0.5 M EDTA. A total of 400 μg of yttrium silicate (Ysi) poly-
Microbiological Assays
Broth microdilution minimum inhibitory concentration (MIC) testing was performed in 96-well microtiter plates according to Clinical Laboratory Standards Institute guideline M7-A7. 17 MIC values were determined for E. coli (ATCC 25922), E. coli tolC mutant (W4573:tolC::Tn10), Enterococcus faecalis (ATCC 29212), Haemophilus influenzae (ATCC 49766), P. aeruginosa (ATCC 47085), P. aeruginosa PAO200 (efflux pump mutant), P. aeruginosa hypersensitive strain (ATCC 35151), Staphylococcus aureus (ATCC 29213), and Streptococcus pneumonia (ATCC 49619).
Time-kill studies were performed according to the CLSI document M26-A 18 using S. aureus and the P. aeruginosa hypersensitive strain for compounds BT02D04, BT08E04, and BT09C11, and S. aureus and H. influenzae for compound BT02C02 based on the MIC assay results. Growth media was haemophilus test medium for H. influenzae and brain heart infusion for P. aeruginosa and S. aureus (Becton, Dickinson and Company, Franklin Lakes, NJ). The same growth media was used in MIC and time-kill studies.
In Vitro Cytotoxicity Test
The toxic effect of each compound on the growth of human cell cultures was carried out as described using human embryonic kidney 293 cells (HEK293). 19 The Trevigen TACS MTT Cell Proliferation Assay Kit (Gaithersburg, MD) was used to assess impacts on human cell proliferation and/or viability. The MTT assays were carried out in triplicate at each compound concentration. The concentration of hit compounds in assays ranged from 25 to 400 μg/mL, and the control staurosporine was serially diluted in assays from 1 to 0.001 μg/mL. Student’s two-tiered t test was utilized to assess statistical significance.
Binding Mode Assay
To determine if the hit compounds competed with either ATP or the amino acid for binding, IC50 values were determined in aminoacylation assays as described. 16 The P. aeruginosa HisRS concentration was set at 0.1 μM. For competition with ATP, IC50 values were determined in reactions containing varying ATP concentrations (25, 50, 100, 250, 500, and 1000 μM). To determine competition with His, the same assays were used with varying concentrations of the amino acid (25, 50, 100, 200, and 300 µM). Compounds were serially diluted in IC50 assays from 200 to 0.4 µM. Background amounts of free [ 14 C]His in the absence of HisRS were insignificant.
Results
Sequence Analysis
The crystal structures of three different bacterial forms of HisRS have been well defined: (1) the structures in apo forms, from S. aureus 9 and Thermus thermophilus (Tth); 20 (2) the structures from T. thermophilus,11,21 E. coli, 12 and Burkholderia thailandensis 22 in a complex with His or tRNA; and (3) the structures from T. thermophilus 11 and E. coli 23 in a complex with the intermediate histidyl-adenylate. When compared with the corresponding enzymes from E. coli, S. aureus, or T. thermophilus ( Fig. 1 ), the amino acid sequence conservation of HisRS from P. aeruginosa is moderately similar, with similarities ranging between 55% and 73% ( Suppl. Table S1 ). However, when compared with human cytosolic or mitochondrial HisRS (hcHisRS or hmHisRS), there is only 30% and 31% sequence similarity, respectively.

Alignment of amino acid sequence of P. aeruginosa HisRS with homologs. The protein sequences were downloaded from the National Center for Biotechnology Information (NCBI). Ec, E. coli; Pa, P. aeruginosa; Sa, S. aureus; Tth, T. thermophilus. Accession numbers for HisRS protein sequences of E. coli, P. aeruginosa, S. aureus, and T. thermophilus are KGL70691, WP_003092797, CAD80177, and YP_143978, respectively. Sequence alignments were performed using Vector NTI Advance 11.5.4 (Invitrogen). Identical residues are indicated by white letters on black background, while similar sequences are black letters on gray background. The three structural motifs I, II, and III and the functional motifs HisA and HisB are indicated. Amino acids that interact with His (●), ATP (▲), and tRNA (*) are indicated above the aligned residues.
The His and ATP binding sites are primarily located in motifs II and III, which is the case with all other class II aaRS enzymes. His binds in a deep pocket formed by two HisRS-specific peptides, HisA and HisB, which are highly conserved among all known HisRS proteins. 11 The His imidazole ring forms hydrogen bonds with Glu-130 and Tyr-264 (E. coli numbering), and Tyr-264 also forms hydrogen bonds to Thr-85, Arg-259, and Tyr-288 to maintain correct orientations for binding the amino acid. 12 The α-amino group of His interacts with both Thr-85 and Tyr-263, and stabilizes Glu-83 by hydrogen bond formation.11,12 These residues are all identical in the four HisRS proteins analyzed ( Fig. 1 ). The polar pocket, together with the hydrogen bonds, excludes any hydrophobic amino acid from binding. 11
Unlike the strict conservation observed in the amino acids around the His binding site, the amino acids forming the ATP binding pocket are more variable. The three phosphates of ATP interact with the guanidinium groups of four arginine residues that are completely conserved in HisRS from the aligned bacteria. The α-phosphate associates with Arg-113 and Arg-259 (E. coli numbering), the β-phosphate interacts with Arg-113, and the γ-phosphate makes contact with Arg-121 and Arg-311. 12 The variation in conservation is seen in some of the amino acids interacting with the ribose group of ATP. In E. coli HisRS, Gln-127, Ala-335, Ser-336, and Gly-365 likely have interactions with the ribose. 8 All four of these amino acids are conserved in HisRS from E. coli and P. aeruginosa, but slight variations occur at Ala-335, Ser-336, and Gly-365 in HisRS from S. aureus and T. thermophilus.
The tRNA binding regions are less conserved than either the His or the ATP binding sites. The acceptor stem of tRNAHis is recognized by interaction of Arg-7, Arg71, Arg-115, and Arg-122 (T. thermophilus numbering) with the tRNA G1 nucleotide; Glu-64, Glu-114 Gln-117, Lys-118, and Arg-120 with the tRNA C70-C72, C74, and A76 nucleotides; and Arg197, Arg-204, and Lys-209 with the tRNA C2-C75 base pair; and Gly-260, Tyr-264, and Ala-278 form nonspecific hydrogen bonds through their main chain amide nitrogens. The absolutely conserved residues Glu-64 and Glu-114 are essential for aminoacylation. However, the other residues contribute less to substrate recognition. 21 The GUG/A triplet anticodon forms specific hydrogen bonds with HisRS. tRNA G34 forms six hydrogen bonds with Glu-386, Asp-387, and Glu-388; U35 forms three hydrogen bonds with Gln-404; and G/A36 forms two hydrogen bonds with Lys-397 and Leu-399. 21
Without the luxury of a crystal structure for P. aeruginosa HisRS, the conservation of the amino acids observed in regions vital for interactions with the substrates when compared with that of the homologs from E. coli, S. aureus, and T. thermophilus indicates that the active sites are similarly constructed. This suggests that a compound that interferes with activity by binding in or near these strategic regions may have broad-spectrum inhibitory activity.
Protein Expression and Characterization
The gene encoding HisRS (hisS) from P. aeruginosa was cloned, expressed, and purified as described in Materials and Methods. The purified protein was greater than 98% homogeneous as visualized by SDS-PAGE ( Suppl. Fig. S1 ). In aminoacylation assays, P. aeruginosa HisRS was shown to be active in attaching radiolabeled His to tRNAHis ( Suppl. Fig. 2A ).
The result of the aminoacylation reaction is the covalent attachment of a specific amino acid to the cognate tRNA, and this process occurs via two steps. In the first step, the amino acid and ATP are condensed, resulting in the formation of an aminoacyl-adenylate intermediate, followed by the release of an inorganic pyrophosphate. This reaction is reversible with most aaRS enzymes in the absence of a cognate tRNA, and the parameters governing the kinetic interaction of the aaRS with the amino acid and ATP substrates may be determined. The kinetic parameters KM, kcat, and kcat/KM for the interaction of HisRS with His and ATP have been determined by measuring the initial rate of the ATP:PPi exchange during this reversible reaction and fitting the data to the Michaelis–Menten steady-state model. To determine the interaction of HisRS with the amino acid, the concentration of ATP was held constant (2 mM), while concentrations of His were varied (3–200 µM) ( Fig. 2B ). From these data, the KM and kcat values for the interaction with His were calculated to be 37.6 μM and 8.32 s−1, respectively ( Fig. 2E ). E. coli HisRS was previously shown to have a similar KM of 30 μM 12 and a turnover of 7–142 s−1, as measured by different research groups.24,25 Alternatively, to determine the kinetic parameters for the interaction with ATP, the concentration of ATP was varied between 100 and 750 μM while maintaining a constant concentration of His (2 mM) ( Fig. 2C ). The KM and kcat values for interaction with ATP were calculated to be 298.5 μM and 16.8 s−1, respectively. This again is similar to the interaction observed with ATP for E. coli HisRS, which has a reported KM of 890 μM 12 and a kcat of 34 s−1. 24
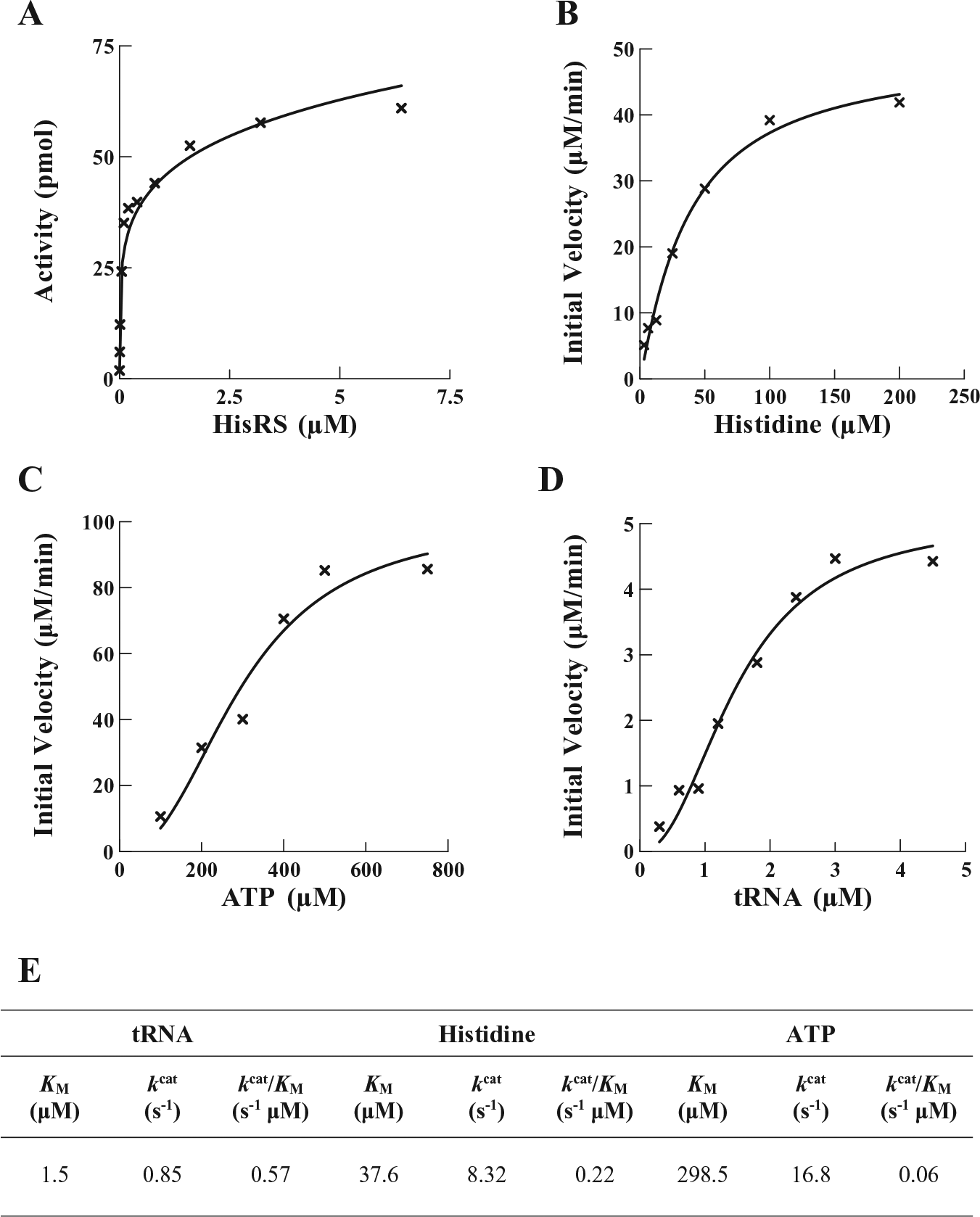
Enzyme activity and determination of kinetic parameters for interactions of P. aeruginosa HisRS with His, ATP, and tRNAHis. (
The formation of an ester bond between the hydroxyl group of the 3′ terminal adenosine of tRNA and the carboxylic acid function of the amino acid constitutes the second step in the aminoacylation process. The ability of P. aeruginosa HisRS to aminoacylate tRNAHis was determined at various concentrations of tRNA while maintaining ATP and His constant at 2.5 mM and 75 μM, respectively ( Fig. 2D ). The KM and kcat values were determined to be 1.5 μM and 0.85 s−1, respectively. The same values reported for E. coli HisRS are 1.4 μM and 2.6 s−1, respectively. 12
Screening Chemical Compounds against the Activity of P. aeruginosa HisRS
Initially, to achieve the maximum signal in the SPAs, all nonenzymatic components of the aminoacylation reaction were individually titrated into the assay to determine the saturating concentration of each (data not shown). Next, tRNA was titrated into the assay to ensure that the assay was within the linear region of the reaction detection time and to determine the concentration of tRNA to be used in the screening assays ( Suppl. Fig. S2 ). Finally, HisRS was titrated into the assay to determine the enzyme concentration yielding maximum sensitivity to enzymatic inhibition ( Fig. 2A ). From these data, a HisRS concentration of 0.2 μM was chosen for screening reactions. Using this optimized assay, the activity of HisRS in the aminoacylation assay was screened against two chemical compound libraries. One contained an 800-compound collection of natural products, including simple and complex oxygen heterocycles, alkaloids, sesquiterpenes, diterpenes, pentacyclic triterpenes, and sterols from Microsource Discovery Systems (Gaylordsville, CT). The second collection included 890 low-molecular-weight, druglike molecules with scaffolds found in antiseptic agents with antibacterial, antifungoid, and antimicrobial activities from TimTec LLC (Newark, DE). The SPA detects the ability of HisRS to aminoacylate tRNAHis and, in the presence of a chemical compound, to measure the effect of the compound on the activity of HisRS. Chemical compounds were dissolved in 100% DMSO, resulting in final DMSO concentrations in screening assays of 4%; therefore, the ability of P. aeruginosa HisRS to function in the presence of increasing amounts of DMSO was determined. No decrease of activity was observed in aminoacylation assays containing up to 10% DMSO ( Suppl. Fig. S3 ). Initial screening assays contained chemical compounds at a concentration of 132 μM and were carried out as single point assays. Compounds observed to inhibit at least 50% of enzymatic activity were reassayed in triplicate. These assays resulted in 15 confirmed hit compounds, all from the synthetic compound library. The IC50 values for the confirmed hit compounds were determined and as a group ranged from 2 to 45.1 μM ( Suppl. Table S2 , Fig. 3 ).
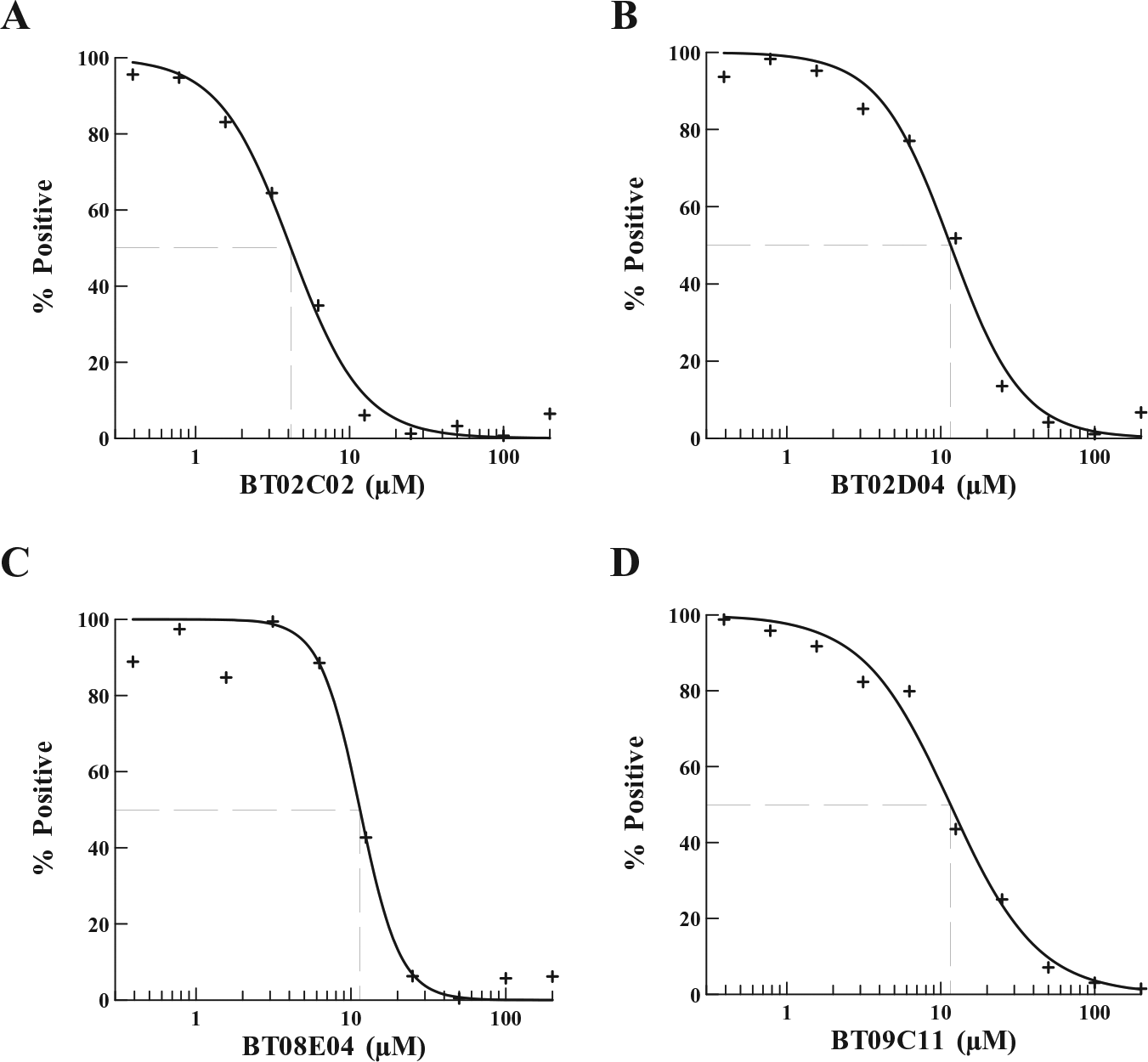
Determination of the IC50 of selected hit compounds against the activity of P. aeruginosa HisRS. The IC50 values for the inhibitory activity of (
Microbiological Assays
The 15 hit compounds were tested in broth microdilution assays to determine MICs against a panel of nine pathogenic bacteria, including efflux pump mutants of E. coli and P. aeruginosa and a hypersensitive strain of P. aeruginosa ( Suppl. Tables S2 and S3 ). Despite the similarity in the IC50 values of the compounds, the ability to inhibit bacterial growth in cultures varied considerably. Based on compound scaffold similarity and broad-spectrum activity against both Gram-positive and Gram-negative pathogens, four of the compounds, BT02C02, BT02D04, BT08E04, and BT09C11 ( Fig. 4 ), were selected for further analysis. BT02C02 inhibited the growth of the S. pneumoniae and E. coli efflux pump mutant at 16 µg/mL, S. aureus at 32 µg/mL, and E. faecalis and H. influenzae at 64 µg/mL. BT02D04 inhibited the growth of S. pneumoniae and S. aureus at 32 µg/mL, and H. influenzae, the E. coli efflux pump mutant, and the P. aeruginosa hypersensitive strain at 64 µg/mL. BT08E04 inhibited the growth of E. faecalis, S. pneumoniae, and S. aureus at 32 µg/mL, and H. influenzae and the P. aeruginosa hypersensitive strain at 64 µg/mL. BT09C11 inhibited the growth of the E. coli efflux pump mutant, S. aureus, H. influenzae, S. pneumoniae, E. faecalis, and the P. aeruginosa hypersensitive strain at 4, 8, 16, 16, 32, and 32 µg/mL, respectively. None of the four compounds were observed to inhibit the growth of wild-type E. coli or wild-type P. aeruginosa at concentrations below 128 µg/mL.
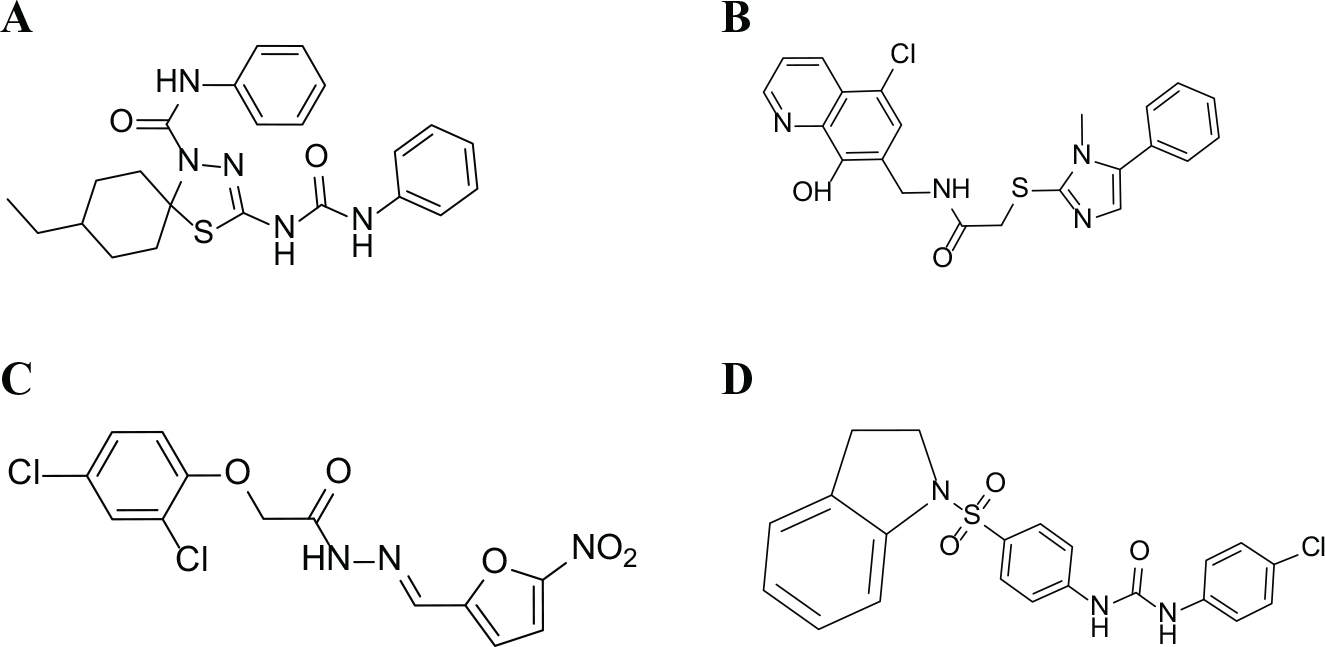
Chemical structure of the hit compounds: (
Next, time-kill studies were performed using the four compounds BT02C02, BT02D04, BT08E04, and BT09C11 to determine the mode of inhibition of bacterial growth in cultures. Based on MIC results, all compounds were tested against cultures of S. aureus, a Gram-positive pathogen. BT02D04, BT08E04, and BT09C11 were also tested against cultures containing the P. aeruginosa hypersensitive strain. However, because of the elevated MIC observed for BT02C02 against the hypersensitive strain of P. aeruginosa, it was tested against H. influenzae, another Gram-negative pathogen. All cultures contained compounds at four times the MIC, and samples were analyzed between 0 and 24 h. All four compounds were shown to be bacteriostatic against the pathogens tested ( Fig. 5 ). In these tests, the bacteria were observed to have constant growth in the presence of compound but a decrease of 1–5 log10 colony-forming units (CFU) compared with the control during the first 6 h.
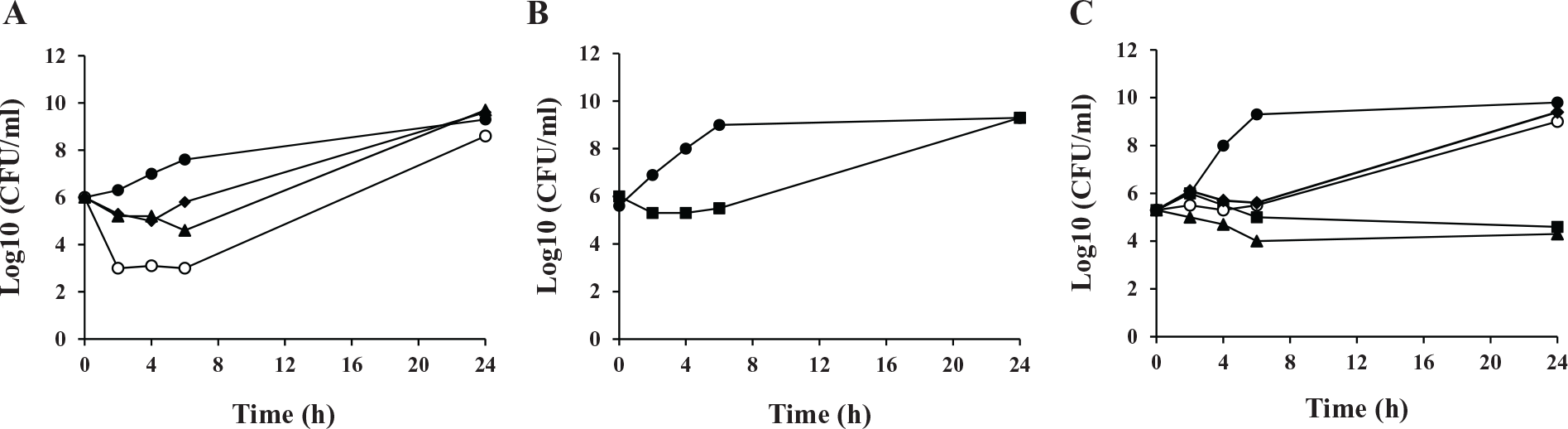
Time-kill kinetics against MIC determined bacterial targets: (
Compound Toxicity in Human Cell Cultures
Antibiotic therapies that specifically target bacterial translation could potentially be toxic to human cells, given that both human cytoplasmic and mitochondria HisRS are contained within the human cell, and each share a modest level of amino acid sequence homology and biochemical similarity with the bacterial HisRS. MTT assays were performed to examine whether the hit compounds BT02C02, BT02D04, BT08E04, and BT09C11 were cytotoxic to human cells. HEK293 cells were treated with 25–400 µg/mL of each compound separately for 24 h under standard tissue culture conditions in triplicate. BT02C02 and BT09C11 were not observed to be toxic to cells at any concentration tested ( Fig. 6A ). Some toxic effects were observed at higher concentrations for BT02D04 and BT08E04 in the HEK293 cell cultures ( Fig. 6C , D ), with the CC50 values for both these two compounds near 200 µg/mL. However, even at these inhibitory concentrations, the cytotoxic potential was 60,000-fold lower when compared with the control staurosporine ( Fig. 6B ). The MIC for BT02D04 and BT08E04 at 32 and 64 μg/mL for susceptible bacteria is three to six times lower than the CC50 of the toxicity studies. The other two compounds, BT02C02 and BT09C11, were observed to have MICs ranging from 4 to 64 μg/mL against susceptible bacteria and were not observed to be toxic in the cell cultures up to 400 μg/mL. This level of difference between MIC and CC50 is advantageous in the development of potential antimicrobial agents. These results indicate that all four compounds may be amenable to development as therapeutics.
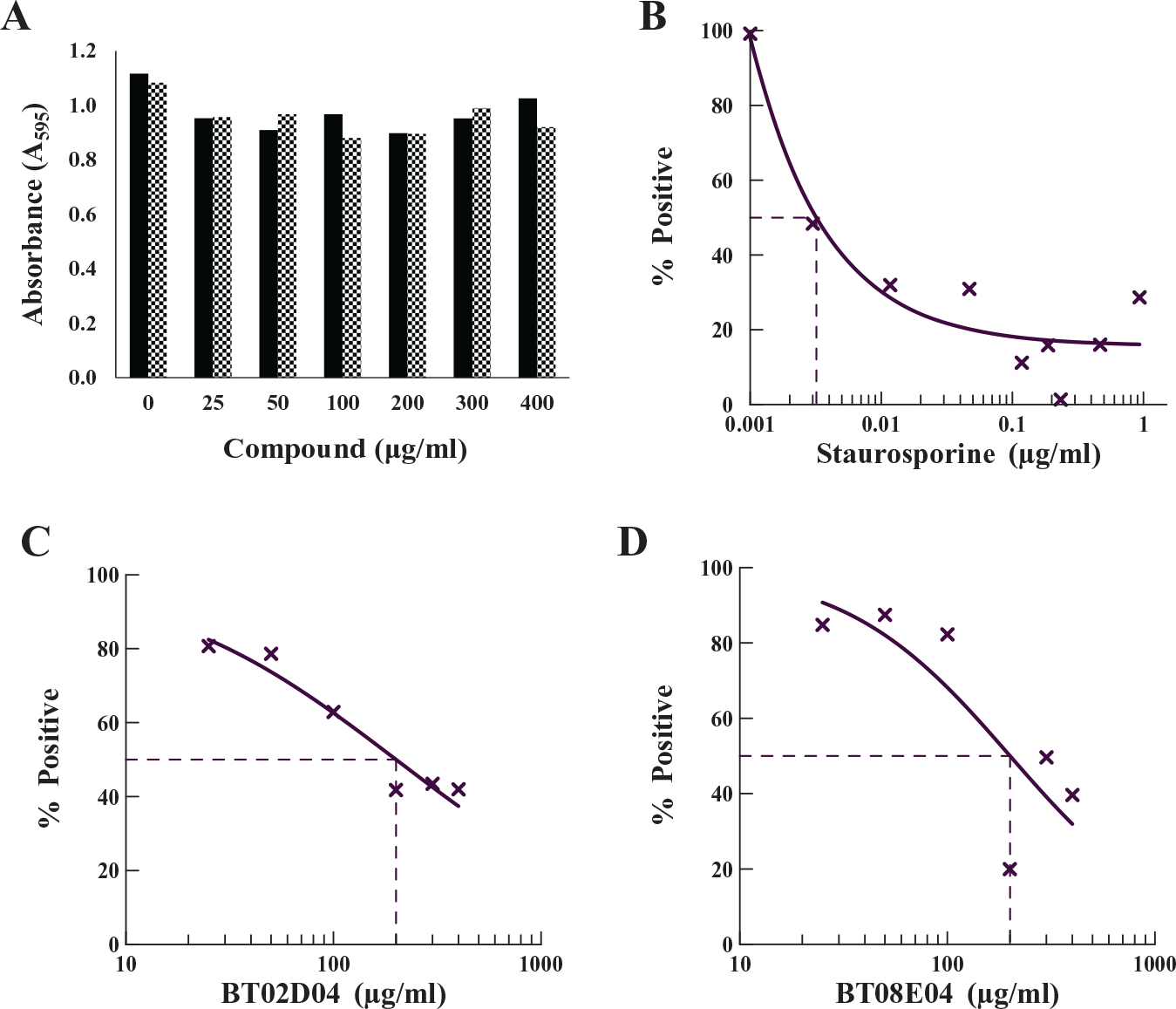
Determination of the compound toxicity in mammalian cell cultures. MTT assays were performed with the indicated dose of drug for 24 h under standard tissue culture conditions, as described in Materials and Methods. (
Mechanism of Action
aaRSs in general have three substrates: tRNA, ATP, and the amino acid. ATP and His are small molecules similar in size to the compounds tested, and both have binding sites in the active site region of the synthetase. Since blocking the binding site of one of these substrates would be a possible mechanism of action of an inhibitor, the ability of each of the four compounds to compete for binding with each of these substrates was determined ( Suppl. Table S4 ). The mechanism of inhibition with respect to ATP was determined using the tRNA aminoacylation assay to determine IC50 values at various ATP concentrations (25, 50, 100, 250, 500, and 1000 µM), while holding the concentration of the amino acid at 100 μM. To determine the mechanism of inhibition with respect to His, the same assay was used, except that ATP was held constant at a saturating concentration (2.5 mM) and the IC50 was determined at different concentrations of His (25, 50, 100, 200, and 300 µM). The IC50 values for BT02C02 remained constant at varying concentrations of ATP. However, an increase was observed at increasing concentrations of His. An increase in the IC50 as the concentration of the substrate increases is characteristic of a competitive inhibitor. 26 The IC50 for BT09C11 remained constant at varying concentrations of ATP, but was observed to decrease as the concentration of His was increased, which is characteristic of an uncompetitive inhibitor. 26 The IC50 values for both BT02D04 and BT08E04 remained constant at all concentrations of both ATP and His, which is characteristic of a noncompetitive inhibitor. 26 These results lead us to conclude that BT02C02 may inhibit HisRS by direct competition with the amino acid, while BT02D04, BT08E04, and BT09C11 inhibit the function of the enzyme by a different mechanism.
Discussion
As reports of resistant bacteria continue to increase, there is a need for the discovery of new antibacterial agents with different modes of action than those of the currently marketed antibiotics. The aaRSs are ideal targets for development of new antibacterial agents for a number of reasons. 5 First, they play a crucial role in protein biosynthesis and are vital for cell viability and growth. Second, the aaRS proteins from prokaryotic origins are conserved in primary structure yet sufficiently divergent from their eukaryotic homologs to make them suitable targets for development of selective broad-spectrum antibacterial agents. Finally, the crystal structures of many of the aaRSs have been solved and provide useful information for rational drug design.
His is one of the more functional amino acids in proteins and often is a catalytic residue or a ligand for metal binding. For these reasons, a high degree of accuracy for attaching the correct amino acid to the cognate tRNA is required. Studies have shown that RNAi silencing to partially suppress HisRS expression had a profound effect on the growth of a bloodstream form of Trypanosoma brucei, decreasing growth by a factor of >103 at day 4 compared with the normal cells. 8 Even with the knowledge of the importance of HisRS, there has been little work to develop inhibitors of this enzyme, and most of the published data are focused on the crystal structures, fidelity mechanism, and tRNA recognition.
In the present work, SPA technology was used to develop a screen for inhibitors of the activity of P. aeruginosa HisRS. The screening assays were robust and resulted in Z′ and Z factors of approximately 0.61 and 0.52, respectively, across all plates. The signal-to-background ratio of the DMSO-positive controls to the EDTA-negative controls was approximately 5:1. From almost 1700 compounds, 15 compounds (exclusively from the synthetic compound library) were identified that inhibited the activity of P. aeruginosa HisRS. Four of the compounds, BT02C02, BT02D04, BT08E04, and BT09C11, inhibited the activity of HisRS with IC50 values of 4.4, 9.7, 14.1, and 11.3 µM, respectively, and exhibited promising MICs against both Gram-positive and Gram-negative bacteria. Time-kill studies indicated that all four compounds were bacteriostatic. This would be the expected mode of action for compounds that inhibit the activity of an aaRS, since inhibition of the aminoacylation activity mimics starvation for amino acids by lowering the ratio of charged to uncharged tRNA, and induces the stringent response resulting in static bacterial growth. 27
Each of the four compounds contains a carboxamide group ( Fig. 4 ), which is present in some well-known antibiotics, as has been noted previously. 13 Tetracycline contains a carboxamide as a side chain. However, in all four of these hit compounds this motif serves as part of the core compound. Compound 08E04 also contains a nitrofuran group. Many compounds containing nitrofurans have been shown to have antibacterial activity and are included in a class of synthetic antibiotics, including furazolidone, furaltadone, nitrofurantoin, and nitrofurazone, 28 that are active against certain Gram-positive staphylococci and streptococci, but inactive against P. aeruginosa. BT09C11 was observed to have the best broad-spectrum activity and lowest MIC values of the four hit compounds. This compound contains a sulfonamide group, which has been shown to have antibacterial activity as a functional group in other compounds. Some compounds containing a sulfonamide group also have been shown to act as a competitive inhibitor of the enzyme dihydropteroate synthase (DHPS), which is involved in folate synthesis. 29 With this in mind, this compound may have more than one mode of inhibitory activity. We searched the PubChem BioAssay Database for biological activity and found no biological information associated with any of the four hit compounds.
In an era of MDR bacteria, compounds with a different mechanism of action, or with different binding sites, would be advantageous. As observed in the competition assays ( Suppl. Table S4 ), as a whole these compounds do not interfere with binding of ATP or the amino acid substrates. This indicates that the compound interactions with HisRS occur at sites other than within the active site for aminoacylation. Structure–activity relationship (SAR) studies will need to be conducted to enhance these interactions, resulting in increased potency while maintaining low levels of toxicity to eukaryotic cells.
Footnotes
Supplementary material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The authors are grateful for the financial support provided by the National Institutes of Health (grant number 1SC3GM098173). The contents of this article/publication/etc. are solely the responsibility of the authors and do not necessarily represent the official views of the National Institutes of Health. A portion of student support was from a Departmental Grant from the Robert A. Welch Foundation (grant BG-0017).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
