Abstract
Highlights
Accessibility, cost, and practical considerations, such as ease of use and clinical utility, were prioritized slightly more than technical validation metrics, such as discrimination and calibration, when deciding to start using a clinical prediction model.
Most breast cancer clinicians valued models with clear inputs (e.g., variable definitions, cutoffs) and outputs; few were interested in the exact model specifications.
Perceptual or subjective factors, including perceived accuracy and peer acceptability, also influenced model adoption but were secondary to practical considerations.
Sociodemographic variables, such as clinical specialization and hospital setting, influenced the importance of factors for model use.
This is a visual representation of the abstract.
Keywords
Breast cancer remains a leading cause of mortality among women worldwide. 1 Adjuvant systemic treatments (e.g., chemotherapy, endocrine therapy) administered after tumor resection can reduce the risk of distant metastases, recurrence, and breast cancer–related mortality, although they may also lead to side effects, including fatigue, nausea, infertility, nerve damage, and osteoporosis, with rare cases of secondary cancers. 2 Clinical prediction models, such as PREDICT, provide individualized survival (or risk) estimates that can aid clinicians in evaluating the added benefits of these treatments.3,4 The use of PREDICT and similar prognostic models can thus improve the identification of women most likely to benefit from additional interventions. 2 Beyond prognostication, models such as BOADICEA can assess an individual’s predisposition to breast cancer and inform preventive interventions and screening strategies. 5 By enabling personalized risk assessments and the application of more tailored treatment strategies, clinical prediction models may help reduce overtreatment, limit low-value care, and improve quality of life.1,2
Despite the proliferation of clinical prediction models over the past two decades, a gap persists between model development and real-world implementation.3,6–8 For instance, there are now more than 60 breast cancer prognostication models, 8 of which only a select few have been widely adopted in clinical practice. 9 Notably, high-performing machine learning–based models are not always extensively used; others lacking rigorous evaluation are sometimes implemented.10,11 This variation in model uptake highlights the complexities of translating models from research into routine clinical practice. Understanding the barriers and facilitators to initial model adoption is essential to informing the design of models that can integrate more effectively into clinical practice and meet the needs of clinicians and patients.
Currently, most research on implementation barriers focuses on patient decision aids, which do not necessarily integrate data-driven algorithms.12–17 In addition, literature on the requirements for implementing clinical prediction models often emphasizes performance metrics, such as discrimination and calibration, while overlooking broader factors that may influence their clinical use. Although some studies have assessed the impact of (widely) used breast cancer prediction models post implementation,18,19 the factors driving their initial clinical adoption remain largely unexplored. In specialized fields such as breast cancer, where integration of validated prediction models can guide complex treatment decisions and improve patient outcomes, understanding what drives clinicians to adopt these tools is essential.
We aimed to 1) identify the key factors influencing breast cancer clinicians’ decisions to initiate the use of clinical prediction models and 2) assess the relative importance of these factors. As a secondary exploratory aim, we examined whether the importance of these factors varied according to sociodemographic characteristics, including age, sex, clinical specialization, years of experience, and healthcare setting.
Methods
Study Design and Ethics Approval
We employed an exploratory sequential mixed-methods design (Figure 1). The study was approved by the Institutional Review Board at the Netherlands Cancer Institute (IRBd22-078). Verbal informed consent was obtained prior to the interviews, while survey participants provided informed consent on the first page of the questionnaire. No personal identifiers were retained during data collection or analysis.

Flow diagram of the sequential exploratory mixed-methods approach applied in the study.
Phase 1: Qualitative Semi-structured Interviews
Participants
We conducted semi-structured interviews to identify key factors influencing clinicians’ decisions to start using clinical prediction models. Using purposive sampling, we recruited medical oncologists, radiation oncologists, surgical oncologists, clinical geneticists, and nurse specialists via the study team’s breast cancer clinician network. All participants were familiar with, but not necessarily using, clinical prediction models. The interview guide (Supplementary File A) was pilot tested with three clinicians prior to finalization.
Data collection
A trained epidemiologist with qualitative research experience (M.A.B.) conducted the interviews between March and November 2022, either in person, online, or by telephone, whichever was most convenient for the participant. The interviews had an average duration of 30 minutes. Data collection continued until we reached theoretical saturation (i.e., no new themes were emerging) and had sufficient diversity in clinical specialization, geographic location, and hospital type. All interviews were conducted in English, transcribed verbatim, and de-identified after transcription. Clinicians did not receive compensation for their participation.
Data analysis
We employed inductive and deductive thematic analysis to summarize the interview transcripts. Two researchers (M.A.B. and E.G.E.) independently double-coded all transcripts using ATLAS.ti versions 9 and, later, 23. 20 The first five interviews served as a training set for developing the initial codebook. Regular meetings were held to review the codes and resolve any discrepancies, with a third researcher (M.K.S.) providing input when needed. As coding progressed, initial codes were consolidated into broader themes representing key factors. Less prevalent themes were retained as subfactors or omitted based on relevance and data support. New themes were added to the codebook as they emerged. Previously coded interviews were reviewed to ensure that the new codes were captured, with recoding performed when necessary. Quotes from the interviews were used to illustrate the themes.
Phase 2: Quantitative Online Survey
Questionnaire development
An online survey was administered to a nationwide network of clinicians to assess the relative importance of factors identified in phase 1. The phase 1 codebook informed the design of the questionnaire. Clinicians were asked to provide sociodemographic information (age, biological sex, years of experience, clinical specialization, hospital type, and region) and rate the importance of each subfactor on a numeric scale from 0 (not important) to 10 (very important) (see Supplementary File B for the full questionnaire). To keep the survey concise, the selection of sociodemographic variables was limited to only key characteristics that we expected might influence initial model uptake. Clinicians who reported never using prediction models were asked about their reasons for non-use. The survey was pilot tested by 2 team members (M.K.S. and A.H.B.) and an experienced medical oncologist (S.C.L.). The anonymous survey was carried out using Castor EDC. 21
Participants and data collection
The survey was conducted from May to October 2023 and recruited clinicians in the Netherlands involved in breast cancer care who self-identified as familiar with prediction models. Invitations, which included details about the study and a link to the survey, were sent via e-mail to the study team’s clinician network, the Netherlands Cancer Institute’s breast cancer working group, the Dutch Society for Radiation Oncology, the Dutch Society for Surgical Oncology, the Hereditary Breast and Ovarian Cancer Research Netherlands, and the Netherlands Comprehensive Cancer Organization.
Data analysis
Surveys were included in the analysis if they were at least 50% complete (i.e., the respondents had rated at least 1 cluster of factors). Descriptive statistics characterized the overall sample and subgroups of model users and non-users. Due to the imbalance between the groups, with only 9 non-users (6%) compared with 137 users (94%), regression analysis was unsuitable, as the limited number of non-users fell below the recommended events per variable threshold for reliable estimation. Although weighting adjustments were considered to address the imbalance, the limited data for non-users made this approach impractical and potentially biased. Consequently, subsequent analyses focused on the more robust group of model users. Nonparametric tests were used to compare score distributions across sociodemographic subgroups: the Mann–Whitney U test for pairwise comparisons between 2 groups and the Kruskal–Wallis test for comparisons across multiple groups. A 10-year cutoff was applied for years of experience, as this is a common threshold in clinical practice that distinguishes between early- and mid-career clinicians, potentially reflecting differences in familiarity with and use of clinical prediction models. A 50-year cutoff was used for age based on established epidemiological significance. To account for multiple testing and reduce the risk of type I errors, a strict significance threshold (P ≤ 0.002, using Bonferroni correction) was used. Data distributions and subgroup comparisons were visualized using density plots and scatterplots. We performed all quantitative analyses using RStudio version 2023.06.0. 22
Results
Phase 1: Qualitative Semi-structured Interviews
We interviewed 16 breast cancer clinicians: 6 medical oncologists, 3 radiation oncologists, 2 clinical geneticists, 2 surgical oncologists, 2 nurse specialists, and 1 radiologist. Ten clinicians worked primarily in academic medical centers, while others were from general teaching hospitals (n = 3) and general hospitals (n = 3). Most participants were from Western Netherlands (n = 13), 2 were from the South, and 1 was from the North. Most (n = 13) were female. The median age was 48 years (range, 31–62 years), and the median number of years of experience in breast cancer care, excluding training, was 14 years (range, 4–35 years).
Key Factors for Model Use
Eight factors (Figure 2) primarily influenced breast cancer clinicians’ decisions to initiate the use of clinical prediction models. Practical/methodological factors included accessibility, cost, understandability, objective accuracy, actionability, and clinical relevance. Perceptual (personal or social) factors included subjective accuracy, acceptability, and risk communication. Below is a general summary of these factors. Illustrative quotes are available in Table 1.

Factors influencing clinicians’ decisions to use clinical prediction models.
Illustrative Quotes from the Interviews

Phase 2: Quantitative Online Survey
We included data from 146 clinicians in the analysis. Of these, 137 (94%) reported using clinical prediction models at least occasionally (Supplementary File C), while 9 (6%) indicated they never used such models. Table 2 summarizes the demographic and professional characteristics overall and by subgroup.
Characteristics of the Survey Participants

Model use was defined as “yes” if clinicians reported using clinical prediction models occasionally, regularly, or always and “no” if they reported never using them.
Model Users
Among the 137 clinicians who reported using models, 7 (5%) seldom, 39 (28%) sometimes, 69 (50%) regularly, and 22 (16%) always used models. The median age was 49 years (range, 28–64 years), and most (n = 106, 77%) were women. Most (n = 100, 73%) had more than 10 years of experience in breast cancer care. The sample comprised medical oncologists (n = 49, 36%), surgical oncologists (n = 24, 18%), radiation oncologists (n = 32, 23%), nurse specialists (n = 26, 19%), clinical geneticists (n = 4, 3%) and radiologists (n = 2, 1%). Clinicians worked across general hospitals (n = 31, 23%), general teaching hospitals (n = 49, 36%), academic medical centers (n = 47, 34%), and radiotherapy centers (n = 10, 7%). Table 3 lists the models with which the respondents were familiar. The PREDICT model was the most recognized (n = 127, 93%), followed by the Memorial Sloan Kettering Cancer Center lymph node metastases nomograms (n = 55, 40%) and the INFLUENCE model (n = 53, 39%).
Breast Cancer Prediction Model Familiarity among Survey Participants
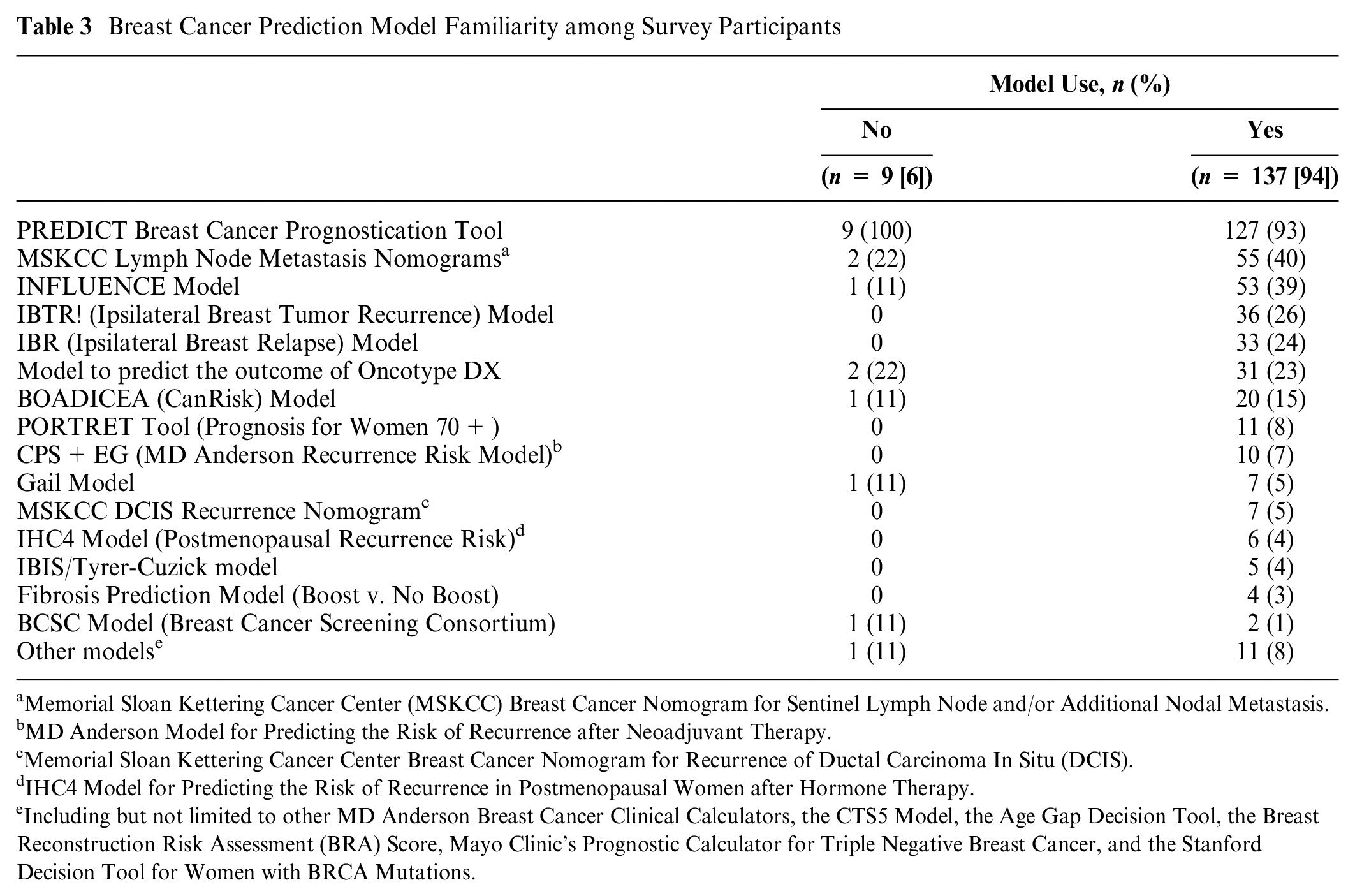
Memorial Sloan Kettering Cancer Center (MSKCC) Breast Cancer Nomogram for Sentinel Lymph Node and/or Additional Nodal Metastasis.
MD Anderson Model for Predicting the Risk of Recurrence after Neoadjuvant Therapy.
Memorial Sloan Kettering Cancer Center Breast Cancer Nomogram for Recurrence of Ductal Carcinoma In Situ (DCIS).
IHC4 Model for Predicting the Risk of Recurrence in Postmenopausal Women after Hormone Therapy.
Including but not limited to other MD Anderson Breast Cancer Clinical Calculators, the CTS5 Model, the Age Gap Decision Tool, the Breast Reconstruction Risk Assessment (BRA) Score, Mayo Clinic’s Prognostic Calculator for Triple Negative Breast Cancer, and the Stanford Decision Tool for Women with BRCA Mutations.
Relative Importance of the Different Factors for Model Use
Figure 3 shows the density distributions of scores (see Supplementary File D for descriptive statistics). The highest-rated factors were accessibility (availability as online tool: median = 9 [interquartile range {IQR} = 8–10]) and cost (free to use: 9 [8–10]; tests included reimbursable: 8 [8–10]). Other factors with strong ratings included actionability (good clinical utility: 9 [8–10]) and understandability (clarity in intended patients: 9 [8–9]; clearly defined predictors: 8 [8–9]; user-friendly interface: 8 [8–9]; clear output for clinicians: 8 [8–9]; clear output for patients: 8 [7–9]). Broad methodological components of accuracy were highly rated (reliable development data: 9 [8–10]; large development sample: 9 [8–9]; external validation: 8 [7–9]), along with performance metrics (good discrimination: 8 [7–8]; good calibration: 8 [7–8], good sensitivity: 8 [8–8]; good specificity: 8 [8–9]). Subjective accuracy (model estimates align with gut feeling: 7 [5–8]) received slightly lower scores among accuracy-related factors. Acceptability (used by colleagues: 8 [8–9]; included in clinical guidelines: 8 [7–8]) and risk communication (aids in information provision: 8 [8–9]; aids in risk communication: 8 [8–9]) were also highly rated. The lowest-rated factors were integration into EHRs (6 [3–8]), inclusion of new biomarkers (6 [5–8])—specifically Oncotype DX (6 [4–8]) and MammaPrint (6 [4–7])—and mobile application accessibility (5 [2–7]).
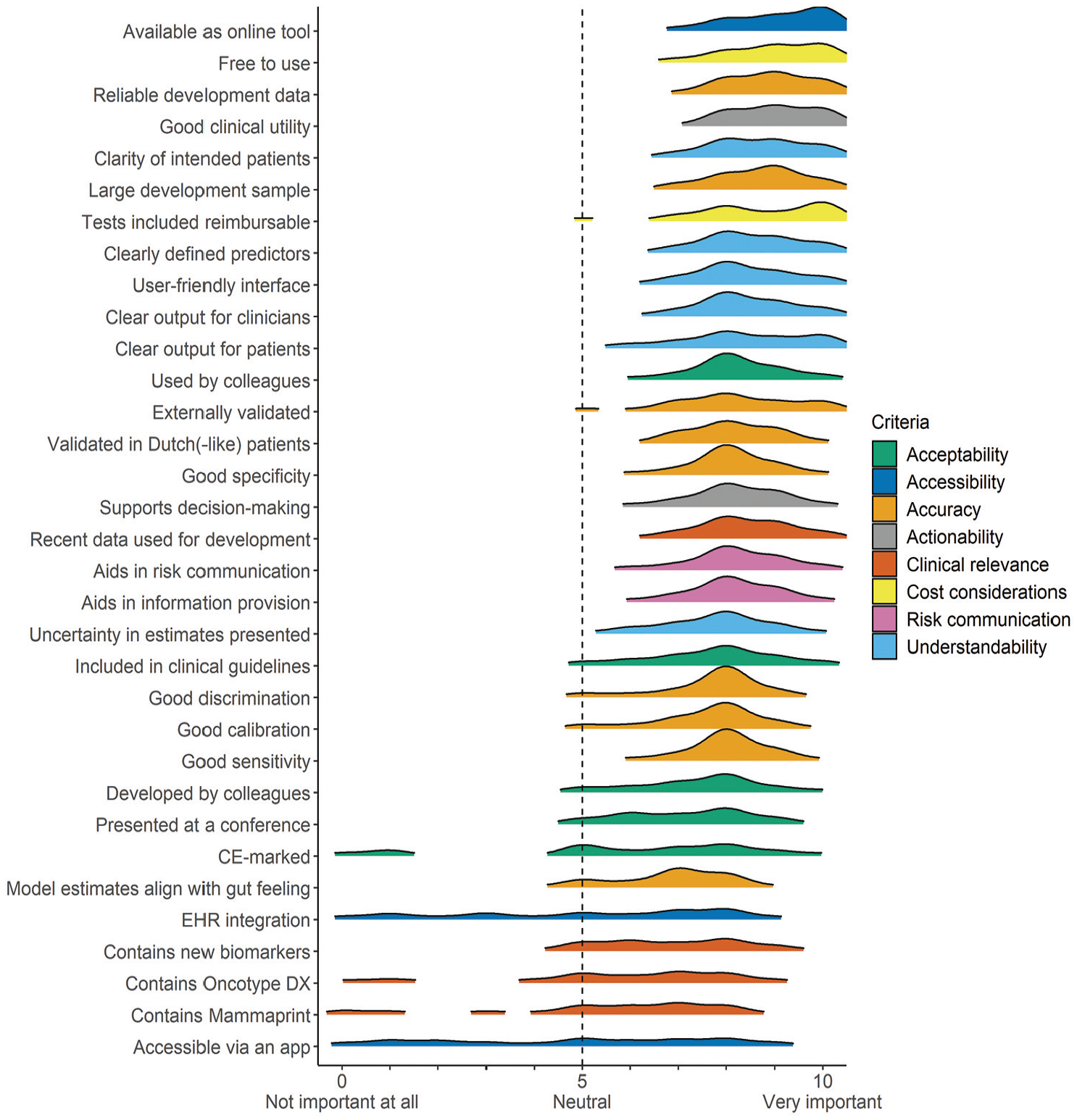
Density plots of the importance scores assigned by breast cancer clinicians who used clinical prediction models (n = 137).
Assessment of Importance Scores across Sociodemographic Characteristics of Model Users
Table 4 summarizes significant differences by sociodemographic characteristics (complete test results in Supplementary Files E–J). Male clinicians rated clear output for clinicians and user-friendly interface as slightly more important compared with female clinicians. Clinicians with more than 10 years of experience placed higher importance on models with clearly defined predictors and app accessibility. Differences were also observed across clinical specializations, such as regarding the importance of CE marking, reimbursement for included tests, and clarity of patient-facing outputs. In addition, clinicians from general hospitals rated test reimbursement and inclusion of new biomarkers higher than those from academic medical centers or radiotherapy centers.
Significant Subgroup Differences in Importance Scores
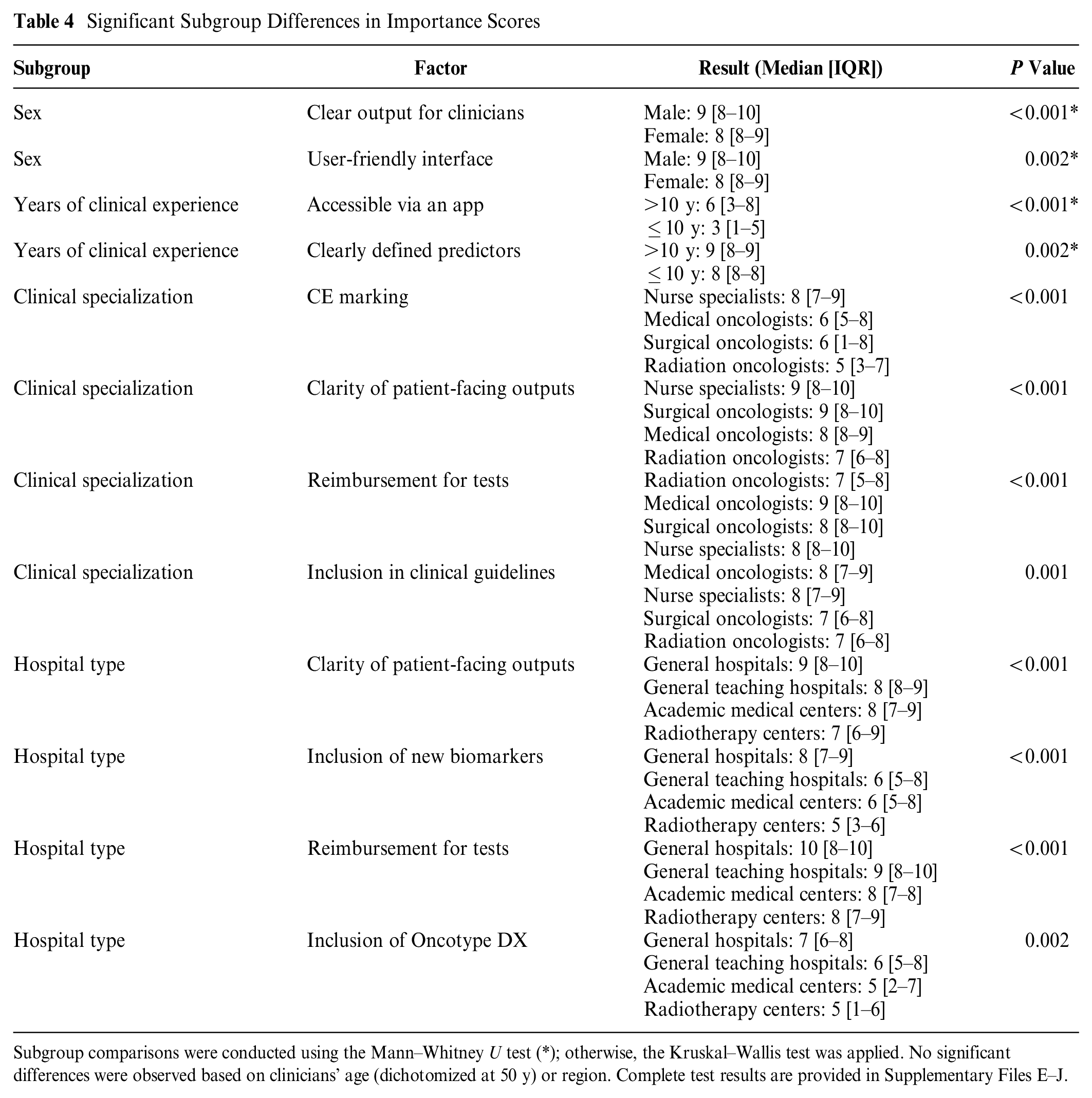
Subgroup comparisons were conducted using the Mann–Whitney U test (*); otherwise, the Kruskal–Wallis test was applied. No significant differences were observed based on clinicians’ age (dichotomized at 50 y) or region. Complete test results are provided in Supplementary Files E–J.
Non-users
Among the 9 who did not use clinical prediction models, 6 (67%) were affiliated with general hospitals. PREDICT was the most recognized model (Table 3). Radiologists found prediction models less relevant to their work, with some considering them redundant given the comprehensive clinical guidelines they follow. One medical oncologist cited frequent discrepancies between patients’ interpretations and their own, which they felt made models inefficient for routine use.
Other Factors for Model Use
Ninety-five (65%) participants reported no additional criteria beyond those covered in the survey. The remaining participants mentioned factors consistent with those included in the survey. No new factors were thus identified.
Discussion
We employed a mixed-methods approach to identify factors influencing breast cancer clinicians’ decisions to initiate use of clinical prediction models and to assess the relative importance of these factors. Our study identified eight key factors, broadly grouped into practical/methodological (accessibility, cost, understandability, objective accuracy, actionability, clinical relevance) and perceptual factors (subjective accuracy, risk communication, acceptability). Sociodemographic characteristics showed potential influence on the perceived importance of these factors.
Our findings indicate that practical factors, particularly accessibility and cost, were the most influential when deciding to use clinical prediction models. Freely accessible online models were strongly preferred, highlighting the importance of digital availability. Nonreimbursable tests were identified as a barrier to model use. While clinicians valued incorporating new markers, such as gene signatures, 23 they emphasized the need to balance their use with out-of-pocket costs, particularly when the added predictive benefit is uncertain.24–26 Notably, accessibility and cost were rated slightly higher than validation metrics, such as discrimination and calibration. 27 The stronger emphasis on practical considerations over technical metrics may stem from clinicians viewing model validation mainly as the responsibility of other stakeholders (e.g., statisticians and leading medical experts), while their primary concern is the effective integration of already validated tools and assessing their utility within clinical workflows. 28 Based on these findings, targeted measures could increase model adoption, such as ensuring health insurance coverage for expensive tests.29–31 In addition, incentivizing developers or funding infrastructure to provide free online access to models could further facilitate their routine use in breast cancer care.12,32
Another key criterion prioritized by clinicians was understandability. Our findings indicate that most clinicians valued model explainability—specifically, understanding model inputs (e.g., included variables/tests) and outputs (e.g., results presentation)—more than algorithmic transparency.33–37 While algorithmic transparency is often promoted to reduce biases and improve health equity, 38 our results suggest that only a few clinicians prioritized knowledge of the specific algorithms underlying models. Those who did were typically academically inclined, recognizing that a global model may perform differently in diverse local settings. 38 For most clinicians, understanding the predictors, definitions, cutoffs, and derivation population was sufficient. 39 To increase model adoption, developers should focus on creating intuitive, user-friendly online tools based on their models. 40 Detailed information on model specifications or algorithm may be added as supplementary material for those interested. 41
Trustworthiness, as highlighted in previous research,37,42,43 is closely linked to model adoption. In our study, we found that trustworthiness was not a standalone factor but was embedded within others, such as understandability (e.g., trust from knowing a model’s inputs and understanding its outputs), accuracy (e.g., trust from evidence of validity and alignment with clinical intuition), and peer acceptability (e.g., confidence from endorsements by trusted colleagues). To foster model trust, several strategies could be employed, including conducting validation studies to assess performance across diverse settings, presenting models at relevant clinical conferences, obtaining endorsements from medical societies, and developing (peer-led) training programs to improve model familiarity. Incorporating clinician feedback to adapt models locally for specific patients can also further build trust and address their specific needs. 44 Such a solution might be more straightforward and cost-effective than continuously deriving new models from scratch.45,46
In the European Union, prediction models intended to support clinical decision making, such as those used to directly guide diagnosis, prognosis, or treatment selection, are classified as medical devices under the Medical Device Regulation or the In Vitro Diagnostic Regulation if they rely on data from in vitro tests.47,48 This classification requires the software or tool based on the model to receive CE certification, which may involve a rigorous, lengthy, and costly process to ensure compliance with harmonized standards (e.g., ISO 13485). 49 Few breast cancer prediction models have managed to obtain this certification. 11 Our findings suggest that most breast cancer clinicians assigned relatively low importance to CE marking. This may be because many already rely on published evidence from validation studies, making CE marking an added formality rather than a crucial indicator of validity or clinical utility.
Our analysis revealed differences in factor prioritization across clinician subgroups. Nurse specialists assigned greater importance to regulatory assurances, such as CE marking, potentially reflecting their emphasis on adherence to protocols and standardized care. In contrast, medical, surgical, and radiation oncologists, who may have greater familiarity with model validation literature, perceived CE marking as less critical. Clinicians in general hospitals prioritized the inclusion of new markers in models more than those in academic settings did. This difference could potentially be attributed to the fact that the clinical utility of many new markers (e.g., gene signatures, tumor-infiltrating lymphocytes) was still under evaluation at the time of data collection. 31 Clinicians in academic or research settings may have had greater involvement in or access to these evaluations, which could have made them more cautious about incorporating new markers. Understanding the distinct needs of specific clinician subgroups could help further optimize the presentation and features of prediction models.
A key strength of our study is that the identified factors were derived directly from clinicians, ensuring the findings represent the views of end-users of clinical prediction models. In addition, our large sample of model users adds robustness to our main findings. There are also some limitations to consider. The reliance on clinician networks for recruitment may have introduced some selection bias. Next, despite a broad recruitment strategy, certain specializations (e.g., clinical geneticists and radiologists) were underrepresented, which may limit the generalizability to these subgroups. Furthermore, the small number of non-users might have restricted our ability to fully understand the barriers for model use. Self-reported data may have introduced social desirability bias, although we attempted to minimize this by ensuring anonymity and neutral phrasing in survey questions. Finally, to keep the survey brief, only specific sociodemographic variables were examined, which may have limited the depth of our subgroup analysis findings. Future studies should aim to confirm our findings with larger and more diverse samples. It is also important to explore whether similar factors are relevant in other medical domains and countries. Evaluating the effectiveness of different implementation strategies could further inform best practices for model implementation.
Conclusion
Clinical prediction models have the potential to support personalized, evidence-based decision making in breast cancer care. To our knowledge, our study is the first to investigate the broad range of factors influencing breast cancer clinicians’ decisions to start using prediction models. Our findings suggest that practical factors, such as ease of access and cost associated with model use, outweigh methodological rigor and technical metrics in determining model use. To bridge the gap between model development and clinical use, improved collaboration among model developers, clinicians, and communication and implementation experts is crucial. Clinician-friendly design that prioritizes usability, utility, and adaptability to local settings could increase model uptake. In addition, advocating for health policy initiatives, such as the reimbursement of expensive tests, providing financial support for translating promising models online, and funding trainings or conference presentations, can help increase familiarity with and confidence in using prediction models, ultimately improving their clinical impact. More research is needed to identify barriers encountered by non-users and to investigate other potential sociodemographic variations, which could allow for a more tailored approach to the implementation of clinical prediction models.
Supplemental Material
sj-docx-1-mpp-10.1177_23814683251328377 – Supplemental material for Bridging the Gap: A Mixed-Methods Study on Factors Influencing Breast Cancer Clinicians’ Decisions to Use Clinical Prediction Models
Supplemental material, sj-docx-1-mpp-10.1177_23814683251328377 for Bridging the Gap: A Mixed-Methods Study on Factors Influencing Breast Cancer Clinicians’ Decisions to Use Clinical Prediction Models by Mary Ann E. Binuya, Sabine C. Linn, Annelies H. Boekhout, Marjanka K. Schmidt and Ellen G. Engelhardt in MDM Policy & Practice
Footnotes
Acknowledgements
The authors would like to thank all of the clinicians who participated in the interviews and online survey as well as the clinicians who helped pilot test the interview guide and questionnaire. We also would like to thank Prof. Dr. Eveline Bleiker for providing feedback on the draft article.
Authors Contributions
EGE, MAB, MKS, and AHB conceived the study and wrote the protocol. All authors revised and approved the protocol. MAB and EGE collected and analyzed the data. All authors helped in interpretation of the results. MAB wrote the first draft of the manuscript. EGE, MKS, AHB, and SCL revised the manuscript. All authors approved the final version for publication.
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: SCL reports grants paid to the institute from AstraZeneca, Eurocept Plaza, Roche, Genentech, Gilead Sciences, GSK, Novartis, and Agendia outside the submitted work; consulting fees from AstraZeneca paid to the institute; educational fees from Daiichi Sankyo paid to the institute; other financial support for attending meetings from Daiichi Sankyo; nonfinancial support from Genentech (drug), Roche (drug), Gilead Sciences (drug), Novartis (drug), Agendia (gene expression tests), and AstraZeneca (drug). SCL has a patent (PCT/EP2022/73958) pending on a method for assessing homologous recombination deficiency in ovarian cancer cells. The other authors declare no conflicts of interest. The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study is part of the PREDICT-NL project (pathways to smart validation and clinical embedding of prediction tools for oncology and beyond) funded by Health~Holland grant No. LSHM19121, received by MKS, EGE, and MAB. Research at the Netherlands Cancer Institute is supported by institutional grants of the Dutch Cancer Society; the Dutch Ministry of Health, Welfare and Sport; and individual donors.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
