Abstract
Reproducibility is fundamental to science, and an important component of reproducibility is computational reproducibility: the ability of a researcher to recreate the results of a published study using the original author’s raw data and code. Although most people agree that computational reproducibility is important, it is still difficult to achieve in practice. In this article, the authors describe their approach to enabling computational reproducibility for the 12 articles in this special issue of Socius about the Fragile Families Challenge. The approach draws on two tools commonly used by professional software engineers but not widely used by academic researchers: software containers (e.g., Docker) and cloud computing (e.g., Amazon Web Services). These tools made it possible to standardize the computing environment around each submission, which will ease computational reproducibility both today and in the future. Drawing on their successes and struggles, the authors conclude with recommendations to researchers and journals.
Keywords
There is broad agreement that reproducibility is generally important for establishing a scientific claim. In this article, we focus on a very narrow form of reproducibility: a researcher’s ability to recreate the results in a published article using the raw data and code of the original author. Following Stodden (2015), we call this “computational reproducibility,” although this same concept has also been called “verifiability” (Freese and Peterson 2017) and “repeatability” (Collberg and Proebsting 2016; Easterbrook 2014). 1
Although computational reproducibility is widely considered to be a desirable goal, it is a standard that a lot of scientific research does not currently meet. There are two main factors that contribute to this shortcoming. First, the data and code used to create results in published articles are often not available to other researchers, despite requirements by some journals, professional organizations, and funding agencies. Second, even if the data and code are available, independent researchers are often not able to reproduce the results. These two patterns have been empirically established in the social sciences (Chang and Li 2018; Dewald, Thursby, and Anderson 1986; Gertler, Galiani, and Romero 2018; Hardwicke et al. 2018; In’nami and Koizumi 2010; McCullough, McGeary, and Harrison 2006; Stockemer, Koehler, and Lenz 2018; Vanpaemel et al. 2015; Wicherts, Bakker, and Molenaar 2006; Wicherts et al. 2011; Wicherts and Crompvoets 2017; Wood, Müller, and Brown 2018), as well as other scientific fields (Alsheikh-Ali et al. 2011; Andrew et al. 2015; Campbell, Micheli-Campbell, and Udyawer 2019; Collberg and Proebsting 2016; Gilbert et al. 2012; Hardwicke and Ioannidis 2018; Ioannidis et al. 2009; Konkol, Kray, and Pfeiffer 2019; Naudet et al. 2018; Ostermann and Granell 2017; Rowhani-Farid and Barnett 2016; Savage and Vickers 2009; Stodden, Seiler, and Ma 2018; Vandewalle, Kovacevic, and Vetterli 2009; Vines et al. 2014). Although the magnitude of the problem differs by field and across time, the limited availability of replication materials and the limited usefulness of the materials that are available are consistent patterns. In other words, a lack of computational reproducibility is itself a reproducible finding.
There are many admirable attempts to improve the computational reproducibility. One of these efforts is the Transparency and Openness Promotion (TOP) Guidelines (Nosek et al. 2015). The TOP Guidelines provide a framework journals can use to assess and improve their efforts to make scientific communication more open. Despite being introduced quite recently, more than 1,000 journals have already adopted the TOP Guidelines in some form (Center for Open Science n.d.).
The TOP Guidelines consist of eight standards, including one called “Data, Analytic Methods (Code), and Research Materials Transparency.” When focused on computational reproducibility, this standard has three levels:
Level 1 (“available upon request”): replication materials available upon request from authors
Level 2 (“access before publication”): replication materials submitted to a trusted archive before publication
Level 3 (“verification before publication”): replication materials verified by journal and then submitted to a trusted archive before publication
For this special issue, we attempted to meet the TOP Level 3 standard (“verification before publication”). Furthermore, in addition to complying with the standard as written,
2
we attempted to give replication materials a more central role in the process of scholarly communication. Our thinking was heavily influenced by what could be called Donoho’s dictum
3
: An article about computational science in a scientific publication is not the scholarship itself, it is merely advertising of the scholarship. The actual scholarship is the complete software development environment and the complete set of instructions which generated the figures. (Buckheit and Donoho 1995)
As we attempted to give replication materials a more central role in the process of scholarly communication, we made three interrelated changes to standard practice. First, inspired by insights from other computational fields, we took a broader view of what should be included in replication materials. In particular, rather than focusing on just data and code, we expanded our focus to include the computing environment (Stodden et al. 2016; Tatman, VanderPlas, and Dane 2018). Second, rather than merely verifying the results in a manuscript, we focused our efforts toward making the replication materials as useful as possible to future scholars. Finally, rather than assessing computational reproducibility after completing peer review of a manuscript, we made computational reproducibility a part of the review process.
We found that computational reproducibility was difficult to achieve in practice. Only 2 of the 14 manuscripts that were submitted to the special issue were computationally reproducible initially, even though we had access to code provided by the authors. After an intensive revision process, 7 of the 12 accepted manuscripts eventually became computationally reproducible. These aggregate results mask important heterogeneity. Some submissions to the special issue used computational approaches that are relatively common in sociology and the other social sciences today, for example, Ahearn and Brand (2019) and McKay (2019). For these submissions, computational reproducibility was relatively easy. Some other submissions, however, drew on more computationally complex approaches from machine learning, for example, Rigobon et al. (2019) and Davidson (2019). These submissions were substantially more difficult to reproduce.
We think this heterogeneity has two important lessons. First, our results suggest that relatively modest efforts could yield large increases in computational reproducibility for the methods that are typically used today. The second important lesson is about the future. We expect that sociologists and other social scientists will increasingly use larger data sets and more computationally intensive methods, such as the methods used in this special issue. Our experience suggests that as research becomes more computationally intensive, the problems with computational reproducibility will likely get worse unless researchers make changes to the process of scientific publishing.
The remainder of this article is organized as follows. In the next section we provide more background on the Fragile Families Challenge and computational reproducibility. Then we describe our process for ensuring computational reproducibility for this special issue. Next we offer recommendations for authors wanting to make their work more computationally reproducible, and we offer recommendations for journals wanting to promote computational reproducibility. We also provide supporting online materials that include a summary of the computational reproducibility status of each article in the special issue; the reproducibility memo that we sent to authors during the peer-review process; and instructions for using the Docker images we created to accompany the articles in this special issue.
Background
Fragile Families Challenge
The Fragile Families Challenge is a scientific mass collaboration designed to yield insights that can improve the lives of disadvantaged children in the United States. The Challenge is described in more detail in Salganik et al. (2019), but we describe it here briefly with a focus on issues that are particularly relevant to computational reproducibility. The Fragile Families Challenge builds on the Fragile Families and Child Wellbeing Study, which collects data about a probability sample of births from 1998 to 2000 in large U.S. cities, with an oversample of nonmarital births (Reichman et al. 2001). During the course of the Fragile Families and Child Wellbeing Study, information has been collected from mothers, fathers, and children, as well as other people in the children’s lives (e.g., teachers). The Challenge set out the goal of using data collected during the child’s first 9 years to predict six outcomes when the child was 15 years old (Figure 1). 4

Fragile Families Challenge data structure. Participants built models predicting the age 15 outcomes using data collected between the time the focal child was born and 9 years old. We provided participants with the data represented by the white boxes. Submissions were scored on the basis of their predictive performance (mean squared error) for the observations represented by the gray boxes, which were available only to organizers. The leaderboard set contained one eighth of all observations and was used to provide instant feedback on submissions. The holdout set contained three eighths of observations and was used to produce final scores for all submitted models at the end of the Challenge.
To this end, the Challenge used a research design from machine learning called the common task method (Donoho 2017). The common task method involves three main elements: a common target for predictive modeling, a common error metric (in our case, mean squared error), and a common data set with both training data available to participants and held-out data used for evaluation. Following this structure, we split the year 15 data for these six outcomes into three groups: (1) a training set that we provided to participants, (2) a leaderboard set participants could access during the Challenge, and (3) a holdout set participants could not access until the first stage of the Challenge was complete (Hardt and Blum 2015) (Figure 1). Participants in the Challenge received the training set and a specially constructed background data file that contained information about the family from birth to age 9; it included 4,242 families and 12,942 variables about each family. 5
Computational Reproducibility
The benefits of computational reproducibility—and increased access to data and code, more generally—have already been articulated many times by researchers in many different fields: archaeology (Marwick 2017), bioinformatics (Mangul et al. 2018), cell biology (Grüning et al. 2018), computational fluid mechanics (Mesnard and Barba 2017), computer systems research (Collberg and Proebsting 2016), economics (Anderson et al. 2008; Koenker and Zeileis 2009; Orozco et al. 2018), epidemiology (Coughlin 2017; Peng, Dominici, and Zeger 2006; Shepherd et al. 2017), geosciences (Claerbout and Karrenbach 1992; Gil et al. 2016; Konkol et al. 2019), high-energy physics (Chen et al. 2018), hydrology (Hutton et al. 2016), mathematical and computational biology (Schnell 2018), machine learning (Hutson 2018; Tatman et al. 2018), neuroscience (Crook, Davison, and Plesser 2013; Eglen et al. 2017; Manninen et al. 2017; Mikowski, Hensel, and Hohol 2018), political science (Alvarez, Key, and Nez 2018; King 1995; Lupia and Elman 2014), psychology (Clyburne-Sherin and Green 2018), signal processing (Vandewalle et al. 2009), sociology (Freese 2007), and statistics (Donoho 2010; Gentleman and Lang 2007; Stodden 2015). We find these arguments convincing. So, for the purposes of this article, we simply assume that computational reproducibility is a desirable goal. 6 We acknowledge, however, that this is a relatively low standard; it does not guarantee correctness of the code or the scientific validity of the claims (Leek and Peng 2015). 7
There are many wonderful efforts under way to increase computational reproducibility, and our approach shares many features with these other efforts. However, there are three things that partially differentiate our approach from other approaches in the social sciences. First, we were not motivated by the “reproducibility crisis” or issues related to “questionable scientific practices.” The structure of the Challenge—with explicit holdout data and a clear prespecified evaluation metric—makes concerns about questionable scientific practices less relevant. Rather, our approach was focused on increasing research impact through reuse. 8
The second way our approach differs from some other approaches in the social sciences is that it focuses on the computing environment, in addition to data and code. Currently, a significant subset of published social science research is based on relatively small data sets using methods that can be deployed in a fairly basic computing environment. In other words, it is not uncommon for many social scientists to do their computing on a laptop computer. However, some social scientists are beginning to use much larger data sources and more resource-intensive computational methods from machine learning. These are potentially exciting developments, but as computational complexity increases, it becomes increasingly difficult to reproduce results using data and code running in a different computing environment (Boettiger 2015; Boisvert 2016; Tatman et al. 2018). Instead, for computationally complex work, reproducibility often requires access to the same computing environment, both hardware and software. Fortunately, software engineers have already developed tools and techniques that make it possible to standardize a computing environment, and for our Challenge reproducibility work, we borrowed liberally from these practices. In particular, for our computational reproducibility work in the Fragile Families Challenge we used software containers (e.g., Docker) to standardize the software environment and cloud computing (e.g., Amazon Web Services) to standardize the hardware environment.
Software containers and cloud computing are not yet common for computational reproducibility in the social sciences, so we will briefly review them. 9 Software containers, such as Docker, are designed to ensure that software that runs on one computer can run in exactly the same way on another computer. Containers ensure portability through encapsulation. Roughly, software containers create an appropriately configured minicomputing environment within a larger computing environment and then make it easy to move this minicomputing environment from one computer to another, independent of the operating systems. This minicomputing environment can include all the supporting software the research code depends on, and it can keep these dependencies separate from all the other code that runs on the host computer. For example, the results in Stanescu et al. (2019) were created with code that requires version 0.7.4 of the R package dplyr. Without a software container, anyone wishing to reproduce the results in the article would need to install version 0.7.4 of dplyr, and all the packages on which it depends, on his or her computer. This installation process is easy with just one relatively new, high-quality package. But when code requires many packages, particularly old and complex packages, installing all the necessary dependencies can be difficult, frustrating, and error prone. These difficulties become even more extreme if the researcher attempting to reproduce these results is also working on another project that requires a different version of the same package. The ugly process of installing dependencies and managing competing dependencies across projects is sometimes referred to as “dependency hell.” Containers prevent dependency hell. They do this by making an appropriately configured minicomputing environment (e.g., with the necessary version of dplyr) that can easily be installed on a new computer and hidden from the other software running on the new machine.
There are two main things a researcher can do with containers: create them and work inside them. To create a container, a researcher starts by making a list of the all the things to be included in the computing environment. With Docker, this configuration file is called a “Dockerfile.” Figure 2 shows the Dockerfile for Stanescu et al. (2019). Then the researcher executes the Dockerfile, and Docker creates a bundle of all the necessary software. With Docker, this bundle of software is called a “Docker image.” The Dockerfile for Stanescu et al. (2019) is relatively simple, but some were much more difficult to create. For example, the Dockerfile for Davidson (2019) required 87 packages. 10
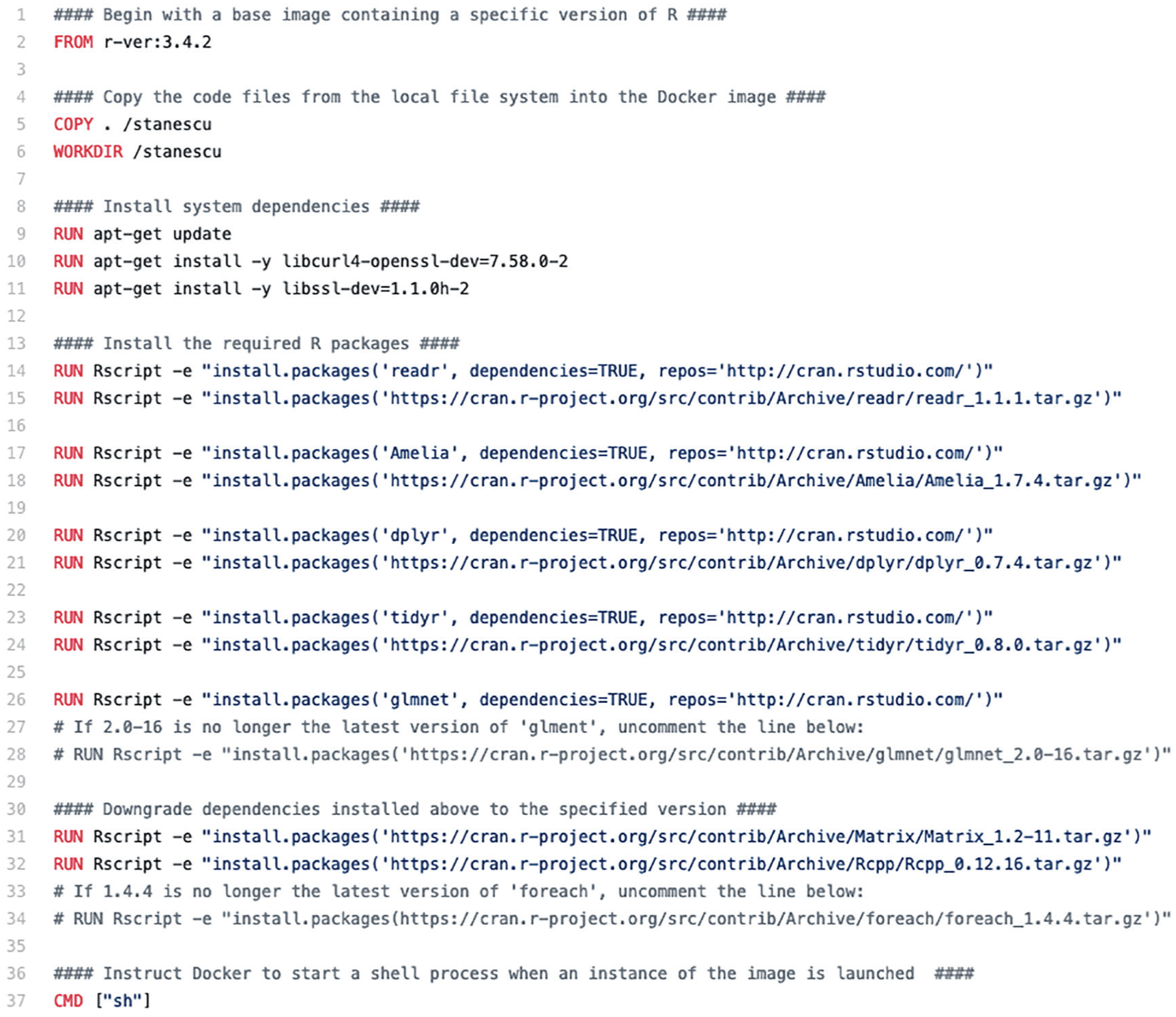
This Dockerfile specifies the dependencies required for Stanescu et al. (2019). In addition to R, we install system packages, such as libssl-dev, and R packages, such as dplyr. For the R packages, we first installed the newest version of the required packages along with the necessary dependencies. However, sometimes these newest versions did not match the versions used by the authors. In these situations, we then overwrote the newest version with the one that was specified by the authors.
Once a Docker image is created and published online, a new researcher can install it to his or her computer and use the image to create an appropriately configured container. The new researcher can then go inside of the container and work in a computing environment that was created to support the replication of the original scientific results. Furthermore, this computing environment is separate from the larger computing environment on the new researcher’s computer. A researcher downloading and using a Docker image does not need to understand exactly how it was created, which means that it is much easier to use an image than create one. Although Docker itself may be replaced in the future by alternative tools, we believe that the idea of software containers will likely be important for computational reproducibility for the foreseeable future.
Although software containers standardize the software environment, cloud computing can be used to standardize the hardware environment. Roughly, a cloud computing environment is a huge collection of computing resources, which can be subdivided so that different users can access different amounts of computing resources. Creating and managing cloud computing environments is extremely complex, but from the perspective of a user, they are relatively simple. At the time of the Fragile Families Challenge, large companies such as Amazon, Microsoft, and Google offered cloud computing environments that enabled people to rent computing resources by the hour. For example, at the time of the Challenge, Amazon Web Services, Amazon’s cloud computing environment, allowed people to rent computational resources with names like “m5.large” and “t3.medium.” These different environments came with different types of processors, random-access memory, and other computing resources. We chose to perform our reproducibility work on Amazon Web Services because we thought this would increase the chance that other researchers will be able to access a very similar hardware environment for the foreseeable future. Having access to a similar hardware environment is helpful because the performance and output of complex software can sometimes depend on the hardware that is used to execute it. 11 Future researches, however, will not be required to use Amazon Web Services or any other cloud computing provider to use the Docker images that we created. Together, software containers and cloud computing make it easier for researchers to have access to a very similar computing environment to the one we used when verifying computational reproducibility, both today and for years to come.
The third and final way our approach differed from common approaches in social science is timing. To the best of our knowledge, most work on computational reproducibility review in social science journals happens after a manuscript has already been conditionally accepted. Our approach, however, involved computational reproducibility review simultaneous to manuscript review. We think this simultaneous review appropriately emphasizes that the published article and code work together to communicate a scientific finding.
Our Process for the Special Issue
Although we had some experience with computational reproducibility in our roles as researchers, we had no experience with computational reproducibility from the perspective of a journal editor. This inexperience meant that we ended up using a process that we now realize was not ideal in several ways. Before offering our recommendations for the future, however, we will describe the process that we used for the special issue.
In our call for papers for the special issue, we stated that manuscripts needed to be accompanied by open-source code that would allow another researcher to reproduce all the figures and tables in the article. 12 However, we did not provide specific guidelines about the form that the code should take. Furthermore, our call for papers had a specific deadline by which manuscripts needed to be submitted (October 16, 2017).
Fourteen manuscript were submitted. While these manuscripts were undergoing peer review, we attempted to reproduce the results in each one. Having computational reproducibility reviews happening at the same time as manuscript peer reviews was consistent with our goal of considering replication materials to be a central part of scholarly communication.
Our process of computational reproducibility review generally involved five steps: (1) create a Docker image with the necessary software and packages, (2) move the authors’ code into the Docker image and adjust the file paths to match those of the container, (3) add the Challenge data file to the container, 13 (4) run the authors’ code inside the container in our cloud computing environment, 14 and (5) compare the results created within our computing environment with the results shown in the manuscript. We considered a manuscript to be computationally reproducible if we could run the author’s code in our computing environment and regenerate (1) all the prediction.csv files (these are the files participants created to record their predictions of all six outcomes for all 4,242 families) within our error tolerance, 15 (2) all the figures that were generated by the author’s code, and (3) all the tables that were generated by the author’s code.
For some manuscripts, this process went smoothly, but for many others it was quite painful. It was often extremely difficult to even run the authors’ code. In some cases, authors sent a lot of unnecessary code, and in other cases, authors did not send all the code that was necessary. Furthermore, the code that was provided was often disorganized and poorly documented. Even if we were able to get the code to run, it was also often difficult to compare the results produced by the code with the results in the manuscript.
During this initial computational reproducibility review, we were able to approximately reproduce the results in 2 of the 14 manuscripts. We had this low rate of success despite being in frequent e-mail contact with some authors and often devoting many hours trying to reproduce the results of each manuscript. 16
Initially, we had hoped to provide each author with individual feedback on his or her project’s computational reproducibility, much as each author receives individual feedback on his or her manuscript. Unfortunately, this proved to be too time consuming for us given our desire, and that of the editors of Socius, to provide rapid feedback to the authors. Ultimately, each author received individual feedback on their manuscript from editors of the special issue and about three anonymous reviewers, and each author received collective feedback on how computational reproducibility could be improved. This collective feedback—reproduced in the supporting online materials of this article—was developed inductively on the basis of our struggles with the initial code submissions, as well as on our review of existing articles about computational reproducibility.
All 14 manuscripts received a decision of “revise and resubmit,” and authors were instructed to improve both their manuscripts and code on the basis of the feedback that was provided. Fortunately, we saw dramatic improvements in code structure and documentation between the initial submissions and the revised versions. We do not know, however, how much of this improvement should be attributed to the guidelines we provided to authors and how much should be attributed to other factors, such as the authors realizing that we were treating computational reproducibility as a part of the review process. 17
After the authors received this first round of feedback, the synchronization of the manuscript review and the computational reproducibility review broke down. In some cases, our computational reproducibility review would proceeded while the manuscript was under review, and in other cases it would proceeded while the authors were working on their revisions. This second stage of the computational reproducibility review was particularly time consuming because it required us to precisely identify, and attempt to eliminate, the discrepancies between the results in the manuscript and the computationally reproducible results. This process was occasionally frustrating for us, and we are sure that it was occasionally frustrating for the authors.
Eventually, the editors of Socius informed us that we needed to bring the review process for the special issue to an end. We gave all authors with outstanding manuscripts a deadline of November 13, 2018, to submit their final manuscripts and final code. In the end, we completely reproduced the results of 7 manuscripts; for the other manuscripts, we ran out of time (although for many of these, we did make substantial progress). For each of the 12 manuscripts, the code will be released along with an accompanying Docker image with the necessary dependencies. Thus, all articles in the special issue reached TOP Level 2 (“access before publication”), and most reached TOP Level 3 (“verification before publication”). The supporting online materials summarize the computational reproducibility of each article in the special issue.
The most important lesson that we take from our experience is that ensuring computational reproducibility was harder than we had anticipated. There was not one specific thing that caused huge difficulties, but many of the difficulties came from a combination of three interrelated factors: many moving parts, long feedback cycles, and communication difficulty. To illustrate these factors, we will briefly summarize our attempts to replicate the results in Altschul (2019), which was neither particularly easy nor particularly difficult.
At first glance, the code for Altschul (2019) appeared straightforward to reproduce because it is a single R file. Yet this code required 13 R packages to be installed, all of which had to be built from source, a more error-prone and time-consuming form of package installation, because our cloud computing environment used Linux. 18 Furthermore, these packages in turn depended on many other packages which also had to be installed. Although each package can be built from source with a high probability of success p, the entire process will fail if just one package fails, and the probability that at least one package fails is 1 – pn, where n is the number of packages. So, as the number of packages increases, the probability of at least one failure increases exponentially. Beyond just installing packages, many of the submissions had a large number of components and a failure of just one component caused a failure of the entire process. This is what we call the “many moving parts problem.”
The difficulty of the many moving parts problem was exacerbated by a long delay between execution and feedback. The process of debugging computational reproducibility issues often involves attempting to isolate the problem and then use trial and error to solve the problem. The longer we had to wait to receive feedback, the longer it would take for us to complete many iterations of this isolation and trial-and-error process. For example, for Altschul (2019) it took about one hour of computing time to build the Docker image and many attempts to isolate and eliminate discrepancies required building a new Docker image and then rerunning the code, which would often have to run for several hours. These long computing times made the process of debugging very slow.
Finally, our slow debugging often had to go through many iterations because of communication difficulties. In the 13 months between initial submission and final publication, we exchanged a total of 28 e-mails with Altschul. These exchanges spanned long periods of time not only because we were also communicating with many other authors at the same time but also because each reply could require hours of additional work. Although not true for Altschul (2019), we also noticed that in some cases, as time went on, it became increasingly difficult for the authors themselves to respond to us quickly because it had been such a long time since they had worked with their own code.
We describe these difficulties replicating Altschul (2019) not because it was a particularly hard case but to illustrate the kind of difficulties that we faced as a result of the many moving parts problem, long feedback times, and communication difficulties. Undoubtedly many of these difficulties were caused by our inexperience with computational reproducibility, and later in the article we offer suggestions for handling these issues more efficiently.
Stepping back, our experience with the Challenge was unusual in a number of ways, and other editors may have different experiences with computational reproducibility in other settings. In fact, we can imagine some reasons to expect that computational reproducibility will be easier in other settings and some reasons to expect the opposite. We think there are three main reasons that ensuring computational reproducibility should be easier in other settings. In sociology and the social sciences more generally, most researchers use simpler data processing and statistical modeling pipelines than were used in the Challenge. We had the easiest time reproducing the results in manuscripts that use methods more common in the social sciences, such as Ahearn and Brand (2019) and McKay (2019). We had more difficulty, however, with manuscripts that included more machine-learning techniques, such as Davidson (2019), Rigobon et al. (2019), and Carnegie and Wu (2019). Given the relatively small number of manuscripts, it is hard to know precisely why those that used more machine-learning methods were harder, but some of the difficulties we encountered included much longer run times and the use of parallelization. Second, the Challenge involved deadlines: there was a hard deadline for submitting to the Challenge and another hard deadline for submitting to the special issue. We think these deadlines, which are uncommon in the social sciences, may have caused code quality to suffer. Finally, we think that in the future, authors and editors will have more experience with computational reproducibility (and we hope that some of the ideas in this article can help with that).
On the other hand, we think achieving computational reproducibility could be harder in other settings for three main reasons. First, in other settings there may be less buy-in from authors and editors; in this case, we think the fact that the Fragile Families Challenge was a mass collaboration helped set the tone of open and reproducible research. Second, in other settings authors may have less experience writing code, which could create barriers to computational reproducibility. Third, in other settings, authors and editors will have to deal with the difficulties related to data access, which did not arise here because all Challenge participants were using the same data set that we had provided. 19
Recommendations for Authors
On the basis of our experience with the special issue—as well as the suggestions of others (Alvarez et al. 2018; Barnes 2010; Benureau and Rougier 2018; Bowers and Voors 2016; Eubank 2016; Fehr et al. 2016; Grüning et al. 2018; Konkol et al. 2019; Marwick, Boettiger, and Mullen 2018; Nagler 1995; Peng 2009; Sandve et al. 2013; Stodden et al. 2016; Tatman et al. 2018; Wilson et al. 2017)—we have some recommendations for authors who wish to improve the computational reproducibility of their results now and in the future. We believe that these recommendations are applicable to all authors, even if they are not using software containers and cloud computing.
We find it helpful to conceptualize the code used to create the results in a manuscript as a research pipeline (Peng and Eckel 2009). The research pipeline accepts the rawest data and produces all the final outputs in the manuscript (Figure 3). To promote computational reproducibility, we recommend that researchers make their research pipelines automated, modular, and friendly. Before describing these three ideas in more detail, and offering a five-item checklist, we want to emphasize that our experience with the Challenge suggest that most researchers do not create automated, modular, and friendly research pipelines without effort. In the Challenge, the code that was most effective had repeatedly gone through a process that software engineers call refactoring. Roughly, refactoring involves taking code that already works and improving its internal structure without changing its external behavior (Fowler 1999). In professional software engineering, refactoring is not considered a sign of poor work; rather it is considered a best practice that acknowledges the reality that software projects evolve over time and that design decisions made early in the project may need to be change as the project evolves. In the context of the computational reproducibility of an academic article, refactoring might include changes such as removing code that is no longer used, improving variable names, and adding a loop or writing a function to eliminate redundant blocks of code. Just as a researcher often writes, rewrites, and re-rewrites a manuscript to communicate ideas as clearly as possible, we think that researchers should also expect to write, rewrite, and re-rewrite their code to end up with a computationally reproducible research pipeline.

Research pipeline based on Figure 1 in Peng and Eckel (2009). When writing code, and refactoring code before submissions, authors can imagine a research pipeline that starts with the rawest form of the data, what Peng and Eckel called the “measured data,” and ends with all the figures, tables, and numerical results in an article. To promote computational reproducibility, we recommend that authors make their research pipelines automated, modular, and friendly.
Three Guiding Principles
Automated
A research pipeline should run from start to finish without human intervention; it should be fully automated. The desire for end-to-end automation is not often common during the research process, because authors are often focused on creating a specific component of the pipeline, such as a specific figure. However, once a manuscript is complete, it is important that all the individual software components seamlessly communicate with each other and can execute from start to finish without any manual intervention to produce all the results that are presented in the manuscript.
Modular
As code becomes more complex, it becomes increasingly important to break it into independent modules, such that each module serves a specific purpose and the inputs and outputs of each module are clear. Modular code offers many benefits during the research process and during the process of preparing the code for others (as well as for your future self). While doing research, modularity benefits authors by making the code easier to understand, maintain, and improve. Furthermore, for analyses that involve long run times, such as many of the research pipelines in the Fragile Families Challenge, modularity lets authors run only specific sections of their code.
Additionally, modularity helps computational reproducibility in three ways. First, it helps future researchers understand how the code works; a well-designed, modular code architecture is a wonderful form of documentation. Second, if computational reproducibility is not successful, having modular code dramatically improves the chances to find and then fix problems. Finally, modular code promotes refinement and reuse by future researchers, because it enables them to easily focus on only the parts of the code that are most important to them.
Friendly
The final characteristic of a good research pipeline is that it is user-friendly. In other words, even thought the code is written to be executed by a computer it should also be written to be read by a person. 20 A user-friendly research pipeline is organized, clean, and clear for someone else without detailed knowledge of the code base. In the special issue, we learned that code that was clear to authors was not always clear to us. To promote friendliness, therefore, we encourage authors to attempt to put themselves in the place of others and ask, “What would I want if I was trying to understand, reproduce, and then improve this code?”
Five-Item Checklist
The three principles that are described above—automated, modular, and friendly—are designed to provide high-level guidance to authors. However, these principles are intentionally abstract, and they do not always provide clear action-guiding advice. Therefore, we now present a simple five-item checklist that is designed to help authors and that could be used by journals. This checklist does not cover every possible situation, but it is designed to be helpful to a wide range of social scientists, impose minimal costs, and be independent of the programming language used by the researcher.
Run-All Script
We recommend that authors include a single “run-all” script that contains a sequence of commands that reproduces all the results in the article. For example, during the Challenge, we suggested that authors write a bash script to call their Python, R, or Stata files. 21 The run-all script serves as a project blueprint; it provides a high-level understanding of the relationships among the scripts, without the need to understand each script individually. The run-all script was one of the first things we reviewed when attempting to reproduce a result, and so it should be considered a key piece of documentation. For example, Figure 4 reproduces the run-all scripts from Filippova et al. (2019). The script first executes an R Markdown file that contains data-cleaning code before executing a series of imputation and then modeling scripts. Without this run-all script it would be nearly impossible for someone trying to reproduce these results to know the proper order of execution of these 17 scripts and the appropriate way to move the output of some scripts into another directory.
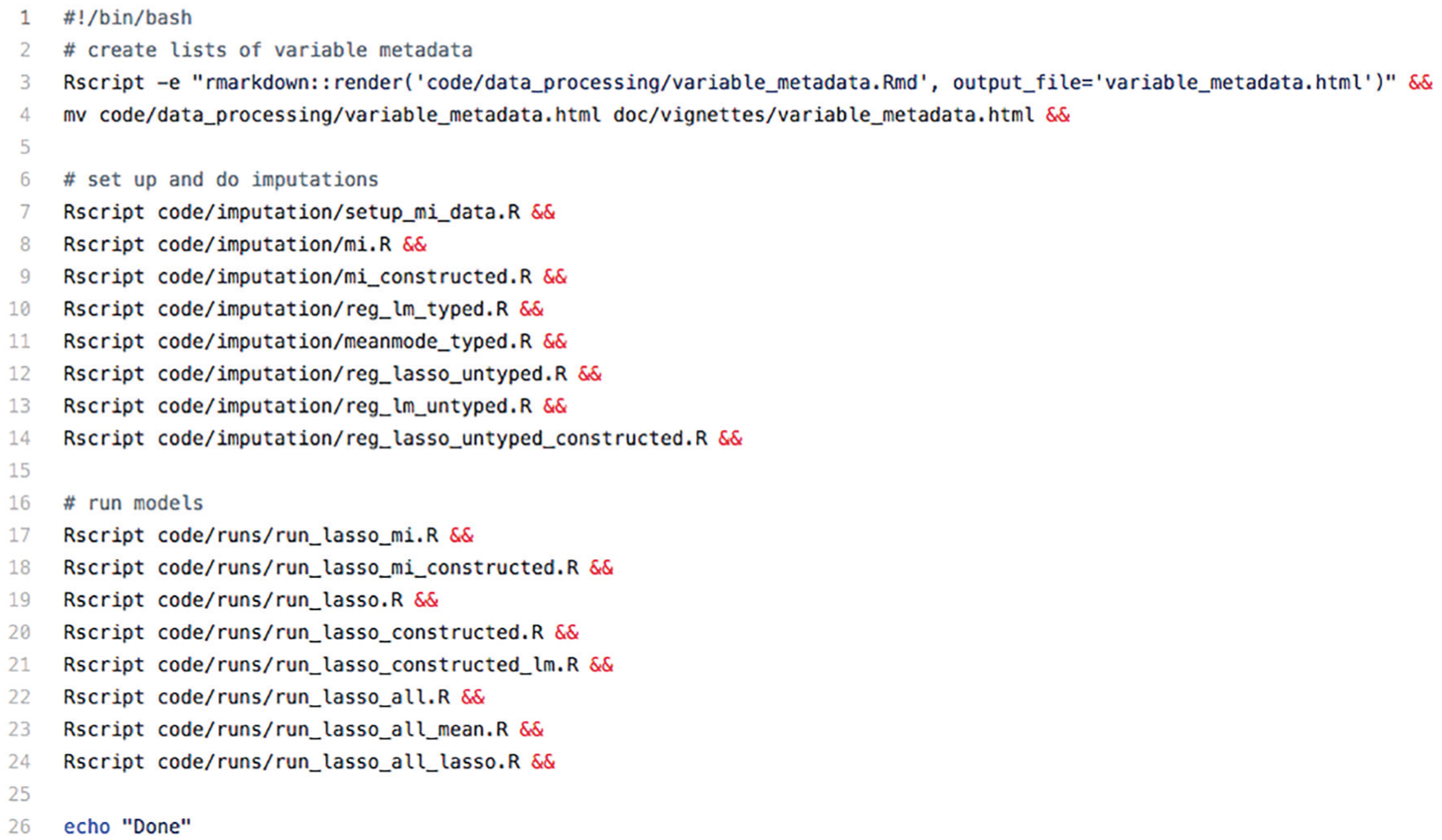
This “run-all” script from Filippova et al. (2019) clarifies the exact commands needed to execute all of the scripts as well as the order these scripts should be executed.
Informative Directory Structure
To supplement the run-all script, we recommend that authors organize their files into a clear directory structure. For example, we asked authors to organize their files in three major directories: code, data, and output. A standardized directory structure is helpful during the execution process because some complex pipelines involve creating multiple intermediate files and the directory structure helps maintain order. Authors can adjust their directory structures to fit the nature of their code. For example, the directory structure in Rigobon et al. (2019), shown in Figure 5, made many common operations more convenient. First, all of the raw Challenge data files, which are not included in the replication materials because of privacy concerns, were meant to be placed in a clearly specified directory (/data). Second, all of the final results created by the script were placed in a single directory named “final_pred” This enabled us to save a copy of these results in “final_pred_author” before rerunning the script, which made it easy for us to compare the two sets of results.

Directory structure for Rigobon et al. (2019). Because all of the prediction files the authors generated resided in a single subdirectory, we were able to easily save a copy of the original results (“final_pred_author”) before generating our own version (“final_pred”) for comparison.
Requirements Files
We recommended that authors include a file listing all of the software and add-on packages used, including the exact version numbers (e.g., “ggplot2 v3.1.0”). Understanding the software requirements is helpful for reproducing results today, and it is necessary for reproducing results in the future. Many of the articles in the special issue used open-source software with a large number of add-on packages, and these packages are regularly updated with improvements. Although these improvements are generally good for users, they are problematic for reproducibility. These new version sometimes introduce what are called breaking changes, which cause code that used to work to suddenly break. For example, the improvements between Python 2 and Python 3 introduced some breaking changes into Compton (2019), which we discovered when we attempted to use Python 3 to import preprocessed data files that had been serialized with Python 2. Furthermore, new versions sometimes cause much more subtle errors. For example, there was a change in Python’s pseudorandom number generator between Python 3.2 and Python 3.3 that could cause some results to fail to be computationally reproducible (Benureau and Rougier 2018).
The deadline for initial submission of manuscript and code to the special issue was October 17, 2017, and the deadline for the final, revised submissions was November 13, 2018. During this 13-month period many of the packages used by participants during the Challenge were improved. However, these improvements also led to problems for computational reproducibility. For example, we when we attempted to reproduce Raes (2019) using the most recent version of all the packages used, the results matched for only 2 of the 11 prediction files. For the other files, the differences were sometimes off by a constant and sometimes off by random noise (Figure 6). However, after matching package versions exactly, we were able to reproduce the results exactly (Figure 6). We close by noting that the problems we had with package versions during the relatively short time window of the special issue will grow over time. Without knowing specific package numbers, we think it will be nearly impossible for researchers in the future to reproduce Challenge results, even if the data and code are otherwise available.
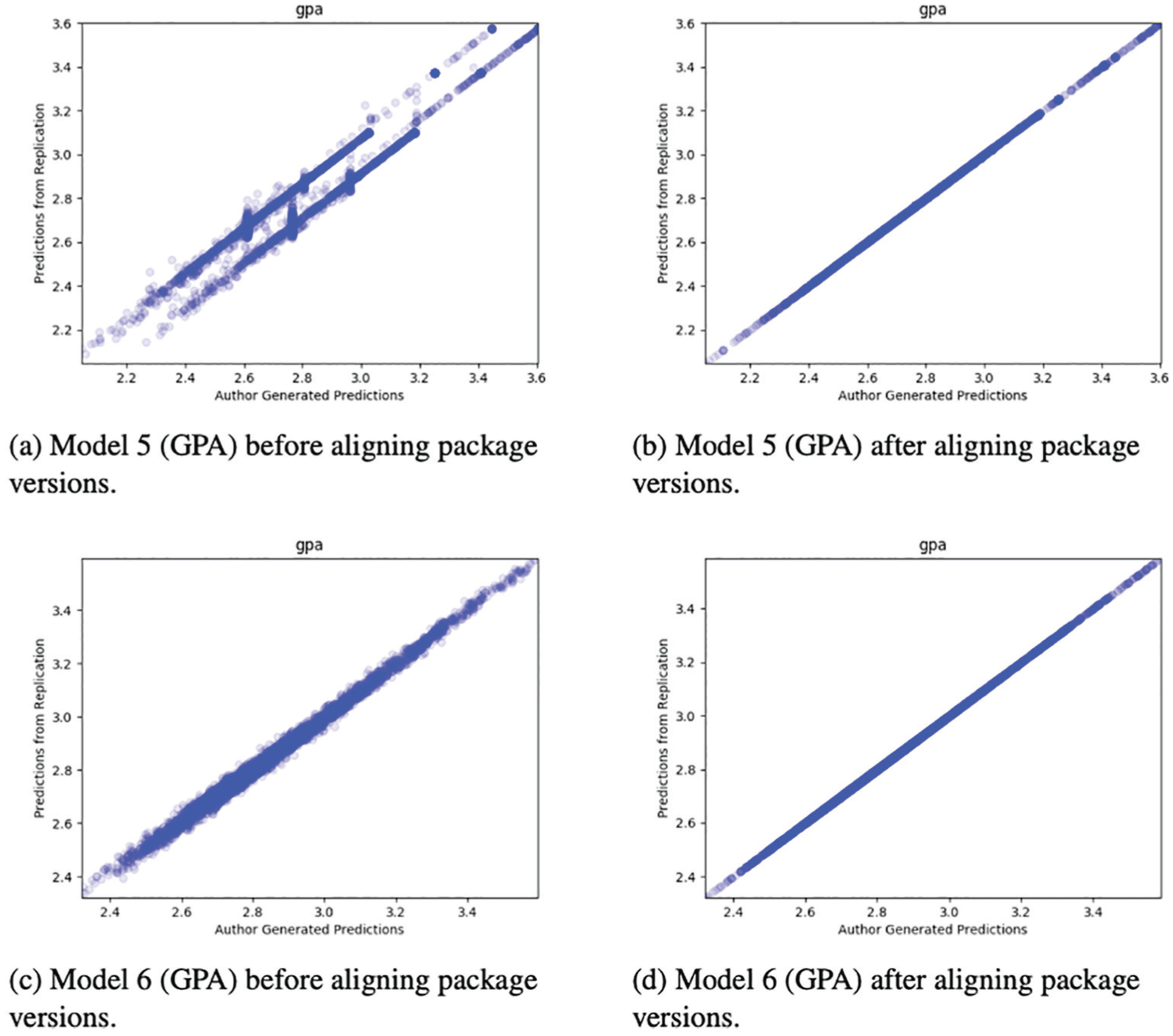
Examples from Raes (2019). Without specifying package versions, we could not replicate the results, even though we were using the exact same data and code. Furthermore, there was no clear pattern to the errors: sometimes they were off by a constant, and sometimes they were off by random noise. However, once we specified the exact package versions in our container, we were able to reproduce the original results exactly.
Helpful Code Headers
We recommended that researchers clearly document their code. Unfortunately, we found that some authors struggled with this very general advice. Therefore, more specifically, we recommend that authors produce code headers that contain the following information:
Purpose (in 280 characters or fewer)
Inputs (i.e., What files does this piece of code require? Is there any code that needs to be run before this code?)
Outputs (i.e., What files, tables or figures does this piece of code create as outputs?)
Machine used (i.e., computer type [laptop, desktop, cluster], operating system)
Expected runtime (e.g., seconds, minutes, hours, days)
For example, Figure 7 presents the code header from the data-cleaning script for Ahearn and Brand (2019). Each piece of information helps the replicator understand what should happen upon execution.
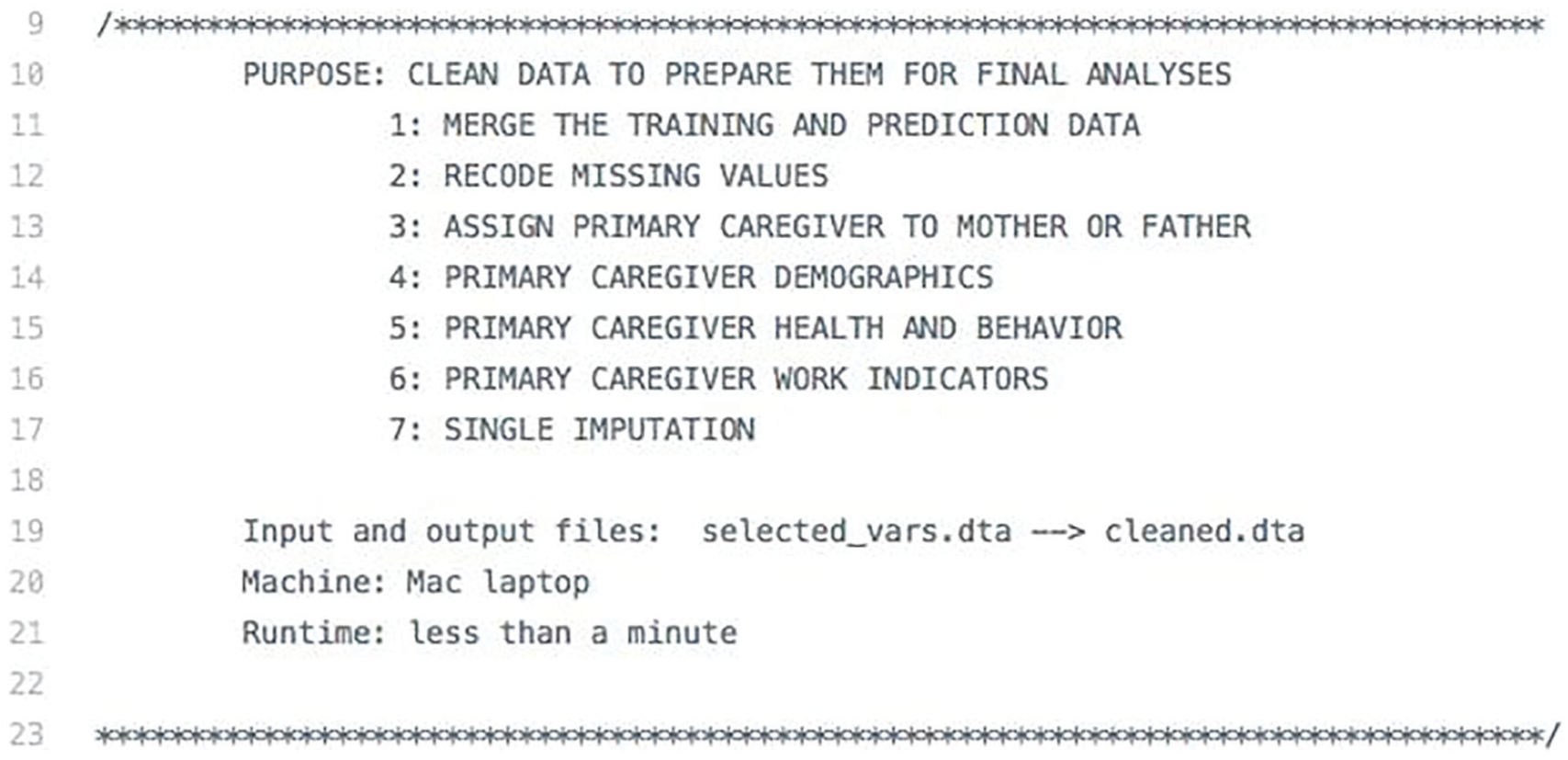
This code header is taken from the data-cleaning script from Ahearn and Brand (2019). The information clearly outlines the code’s structure and characteristics.
Run Twice
Once authors have completed the other four items on their checklist, we recommend that they run the code twice and ensure that it produces the same results. Running the code twice is necessary if it uses a pseudorandom number generator. Roughly, every time a computer generates a sequence of random numbers, the results will be different. But, these results are not truly random; rather they are a pseudorandom sequence of numbers that are determined by a single seed value that was used to initiate the sequence. If the code sets the seed value explicitly, then the results will be reproducible, but if it does not explicitly set a seed value, the exact same code, using the exact same data, and run on the exact same computer can produce different results. 22 For example, initially we were unable to reproduce the results of Stanescu et al. (2019), because their code did not explicitly set a seed value. However, after they explicitly seeded their code, we were able to reproduce their results. For some more computationally complex submissions, multiple seed values had to be set for different packages that used different seeds. For example, Davidson (2019) required five different seeds, ranging from the Python standard library’s seed to TensorFlow’s own graph-level seed. Thus, for some complex submissions, such as Davidson (2019) and Roberts (2019), it was often difficult to detect if the code had been properly seeded, and an empirical test, such as testing for consistency across runs in the same computing environment, was most effective.
We hope that these three principles and this five-item checklist can help authors who want to make their work more computationally reproducible for other researchers. There are also steps that journals can take to promote reproducibility, and we offer some of those recommendations next.
Recommendations for Journals
Although we believe that individual authors have strong incentives to practice open and reproducible research, journals and other institutions also play a key role in accelerating progress toward open and reproducible science (Freese and King 2018). Building on our experience with computational reproducibility in this special issue, we offer a menu of four options that journals can take to promote computational reproducibility, we make specific suggestions related to TOP Level 3, and we present our preferred process.
Menu of Options
Journals can promote computational reproducibility with sticks and carrots. That is, journals can force authors to comply with TOP Level 2 or TOP Level 3 (sticks). Or they can induce authors to comply these guidelines with benefits, such as faster review time or increased probability of acceptance (carrots). Of the carrots available to journals, we believe that a badging system is a particularly promising option. Badges are small symbols that appear on published articles that quickly and clearly indicate the open-science practices associated with those articles (Figure 8). Badges reward authors for following open-science practices, and there is some evidence that badging systems have had their intended effect of promoting open and reproducible research (Kidwell et al. 2016; Rowhani-Farid, Allen, and Barnett 2017). Social scientists are probably most familiar with the badging system developed by the Center for Open Science and first used by the journal Psychological Science starting in January 2014 (Eich 2014; Kidwell et al. 2016; Rowhani-Farid et al. 2017). However, similar badging systems have also been developed in other fields. The journal Biostatistics introduced a badging system in 2009 (Peng 2009, 2011), and the Association for Computing Machinery, a prominent professional association of computer scientists, recently developed a badging system specifically focused on computational reproducibility (Boisvert 2016). Badging systems are often opt-in (i.e., authors can choose to participate or not), which makes them easier to implement than other systems. 23

Example badges from the Center for Open Science (Kidwell et al. 2016) and the Association for Computing Machinery (Boisvert 2016). Badges are small symbols that appear on published articles that indicate the open-science practices that apply to this particular article. They are an inducement journals can use to promote different research practices.
Given the two main levels of computational reproducibility—TOP Level 2 (“access before publication”) and TOP Level 3 (“verification before publication”)—and two main policy options—requirements and badges—we think that journals could consider four main options: (1) badge to TOP Level 2, (2) require TOP Level 2, (3) require TOP Level 2 and badge to TOP Level 3, and (4) require TOP Level 3. In this special issue, we attempted to require TOP Level 3, and we were not able to achieve this goal in the time available. Therefore, we ended up requiring TOP Level 2, although we note that many of the articles in this special issue actually did achieve TOP Level 3.
Suggestions Regarding TOP Level 3
For journals considering either requiring TOP Level 3 or badging to TOP Level 3, we have five suggestions. First, journals should have a clear plan for who is going to do the additional work that is required to ensure computational reproducibility; we think the most likely possibilities include authors, reviewers, editors, existing journal staff members, and new journal staff members. 24 In our case, we did not realize how much extra work would be required, and we did not have a clear plan of who would be responsible for it. Therefore, we ended up doing a lot of work helping authors make their work more reproducible. This was in part because we did not want to reject any articles that failed to achieve computational reproducibility. In retrospect, it is clear that we should have done a better job clarifying the allocation of responsibilities with the authors.
Second, we believe that badging to TOP Level 3 will be substantially easier than requiring TOP Level 3. In our case, it was relatively easy to verify that some of the manuscripts were computationally reproducible but others were extremely difficult. We ended up spending roughly 80 percent of our time on 20 percent of the manuscripts. If we had chosen to badge to TOP Level 3, we expect that authors interested in computational reproducibility, who generally had results that were easier to reproduce, would have attempted to earn the badge and others would not. Furthermore, a badging system would have enabled us to set clear time limits on the amount of time we would spend on each submission, a process sometimes called “timeboxing.” For example, we could have told authors that we would spend six hours attempting to reproduce their results, and if we could not do it within that time, the authors would not receive a badge. We think that a timeboxed system would help promote best practices by authors and would save a lot of time for people verifying reproducibility.
Third, we think that journals, or perhaps publishers, should invest in tools that can facilitate the assessment of computational reproducibility. The online manuscript-handling system used by Socius does not currently provide tools to support sharing and review of code, so we did most of our computational reproducibility work through e-mail, a process that introduced numerous inefficiencies. As a first step, journals could build connectors to existing tools such as GitHub and create simple computational reproducibility checklists that would become a part of the manuscript submission process. Efforts to automate and standardize the process of assessing computational reproducibility will, in the long run, decrease its cost and increase its probability of success.
Fourth, we think that journals just starting to deal with computational reproducibility should draw insights from other journals that already have developed computational reproducibility and open-science policies, such as Biostatistics (Peng 2009), Psychological Science (Eich 2014), ACM Transactions on Mathematical Software (Heroux 2015), International Interactions (Colaresi 2016), the Quarterly Journal of Political Science (Eubank 2016), the Journal of the American Statistical Association (Baker 2016), the American Journal of Political Science (Jacoby, Lafferty-Hess, and Christian 2017), ReScience (Rougier et al. 2017), Political Analysis (Alvarez et al. 2018), and Methods in Ecology and Evolution (Freckleton 2018). We made a mistake by not including detailed instructions about computational reproducibility in our initial call for papers, and we could have easily developed such instructions based on those used by these other journals.
Finally, for journals thinking of requiring TOP Level 3, we would encourage editors to consider whether they have full support from all important stakeholders, including the publisher, the journal’s editorial board, and the pool of potential reviewers and authors. One way to assess this support concretely is with the following question: “Are we willing to reject a manuscript that we would otherwise publish because it is not computationally reproducible?” If the answer is not a resounding yes, we would not recommend attempting to require TOP Level 3. Furthermore, editors should assess whether stakeholders are willing to bear the costs that may come with requiring TOP Level 3, which could be assessed with questions such as “Are we willing to have our turnaround time for manuscripts increase by one week (or one month or three months)?” and “Are we willing to have the number of submissions decrease by 10 percent (or 20 percent or 50 percent)?” In our case, a core value of Socius is “rapid dissemination of high-quality, peer-reviewed research, produced in time to be relevant to ongoing research and public debates” (Keister and Moody 2015), and that value did not always align with the time requirements of our computational reproducibility verification process.
Our Preferred Model
For journals that currently have no policies on computational reproducibility, we think that a possible first step would be to require TOP Level 2 and badge to TOP Level 3. Badging to TOP Level 3 would require code review, so we briefly sketch our preferred workflow, one that elevates the code to a first-class status alongside each article (Figure 9). When authors submit manuscripts, they also submit the code, data, and computing environment needed to reproduce all the results in their manuscripts. The manuscript and replication materials would then be sent to several peer reviewers and to a reproducibility reviewer, who could be a student, a peer, or a professional software engineer. The peer reviewers would write reports, much as they do now; they would be free to assess the replication materials or not. The reproducibility reviewer would, however, focus entirely on the replication materials. He or she would spend a fixed maximum amount of time, say 6 hours, attempting to reproduce the results in the manuscript. Then the reviewer would write a reproducibility report, which would go to the editor and be considered along with the manuscript reviews. If an author is encouraged to submit a revised manuscript, he or she would need to reply to the comments of the peer reviewers and reproducibility reviewer. We think that adding a reproducibility review into the editorial process appropriately draws attention to the code and allows community standards to vary by field and evolve over time. Furthermore, the introduction of a reproducibility reviewer who will work for a fixed maximum amount of time on each submission makes the costs of reproducibility reviews predictable and manageable.
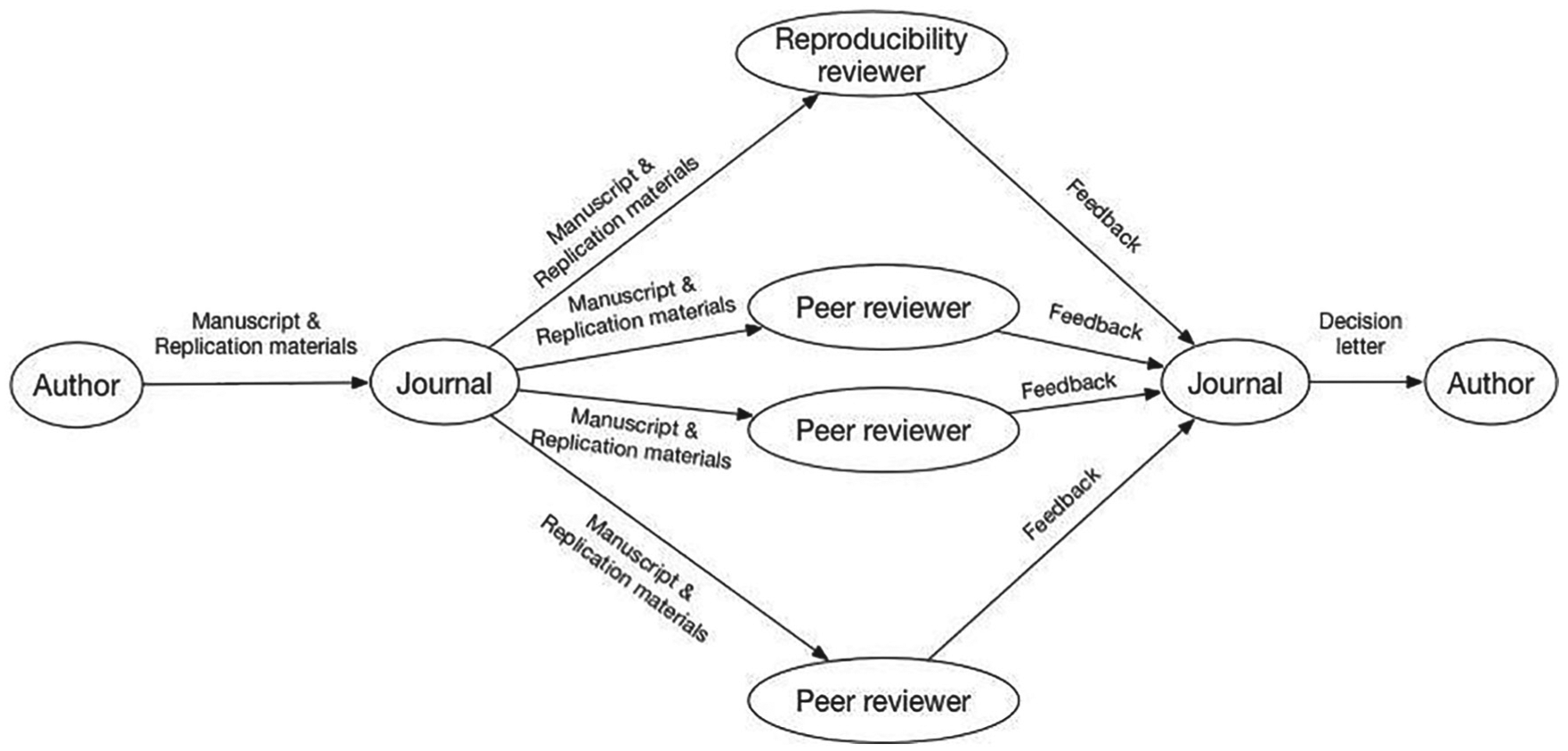
Schematic of our preferred workflow for journals that are badging to Transparency and Openness Promotion Level 3 (“verification before publication”). Adding a reproducibility review into the editorial process elevates the code to a first-class status alongside each article and allows community standards to vary by field and evolve over time.
Conclusions
In this article we have described our experience with promoting computational reproducibility for the special issue of Socius on the Fragile Families Challenge. Our approach was inspired by previous efforts, but it differed from other approaches in the social sciences in terms of focus, motivation, and timing. On the basis of our success and struggles, we have offered recommendations to authors who want to increase the computational reproducibility of their work and to journals that want to promote computational reproducibility.
One important aspect to our approach to computational reproducibility was to expand the focus and include the computing environment in addition to code and data. As sociologists and other social scientists increasingly publish articles based on larger data sets and more computationally intensive methods, we expect the need to have this broader focus will increase. In addition to the tools that we chose to use, there are a number of other tools that may be useful to future researchers interested in computational reproducibility. 25
Our work also raises questions about the costs of computational reproducibility. Our experience suggests that ensuring computational reproducibility of complex computational approaches is going be time consuming for authors and journals. Who should bear these costs? Should we have different computational reproducibility standards for different kinds of research? In what situations would access to code be sufficient, even if that code cannot be rerun or if it could be rerun but produces slightly different results? What about research that is computationally reproducible, but where the code is extremely hard to understand? What if results are computationally reproducible in some computing environments but not others? How long after publication should we reasonably expect results to be reproducible? Ultimately these are questions that have to be answered by the scientific community, and we expect that the answers will vary from field to field. Despite these important unresolved questions, we hope the approach that we took to promote computational reproducibility will increase the impact of the work published in the special issue, further the research goals of the Fragile Families Challenge, and contribute to broader efforts to create a more transparent and open system of scientific publishing.
Supplemental Material
Liu_Successes_2020_Supplementary_Material_Revised – Supplemental material for Successes and Struggles with Computational Reproducibility: Lessons from the Fragile Families Challenge
Supplemental material, Liu_Successes_2020_Supplementary_Material_Revised for Successes and Struggles with Computational Reproducibility: Lessons from the Fragile Families Challenge by David M. Liu and Matthew J. Salganik in Socius
Footnotes
Acknowledgements
We thank the Board of Advisers of the Fragile Families Challenge. Research reported in this publication was supported by the Russell Sage Foundation and the Eunice Kennedy Shriver National Institute of Child Health and Human Development of the National Institutes of Health under award number P2-CHD047879. Funding for the Fragile Families and Child Wellbeing Study was provided by the Eunice Kennedy Shriver National Institute of Child Health and Human Development through grants R01-HD36916, R01-HD39135, and R01-HD40421 and by a consortium of private foundations, including the Robert Wood Johnson Foundation. We thank Caitlin Ahearn, Drew Altschul, Nicole Carnegie, Connor Gilroy, Brian Goode, Seth Green, Jake Hofman, Alex Kindel, Dawn Koffman, Daniel Rigobon, Julia Rohrer, and Janet Xu for helpful feedback on drafts of this article. This article draws on David Liu’s senior thesis for the Department of Computer Science at Princeton University (![]() ). The content of this article is solely the responsibility of the authors and does not necessarily represent the views of anyone else.
). The content of this article is solely the responsibility of the authors and does not necessarily represent the views of anyone else.
1
For more on the somewhat inconsistent terminology in this area, see Goodman et al. (2016), Patil et al. (2016), Clemens (2017), Plesser (2018), Barba (2018), and ![]() .
.
2
The introductory paragraph for the “Data, Analytic Methods (Code), and Research Materials Transparency” standard is as follows: “The policy of the [Journal of Research] is to publish papers only if the data, methods used in the analysis, and materials used to conduct the research are clearly and precisely documented and are maximally available to any researcher for purposes of reproducing the results or replicating the procedure. All materials supporting the claims made by the author must be made available to the journal prior to publication. The journal, or an entity acting on behalf of the journal, will verify that the findings are replicable using the authors data and methods of analysis. Failure to replicate at this stage may result in the paper not being published.” See ![]() for the most recent version.
for the most recent version.
3
Although this quotation is attributed to David Donoho, Donoho was himself influenced by Jon Claerbout and colleagues (Claerbout and Karrenbach 1992). See ![]() for more historical background.
for more historical background.
4
Participants in the Challenge attempted to predict six outcomes: grade point average (GPA) of the child, grit of the child, material hardship of the household, whether the household was evicted from their home, whether the primary caregiver participated in job training, and whether the primary caregiver lost his or her job. The choice of these outcomes was driven by our scientific goals and ethical considerations; each outcome is described in more detail elsewhere (Lundberg et al. 2019; ![]() ).
).
6
This is not to say that access to data and code is always desirable (![]() ). There are of course exceptions, such as data that contains personally identifiable information or code that contains proprietary business information. However, these exceptions do not call into question the general idea that for scientific research, computational reproducibility is a desirable goal.
). There are of course exceptions, such as data that contains personally identifiable information or code that contains proprietary business information. However, these exceptions do not call into question the general idea that for scientific research, computational reproducibility is a desirable goal.
7
8
To illustrate the value of reuse, we briefly sketch three research projects that are enabled by the computational reproducibility of the articles in the special issue. First, one could imagine trying to create a super-model that combines the best pieces of each of the individual papers. For example, imagine combining the imputation approaches of Goode et al. (2019), the approach to missing data of Carnegie and Wu (2019), the approach to adding expert knowledge of Filippova et al. (2019), and the machine-learning approach in Rigobon et al. (2019). Might this super-model, or some other combination of existing strategies, perform better than the best strategy used during the Challenge? Second, rather than combining modeling components, future researchers could explore different ways to ensemble predictions. Fragile Families Challenge Team (2019) shows that one such approach to combining predictions—stacking (Breiman 1996; Wolpert 1992)—did not lead to large improvements. Might other approaches, particularly approaches that require running code, lead to better ensembling results? Third and finally, the predictions submitted in the Challenge did not include any estimate of uncertainty. However, as predictive models get deployed for high-stakes social decisions (Rudin 2018), it would be desirable for them to also produce estimates of uncertainty. Using existing ideas from the literature (e.g., King, Tomz, and Wittenberg 2000), would it be possible to produce confidence intervals from these models? What might there coverage properties be? We emphasize that these are just three possibilities, and we expect that many other ideas might occur to other researchers.
9
For more about software containers and virtual machines (a closely related concept), we recommend Merkel (2014), Boettiger (2015), and Clyburne-Sherin and Green (2018). For more about cloud computing in the context of scientific research, we recommend Dudley and Butte (2010) and ![]() .
.
10
The 87 required packages and version numbers for ![]() are as follows: absl-py v0.6.1, appnope v0.1.0, astor v0.7.1, backcall v0.1.0, bleach v3.0.2, cloudpickle v0.6.1, cvxpy v1.0.10, cycler v0.10.0, dask v0.20.0, decorator v4.3.0, defusedxml v0.5.0, dill v0.2.8.2, ecos v2.0.5, entrypoints v0.2.3, fancyimpute v0.4.0, fastcache v1.0.2, future v0.17.1, gast v0.2.0, grpcio v1.16.0, h5py v2.8.0, ipykernel v5.1.0, ipython v7.1.1, ipython-genutils v0.2.0, ipywidgets v7.4.2, jedi v0.13.1, Jinja2 v2.10, jsonschema v2.6.0, jupyter v1.0.0, jupyter-client v5.2.3, jupyter-console v6.0.0, jupyter-core v4.4.0, Keras v2.2.4, Keras-Applications v1.0.6, Keras-Preprocessing v1.0.5, kiwisolver v1.0.1, knnimpute v0.1.0, lime v0.1.1.32, Markdown v3.0.1, MarkupSafe v1.1.0, matplotlib v3.0.1, mistune v0.8.4, multiprocess v0.70.6.1, nbconvert v5.4.0, nbformat v4.4.0, networkx v2.2, notebook v5.7.0, np-utils v0.5.5.2, osqp v0.4.1, pandas v0.23.4, pandocfilters v1.4.2, parso v0.3.1, patsy v0.5.1, pexpect v4.6.0, pickleshare v0.7.5, Pillow v5.3.0, prometheus-client v0.4.2, prompt-toolkit v2.0.7, protobuf v3.6.1, ptyprocess v0.6.0, Pygments v2.2.0, pyparsing v2.3.0, python-dateutil v2.7.5, pytz v2018.7, PyWavelets v1.0.1, PyYAML v3.13, pyzmq v17.1.2, qtconsole v4.4.2, scikit-image v0.14.1, scikit-learn v0.20.0, scipy v1.1.0, scs v2.0.2, Send2Trash v1.5.0, six v1.11.0, sklearn v0.0, statsmodels v0.9.0, tensorboard v1.12.0, tensorflow v1.12.0, termcolor v1.1.0, terminado v0.8.1, testpath v0.4.2, toolz v0.9.0, tornado v5.1.1, traitlets v4.3.2, wcwidth v0.1.7, webencodings v0.5.1, Werkzeug v0.14.1, and widgetsnbextension v3.4.2.
are as follows: absl-py v0.6.1, appnope v0.1.0, astor v0.7.1, backcall v0.1.0, bleach v3.0.2, cloudpickle v0.6.1, cvxpy v1.0.10, cycler v0.10.0, dask v0.20.0, decorator v4.3.0, defusedxml v0.5.0, dill v0.2.8.2, ecos v2.0.5, entrypoints v0.2.3, fancyimpute v0.4.0, fastcache v1.0.2, future v0.17.1, gast v0.2.0, grpcio v1.16.0, h5py v2.8.0, ipykernel v5.1.0, ipython v7.1.1, ipython-genutils v0.2.0, ipywidgets v7.4.2, jedi v0.13.1, Jinja2 v2.10, jsonschema v2.6.0, jupyter v1.0.0, jupyter-client v5.2.3, jupyter-console v6.0.0, jupyter-core v4.4.0, Keras v2.2.4, Keras-Applications v1.0.6, Keras-Preprocessing v1.0.5, kiwisolver v1.0.1, knnimpute v0.1.0, lime v0.1.1.32, Markdown v3.0.1, MarkupSafe v1.1.0, matplotlib v3.0.1, mistune v0.8.4, multiprocess v0.70.6.1, nbconvert v5.4.0, nbformat v4.4.0, networkx v2.2, notebook v5.7.0, np-utils v0.5.5.2, osqp v0.4.1, pandas v0.23.4, pandocfilters v1.4.2, parso v0.3.1, patsy v0.5.1, pexpect v4.6.0, pickleshare v0.7.5, Pillow v5.3.0, prometheus-client v0.4.2, prompt-toolkit v2.0.7, protobuf v3.6.1, ptyprocess v0.6.0, Pygments v2.2.0, pyparsing v2.3.0, python-dateutil v2.7.5, pytz v2018.7, PyWavelets v1.0.1, PyYAML v3.13, pyzmq v17.1.2, qtconsole v4.4.2, scikit-image v0.14.1, scikit-learn v0.20.0, scipy v1.1.0, scs v2.0.2, Send2Trash v1.5.0, six v1.11.0, sklearn v0.0, statsmodels v0.9.0, tensorboard v1.12.0, tensorflow v1.12.0, termcolor v1.1.0, terminado v0.8.1, testpath v0.4.2, toolz v0.9.0, tornado v5.1.1, traitlets v4.3.2, wcwidth v0.1.7, webencodings v0.5.1, Werkzeug v0.14.1, and widgetsnbextension v3.4.2.
11
For example, the runtime typically depends on the amount of random-access memory, the number of processors, and the type of processor (e.g., central processing unit vs. graphics processing unit). For more about how the underlying computing hardware can affect the results of scientific computing, see ![]() .
.
12
The full call for papers is included in ![]() , and here is the part most related to computational reproducibility: “What are the requirements for the open source code? The code must take the Fragile Families Challenge data files as an input and produce (1) all the figures and tables in your manuscript and supporting online materials and (2) your final predictions. The code can be written in any language (e.g., R, Stata, Python). The code should be released under the MIT license, but we will consider other permissive licenses in special situations.”
, and here is the part most related to computational reproducibility: “What are the requirements for the open source code? The code must take the Fragile Families Challenge data files as an input and produce (1) all the figures and tables in your manuscript and supporting online materials and (2) your final predictions. The code can be written in any language (e.g., R, Stata, Python). The code should be released under the MIT license, but we will consider other permissive licenses in special situations.”
13
14
As our computational reproducibility review improved, we would run the same code twice and compare the results. If these results did not match, we did not even attempt to compare with the manuscript.
15
We considered two numbers the same if they differed by less than 10−10. We acknowledge that this is an arbitrary standard, but this threshold was effective for us because most differences were either much larger or smaller. In other situations a different threshold might be reasonable, and we hope that community standards develop for when two numbers should be considered the same.
16
This low success rate roughly matches the rate for the Quarterly Journal of Political Science. From September 2012 to November 2015, the replication materials for 20 of the 24 accepted empirical articles were found to require some kind of modification. Furthermore, 14 of 24 accepted empirical articles were found to have discrepancies between the results generated by authors’ code and those in their articles (Eubank 2016). However, the journal International Interactions reports higher success rates. Between 2014 and 2016, they successfully replicated the results in 22 of 33 articles without needing to contact the authors (![]() ).
).
17
18
Precompiled R package binaries are available only for computers using Windows and MacOS.
19
For more on data sharing, privacy, and reidentification risk, we recommend Lundberg et al. (2019), Meyer (2018), Salganik (2018), and ![]() .
.
20
23
Given that defaults can have a big impact on behavior (![]() ), we think that journals could also consider a badging system that is opt-out. In other words, authors would be assumed to follow open-science practices, and badges would mark articles for which these practices are not followed. To the best of our knowledge, no journal currently uses this kind of opt-out badging system.
), we think that journals could also consider a badging system that is opt-out. In other words, authors would be assumed to follow open-science practices, and badges would mark articles for which these practices are not followed. To the best of our knowledge, no journal currently uses this kind of opt-out badging system.
24
We wish to emphasize that the people needed to do this additional work related to computational reproducibility will need skills that are already in high demand in research and industry. For example, doing the computational reproducibility work for this special issue required basic familiarity with bash scripting, Git, Docker, and cloud computing; in-depth familiarity with R and Python; and a strong background in statistics and machine learning. People with this skill set are currently in high demand, and we expect that for the foreseeable future, people who have the skills needed to computationally verify scientific research are likely to have many other demands on their time.
25
Some examples of tools that might be useful to other researchers are GUIdock (Hung et al. 2016), Algo-Run (Hosny et al. 2016), ReproZip (Chirigati et al. 2016)/ReproServer (Rampin et al. 2018), Singularity (Kurtzer et al. 2017), Rocker (Boettiger and Eddelbuettel 2017), Jug (Coelho 2017), archivist (Biecek and Kosiski, 2017), encapsulator (Pasquier et al. 2018), Binder (Project Jupyter et al. 2018), Packrat (Ushey et al. 2018), and Whole Tale (![]() ).
).
Supplemental Material
Supplemental material for this article is available online.
Author Biographies
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
