Abstract
The analysis of human microbiome is an exciting and rapidly expanding field of research. In the past decade, the biological relevance of the microbiome for human health has become evident. Microbiome comprises a complex collection of microorganisms, with their genes and metabolites, colonizing different body niches. It is now well known that the microbiome interacts with its host, assisting in the bioconversion of nutrients and detoxification, supporting immunity, protecting against pathogenic microbes, and maintaining health. Remarkable new findings showed that our microbiome not only primarily affects the health and function of the gastrointestinal tract but also has a strong influence on general body health through its close interaction with the nervous system and the lung. Therefore, a perfect and sensitive balanced interaction of microbes with the host is required for a healthy body. In fact, growing evidence suggests that the dynamics and function of the indigenous microbiota can be influenced by many factors, including genetics, diet, age, and toxicological agents like cigarette smoke, environmental contaminants, and drugs. The disruption of this balance, that is called dysbiosis, is associated with a plethora of diseases, including metabolic diseases, inflammatory bowel disease, chronic obstructive pulmonary disease, periodontitis, skin diseases, and neurological disorders. The importance of the host microbiome for the human health has also led to the emergence of novel therapeutic approaches focused on the intentional manipulation of the microbiota, either by restoring missing functions or eliminating harmful roles. In the present review, we outline recent studies devoted to elucidate not only the role of microbiome in health conditions and the possible link with various types of diseases but also the influence of various toxicological factors on the microbial composition and function.
Keywords
The microbiome and its importance in human health
We are close to the truth if we state that our bodies contain almost the same number of microbes, or even more based on an old measurement, as the number of human cells. 1,2 In fact, we live in symbiosis with a vast population of bacteria, archaea, eukaryotes, and viruses, collectively called microbiota, that colonize different bodily parts. 3 The “microbiome” represents all the genes and genomes of the microbiota, as well as their metabolites and protein products and those of the host. 4
Research has now started to focus on the roles played by fungal and viral members of the microbiota community in health and disease; however, scientific interest to date has mainly been focused on bacterial populations and their interactions with the human host. It is the latter area this review will focus on.
In the human body, the major niches colonized by microbes are the gut, mouth, genitals, skin, and airways. The composition of this bacterial population varies depending on the body part. For example, the human intestine is composed predominantly of Bacteroidetes and Firmicutes (90%), and complemented by Actinobacteria, Proteobacteria, and Verrucomicrobia, 5 –7 whereas the microbiome of the retroauricular skin crease is composed mainly of Actinobacteria followed by Firmicutes, Proteobacteria, and Bacteroidetes 8 (see Figure 1, Supplementary Figure 1, and Supplementary Table 1 for general classification).
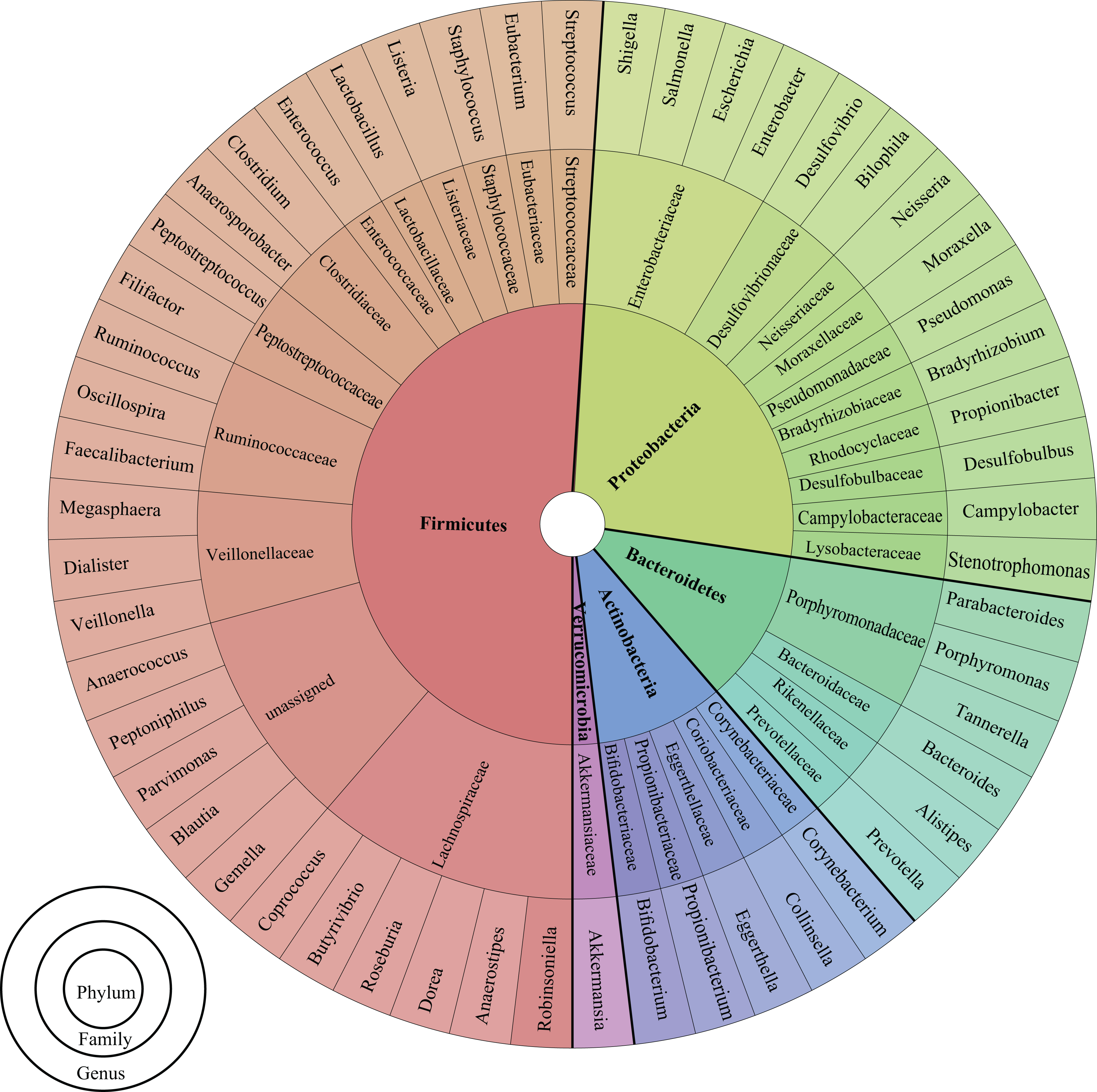
Taxonomical classification chart. 9 The pie chart reports the classification for all genus discussed in the review. The chart shows, for all genus (outer layer), the corresponding family (middle layer) and phylum (inner layer), omitting the class and the order classification rank. See Supplementary Figure 1 and Supplementary Table 1 for a more extensive interactive chart, and a table with the complete genus list and classification, respectively.
The microbiome can affect many physiological processes in our bodies, including immune system development, the ability to process dietary polysaccharides, vitamin and hormone production, pH regulation, processing and detoxification of environmental chemicals, and maintenance of the skin and mucosal barrier function 10,11 (Figure 2).
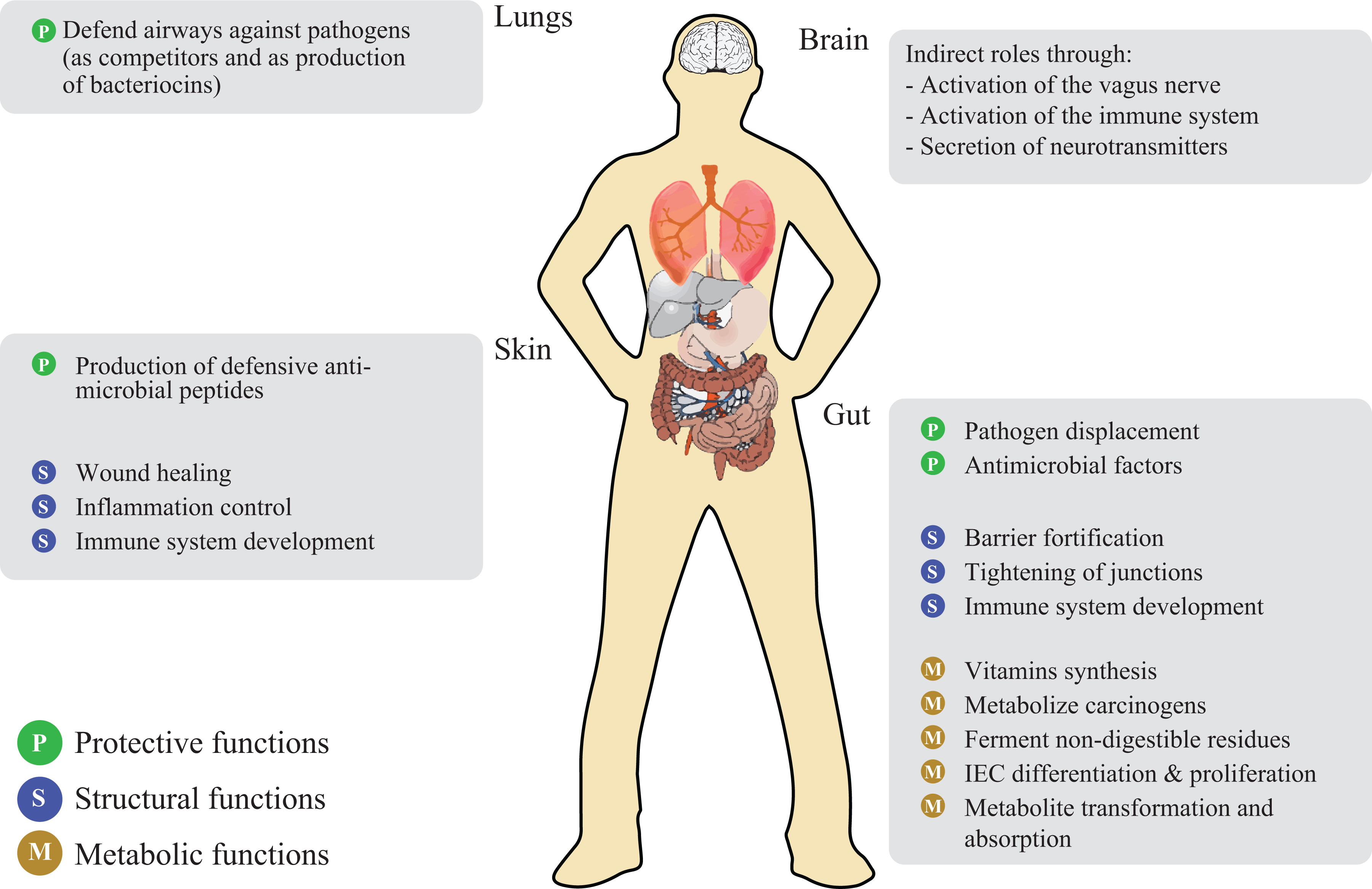
Main functions of bacteria in the human body. IEC: intestinal epithelial cell.
In the intestine, for example, a healthy microbial population maintains local homeostatic immune responses via exposure to bacterial structural ligands (e.g. lipopolysaccharide, LPS, and/or peptidoglycan). 12 –15 The importance of gut microbes in maintaining immune system homeostasis is highlighted by the fact that the complete absence of this population in mice leads to large defects in the development of gut-associated lymph tissues, low levels of intestinal secretory IgA antibodies, and fewer and smaller mesenteric lymph nodes. 13,16
Intestinal bacteria are also essential for the breakdown of dietary fibers, such as inulin, pectin, xylans, and mannans. This process yields energy, which is important for the growth and maintenance of the microbial community and for the production of metabolic end products that are beneficial to the host. 17,18 The principal end products of carbohydrate processing are gases, such as CO2, H2, and CH4, and short-chain fatty acids (SCFAs), such as acetate, butyrate, and propionate, which can serve as (1) important metabolites that create a direct energy source for intestinal epithelial cells, (2) inhibitors of inflammation, or (3) modulators of insulin secretion. 17,19 –23 In fact, while butyrate is an important energy source for colonic epithelial cells and an inhibitor of the NF-κB signaling that supports mucosal barrier integrity, acetate and propionate can be utilized by the liver for lipogenesis and gluconeogenesis. 7,18,24
Intestinal microbiota can also perform multiple metabolic activities ranging from the catabolism of certain oral drugs 25 to the synthesis of a wide range of compounds that have various effects on the host, such as several classical neurotransmitters like γ-aminobutyric acid (GABA) as well as bile acids. The latter compounds, synthesized in the liver, can undergo a second conversion in the intestine via microbiota processes to generate secondary bile acids. Ultimately, these acids can be taken up into the bloodstream, where they can potentially modulate host metabolism and other functions, including behavioral and neural functions. 26 –30
Thus, a complex symbiosis exists between the human body and its microbiome, the disruption of which can have detrimental effects on both. Indeed, several physiological changes (via antibiotic use and other lifestyle factors, including hygiene and diet)
8,31
can strongly impact microbial composition. The resulting dysbiosis (alteration of the microbial composition) may be unfavorable and associated with the development of the following diverse diseases
14
(Figures 3 and 4 and Table 1):
Digestive pathologies (celiac disease, inflammatory bowel disease (IBD), and nonalcoholic hepatitis); Metabolic diseases (obesity, cardiovascular diseases, and type 2 diabetes (T2D)); Neurological disorders (neurodegenerative diseases and other brain disorders); Skin problems (eczema, acne, and dermatitis); Oral diseases (caries and periodontitis); Pulmonary pathologies (chronic obstructive pulmonary disease (COPD), asthma, and cystic fibrosis (CF));
33
Cancers (e.g. colorectal cancer and colorectal adenoma, gastric and esophageal cancers, hepatobiliary cancers, pancreatic cancer, and lung cancer);
14,34
–38
and Immune-related diseases (e.g. food allergies, asthma, celiac disease, and rheumatoid arthritis).
39
–45
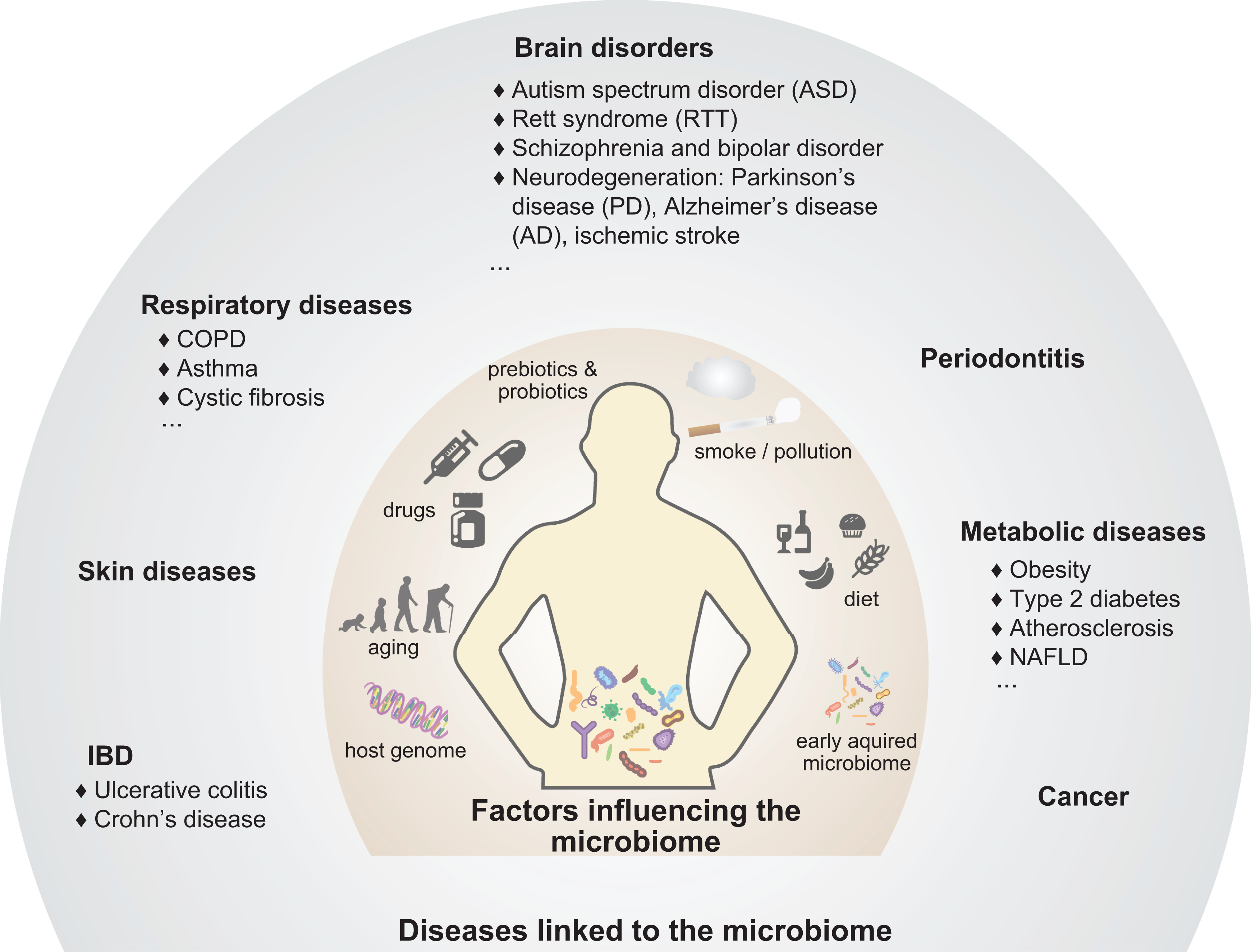
What can influence the microbiome? Factors influencing the intestinal microbial composition and the effects of dysbiosis on host health.
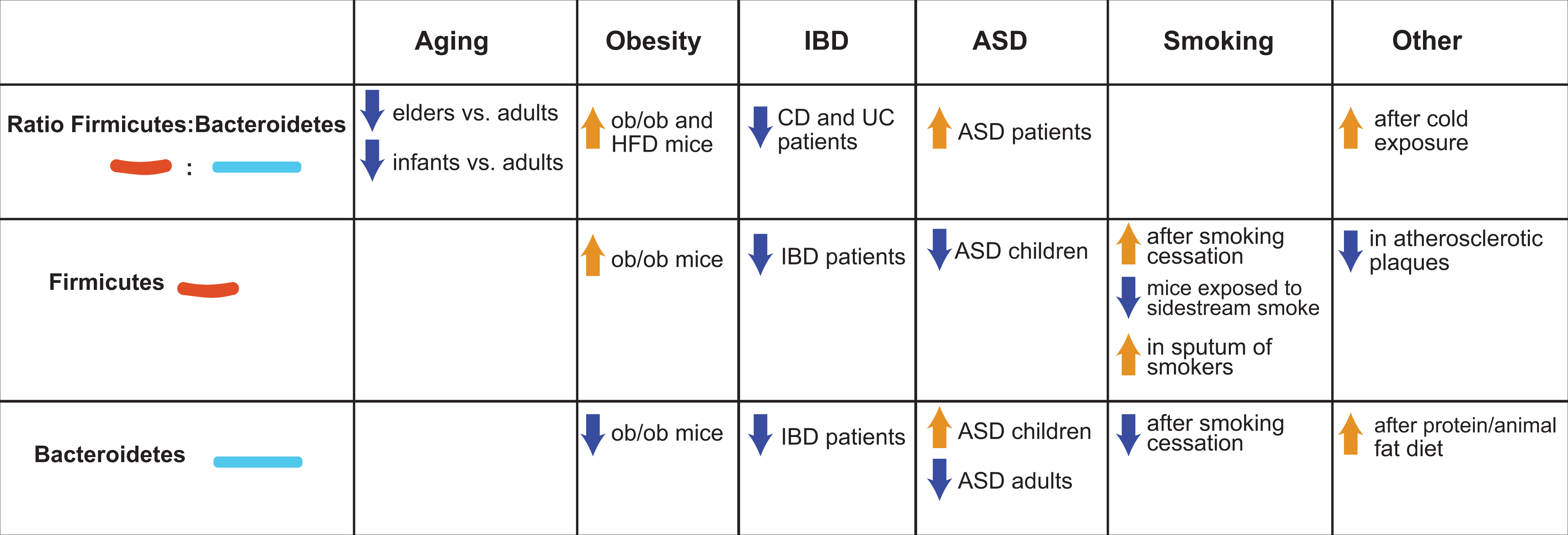
Summary of the main features of Firmicutes and Bacteroidetes.
Overview of selected potentially harmful and potentially beneficial bacteria present in our body.

Source: Adapted from “Influence of diet on the gut microbiome and implications for human health,” 2017. 32
BCAA: branched-chain amino acids; BMI: body mass index; CD: Crohn’s disease; COPD: chronic obstructive pulmonary disease; CS: cigarette smoke; IBD: inflammatory bowel disease; IR: insulin resistance; PD: Parkinson’s disease; RTT: Rett syndrome; SCFA: short-chain fatty acids; T2D: type 2 diabetes; TLR: Toll-like receptor; UC: ulcerative colitis.
The microbiome and lifestyle factors
In combination, multiple factors, such as genotype, dietary composition and mode of delivery, antibiotic therapy, pre and probiotic treatment, lifestyle (e.g. smoking and physical activity), social interactions, and environmental exposure to various xenobiotics, shape the gut microbiota, making every person microbially unique (Figure 3).
Diet
Diet plays an important role in shaping the gut microbiome. An increasing body of evidence from humans and mouse models suggests that diet supersedes host genetics in terms of the gut microbiota composition. 46,47 Studies on the influence of diet on the human gut microbiome that have involved the intake of specific dietary components (e.g. proteins, fats, carbohydrates, polyphenols) have shown how certain bacteria respond to nutrient-specific challenges. 32,48 A shift in the microbiome composition may have secondary effects on host immunologic and metabolic markers. For instance, a plant-based diet is reported to be associated with a higher abundance of Bacteroides (Bacteroidetes) and Firmicutes, an animal-based diet has been linked with a higher abundance of Bacteroides, and a high fiber diet is linked with higher levels of Prevotella (Bacteroidetes). 49
Diets rich in saturated fat or in polyunsaturated fat seem to profoundly affect the gut microbiome composition and the host immune system, 50 and in one study, mice fed fish oil showed increased levels of Lactobacillus and Akkermansia while those fed on lard had increased levels of Bilophila (which has been shown to be correlated with colitis 51 ). The high level of white adipose tissue inflammation observed in the lard-fed mice could be caused in part by gut–microbiome metabolism, whereby bacterial products, like LPS, can activate inflammatory immune processes, such as the Toll-like receptor (TLR) signaling.
Different diets can also affect microbial gene richness (determined simply by gene count) and microbial gene diversity (which also takes into account the degree to which the numbers of genes are evenly distributed). A comparison of three different diets revealed that the healthiest diet (with the lowest consumption of sugary drinks and confectionary and the highest consumption of fruit, yogurts, and soups) is associated with the highest microbial diversity in overweight or obese people, whereas a high-fat diet (HFD) is linked to lower microbial gene richness. 52
Diet-induced microbial changes in the gut can be gender-specific 53 and affected by the dietary history (any effect due to diet change depends on the initial individual’s microbiota composition 46,54 ) or the timing of the diet change because the composition of the gut microbiota appears to be stable up to 10 days after switching to a new diet. 55 Some of the results in this area have been summarized in recent reviews. 31,32,49,56
Smoking
Smoking is addictive and a major risk factor for several diseases, like cardiovascular disease (CVD), lung cancer, and COPD. 57 –59 Two studies have highlighted the strong impact of smoking on gastrointestinal microbiota. 60,61 Significant changes were also observed in the fecal microbiota of healthy individuals undergoing smoking cessation that included an increase in the relative abundance of Firmicutes and Actinobacteria and a reduction of Bacteroidetes and Proteobacteria. 60,61 These changes were similar to the gut microbial changes observed in obese people compared with leaner people, suggesting a potential link between smoking cessation and weight gain. Currently, only a few studies in nonhuman animals have been published, but they do align with observations in humans, thereby highlighting the cigarette smoke (CS)-dependent shift in the gut microbiota structure. 62 –65 Four weeks of CS exposure induced a decrease of the Bifidobacterium population in the rat cecum. 64 In mice, sidestream smoke exposure increased Clostridium spp. but decreased the Firmicutes phylum, the Enterobacteriaceae family, and the segmented filamentous bacteria in the cecum. 63
The first comprehensive analysis of bacterial colonization of the upper respiratory tract of healthy adult cigarette smokers 66 found that the microbial communities in this tract differed significantly from those of nonsmokers, suggesting that degradation of normal community structure in the smokers had occurred. In nonsmoking subjects, the nasopharynx bacterial population mainly comprised Firmicutes (73%), Proteobacteria, Bacteroidetes, and Actinobacteria phyla. However, smokers also had greatly increased levels of Megasphaera spp. (of Firmicutes phylum), which are known to reside in the oral cavity and to be associated with periodontitis. 67 Moreover, studies analyzing the oral and lung microbiome of smokers and nonsmokers have shown significant differences between them. 68 In particular, in the oral cavity of smokers, the abundance of Neisseria, Porphyromonas, and Gemella species decreased, while no significant differences were found in the lungs of smokers versus nonsmokers.
Physical activity
There is new evidence suggesting that physical activity may modify the microbiota of mice 69 and humans. 70,71 In humans, it has been shown that athletes have a higher diversity of gut microbiota (with higher proportions of the genus Akkermansia levels), which in turn is positively correlated with improved protein consumption and higher creatine kinase levels. 70
Drugs
Widespread antibiotics use has led to deleterious consequences for microbiome diversity in humans. 8 For example, a longitudinal study found that ciprofloxacin use in adults led to decreased bacterial diversity in the gut, which had not recovered 6 months after treatment. 72 Similar results have been seen in mice where antibiotic treatment depleted the gut microbiota. 73 In humans, obesity development is also associated with antibiotic treatment (see under paragraph “Metabolic diseases and the microbiome”).
In addition to the disruption of the ecology of the human microbiome, inappropriate use of antibiotics has led to heavy selective pressure and, consequently, to the development of human pathogens able to survive antibiotic treatment. Antibiotic resistance is quickly becoming a significant health-care problem and we refer the reader to recent reviews on this important topic. 74 –76
Prebiotics and probiotics
Prebiotics are food supplements ingested by the host but metabolized by the host’s gut bacteria. These supplements favor specific changes in the activity and composition of the gut microbiome, which in turn benefits the host’s health and wellness. The ability of prebiotics to promote human health has been extensively studied in animals and clinical studies.
48
Prebiotics are mainly short-chain nondigestible carbohydrates like inulin-type fructans, fructo-oligosaccharides, and galacto-oligosaccharides. Interestingly, one of the first prebiotics a human receives is in the form of oligosaccharides from mother’s milk (human milk oligosaccharides, HMO). HMOs are a group of complex and diverse glycans that are resistant to gastrointestinal digestion
77
and are used as energy sources for the development of Bifidobacteria.
49
Furthermore, HMOs act as antiadhesive molecules that block the attachment of viral, bacterial, or protozoan parasite pathogens to epithelial cells, thereby preventing infectious diseases. HMOs also have bacteriostatic and bactericidal activities and they alter host epithelial and immune cell responses with benefits for the neonate.
In general, the prebiotics are only nutrients for Bifidobacterium and Lactobacillus, and these genera are not the only important microbial contributors to human health. Recently, discussion around the development of new prebiotics with wider ranges of action and the capacity to have positive effects on the growth of other potentially beneficial bacteria like Ruminococcus bromii, Roseburia intestinalis, Eubacterium rectale, and Faecalibacterium prausnitzii 48 has been ongoing.
Unlike prebiotics, which are nutrients for the microbes already present in the host, probiotics are specific bacterial strains administered with the aim of improving the health of the recipient. 79 –81 The most commonly used probiotics are Lactobacillus and Bifidobacterium, although other genera like Bacillus, Enterococcus, and Streptococcus, as well as yeast, such as Saccharomyces, have also been included in the probiotics category. 82
The principal roles for probiotics in promoting health are as follows: (1) reinforcing mucosal barrier functioning, (2) reducing the mucosal transfer of luminal organisms and metabolites to the host, (3) improving the mucosal antibody production, and (4) the direct antagonism of pathogens.
The results from human studies on prebiotics are variable, and this probably relates to the methodological differences employed (e.g. dosing, duration of administration, sample collection) and differences in the chosen cohorts (age, health status). 48 For instance, in the treatment of allergic diseases, while a few clinical trials show outstanding support for prebiotic use, some studies also report no effects. 83 Other studies have reported on the significant cholesterol-lowering effect of probiotic treatment, while others have found no effects. 84
Both prebiotics and probiotics target or augment specific genera (Lactobacillus and Bifidobacterium), but the overall compositional change only occurs over the treatment duration. Definitive proof linking transient compositional alterations with improved health in the host remains elusive.
Age
The host’s age has a significant effect on the microbiota composition. Emerging evidence shows that, unlike what was once believed, the in utero environment is not sterile. Indeed, traces of Enterococcus faecalis, Staphylococcus epidermis, and Escherichia coli have been found in the newborn meconium. 85 Wider microbiota colonization appears immediately after birth and varies depending on the route of delivery (vaginal or caesarian). 86,87 In fact, the neonate gut microbiome is colonized first by the mother’s vaginal or skin bacteria population based on the route of childbirth. 85 Later on, the microbes present in the colostrum add to this community, thereby increasing the child’s microbiota complexity. 49 The first bacteria to appear in the intestines are aerobic strains, such as Proteobacteria. These bacteria decrease the oxygen concentration and allow colonization by anaerobic strains, such as Bacteroides, Actinobacteria, and Firmicutes.
During the first year, the microbiota composition continues to change and starts to resemble that of the adult at 1 to 2 years of age. 14 The composition of gut microbiota appears to be more stable during adulthood with small changes occurring in adolescents. 85 Later on, a final set of changes in gut microbiota composition and function occurs, as characterized by a shift in the ratio of Firmicutes to Bacteroidetes, a decrease in Bifidobacteria, an increase in anaerobic species, an increase in the potentially toxic Clostridium perfringens, and production changes in the gut-related metabolites of choline. 48,81,85,88 The reasons for these microbial changes have been proposed to include increased frailty, decreased diet diversity and health, and increased inflammatory marker levels. 89
Various studies have shown that ageing is correlated with a shift in the intestinal microbe composition in Drosophila and mice also. 90 Very recently, a study by Thevaranjan and co-workers reported that ageing induces intestinal permeability changes in mice. 91 This type of disruption results in microbes colonizing nonpermissive areas with consequent inflammation, mostly via tumor necrosis factor (TNF) involvement. This inflammation status can also affect macrophage functions, and antipneumococcal immunity, in particular. This could explain why elderly people are often hospitalized with pneumonia. 92
Environmental factors
Different environmental factors can affect our bodily microbial composition. High humidity and low temperature are associated with higher gram-negative bacteria in the skin’s microbiome, in particular on the back and the feet. 87 Our ethnicity and cultural habits are additional environmental factors influencing the gut microbiome. 49 Not surprisingly, different studies have reported changes in taxonomic/phylogenetic composition of fecal samples from different geographical areas. 93,94 There are also well-documented disparities in the genetic background, body mass index (BMI), diet, sanitary and hygiene (including the use of antibiotics and vaccines), variations in the incidences and concentrations of foreign (xenobiotic) metabolites, and differences in the socioeconomic status between geographically dispersed populations. 10 Even traveling overseas can alter the composition of the microbiome, and contracting gastrointestinal tract infections while abroad can also affect the microflora content. 48
A variety of environmental chemicals are metabolized by the enzymatic activities of the gut microbiota. They are classifiable into five core enzymatic families (azoreductases, nitroreductases, β-glucuronidases, sulfatases, and β-lyases) that are capable of metabolizing 430 environmental contaminants. Conversely, environmental contaminants can alter the composition and/or the metabolic activities of gastrointestinal bacteria, thereby probably contributing to their toxicities. Such contaminants include pesticides, heavy metals (e.g. cadmium), persistent organic pollutants, such as the aryl hydrocarbon receptor ligand 2,3,7,8-tetrachlorodibenzofuran, and artificial sweeteners. 95
Laboratory animal studies have revealed a correlation between xenobiotic exposure and gut microbiota composition changes. Arsenic-treated mice experienced reductions in Firmicutes but not Bacteroidetes in the gut. 96 The same mice also exhibited bidirectional changes in key metabolites, including those related to bile acids, lipids, amino acids, and isoflavones. 97 Further evidence in the same mice pointed to a reduction in the gut bacterial abundance after ingesting polychlorinated biphenyls. 96 Similar results have been obtained in studies on humans. 98,99 A recent longitudinal agricultural community cohort study investigated the effect of pesticides on the oral microbiome. In this study, the farmworkers in whom organophosphate pesticide azinphos-methyl was detected in the blood showed a significantly reduced abundance of seven common taxa of oral bacteria, including Streptococcus and Halomonas. This pesticide-associated spring/summer general reduction in bacterial diversity lasted until the winter season. 100
Metabolic diseases and the microbiome
Metabolic diseases, such as obesity, CVD, and T2D, are multifactorial in their etiologies and chronically persistent. The prevalence of two closely linked metabolic disorders, obesity and T2D, is increasing worldwide. These disorders pose a major public health risk in European countries and in the United States, and also in rapidly developing countries such as China and India. 101 –104 Their rise has been attributed mainly to socioeconomic factors, dietary changes, and sedentary lifestyles. Recent evidence suggests that the gut microbiome also influences the whole metabolic activity of the body and its immune function and could, therefore, play a role in these disorders. The complex metabolic interplay between the gut microbiome and the host has encouraged a detailed analysis of the potential role of the microbial population in metabolic conditions to be undertaken by many researchers. Below, we provide an overview of the most recent reviews on gut microbiota and metabolic diseases. 31,56,105 –110
Obesity
Gut microbial composition alterations have been extensively described as either potentially causal or protective toward weight gain 111,112 both in human and mouse studies. An analysis of the metabolic phenotypes of eight genetically distinct inbred mouse strains in response to a HFD revealed several variations in their metabolic-related phenotypes and gut microbiota. 113 Clostridiaceae show the strongest positive correlation with plasma insulin levels and weight gain, whereas Bacteroidaceae are negatively associated with these parameters. In a follow-up microbiota transplant experiment, the composition of the murine gut microbiome was shown to be able to modify the host’s susceptibility to diet-induced metabolic diseases, 113 a finding that is possibly related to an enhanced capacity to process dietary sugar and to produce hydrophobic bile acids. Furthermore, genetically obese mice (ob/ob mice) had an altered microbial composition with increased cecal levels of Firmicutes and reduced levels of Bacteroidetes when compared with their lean wild-type, heterozygous littermates. 114 An increase in the Firmicutes to Bacteroidetes ratio was also observed in obese rats 115 and pigs, 116 when compared with lean animals. However, a more recent paper reported that these changes, in mice, are caused primarily by high-fat feeding rather than genetically induced obesity. 117 Therefore, high-fat feeding, independent of the genetic background, seems to be the primary determinant of gut microbial alterations.
The important role of the gut microbiome in obesity development was also confirmed by a study done with germ-free HFD-fed mice. In this study, germ-free mice fed an HFD displayed reduced adiposity compared with conventionally raised animals that were also fed an HFD, despite the increased caloric intake and decreased energy expenditure. After colonization of germ-free mice with the fecal content from conventionally raised mice, the originally germ-free mice rapidly gained weight and had an increased fat mass without any change in caloric intake, 118 –120 thus illustrating a cross talk between gut microbiota and host tissue homeostasis. This finding also indicates that the gut microbial ecosystem is transmissible and that gut microbiota composition can affect obesity in mammals.
Interestingly, weight gain in germ-free mice can also be affected if they are colonized with the fecal microbiota from adult humans placed on a Western diet. 121 Indeed, when feces from human twin pairs discordant for obesity were transferred to germ-free mice, 122 different results were obtained. The mice that received feces from the obese twin developed metabolic alterations and gained more weight than those that received feces from the lean twin. The authors of this study showed that the microbiota from the lean twin had a higher SCFA fermenting efficiency, which promoted good metabolic health. Moreover, when the two groups of mice were co-housed, microbial transfer occurred by coprophagia; consequently, the mice that received the microbiota from the obese twin showed changes in their microbiota profile that were accompanied by improvements in their metabolic phenotype. The improved phenotype in the obese mice was, however, only achieved upon feeding them with a healthy diet that was high in fiber and low in saturated fat. Additionally, following consumption of a low-fat, high-fiber diet, the “obese-microbiota” failed to colonize lean mice as efficiently as they had in mice that had consumed an HFD. This emphasizes, once again, the importance to health of dietary components that may favor the growth of certain bacterial strains. 123,124
An interesting line of research has shown that the pharmacological removal of gut microbiota using broad-spectrum antibiotic cocktails prevented fat accumulation even in ob/ob mice, highlighting the importance of the gut microbiome in mediating diet-induced obesity. 125 However, such antibiotic administration was shown to have the opposite effect if administered in a different situation. As discussed in the paragraph “The microbiome and lifestyle factors,” for instance, the antibiotic treatment in early life can induce disruption of the normal microbial balance and increase the risk of developing obesity. 126
Analyses of the gut microbiota composition in humans and mice have shown that the presence of Akkermansia muciniphila and F. prausnitzii is inversely correlated with weight gain, obesity, and diabetes. 127,128 In addition, inoculation with Enterobacter cloacae can induce fully developed obesity phenotypes in germ-free mice. 129 The following compositional differences have been found in obese people relative to lean people: Bacteroides, Parabacteroides, Ruminococcus, Campylobacter, Dialister, Porphyromonas, Staphylococcus, and Anaerostipes are more prevalent in people with obese phenotypes, while Faecalibacterium, Bifidobacterium, Lactobacillus, Butyrivibrio, Alistipes, Akkermansia, Coprococcus, and Methanobrevibacter are more prevalent in people with lean phenotypes. 130 A positive association has also been found between weight gain and the presence of Lactobacillus reuteri, whereas the opposite has been shown for R. intestinalis. 109 Elsewhere, a strong correlation has been shown to exist between a low BMI in humans and an increased abundance of the Christensenellaceae family. 131 This finding was supported by an association study involving 416 twin pairs where the Christensenellaceae family showed an increased abundance in individuals with low BMI. After being transplanted to germ-free mice, Christensenella minuta (DSM22607) was able to reduce weight gain and alter the microbiome of the recipient mice. 132 An association has been found between weight gain and the presence of L. reuteri, 133 whereas the opposite has been found for R. intestinalis. 109
When advocating the positive or negative health benefits of different bacterial phyla, one should be careful, as a recent study has revealed that Bifidobacteria and Lactobacillus may have different properties according to their species. For example, Bifidobacterium animalis, Lactobacillus plantarum, and Lactobacillus paracasei are associated with lean phenotypes, whereas, as previously mentioned, L. reuteri is associated with obesity in humans and other animals. 133,134
In a prospective study of a population of children that were followed for several years, the authors found that microbiota alteration preceded weight gain and that Staphylococcus aureus could have an important role in triggering the low-grade inflammation that contributes to obesity development. 135 A new concept is now emerging: rather than the loss of a specific microorganism, it is the loss in microbial richness that is related to the development of obesity and other metabolic diseases. 123,129
Beyond pure association studies, recent work has elucidated some of the mechanisms by which microbiota may influence obesity development. Although the main cause of obesity remains an excess calorific intake over energy expenditure, it has been shown recently that differences in gut microbial composition may be an important factor affecting host energy homeostasis and lipid storage. In particular, gut microbiota composition has been linked with the low-grade inflammatory status present in obesity, which may be caused by bacterial LPS entering the systemic circulation via gut barrier dysfunction (i.e. disruption of the tight-junction proteins that link epithelial cells together). 136 Several factors can perturb gut permeability: for example, genetic factors can create a weaker mucosal barrier, as can gastroenteritis, detergent ingestion, and emotional stress. 137,138 Furthermore, recent evidence from animal models suggests that mucolytic bacterial species, such as A. muciniphila and Bacteroides thetaiotaomicron, can degrade the mucosal layer, thus reducing the barrier function of the gut. 139
Another important mechanism by which the gut microbiome can promote excessive fat accumulation in obesity is dysregulation of the important genes involved in host lipid metabolism (e.g. acetyl-CoA carboxylase 1 and fatty acid synthase), which is mediated by suppressing the intestinal expression of a circulating lipoprotein lipase inhibitor. 120
Other lines of research have investigated the effects of microbial metabolism on energy balance. 36,119,120,122,140 For example, in a recent paper, Trajkovski et al. showed that the microbiota plays a key role in mediating tight control of energy homeostasis by helping the host to withstand periods of high energy demand. 141 They found that exposure to cold temperature leads to dramatic changes in the microbiota composition, referred to as “cold microbiota,” and this was characterized by an increase in the Firmicutes to Bacteroidetes ratio with almost complete depletion of the Verrucomicrobia phylum. These changes favored enhanced energy extraction during cold and were mediated by an increase in the intestinal absorptive surface via a marked increase in the number of intestinal villi and microvilli length. This increased absorptive surface is a general adaptive mechanism that promotes caloric uptake when food is available. Transplantation of cold microbiota to germ-free mice was sufficient to induce cold tolerance and increase insulin sensitivity, which is in part mediated by browning of their white fat depots. This effect was diminished by co-transplantation with A. muciniphila, the most abundant Verrucomicrobia species and the phylum most negatively affected by cold exposures.
Type 2 diabetes
T2D is a chronic metabolic disorder where the body either does not produce enough insulin or cannot effectively metabolize glucose despite insulin production. Alterations in the intestinal microbiota composition have been shown to modulate insulin sensitivity and thus play a role in diabetes susceptibility. 142 The fact that germ-free mice, after receiving microbiota from conventionally raised mice, increased their body fat content and became insulin resistant highlights the importance of the microbiome in the regulation of insulin and glucose homeostasis. 120
Two recent quantitative gut metagenomics studies on patients with T2D (unstratified for treatment) yielded divergent conclusions about gut microbial dysbiosis. The first metagenome-wide association study on a large cohort of Chinese patients with T2D revealed a decrease in butyrate-producing bacteria like R. intestinalis and F. prausnitzii in them, together with an increased prevalence of opportunistic pathogens, such as Bacteroides caccae, various Clostridiales and E. coli, and decreases in the known mucin degrading species A. muciniphila and in the sulfate-reducing genus Desulfovibrio. 143 In the second study, a similar analysis was conducted in a cohort of Scandinavian women with T2D, but the authors instead reported on an enrichment of several Lactobacilli species; 127 this apparent contradiction may be explained by the fact that the antidiabetic medication (metformin) used by this group confounded the results. The following year, the role of A. muciniphila in mediating an improvement of the metabolic profile of T2D mice was elucidated in a published study showing that oral administration of A. muciniphila to mice reversed metabolic disorders, including insulin resistance (IR). 128
As previously mentioned, metformin, one of the most widely prescribed anti-diabetic drugs, can have a profound effect on the microbiome composition, leading to an improvement in the gut microbial profile. In patients with T2D taking the drug, an increase in SCFA production, butyrate and propionate production, as well as increased Escherichia abundance (associated with known side effects of metformin, such as bloating) were observed. A similar effect was shown in HFD-fed mice that exhibited a higher abundance of the mucin-degrading bacterium Akkermansia spp. after antidiabetic treatment. 144,145 Other studies have found that the IR-associated metabolome is linked with gut microbiome-encoded functions, such as production of LPS and branched-chain amino acids (BCAAs). Positive correlations between these microbial functions and IR are largely driven by Prevotella copri and Bacteroides vulgatus, suggesting that they may directly impact host metabolism. In mice, P. copri led to increased serum levels of BCAAs and IR. 146
Despite these studies, understanding the precise mechanisms underlying the association between microbiome dysbiosis and T2D remains elusive. Multiple mechanisms of action have been proposed in mediating the T2D-associated inflammation such as gut permeability, metabolic LPS-mediated endotoxemia, and modifications to incretin secretion and butyrate production. 145
Cardiovascular disease
Some studies have suggested a linkage between the gut, oral microbiota, and several facets of CVD such as atherosclerotic plaque formation, myocardial infarction, and heart failure. Patients with atherosclerosis have a reduced fecal abundance of butyrate-producing Roseburia and Eubacterium, which are also known to be decreased in patients with T2D. 143,147,148 The gut microbiome from CVD patients is enriched in genes encoding peptidoglycan synthesis and depleted in phytoene dehydrogenase, which contributes to a pro-inflammatory status. 148
Interestingly, bacterial DNA was also discovered in atherosclerotic plaques and some of the species identified were similar to those found in the microbiota of the oral cavity, with high levels of Proteobacteria and low levels of Firmicutes being identified. 149 Moreover, the abundance of bacterial DNA in the atherosclerotic plaques was correlated with CVD risk factors, 150 such as serum LDL-cholesterol. 149
The metabolic activity of gut microbiota can also have a direct effect on atherosclerosis development, and a key example of this is the metabolism of dietary phosphatidylcholine. This phospholipid (which is present mostly in meat, eggs, and fish) is hydrolyzed by gut bacteria into trimethylamine, which is further oxidized in the liver by flavin monooxygenase into circulating trimethylamine N-oxide (TMAO), 151 –153 whose levels are associated with cardiovascular risk. 154 –156 Intriguingly, it has also been found that inhibiting gut microbial TMAO production using nonlethal inhibitors may serve as a potential therapeutic treatment for atherosclerosis. 157 This study’s findings may explain the link between certain dietary habits and CVD development.
Nonalcoholic fatty liver disease
Nonalcoholic fatty liver disease (NAFLD), as characterized by lipid deposition in the hepatocytes, is considered the hepatic manifestation of metabolic syndrome. NAFLD is initiated by hepatic steatosis and may progress to nonalcoholic steatohepatitis (NASH). While most patients with NAFLD remain asymptomatic, 20% develop chronic hepatic inflammation, which in turn can lead to cirrhosis, portal hypertension, and hepatocellularcarcinoma. 158
During the last decades, researchers have characterized multiple processes that play crucial roles in the so-called gut–liver axis. These mechanisms may explain the multiple processes that occur from fatty liver accumulation to inflammation and fibrosis. In fact, the gut–liver axis is the route by which bacteria and their potential hepatotoxic products, like LPS, can easily reach the liver. 159 –161 Ultimately, pro-inflammatory cytokine (e.g. interleukin: IL-1β and IL-8) production plays a pivotal role in the induction and progression of nonalcoholic liver disease to NASH and cirrhosis.
In this context, the microbiome plays an important role in the development of NAFLD, and multiple molecular pathways have been postulated to explain the relationship between NAFLD and dysbiosis. 162 Several human and animal studies that have shown a link between the gut microbiome, NAFLD and pathogenesis have been reviewed in some recent papers. 107,158,163 –165 Zhu et al. showed that children and adolescents with NASH have different microbiome patterns respect to the healthy subjects. In fact they have increased Bacteroidetes (Prevotellaceae (Prevotella, Porphyromonas)) and Proteobacteria (Enterobacteriaceae (Escherichia), Alcaligenaceae) numbers, and decreased Firmicutes (Lachnospiraceae (Blautia, Coprococcus, Eubacterium, Roseburia), Ruminococcaceae (Faecalibacterium, Oscillospira, Ruminococcus)), and Actinobacteria (Bifidobacteriaceae (Bifidobacterium)) numbers in their feces. 166 Supporting these studies, pediatric patients with NAFLD had increased numbers of Bradyrhizobium, Anaerococcus, Peptoniphilus, Propionibacterium acnes, Dorea, and Ruminococcus, and decreased numbers of Oscillospira and Rikenellaceae. 167
In another study with adults, patients with NASH harbored a lower abundance of Faecalibacterium and Firmicutes (Clostridiales family, Anaerosporobacter) but a higher abundance of Parabacteroides and Allisonella in their fecal microbiomes. 168 Crucially, it has been reported that improved intrahepatic triglyceride content is related to a lower abundance of Firmicutes and a higher abundance of Bacteroidetes. 168 In contrast with this, obese individuals with NAFLD have increased members of the Firmicutes phylum (Lachnospiraceae (Dorea, Robinsoniella, and Roseburia)). 169
Conversely, a recent study showed a reduction in Bacteroidetes in NASH patients compared with the other groups, 170 while Qin et al, when characterizing the gut microbiome in liver cirrhosis, showed that at the phylum level, patients with this disease had fewer Bacteroidetes but more Proteobacteria and Fusobacteria than the controls 171 (without distinguishing patients with NASH or virus-related cirrhosis). Similar to other microbiome studies, discrepancies can be caused by variations in the study design (factors such as age, concomitant medications, health status, and lifestyle are heterogeneous among all studies). An exhaustive table of the intestinal microbiota composition in NAFLD patients can be found in the review by Mokhtari et al. 164
Possible explanations for the correlation found between the microbiome and NAFLD could relate to changes in SCFA metabolism, release of LPS from gut microbiota, endogenous ethanol production, or a decrease in choline and trimethylamine levels. Dysbiosis might also promote de novo hepatic lipogenesis, increase intestinal permeability, and lead to translocation of both bacteria and endotoxins from the intestinal lumen to extraintestinal sites. 158 More recently, it has been shown that diet fructose can induce alterations of the tight junction proteins, altering the gut permeability, which contributes to an inflammatory status promoting the development of NAFLD. 172,173
The use of microbiome-based therapies as possible treatments for NAFLD is still under investigation. However, a recent study showed that fecal microbiota transplantation (FMT) can alleviate the steatohepatitis induced in mice by a HFD, which was accompanied by an observed increased in beneficial bacteria like Christensenellaceae and Lactobacillus, decreased hepatocyte lipid accumulation, and decreased pro-inflammatory marker levels. 174
Links between metabolic diseases, the microbiome, and lifestyle factors
Both genetic susceptibility and environmental factors (e.g. intrauterine conditions, physical inactivity, smoking, and unhealthy dietary habits) are involved in the pathogenesis of metabolic disorders. 110 Recently, studies have suggested that the environmental component in the development of metabolic syndrome is partially mediated through an altered gut microbial structure and function.
Diet
An individual’s diet shapes the diversity of their gut microbial community. As mentioned previously, an example of this is where HFD-fed mice experienced an increased Firmicutes to Bacteroidetes ratio. However, although human studies have not been equally consistent on the link between diet and microbial content, 110,175 –177 it is clear that the composition and the functional capabilities of human and rodent gut microbiota rapidly adapt to changes in the macronutrient content of the diet. 110
Western-style diets have a strong impact on gut microbiota, particularly on its structure and function. 178 When trying to understand the impact of diet in determining the influence of the microbiome in the development of metabolic diseases, the following points should be considered: (1) the amount, the type (e.g. unsaturated vs. saturated fatty acids), and the mixture of dietary fats can affect the gut microbial composition; (2) the effect of a HFD on the gut microbiome can be rapid (occurring within 24–48 h) and sustained if the dietary habits persist, and may not be recoverable without a change in diet, reintroduction of the extinguished microbial strains, or dietary supplementation.
A HFD induces complex interactions between microbes and the host via various mechanisms such as those mediated by angiopoietin-like 4 (Angptl4), AMP-activated protein kinase, and TLRs.
Physical activity
Physical activity has been shown to affect gut microbiota composition and diversity in murine models of obesity and hypertension, with some results indicating that the effects demonstrated are independent of diet. 110,179,180 Following exercise, obese and hypertensive rats both experienced increased microbiota diversity, and this was associated with an increase in the relative abundance at the genus level in all the rat models that were studied.
Drugs
Broad-spectrum antibiotic use has been proposed to possibly contribute to the obesity epidemic and to a decreased gut microbiome diversity. This theory is supported by a few epidemiological studies showing that antibiotic intake in infancy or exposure during prenatal life increases the risk of being overweight in childhood. 181 –183
Additionally, a few clinical studies in adults have identified an increase in BMI following antibiotic treatment, especially after Helicobacter pylori eradication in patients with gastric ulcers. In contrast, individuals with metabolic syndrome when treated with vancomycin have reduced peripheral insulin sensitivity, while amoxicillin treatment has no such effect. 184 This probably results from the preferential targeting of gram-positive butyrate-producing bacteria by vancomycin. However, the decrease was modest, and the study did not include a control group. In a more recent study, researchers analyzed the effects of vancomycin and amoxicillin (compared with a placebo-treated group) on gut microbiota and metabolism in obese people. 185 Here, Reijnders et al. reported that in contrast to amoxicillin, vancomycin decreased bacterial diversity in the subjects; however, neither antibiotic had an effect on insulin sensitivity, calorie expenditure, or body weight.
In experimental animal studies, the effect of antibiotic treatment on metabolic features has led to controversial results. An increased SCFA content and lower caloric output have been observed in the feces from antibiotic-treated newborn mice, despite having a similar caloric intake as the control mice, which suggests a possible link between obesity and antibiotic treatment. 126 By contrast, another mouse study has shown improved glucose tolerance that was independent of weight changes, along with lower LPS levels and a lower bacterial count following antibiotic treatment, thereby suggesting an improved metabolic state. 186,187 In an animal model of NAFLD, chronic oral antibiotic administration was able to attenuate hepatic inflammation and fibrosis. 188 This difference could probably be ascribed to the different age, genetic background, and diet used in the two studies.
Prebiotics and probiotics
In a series of studies in rodents and humans, prebiotics have been shown to reduce energy intake and body weight, concomitantly reducing IR and hyperglycemia.
110,178
These effects appear to be mediated by Increased release of anorexigenic gut hormones such as glucagon-like peptide (GLP)-1, GLP-2, and peptide tyrosine; Reduced release of ghrelin (an orexigenic peptide); Improved mucosal barrier function with consequent reduced levels of inflammatory markers and decreased endotoxemia; and Increased butyrate production
Several studies in murine models of obesity and diabetes have demonstrated an improved metabolic profile following probiotic administration, with administration of Bifidobacterium adolescentis, Lactobacillus acidophilus, Lactobacillus casei, L. plantarum LG42, Lactobacillus gasseri BNR17, and Lactobacillus rhamnosus. 56,110,178
Most human studies have reported the beneficial metabolic effect of probiotics, with only a few smaller studies failing to show an improvement in cardio-metabolic variables. 110 Probiotics have been also used in NAFLD animal models. Several animal studies have reported on the profound effect of probiotics on NASH, showing reduced liver damage and de novo fatty acid synthesis, decreased metabolic endotoxemia and inflammation, 189 –192 as well as improved aminotransferase concentrations. 193 –196 Despite these findings, the potential effectiveness of probiotics for patients with NAFLD is still unclear. Loguercio et al. provided the first evidence that probiotic treatment with “VSL#3” (a mixture containing 450 billion bacteria (including Streptococcus thermophilus, Bifidobacterium breve, Bifidobacterium longum, Bifidobacterium infantis, L. acidophilus, L. plantarum, L. casei, and Lactobacillus bulgaricus) could reduce the serum transaminase level and improve some liver function parameters in a group of patients with different types of chronic liver disease. 197 Other studies have reported an improvement of biochemical parameters in patients with NAFLD after probiotic treatments (in particular, B. longum or Lactobacillus spp.). 193,198 However, few randomized controlled trials have found support for the therapeutic use of probiotics in humans. Some recent reviews have summarized the findings of studies showing that probiotic supplementation in animal models and humans improves the inflammatory status and clinical manifestations of NAFLD. 164,199
IBD and the microbiome
IBD is a group of disorders characterized by chronic inflammation of the gastrointestinal tract, of which the two main disease manifestations, ulcerative colitis (UC) and Crohn’s disease (CD), each have distinctive clinical and pathological features. 200 –203 Both CD and UC result from a complex interplay among environmental factors, genetics, and intestinal microbiota composition. 204,205
The important role of the microbiome in IBD pathogenesis has been elucidated by a study where germ-free animals showed less inflammation than controls in response to dextran sulfate sodium (DSS)-induced colitis. 13,206 The fact that the gut microbiota shapes intestinal immune responses has been demonstrated in germ-free mice that showed impaired gut immunity. Evidence for the involvement of the microbiome in IBD comes from experiments where FMT of the microbiota from mice with colitis induced colitis in healthy mice. 207
Another study has found that antibiotic administration in wild-type mice may protect them against colitis. 208 Antibiotics can induce changes in T-cell subpopulations, including reductions in colonic lamina propria, and T cell and CD4+ T cell numbers. Antibiotic treatment conferred protection against DSS-induced colitis, and this effect was transferable by FMT indicating the protective role of the microbial community.
Multiple clinical studies have shown that the intestinal microbiome composition of IBD patients differs from that of people without IBD. Dysbiosis in IBD is characterized by a depletion of Bacteroidetes and Lachnospiraceae (phylum Firmicutes), an increase in Actinobacteria and Proteobacteria, 209 and decreased microbial diversity overall. 209,210 Moreover, gut samples from patients with IBD were depleted of Lachnospiraceae members, in particular, group IV and XIVa Clostridia. 210 At the species level, in patients with CD postsurgery, a lower proportion of F. prausnitzii bacteria were found in their ileum samples, 211,212 while the feces samples from UC patients were associated with an abundance of butyrate-producing Roseburia hominis (phylum Firmicutes). 213 It is important to realize that most of these studies were cross-sectional, which makes it difficult to determine a cause and effect relationship between disease and microbial composition. However, a more recent longitudinal study that analyzed the microbiome in patients at the initial occurrence of disease found an increased abundance of Enterobacteriaceae and Pasteurellaceae (Proteobacteria), Veillonellaceae (Firmicutes), and Fusobacteriaceae, and a decreased abundance in Erysipelotrichales (Firmicutes) and Bacteroidales, and the presence of Clostridia was strongly correlated with the disease status. 214
Discordant results have been found in IBD microbiome studies when different sample types and diverse collection sites (e.g. fecal or surgical sample, remission or inflamed condition, or different portions of the gastrointestinal tract) are taken into account. Indeed, the composition and abundance of both lumen and mucosa-associated microbiota vary along the gastrointestinal tract and both can be significantly impacted by intestinal inflammation in a number of different ways. Ultimately, this could represent a confounding factor for data interpretation.
FMTs are being extensively investigated as therapies for treating IBD.
215
The availability of different mouse models of IBD has led to multiple studies focusing on discerning the mechanisms underlying linkage between the microbiome and IBD. The following immunoinflammatory mechanisms have been proposed: An increased polysaccharide A production and suppressed production of pro-inflammatory cytokine IL-17 and IL-10-producing CD4+ T cell induction;
216
The complexity of the interplay between gut microbiota and luminal IgA responses;
217
–219
An increase in glycerophospholipid and lipopolysaccharide metabolism;
220
A decreased SCFA production with compromised intestinal and immune homeostasis;
213,221
An increase in sulfur-reducing bacteria (observed in UC patients) with consequent production of hydrogen sulfide, a known genotoxic substance able to modulate gene expression in the cell cycle and induce inflammatory responses;
220
and An increased representation of genes involved in cell wall degradation and the exotoxins that facilitate the passage of inflammatory gut mediators into the blood circulation.
Recently, Chu et al. found new gene–microbiota interactions that can contribute to IBD development. 222 In their study, the human gut microbe Bacteroides fragilis was found to produce immunomodulatory molecules that are released via outer membrane vesicles (OMVs). The OMVs triggered the ATG16L1-mediated and the nucleotide-binding oligomerization domain-containing protein 2-mediated noncanonical autophagy pathway in the host dendritic cells (DCs). The OMV-primed the DCs and, in turn, induced the intestinal regulatory T cells that protect against colitis. Immune cells from humans at high risk of IBD have polymorphisms in ATG16L1 and they are defective in their response to OMVs; consequently, this promotes disease through defects in “sensing” the protective signals from the gut microbiome.
IBD, microbiome, and lifestyle factors
Diet
A person’s diet can influence intestinal inflammation (see paragraphs “The microbiome and lifestyle factors” and “Metabolic diseases and the microbiome”) and increase their risk of developing IBD. In human cohort studies, this risk is positively correlated with dietary fat intake. Different mouse models of intestinal inflammation (i.e. TNFdeltaARE, CEABAC10, and IL-10−/− mice) have also identified the same risk. Moreover, the presence of iron in the drinking water, detergents, and artificial sweeteners in drinks and processed foods has also been associated with IBD. 223 Additionally, different diet modification strategies have been shown to have positive effects in IBD patients, including exclusive enteral nutrition (where a precisely defined liquid diet is used exclusively for nutrition), partial enteral nutrition, and semi-vegetarian diets. In addition, probiotic and prebiotic administration showed beneficial effects in treating IBD. 224
Smoking
Extensive research has shown the strong impact of environmental factors on gut microbiota, and smoking has recently been studied as a potential factor involved in shaping microbiota communities. Interestingly, in the two main subtypes of IBD, CD and UC, smoking has divergent effects on the disease course. While CS is the most prominent environmental risk factor for developing CD, it exerts a protective role in UC. 225,226
The molecular and cellular mechanisms by which CS interferes with the pathogenesis of CD and UC are poorly understood. Several potential mechanisms have been proposed and, most probably, the causes of CS-associated changes in the composition of the gut microbiota are a combination of environmental, host, and microbial changes. Among them, the following should be mentioned: epigenetic susceptibility, modulation of mucosal immune response, alterations in intestinal cytokine and eicosanoid levels, modification of gut permeability with consequent impaired clearance of pathogen alteration in the H+/K+-ATPase pump with consequential acidification of gastric contents, and ingestion of the bacteria present in cigarettes. 227 –230,231,232
Human studies targeting selected bacterial groups have reported that smoking patients with active CD have increased abundances of Bacteroides and Prevotella 233 (both Bacteroidetes) and decreased Firmicutes to Bacteroidetes ratios. 60 Similar results were obtained in healthy CS controls, suggesting that the association may not be related to intestinal inflammation but may instead reflect the direct impact of CS on the microbiota. Smokers also display a decreased abundance of Bifidobacterium spp., 60 and hence may lose the anti-inflammatory effects that are often associated with this genus. Despite its small sample size, a recent study partially confirmed and further expanded on the aforementioned studies using a more comprehensive approach (metagenomics) in which the researchers were able to sequence and evaluate the whole gut microbiota of CD patients. 234
Despite the species level differences between human and rodent microbiota, 114,235 a few studies with rats and mice have aligned with the observations from humans, highlighting the CS-dependent shift of the gut microbiota composition. 62 –64 A recent paper reported that 24 weeks of CS exposure increased the activity of Lachnospiraceae spp. (Firmicutes) in the colon, with alteration in inflammatory gene expression. 231 These data highlight a possible role for CS in gut microbiome shaping with unknown consequences in the evolution of inflammation-related disorders such as IBD.
COPD and the microbiome
COPD is characterized by a slow, progressive, irreversible airflow obstruction, and loss of lung tissue leading to emphysema and remodeling of the tissue (fibrosis), both of which contribute further to lung function decline, reduced quality of life, and high mortality. 236,237 Lung infections triggered by pathogenic bacteria and viruses can lead to acute worsening of COPD (exacerbation). Exacerbations are an additional major factor in the morbidity and mortality caused by COPD, and the major source of health-care costs associated with the disease. 238,239
Recent studies, using high-throughput next-generation sequencing (NGS) techniques, have reported that the airways are not sterile (even in healthy hosts) and harbor diverse microbial communities. 240 Nevertheless, the enclosed nature of the lungs presents difficulties for microbiome sampling. Biofluid sampling is possible via bronchoalveolar lavage (BAL) or sputum collection. Recently, it has been suggested that sputum and BAL samples offer spatially distinct representations of the lung microbiome: BAL samples appear to represent the lower bronchial mucosal flora and sputum samples the upper bronchial tract. 241 Therefore, this should be taken into consideration when interpreting COPD microbiome studies in view of the differences between spatially distinct regions of the lungs.
Several investigations have shown that changes in the composition of the airway microbiota are associated with developing chronic lung diseases, such as asthma, COPD, and CF. 242 –244 The heterogeneous nature of COPD implies that for this disease the concept of a “core” lung microbiome is difficult to establish. Candidate genera that constitute the lung core microbiome from a number of COPD lung microbiome studies include Pseudomonas, Streptococcus, Prevotella, and Fusobacteria. 245
Studies have found that the lung microbiome plays an important role in COPD severity. 246,247 Initial studies have suggested that the lung microbiome of patients with moderate and severe COPD is less diverse than those of healthy controls, 244 although other work has suggested that this conclusion may have been based on an underestimation of bacterial diversity. 248 Thus, using a larger cohort of moderate and severe COPD patients, more recent work has suggested that there is increased microbial diversity in more severe COPD cases. 249
Metagenomic sequencing of the sputum microbiome of COPD patients and healthy smokers (considered as controls), which was conducted to elucidate its taxonomic composition, revealed an increase in four bacterial species, all pathogens (S. aureus, Stenotrophomonas maltophilia, Streptococcus agalactiae, and Streptococcus pyogenes). 250 Cameron et al. analyzed the lung microbiome in a large COPD patient cohort and found interesting changes that were associated with multiple characteristics of COPD, including specific exacerbation phenotypes, treatment regimen differences, and differences in the levels of key sputum and serum mediators. The COPD exacerbation events appeared to be associated with decreased microbial diversity and an increased proportion of Proteobacteria. There was also a marked proliferation of Moraxella in a subgroup of subjects during their disease exacerbations. A reduction in microbial diversity and an increased Proteobacteria to Firmicutes ratio toward recovery were observed in the subjects treated with steroids alone, whereas the trend was reversed in the subjects who received antibiotics. However, this study, despite its large cohort size, was focused exclusively on COPD patients undergoing exacerbations. Healthy control subjects and those who did not experience exacerbations were excluded. 250
Links between COPD, the microbiome, and lifestyle factors
Smoking
Smoking is the leading risk factor for COPD. Changes in the immune system, triggered by the noxious particles present in tobacco smoke, lead to an inflammatory cellular infiltrate and to a pronounced and chronic lung inflammation. This, in turn, induces other pathological changes, including chronic obstructive bronchitis with fibrosis and obstruction of the small airways, emphysema, destruction of lung parenchyma, and the loss of lung elasticity. 251,252 CS also leads to lung infections and to consequential exacerbations.
Opinion on the effect of smoking on the COPD microbiome is controversial. Erb-Downward et al. recently failed to report any difference in the bacterial communities of the lower respiratory tract microbiome from a very small cohort of smokers, nonsmokers, and COPD patients through analysis of their BAL fluids. 244 Contrastingly, an analysis of sputum microbial composition revealed that CS is a major environmental factor that not only drives the difference in microbial community structure between samples but also affects the abundance of specific microbial taxa. 253 In particular, smokers showed an increased abundance of specific members of sputum microbiota, such as Veillonella and Megasphaera (both Firmicutes) and all the taxa to which these genera belong (Firmicutes, Clostridia, Clostridiales, and Veillonellaceae). This respiratory microbiota alteration can lead to an inflammatory condition or an environment in which pathogenic bacteria can thrive and can also subsequently contribute to the development of smoking-related lung diseases, such as COPD. Similar to what occurs in the gastrointestinal tract, an interesting paper has reported that CS can induce the loss of lung barrier integrity, and this allows bacterial translocation and increased tumor-associated inflammation. 254
Probiotics
Cigarette smoking impairs human natural killer (NK)-cell cytotoxic activity and cytokine release. 255 However, it has been found that the daily intake of the L. casei Shirota strain increases NK cell activity in smokers. This suggests that probiotics may be useful for patients with COPD, particularly those experiencing frequent viral infections. 256
Drugs
Currently, there is no cure for COPD. Therefore, the goals of COPD treatment are mainly focused on relieving the symptoms and preventing or treating the complications of it. The main drugs used are bronchodilators that can be used in combination with glucocorticosteroids and antibiotics in cases with complications. These treatments can also affect the composition of the gut microbiota. With disease exacerbations, antibiotic treatment induces a reduction in Proteobacteria, and prolonged suppression of some microbiota members has been observed. 257 Conversely, corticosteroid treatment increases the abundance of Proteobacteria and other phyla members.
Environmental factors
Research on the alterations in lung microbiota resulting from environmental exposure to a range of pollutants is a nascent field with the potential to explain the pathophysiological mechanisms of lung disease. A clinical study in a healthy adult population in Malawi revealed a correlation between high levels of particulates from the inhalation of smoke from biomass fuel and changes in their lung microbiomes. Exposure to other environmental sources of particulates caused a higher abundance of potentially pathogenic bacteria (Streptococcus, Neisseria) within their lung microbiomes. 258
Periodontitis and the microbiome
Periodontitis, a chronic inflammatory disease affecting both soft (gingiva) and hard (alveolar bone) tissues, 259,260 can destroy periodontal tissues and cause tooth loss. Around 40% of people in low-income countries are affected by periodontal diseases, and the percentage remains high even in high-income countries, where 30% of the people are reported to have oral health problems. 261 Periodontitis development is associated with an increase in microbial load and dramatic shifts in the microbial community structure, with the primary health-associated species remaining part of the periodontal communities, albeit in low proportions, and a diverse range of periodontitis-associated taxa becoming numerically dominant. 262 –264
The human oral cavity contains a number of different habitats, including the teeth and oral mucosa, which are colonized by bacteria. Saliva, which is constantly in contact with the bacterial flora attached to the surfaces, contains the whole representative population of the oral cavity. 265 The commensal populations (normal microflora) may keep pathogenic species in check by creating a biofilm and not allowing them to adhere to mucosal surfaces. In effect, the bacteria do not become pathogenic and cause infection and disease until they breach the commensal barrier. 266 The bacterial tropism and distribution in the oral cavity varies depending on the physical location, oral cavity physiology, immunity, metabolism, and habits. 267 –270 Disruption of bacterial plaque homeostasis and an increased bacterial load are associated with pathological outcomes such as dental caries, gingivitis, and periodontitis. 271,272
The oral microbial composition in patients with periodontitis undergoes complex changes and more than 400 microbial phenotypes are associated with periodontal pockets. 272 Different bacterial species combinations are associated with different pathogenicity grades. For example, Porphyromonas gingivalis, Tannerella forsythia, and Treponema denticola (all Bacteroidetes) are considered among the most pathogenic when complexed together. 273,274 Other Bacteroidetes species, like Eubacterium saphenum, Porphyromonas endodontalis, Prevotella denticola, Parvimonas micra, Peptostreptococcus species, Filifactor alocis, Desulfobulbus species, Dialister species, and Synergistetes are also associated with periodontitis. 67,275 –277
Links between periodontitis, the microbiome, and lifestyle factors
Diet
Until now, very little information has been available about the association between diet and the oral microbiome. Individuals fed on high-carbohydrate diets have higher abundances in their oral cavities of acidogenic and aciduric bacteria such as Lactobacilli and Streptococcus mutans (both Firmicutes) whose metabolites are the primary cause of dental caries. 278 Consistent with these findings, it has been found that high levels of salivary glucose (derived from an unbalanced diet) are associated with a reduced bacterial count and modified bacterial frequencies. 279 Fatty acid and vitamin intake can also modify the oral microbiome, with saturated fats being associated positively with a high Betaproteobacteria and Fusobacteria abundance, whereas vitamin C supplementation is associated with the presence of Fusobacteria, Leptotrichiaceae, and Lachnospiraceae; however, these findings involve quite modest relative changes. 278
Oral bacteria can activate alcohol and convert it to the genotoxic and carcinogenic compound, acetaldehyde. For this reason, oral exposure to acetaldehyde can lead to carcinogenic effects in the oral and gastrointestinal tract. Moreover, researchers have found that pretreatment with the antibacterial chlorhexidine prior to ethanol exposure can reduce acetaldehyde levels in the saliva. 280 The continued use of alcohol and tobacco is known to lead to reduced bacterial richness (decrease in Neisseria, Fusobacteria, Granulicatella, Peptostreptococcus, and Gemella abundance), with possible consequences for human oral diseases. 281
Smoking
Several conditions are risk factors for periodontitis, with smoking identified as a major contributor. 282,283 Depending on the definition of disease and the exposure level to smoking, the risk of contracting a destructive periodontal disease is 5- to 20-fold higher for a smoker than for someone who has never smoked. 284 Ge et al. 285 reported that smoking was accountable for bacterial variations in both healthy and diseased pockets in patient with periodontitis. 285 They also found that periodontal disease may be associated with a consortium of bacteria rather than few specific species. However, the composition of oral bacteria in smokers versus nonsmokers varies largely among studies. 66,285 –288 This might be related to different sample sizes, different sampling sites in the mouth, and/or the use of different methodologies and analysis tools. Generally, smokers have a variable, pathogen-rich, and commensal-poor anaerobic microbiome, resembling more closely a disease-associated state than a clinically healthy community. 66,288 –290
Smoking-induced perturbations may, for example, be attributed to oxygen deprivation, the effects of antibiotics, decreased saliva pH, and impaired host immunity. 291 –293 Several studies have investigated the changes occurring in the oral bacterial composition in smokers, nonsmokers, and quitters. Higher abundances of species belonging to Bacteroides, Campylobacter, Fusobacterium, Parvimonas, and Treponema, and lower abundances of Veillonella and Streptococcus (both Firmicutes) as well as Neisseria, have been detected in smokers with periodontitis compared with those who had never smoked. 294 Streptococcus prevents pathogen colonization 295,296 but its commensalistic relationship with Parvimonas is impaired by CS. 297
A longitudinal 12-month smoking cessation study in patients with periodontitis showed that the changes occurring in the microbiome of quitters were related mainly to shifts in the relative proportions of bacterial species rather than in the number of species. 297 In a similar study, Delima et al. observed a decreased presence of P. endodontalis, Dialister pneumosintes, P. micra, F. alocis, and T. denticola, and a re-colonization with healthy bacterial species, such as Veillonella parvula, in patients with periodontitis who quit smoking. 291 Interestingly, members of the Veillonella genus (Firmicutes) form a major component of the subgingival microbiome when the periodontal area is healthy. Finally, Wu et al. showed that the oral microbiome of smokers versus nonsmokers and former smokers differed substantially and that smoking cessation reverted the microbiome to a composition similar to that of nonsmokers. 298
The impact of electronic nicotine delivery systems (ENDS) on the microbial physiology is poorly understood. Recently, Kumar et al. compared the effects of ENDS and CS, reporting that the oral microbial composition of ENDS users differs from smokers and those who had never smoked, with its composition sharing no more than 15% of functionally annotated genes with controls and current smokers (abstract from IADR 2017—Using e-cigarettes to quit smoking is not helping your microbiome—Kumar et al.).
Drugs
Antibiotic use in dentistry has a profound impact in the clinical setting, as antibiotic-resistant bacteria cause difficult to treat oral infections or cause treatment failures. Metagenomics insights have provided evidence of multiple antibiotic resistance genes in the human oral microbiome. This type of antibiotic resistance is acquired through horizontal gene transfer and the oral biofilm is probably the perfect environment for this transfer through the close physical contact between phylogenetically distant bacteria. 299 Recently, a metagenomic approach was used to develop “genome-inspired personalized medicine,” whereby analyzing the oral microbiome after antibiotic treatment would allow the prescription of an antibiotic with an appropriate dosage and spectrum of activity to be tailored to the targeted bacteria. For instance, in a clinical study on a healthy population, Zaura et al. found that after one treatment with antibiotics the oral microbiome became more ecologically stable than the gut microbiome in terms of its species composition. 299,300
Probiotics
Diet supplementation with Lactobacilli has negative effects on the growth of salivary S. mutans, which is a major contributor to tooth decay. 301 Similar results have been found with people who have a daily intake of yoghurt containing Bifidobacteria and tablets containing L. reuteri. One recent model for probiotic treatment is represented by the ingestion of Streptococcus salivarius, a nonpathogenic oral species that produces broad-spectrum bacteriocins (antimicrobial peptides produced by bacteria) and also harbors mega-plasmids containing bacteriocin-encoding genes. 302
Environmental factors
Personal hygiene products, such as soaps and toothpastes, that contain the antibiotic triclosan do not seem to have a major influence on microbial communities or endocrine function, according to a small, randomized trial. 303
Brain disorders and the microbiome
Mental health problems such as anxiety and psychotic disorders are not just diseases caused by psychological stressors added to genetic vulnerability, but rather full-body, inflammatory conditions related to the immune state. 304 –307 Similarly, even neurodegenerative diseases, such as Alzheimer’s disease (AD), are marked not only by age-related brain changes but also by disturbed immune function and increased oxidative stress. In rodent models, these factors have been shown to be influenced by diet and the gut microbiota. 308 Over the past few years, accumulating evidence has pointed to the critical role played by bidirectional communication between the gut and the brain in neurological disorders.
The gut microbiota makes a critical contribution to the control of the central nervous system (CNS) activities through neural, endocrine, and immune pathways, 309 including a direct interaction between the gut microbiota and enteric neurons, 310,311 regulation of the hypothalamic–pituitary–adrenal axis (HPA axis), 312 growth and function of CNS cell populations, 313 and the production of many chemicals important for normal brain functioning (e.g. serotonin, dopamine, kynurenine, γ-aminobutyric acid, SCFAs, p-cresol). 314,315 This gut–brain axis will be discussed further in the paragraph “Microbial community cross talk.”
Alterations in the gut microbiome are also associated with neuropsychiatric disorders, such as schizophrenia, bipolar disorder, autism, major depressive disorder, chronic fatigue syndrome, anxiety, and stress, 316 –319 as well as neurodegenerative diseases (Parkinson’s disease (PD), AD, dementia, and stroke). 313
Autistic spectrum disorders
Autistic spectrum disorders (ASDs) are neurodevelopmental disorders characterized by alterations in the social interactions associated with communication as well as behavioral impairment. It has been found that ASD subjects frequently have problems related to a dysfunctional bowel with an aberrant intestinal barrier function. 320,321 While analyzing the microbiota of children with ASDs, it was found that they usually have a higher abundance of Proteobacteria and Bacteroidetes and a lower abundance of Firmicutes and Bifidobacteria, when compared with the healthy controls. 322 –324 Interestingly, of the many classes of bacteria within the Firmicutes phylum, one class, in particular, the Clostridia, are present in higher numbers in autistic children with a history of gastrointestinal problems. 325 Moreover, despite the overall abundance of Bacteroidetes in autistic subjects, lower counts of Prevotella are seen. 49
A recent study analyzed a cohort of autistic individuals and the authors found a different composition of bacterial gut microbiota compared with the controls, 326 typified by a significant increase in the Firmicutes to Bacteroidetes ratio in these subjects via a reduction the relative abundance of Bacteroidetes. At the genus level, there was a decreased relative abundance of Alistipes, Bilophila, Dialister, Parabacteroides, and Veillonella in the ASD cohort, while Collinsella, Corynebacterium, Dorea, and Lactobacillus levels were significantly increased. Additionally, the same authors analyzed the ASD subjects with constipation, one of the common gastrointestinal problems in this group and found high levels of bacterial taxa belonging to Escherichia, Shigella, and Clostridium cluster XVIII. In a recent small study of 18 children with autism, positive changes in gastrointestinal symptoms and neurological symptoms have been noted after microbiota transfer therapy. 327
Rett syndrome
Rett syndrome (RTT), a severe and progressive neurological disorder, is linked primarily with a mutation in the gene encoding methyl-CpG-binding protein 2, which is a fundamental mediator of synaptic development and plasticity. This syndrome is commonly associated with gastrointestinal disorders such as constipation. Indeed, RTT subjects are characterized by a reduction in gut microbial richness and an increase in microbial taxa belonging to Bifidobacterium, several Clostridia (Anaerostipes, Clostridium XIVa, and Clostridium XIVb) as well as Erysipelotrichaceae, Actinomyces, Lactobacillus, Enterococcus, Eggerthella, Escherichia, and Shigella. 328
Schizophrenia and bipolar disorder
Schizophrenia and bipolar disorder are two human psychiatric disorders with uncertain etiologies. To date, few studies that have analyzed a possible link between the microbiome and these two mental illnesses have been published, but the topic has been reviewed very recently by Dickerson et al. 311 Dysbiosis, increased gastrointestinal inflammation, and the association between antibiotic treatment and the incidence of psychiatric disorders are the main features found in the different clinical studies that have analyzed patients with psychiatric disorders. 311
Neurodegeneration
Parkinson’s disease
PD is the second most common adult neurodegenerative disorder. Its clinical signs include abnormal movement, tremor at rest, rigidity, slowness or absence of voluntary movement, postural instability, and movement freezing, all of which are caused by defects in motor control, but nonmotor symptoms are also common (depression, lack of motivation, passivity, and dementia). 329 Despite improvements in treatment, the etiology of PD remains unclear and there are no therapies to delay or prevent it.
The role of the gut microbiome in the pathogenesis of PD is beginning to emerge. Braak et al. hypothesized that the disease begins in the gut and spreads to the brain via the gut–brain axis (i.e. the vagus nerve and spinal cord). Indeed, the parasympathetic fibers of the vagus nerve that innervate the intestine among other regions arise from the dorsal motor nucleus. Lewy bodies (aggregated proteins, mainly alpha-synuclein and ubiquitin), which are the hallmark of PD, were found in the enteric nervous system in postmortem cases of early PD. 330
In one study, PD was reported to be associated with alterations in Prevotella and Enterobacteria populations. 331 Prevotella is known to break down complex carbohydrates, providing SCFAs as well as thiamine and folate as by-products that promote a healthy intestinal environment. Decreased Prevotella numbers are likely to result in reduced production of these important micronutrients, and this might lead to reduced production of essential vitamins and impaired secretion of gut hormones. However, the study did not evaluate whether the patients had a history of gastrointestinal disturbances or significant inflammation. 332
Several papers published by Sampson et al. over the last few years showed that in animal studies the connection between the gut microbiota and the pathogenesis of PD seems to be mediated by the regulation of the activation status of microglia 333 and that gut microbiota could affect the disease severity. 334,335 It is clear that identifying the gut microbiota-generated compounds that can affect the immune response of microglia in the brain and the development of PD will be important in future investigations.
Alzheimer’s disease
The microbiome may play a role in the formation of beta-amyloid plaques in the mouse brain during AD development. 336 Germ-free AD transgenic mice have significantly lower levels of beta-amyloid in their brains than conventionally raised transgenic mice. Moreover, AD transgenic mice have a significantly different microbiome than that of wild-type mice and, at the phylum level, Firmicutes and Bacteroidetes levels were significantly altered, whereas at the genus level, Allobaculum and Akkermansia levels were lowered, while unclassified Rikenellaceae and S24-7 genera levels increased. Furthermore, fecal transplants from transgenic mice were found to induce significant upregulation of beta-amyloid production in the brains of the germ-free AD transgenic mice. 336
Ischemic stroke
In three separate mouse models of microbiota disruption, the microbiome was shown to impact the outcome of ischemic stroke. Depletion of the microbiota with a cocktail of antibiotics decreased survival in a middle cerebral artery occlusion model of murine stroke and severe colitis in mice. 337 In another model in which the mouse microbiota was altered, but not depleted by antibiotic treatment, the infarct volume was significantly reduced compared with mice with an unaltered microbiota. 338 This microbiota-stroke effect may be bidirectional, as several changes in the microbiome have been identified in different experimental stroke models (reviewed in literature 339 ). For instance, Singh et al. found that particularly large infarcts can cause gut dysbiosis, possibly potentiating the neuroinflammatory effects within the CNS. 340 Interestingly, Benakis et al. reported that gut microbiota confers a neuroprotective effect by modulating immune cells in the small intestine. 338 After antibiotic treatment, the authors found that the consequent dysbiosis induced an altered DC activity associated with an expansion of regulatory T cells which secrete IL-10. This cytokine is able to suppress the differentiation of γδ T cells into IL-17-producing γδ T cells. IL-17+ γδ T cells are known to have a deleterious effect after stroke because they can migrate to the meninges and aggravate ischemic brain injury by secreting IL-17 and promoting neutrophil infiltration. The decrease in IL-17+ γδ T cells, observed after antibiotic treatment, is essential for the reduction of post-ischemic chemokines (Cxcl1 and Cxcl2) leading to a decrease in ischemic brain injury. 338 A recent study discovered the molecular mechanism in endothelial cells that underlies the formation of cerebral cavernous malformations (CCMs), which are clusters of dilated, thin-walled blood vessels in the brain that can cause strokes and seizures. This molecular pathway is activated by TLR4, a receptor for the bacterial molecule LPS, which is present on brain endothelial cells and is capable of vastly accelerating CCM formation. 341
Links between brain disorders, the microbiome, and lifestyle factors
Prebiotics and probiotics
Recently, the term “psychobiotics” has been introduced into the scientific vocabulary. They are a family of probiotics and prebiotics that are capable of modulating the gut–brain axis and have a positive mental health benefit. In animal models, it has been extensively shown that psychobiotics can alleviate neuropsychiatric disorders via different neurochemical and humoral mechanisms. These mechanisms include modulation of the expression of brain-derived neurotrophic factor, increased hippocampal neurotrophin levels, regulation of GABA transcription, activation and/or inhibition of specific neurons, and regulation of the immune system in CNS autoimmunity. 316 Prolonged administration of L. rhamnosus reduced the stress-induced corticosterone levels and signs of depression and anxiety in mice. 342 These effects were probably induced by alterations in GABA (B1b) receptor gene expression in the brain. However, these effects disappeared in mice in which the vagus nerve had been removed, suggesting once again that the vagus nerve is a key mediator of gut–brain communication. In contrast, similar administration of a combination of L. rhamnosus and B. animalis in patients diagnosed with schizophrenia saw no difference in the psychiatric symptom severity between the placebo and probiotics groups. 311 Contradictory results have emerged from depressed patients who consumed probiotics; while in some studies the use of probiotics seemed not to be associated with lower rates of depression, 318 in other studies the probiotic treatment produced a reduction in network-level neural reactivity to negative emotions, anxiolytic activity, suppression of psychological stress, and reduction of the negative thoughts associated with low mood. 343
Skin diseases and the microbiome
The skin is the largest organ in the human body and forms an immense interface between the host and its environment. As such, it is an important site for interactions between the immune system and its microbial inhabitants. Skin symbiotic organisms play essential roles in lipid metabolism and colonization resistance to transient organisms. 86,344
Generally, the four dominant bacterial phyla residing on the skin are Actinobacteria, Proteobacteria, Firmicutes, and Bacteroidetes, 345 whereas, in terms of the different genera, the most abundant are Staphylococcus, Propionibacterium, and Corynebacterium. A feature of the skin is the topographical diversity of the bacterial populations, with the composition and diversity of the bacteria colonizing the skin depending on the microenvironment. Compared with other sites in the human microbiome, variability between individuals is similarly high. 346,347 Recently, it has been shown that despite the skin’s exposure to the external environment, its bacterial community is stable at the strain level. The nature and degree of this stability is highly individual and site-specific and is driven primarily by the maintenance of individual strains over time. 348
Emerging evidence has established a link between the development of the resident immune system of the skin and its microbiota by identifying the direct contact between the two (reviewed in Belkaid and Segre 344 ). A clinical study showed that patients with primary immunodeficiency suffer from atopic dermatitis-like eczema and increased risk of skin infections. 349 Moreover, a study in mice has shown that complement signaling antagonism (through the C5a receptor) resulted in a shift in skin microbial composition and diversity. 350 A different study found that a molecule released by staphylococcal bacteria, lipoteichoic acid, inhibits skin inflammation through a TLR2-dependent pathway after injury. 351 Collectively, these findings suggest that the microbiota modulates cutaneous inflammation and immunity, thereby maintaining the delicate balance between the host and microorganisms. Disrupting this balance can cause skin disorders and infections. Classical microbiological and dermatological studies have reproducibly pointed to the strong association between P. acnes and acne vulgaris, 352 between S. aureus and atopic dermatitis, and between Malassezia species and seborrheic dermatitis. 86,352
The advent of NGS technology, as applied to research on the microbial communities of the skin, has led to further information about the microbial biofilm nature of certain skin diseases that were difficult to analyze using classical microbial techniques. In atopic dermatitis, for example, a decrease in microbial diversity was discovered in the lesions. 353
Links between skin diseases, the microbiome, and lifestyle factors
Diet
Acne vulgaris is known to be fueled by the high glycemic load typical of a Western diet, which stimulates lipid production in hair follicle sebaceous glands, leading to overgrowth of P. acnes. 354 In people with acne, the activity of the transcription factor FoxO1 and the kinase mTORC1 have been found to be aberrant, leading to an overproduction of monounsaturated fatty acids and triglycerides in the sebum, thereby allowing the growth and colonization of P. acnes. 355 –357 Treatment with metformin, an mTORC1 inhibitor, produced positive results in male subjects who did not respond to common acne treatments. 358
Prebiotics and probiotics
The beneficial effects of pre and probiotics are achievable either by ingestion or by topical application. Prebiotics applied topically can enhance the growth of beneficial “normal” resident skin microbiota. A cosmetic product with specific prebiotic extracts including ginseng, black currant, and pine was shown to reduce colonization by P. acnes in patients with acne. 359 Ingested probiotics can be effective at reducing acne and atopic dermatitis, probably by inhibiting pro-inflammatory cytokine production. Topical applications of probiotics have a direct result at the application site and improve the skin’s natural barrier defenses. In fact, probiotics can produce antimicrobial peptides that enhance the skin’s immune responses thereby eliminating pathogens. 82
Topical probiotics and their lysates have been shown to be of value in treating acne. A topical product containing a 5% extract of L. plantarum was found to reduce erythema, acne lesion size and improve the skin barrier of patients with acne. 360 A lactic acid bacterial strain, E. faecalis SL-5, seems to have antimicrobial activity against P. acnes. 361 Furthermore, innovative nutritional products called synbiotics, which contain prebiotics and probiotics, have positive therapeutic results for atopic diseases. 82
Microbial community cross talk
In this ever-expanding field, research has focused particularly on how local microbiota influences immunity at distal sites, especially on how gut microbiota influences other organs, such as the brain, liver, or lungs. This has led to the coining of terms such as “gut–lung axis,” “gut–brain axis,” “gut–brain–β-cell axis,” and “gut-skin axis.”
The gut–lung axis
Chronic lung diseases, such as asthma and COPD, often occur together with chronic gastrointestinal tract diseases, such as IBD or IBS. 362,363 Almost 50 years ago, the first evidence of pulmonary-intestinal cross talk was reported by Turner-Warwick. 364 In 1976, Kraft et al. 365 also noted the development of severe, chronic bronchopulmonary disease in patients with IBD, years after being diagnosed. In a large population-based Canadian study performed 15 years ago, it was reported that pulmonary complications were the most common concomitant chronic disorder in patients with IBD 366 and that pulmonary involvement was more pronounced in patients with active disease compared with those in remission. 367
Today, it is believed that almost half of patients with UC 368,369 and half of those with CD 369,370 have subclinical pulmonary abnormalities, and COPD is now even considered to be a significant mortality factor among patients with CD. 371,372 Evidently, it seems that a healthy gut microbiota is beneficial for lung health and that the existence of co-morbidities in many diseases should not be ignored.
From a mechanistic perspective, we now know that cross-regulation of gut–lung immunity exists and that intestinal microbiota have been shown to directly modulate the pulmonary immune responses to invading pathogens. 373,374 When the gut microbiota is disturbed during infection or antibiotic exposure, for example, the normal microbiota-derived signals are altered, which changes in the immune response. This means that dysbiosis of the gut microbiota, for example, through exposure to CS, can cause systemic inflammation with outgrowth and secretion in the systemic circulation of opportunistic pathogens, which in turn can further negatively perturb the chronic inflammation already present at distal sites (e.g. the lungs). Considerable evidence suggests that host epithelia and other structural and immune cells assimilate information directly from microorganisms and from the concomitant local cytokine response to adjust inflammatory responses and that this shapes immune responses at distal sites, such as in the lungs. 375,376 It is not clear if there is a direct transfer of microorganisms between sites, although the translocation of bacteria from the gastrointestinal tract to the lungs has been observed in sepsis and acute respiratory distress syndrome, where the barrier integrity has been compromised. 377
Since 2015, few clinical studies have tested whether altering the gut microbiota with probiotics or antibiotics can influence lung health. 378 Research conducted by Benoit Guery (Lausanne University Hospital, Switzerland) is now exploring “narrow-spectrum” antibiotics to treat lung infection without damaging the gut microbiota. 378
Thus far, in gut–lung microbiota studies, it is difficult to ascertain whether changes in the gut microbiota are a cause or a consequence of COPD. Most likely, both are true and operate simultaneously or at different stages of the disease. Clearly, longitudinal studies in humans and other animals are now required to determine whether changes in the gut or lung microbiota are associated with the development and severity of COPD or IBD, respectively. Undoubtedly, diseases should no longer be studied in isolation, but rather all comorbidities should be taken into account so that treatments are adapted to avoid a negative impact on overall health.
The gut–brain axis
The role of the gut–brain axis is to monitor and integrate gut functions as well as to link the emotional and cognitive centers of the brain with the peripheral intestinal functions and mechanisms such as immune activation, intestinal permeability, enteric reflex, and enteroendocrine signaling. The mechanisms involve neuro-immuno-endocrine mediators, including also the glucose produced in the liver through gluconeogenesis. 379 –382 This bidirectional communication network includes the CNS, the brain and spinal cord, the autonomic nervous system, the enteric nervous system, and the HPA axis. Recent advances in research have described the importance of gut microbiota in influencing these interactions.
The gut–brain microbiota axis is a bidirectional communication system enabling gut microbes to communicate with the brain and vice versa. 383 Different studies have pointed out the correlation between gastrointestinal and brain disorders. For example, IBD has been shown to be a risk factor for PD, 384 for dementia 385 and for AD. 386 Moreover, IBD was shown to precede glaucoma, a progressive neurodegeneration of the optic nerve, in two different cohorts. 387
The mechanisms of signal transmission are complex and not fully elucidated, but we are aware of the involvement of endocrine-, neurocrine-, and inflammation-related signals generated by the gut microbiota that can affect the brain. In turn, the brain can influence the microbial composition and function via endocrine and neural mechanisms 388 –390 (Figure 5). An intestinal change in the symbiont (commensal bacteria with beneficial potential) balance and pathobionts (commensal bacteria with pathogenic potential), favoring pathobiont overgrowth, results in dysbiosis, which can induce inflammation. During inflammatory responses, macrophages contribute to pathogenesis via their inappropriate responses to enteric microbial stimuli, their inefficient clearance of microbes from host tissues, their impaired pro- and anti-inflammatory responses, and the loss of barrier function (leaky gut). This promotes the increased translocation of pathogenic bacterial components from the intestinal mucosa to the systemic circulation, where they activate innate immunity, as characterized by the production of pro-inflammatory cytokines, and resulting in systemic inflammation and abnormal gut function. These mechanisms potentially lead to impaired CNS function such as alterations in neurochemistry, cognition, behavior, stress responses, and visceral pain. Conversely, pathological stress at the level of the CNS can affect the gut microbiota function and lead to perturbations of the barrier function and gut motility.
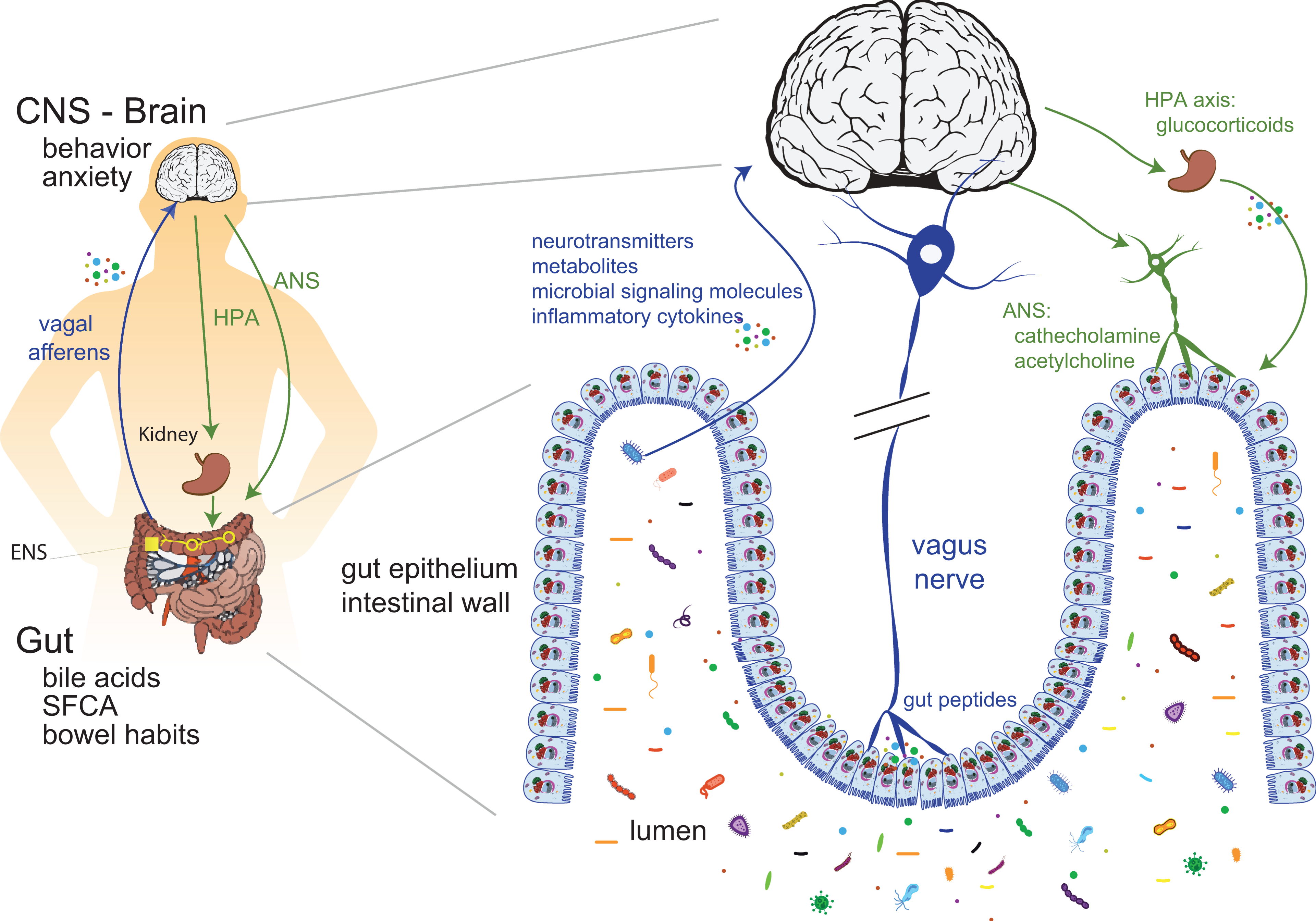
Gut–brain axis. The principal components of the gut–brain axis: endocrine, neural, immunological, and metabolic pathways by which the microbiota influences the brain and vice versa. The afferent arm pathways are represented in blue. (1) inflammatory cytokines released by lymphocytes in the gut lumen may have endocrine or paracrine actions, (2) vagus nerve can be activated by gut peptides released by enteroendocrine cells, (3) gut microbiota can also release neurotransmitters or its precursors with endocrine or paracrine actions, (4) microbial metabolites like SCFA could influence directly or indirectly brain function and behavior. On the opposite direction, there is the efferent arm (in green), (5) glucocorticoids, released as results of the HPA axis activation, modulates gut microbiota composition, (6) neuronal efferent activation of ANS may include the release of acetylcholine and catecholamine. These pathways are so-called “anti-inflammatory cholinergic reflex” and sympathetic activation respectively and they can alter the gut microbiota composition. Adapted from “The microbiota–gut–brain axis: neurobehavioral correlates, health and sociality,” 2013. 391 SCFA: short-chain fatty acids; HPA: hypothalamic–pituitary–adrenal; ANS: autonomous nervous system.
Several animal studies have elucidated various details about the gut–brain axis. For example, the prevention of gut leakiness with Lactobacillus farciminis and blocking LPS translocation using a specific myosin light chain kinase inhibitor have been found to attenuate the HPA response to stress. 392 Animal models have also shown that chronic mild stress alters the microbiota profile associated with stress-induced despair behavior 393 and that in germ-free mice there is an exacerbation of the HPA axis response to acute stress. 394 In vivo studies have opened new avenues to future therapeutic approaches. In mice, a specific probiotic therapy attenuated intestinal dysbiosis and its consequent visceral hypersensitivity. 395 Similarly, the ingestion of L. rhamnosus (JB-1) in mice reduced stress-induced anxiety and depression-like behavior through a vagal pathway. 396 The effect of probiotics on depressive clinical signs has been extensively studied in clinical studies and is reviewed by Wallace and Milev. 397 From a mechanistic standpoint, preclinical studies have implicated the vagus nerve as a key route of neural communication between gut microbes and centrally mediated behavioral effects. As mentioned above, this connection was revealed by the elimination of the CNS effects of L. rhamnosus after vagotomy 396 and by a decreased risk of certain neurologic disorders in humans who underwent vagotomy. 398
The gut microbiota also regulates key neurotransmitters, such as serotonin, by altering the precursor levels; for example, B. infantis has been shown to elevate plasma tryptophan levels and thus influence central serotonin transmission. 399 Intriguingly, the synthesis and release of neurotransmitters from bacteria has been reported. Lactobacillus and Bifidobacterium spp. can produce GABA, Escherichia, Bacillus, and Saccharomyces spp. can generate noradrenaline, Candida, Streptococcus, Escherichia, and Enterococcus spp. can produce serotonin, Bacillus can produce dopamine, and Lactobacillus acetylcholine. 400,401 These microbial-synthesized neurotransmitters can cross the mucosal layer of the intestine. However, it is highly unlikely that they directly influence brain function because they are incapable of crossing the blood–brain barrier (BBB). Instead, their impact on brain function is likely to be indirect, acting on the enteric nervous system. Inflammatory cytokines produced in the gut can travel via the bloodstream to the brain. Under normal physiological circumstances, they cannot signal across the BBB, but they can influence brain areas, such as the hypothalamus, where the BBB is more permeable. It is through the latter mechanism that IL-1 and IL-6 cytokines activate the HPA axis, leading to the release of cortisol, which is the most potent activator of the stress system. In conclusion, the HPA axis, immune system, and vagus nerve represent key players in the neuroendocrine system of the gut and have a significant impact on the gut–brain–microbiota axis. 316,402
The gut–brain–β cell axis
Recent publications have highlighted the previously unknown role for gut microbiota in stimulating insulin secretion by signaling to the brain, through an axis called the “gut–brain–β-cell axis.” 22,403,404 Upon exposure of rodents to a high-calorie diet, the microbiota increases the acetate turnover. The resulting augmentation of acetate production leads to activation of the parasympathetic nervous system. This neurological activity triggers in turn both secretion of the hormone ghrelin, which increases food intake, and augments insulin levels, thereby promoting calorie storage and fat gain. This generates a positive feedback loop, resulting in hyperphagia (via increased ghrelin secretion), increased energy storage as fat (via increased glucose-stimulated insulin secretion) and liver and muscle IR. Consequently, the gut microbiota–brain–β cell axis promotes obesity and concomitant hyperlipidemia, fatty liver disease, and IR. Thus, a high-fat enriched diet can induce obesity not only by providing excess nutrients but also by increasing the microbiota-mediated acetate production that stimulates the brain to promote hyperinsulinemia and energy storage.
The gut–skin axis and gut–brain–skin axis
Recently emerging evidence from interdisciplinary research groups supports the existence of a gut–skin axis that communicates via metabolites, the neuroendocrine system, diet, and the central nervous system. 354 In healthy state, different types of interactions occur between the gut and skin. Gut microbiota produce metabolites, hormones, and neurotransmitters that can enter the blood circulation and affect the skin function. Dietary components can also have access to the skin either directly or via processing by the microbiota. On the other hand, the skin is able to synthesize vitamin D, which can modulate the gut function. 405
In a disease state, under the conditions of dysbiosis, the toxins produced by the gut microbiota can reach the skin through a leaky gut barrier. The inefficient handling of gut commensal bacteria and bacterial components by the liver can also cause the induction of pro-inflammatory cytokines that can reach the skin. This alteration in the skin’s immune environment, together with toxin production, can then contribute to the development of skin pathologies.
Only recently has more attention been focused on the interaction of the gut–skin axis with the CNS, the so-called gut–brain–skin axis. 406,407 Several studies have demonstrated a correlation among these organs such as pathological changes in the visceral organs that can lead to “zones of referred pain” (Head’s zones) in the skin. 408 Arck et al. found that when mice ingested a Lactobacillus strain, reduced neurogenic skin inflammation and hair growth inhibition was induced by a perceived stress. 406,407 Several questions remain open for the scientific community to answer, including the nature and the effects of the molecules present in the gut on the skin and the response of the skin microbiota to gut changes.
Conclusions and outlook
According to the current scientific literature, the composition and metabolic activity of the microbiota has an association with many diseases, including the disorders associated with obesity, chronic inflammatory diseases, CVD, cancers, stress, and even neurodegenerative disorders. Our biology seems to be a complex interplay between the gene products encoded in our DNA and those originating from the microbes within us.
During the past decade, microbiome research has expanded massively. Innovative sequencing techniques now allow us to detect bacteria and other organisms that are unable to grow in the laboratory and to reveal the nature of the microbial communities present on every bodily surface. Studies involving germ-free mice or antibiotic-treated mice have also uncovered connections between microbiota and human health.
The increasing need for population-scale data to better understand the role of these bacterial communities has led to the creation of different global projects. The US NIH-funded Human Microbiome Project Consortium has brought together several scientific experts to evaluate the role of the microbiome in health and disease and its relationship with the human host. 409 Similarly, the American Gut Project aims to analyze the human gut microbiome to determine what makes a healthy or sick gut. 410
One important factor emerging from these research advances is the importance of microbial diversity. The gut microbiota of an individual is more diverse under healthy conditions than during disease when the diversity is reduced. In fact, low microbiome richness is associated with metabolic dysfunction, skin disorders, gastrointestinal disorders, and low-grade inflammation. 130,411,412 It is believed that for pathology to develop, the ratio of potentially pathogenic to beneficial commensal microbes determines outcome, rather than the presence of a specific organism or group 413 (Table 1 and Figure 2). Microbial diversity is influenced not only by unhealthy conditions but also by diet and exercise, with diversity levels found to be decreased in overweight and obese patients. 130,412
Promising results are also emerging from metabolomics analyses in various biological matrices (plasma, feces, urines, or tissues). The metabolite profiles obtained from the microbiota, the host, or their co-metabolites have a high-resolution power that enables discrimination of healthy from diseased subjects.
414,415
IBD: A complete list of the altered metabolites can be found in the review by Smirnov et al.
414
The most significantly affected metabolites are certain amino acids (alanine, isoleucine, leucine, lysine, valine, and phenylalanine), SCFAs, and some acids (propanoic, butanoic, oleic, stearic, palmitic, linoleic, and arachidonic). However, most of these effects are a consequence of the inflammation status during IBD with malnutrition and decreased nutrient absorption. Obesity: Different studies have discovered altered levels for urine hippurate, 4-hydroxylphenylacetic acid and phenylacetylglycine, acetate, lactate, and branched chain amino acids.
415
NAFLD: Metabolites like 1-butanol, 2-butanone, and 4-methyl-2-pentanone have been found to be perturbed in people with NAFLD;
415
Neuropathology: Modified levels of free amino acids and SCFAs, tryptophan-nicotinic acid, sulfur metabolic pathways, and urinary metabolites have been recorded in autistic children.
415
Microbial metabolites should be adequately quantified using different methods in order to cover as much as possible of the metabolome and be further correlated against the gene expression levels and the corresponding enzymatic activity of each gene. Preferably, these metabolomics results should be combined and integrated with different “omics” techniques applied to DNA, RNA, and protein to provide a full microbiome analysis.
Despite the growing interest and exponential number of studies on this topic, many questions, including those that follow, require clarification: What is the precise composition of the different microbial communities living in our bodies? We still do not really know what a healthy microbiota composition looks like and how much variation exists between people. The microbiota are constantly under the influence of many factors, such as the delivery route during birth, genetics, metabolism, immunity (innate and adaptive), nutrition, and many different lifestyle factors. These factors have made it difficult to define the core healthy microbiome associated with a “normal” healthy individual. What are the causative and correlative effects between the microbiome and disease? It is typically unclear whether the changes observed in the microbiota are the cause or the effect of different diseases. Comparing health and disease situations without clearly demonstrating causality, or at least having some strong evidence of any link, is a major issue. In most studies, the overall assumption that the gut microbiota are causally linked with the onset or progression of a disease is often made based on a unique analysis of the composition of the fecal microbiota at a specific time point. Hence, being able to determine whether microbiome alterations precede or are caused by a particular disease would be essential. However, in terms of what came first, a disease or a change in the microbiome, there probably will not be a single answer, but it would be very informative to be able to characterize the disease-associated changes in the microbiome before and after the disease diagnosis. Thus, variations in scientific results may be reduced by performing long-term longitudinal studies monitoring the disease progression and by characterizing the changes in the taxonomic and functional composition of the microbiome, thereby defining the disease state. The further investigations that follow would need to confirm that in vivo replacement of the missing microorganisms could at least mitigate and/or cure the disease, which is the ultimate objective to reach. What are the functional pathways by which the gut, lungs, brain, and skin microbial populations can communicate inside the human body? The ability to modify the function of one organ by manipulation of the other now depends on concerted interdisciplinary efforts focused on better understanding of the targetable pathways by which the gut, skin, lungs, oral cavity, and brain communicate with each other.
Evidently, an enormously complex interplay exists among our different bodily organs and the different microbial communities we harbor. We are just at the beginning of our understanding of the microbiome and interdisciplinary effort will be essential for us to answer some of the open questions about the role of the microbiome in health and disease.
Footnotes
Declaration of conflicting interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: All authors are employees of, or contracted and paid by, Philip Morris International.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Philip Morris International is the sole source of funding and sponsor of this article.
Supplemental material
Supplementary material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
