Abstract
Background
Bladder cancer is one of the most prevalent malignancies, and the mechanisms underlying its progression and the role of the tumor microenvironment (TME) are unclear. Recent advancements in single-cell RNA sequencing (scRNA-seq) and spatial transcriptomics (ST) enable detailed analysis of the cellular heterogeneity, gene expression, and cell–cell interactions in bladder diseases.
Methodology
We conducted a comprehensive search for recent articles that have investigated bladder diseases using scRNA-seq and ST.
Results
scRNA-seq and ST have led to significant discoveries in bladder disease research. These technologies have enabled the identification of multiple molecular subtypes within individual tumors and of the mechanisms of treatment resistance. Additionally, molecular differences based on gender have been explored, explaining the heterogeneity of the incidence and progression of bladder cancer. These findings deepen our understanding of the pathology of bladder diseases and highlight the transformative potential of scRNA-seq and ST in identifying novel biomarkers and therapeutic targets.
Conclusions
Integrating scRNA-seq and ST has considerably enhanced our understanding of tumor heterogeneity and the tumor microenvironment within tissues. These insights may lead to the development of personalized therapies and the improvement of patient outcomes. Several challenges, such as technical limitations and access difficulties, need to be addressed for the future clinical application of these technologies.
Keywords
Introduction
Bladder cancer is the tenth most common cancer worldwide. 1 The majority of bladder cancer cases (approximately 90%) originate from urothelial carcinoma, with other histological types, such as squamous cell carcinoma and adenocarcinoma, being less common. 2 Bladder cancer is about four times more common in men than in women. 3 Risk factors include smoking, family history, and exposure to chemical substances. 4 Bladder cancer is classified into non-muscle invasive bladder cancer (NMIBC) and muscle-invasive bladder cancer (MIBC), with the former making up about 75% of newly diagnosed cases. 5 Furthermore, 30–50% of patients with NMIBC experience recurrence or progression to MIBC. 6
The treatment options for localized bladder cancer include transurethral resection of bladder tumor (TURBT) and radical cystectomy. In metastatic cases, chemotherapy or immune checkpoint inhibitors (ICIs) are used, but the response rates are low and there is a need for further improvement. 7 Recurrence is an important issue in NMIBC, requiring patients to undergo regular follow-up with cystoscopy. TURBT is necessary again if recurrence is detected, imposing physical and psychological burdens on the patient. In cases of metastatic MIBC, cisplatin-based chemotherapy or ICIs are options. A large proportion of patients with bladder cancer experience side effects, and the response rate is low after completion of treatment, presenting another challenge.
The mechanisms underlying the development and recurrence of bladder cancer are not yet fully understood. Genetic alterations and epigenetic modifications influence the development of bladder cancer, and in recent years, molecular approaches have been increasingly employed in its diagnosis and pathological studies. DNA methylation is an epigenetic mechanism that alters gene expression without changing the underlying DNA sequence and represents an oncogenic process.8–10 In recent years, with the rise of next-generation sequencing (NGS), studies using scRNA-seq and ST have been increasing. scRNA-seq provides transcriptional profiles of individual cells, enabling more comprehensive analysis of tumor heterogeneity than conventional bulk RNA-seq.
11
However, because scRNA-seq lacks spatial information, ST was developed to enable comprehensive transcriptome analysis while preserving the spatial context. ST was awarded ‘Method of the Year 2020’ by
Overview of single-cell RNA sequencing and spatial transcriptomics
Single-cell RNA sequencing
scRNA-seq has revolutionized our understanding of cellular heterogeneity at the gene expression level, largely as a result of advances in next-generation sequencing technologies. 11 In addition to bladder cancer, other types of cancer exhibit marked cellular heterogeneity. 13 scRNA-seq enables visualization of the diversity of cells within the tumor microenvironment (TME) and assessment of cell–cell interactions. The TME is composed of cellular components such as immune infiltrating cells, stromal elements, and endothelial cells, as well as non-cellular components like the extracellular matrix and signaling molecules. 14 The interactions among these factors influence tumor growth and invasion. Unlike bulk RNA sequencing, which analyzes RNA extracted from whole tissues or cell populations and provides only the average gene expression profile, scRNA-seq enables the analysis of gene expression and signaling in small, distinct cell populations. 15 This has led to a sustained increase in research over the past 10 or so years. The scRNA-seq workflow involves the isolation of single cells, reverse transcription, cDNA amplification, and sequencing. Isolation methods include micropipetting, fluorescence-activated cell sorting (FACS), and microfluidic techniques. A frequently used scRNA-seq platform is 10x Chromium, which is based on microfluidic technology. This technique creates droplets known as gel beads in emulsion (GEMs), each containing a single cell. GEMs are mixed with reverse transcriptase, enabling mRNA capture and cDNA synthesis, followed by PCR amplification and library construction. 14
Multimodal approaches combining scRNA-seq with other sequencing methods, such as scATAC-seq or CITE-seq, have also been developed. 16 scATAC-seq assesses chromatin accessibility, providing insight into the epigenetic landscape of individual cells. CITE-seq simultaneously analyzes cell-surface proteins, facilitating the identification and classification of immune cells. 17 A limitation of scRNA-seq is the loss of spatial information regarding the tissue origins of the cells, but this can be addressed by integrating ST data.
Spatial transcriptomics
ST enables the visualization of gene expression within tissue sections, facilitating analysis of cellular composition and providing transcriptomic information in precise spatial locations. ST was introduced in 2016 by Lundeberg
Sequencing-based methods include those using
GeoMx Digital Spatial Profiling (DSP) by NanoString, a detection method based on ultraviolet (UV) irradiation, was launched in 2019. The tissue is visualized using fluorescent morphological markers, and regions of interest (ROIs) are defined. UV irradiation of the ROIs causes the release of oligonucleotides, which are collected and analyzed by next-generation sequencing (NGS) for molecular barcodes. DSP enables the analysis of the whole transcriptome as well as the proteome. 20
Xenium by 10× Genomics, which is based on
In imaging-based methods such as serial FISH, genes corresponding to mRNA molecules can be identified by repeatedly observing individual molecules under a microscope and attaching barcodes to the tissue. By detecting fluorescent dyes or color markers attached to these barcodes under a microscope, spatial information can be obtained. MERSCOPE by MERFISH has been commercialized and enables the detection of numerous RNA molecules at the single-cell level. However, it requires prior knowledge of probe design and has the limitation of not covering the entire transcriptome. 22
ST is useful for tissue analysis, and its integration with scRNA-seq enables localization at the single-cell level within tissue sections. ST has been used in research on breast, lung, colorectal, and prostate cancers.23–26 The main technical characteristics of these ST techniques are summarized in Table 1. This review focuses on how scRNA-seq and ST have been used to investigate bladder diseases (Figure 1). We searched PubMed for information on bladder diseases using the terms “single cell RNA sequencing or spatial transcriptomics” and “bladder cancer or urothelial cancer,” as well as “spatial transcriptomics” and “cystitis.” We extracted information on the ST tools used and the topics of the studies; the results are listed in Table 2.
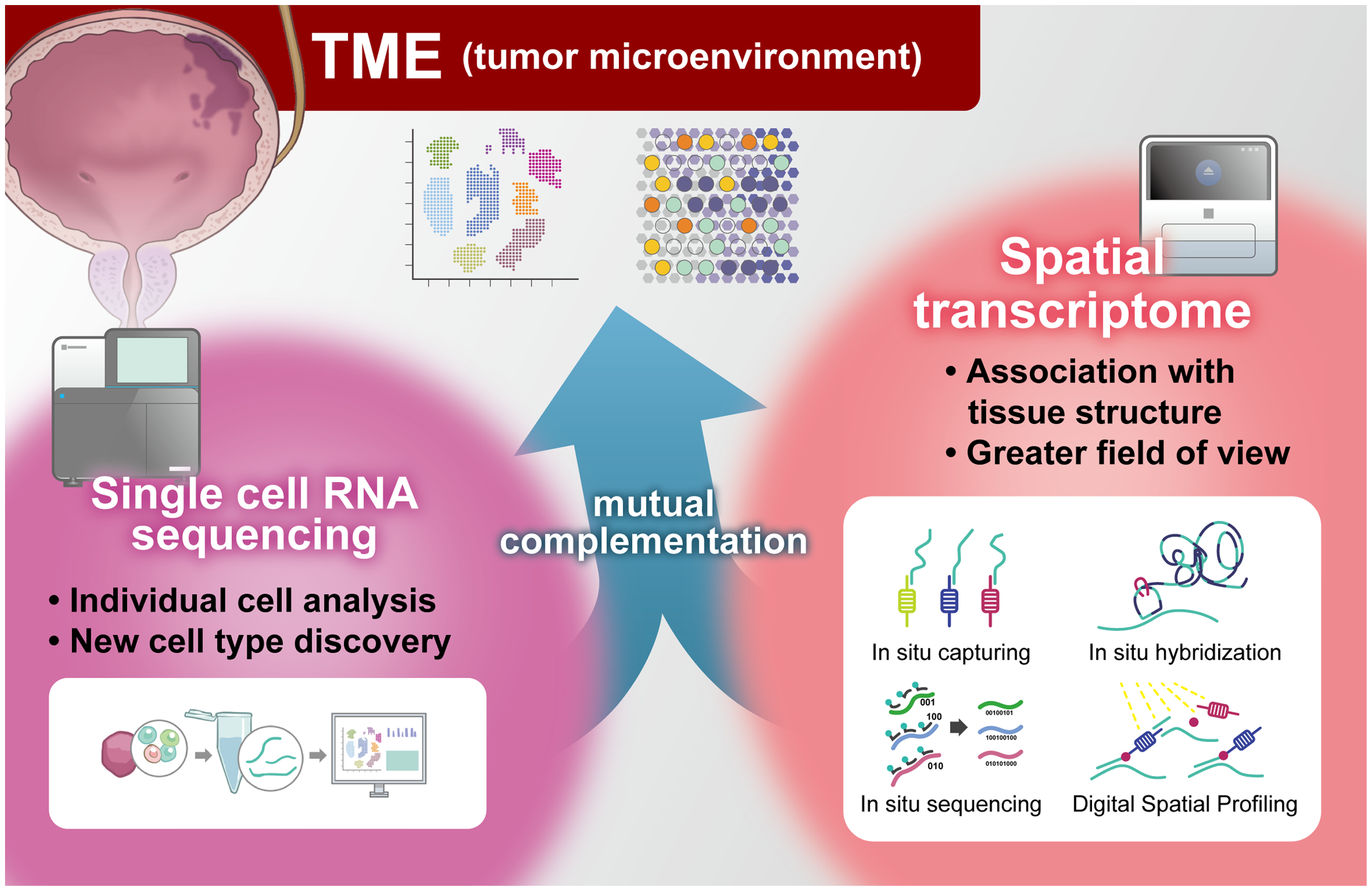
Integration of scRNA-seq and spatial transcriptomics to investigate the tumor microenvironment (TME). scRNA-seq and spatial transcriptomics (ST) enable investigation of the TME. scRNA-seq allows for the analysis of individual cells, providing insight into tumor heterogeneity, and ST allows analysis of gene expression in relation to tissue structures. Sequencing-based ST approaches include
Summary of advantages and disadvantages of spatial transcriptomics technology.
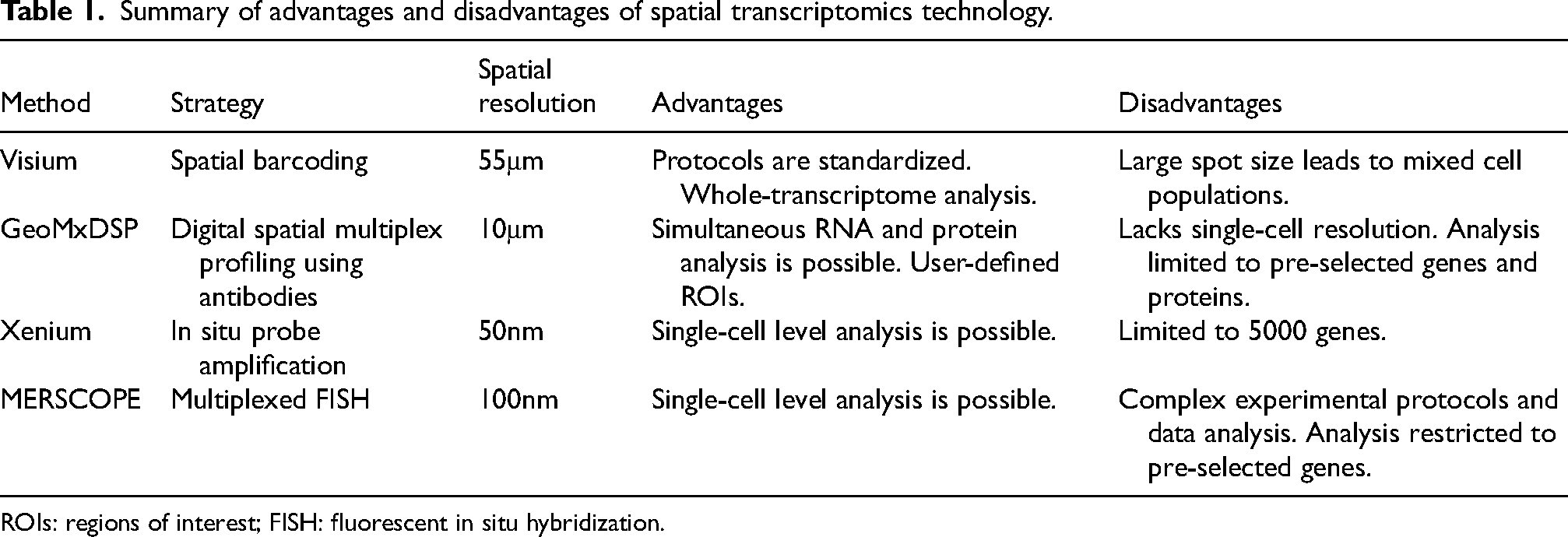
ROIs: regions of interest; FISH: fluorescent in situ hybridization.
Summary of spatial transcriptomics technology applications in bladder diseases.

IC/BPS: Interstitial Cystitis/ Bladder Pain Syndrome; NMIBC: Non-Muscle Invasive Bladder Cancer; MIBC: Muscle Invasive Bladder Cancer; DSP: Digital Spatial Profiling; Pem: Pembrolizumab; CAF: Cancer Associated Fibroblast; NAC: Neoadjuvant Chemotherapy; pCR: partial Complete Response.
Benign bladder diseases—interstitial cystitis/bladder pain syndrome
Interstitial cystitis/bladder pain syndrome (IC/BPS) is characterized by chronic pain related to the bladder and lower abdominal pressure or discomfort, accompanied by frequent urination and urgency, with the exclusion of other diseases. 27 IC/BPS is classified into Hunner-type interstitial cystitis (HIC), which is associated with Hunner lesions observed via cystoscopy, and non-lesion bladder pain syndrome. The prevalence of IC/BPS ranges from 0.01% to 2.3%, and women are five times more likely to develop the condition than men. 28 Histologically, HIC is characterized by urothelial exfoliation, significant infiltration of inflammatory cells (including lymphocytes and plasma cells), and clonal expansion of infiltrating lymphocytes. 29 The differences between HIC and non-Hunner interstitial cystitis (NHIC) have gained attention. The bladder tissues of patients with IC/BPS show activated Th1 responses and a lower number of Tregs, promoting the conversion of Tregs to Th1 cells; furthermore, in HIC, there are higher numbers of M2 macrophages and the presence of inflammatory fibroblasts, with the main cell–cell interaction being the CXCL2-ACKR1 pathway. 29
The use of scRNA-seq and ST in combination has revealed that immune cells in patients with IC are located near the urothelial regions or fibroblasts. By contrast, in normal bladder mucosa, these cells are around the urothelium and smooth muscle cells. 30 Additionally, 95% of patients with IC are positive for human polyomavirus. A number of genes associated with infection with several viruses, such as HIV and HPV, have been identified in the epithelial cells of patients with IC. This suggests that viral infections may be a cause of IC.
Bladder cancer cells: subtypes and treatment classification
Bladder cancer subtypes
Bladder cancer is classified into NMIBC and MIBC. Due to high tumor heterogeneity, molecular-level subtypes have been developed to compare among patients. The widespread use of NGS has enabled a molecular-based approach to tumors, rather than the conventional histopathological classification.
In UROMOL2021, NMIBC was classified into Class 1, 2a, 2b, and 3 based on bulk RNA sequencing. Class 2a was significantly associated with T1 tumors, high-grade tumors, and carcinoma
Subtypes of MIBC based on bulk RNA-seq have been established. The consensus molecular classification divides MIBC into six subtypes: luminal-papillary (LumP), luminal-non-specified (LumNS), luminal-unstable (LumU), stroma-rich, basal/squamous (Ba/Sq), and neuroendocrine-like (NE-like). LumP tumors often occur in younger patients and are less aggressive, with frequent mutations in FGFR3 and KDM6A. LumNS is common in older patients, has abundant fibroblast stromal infiltration, and shows a high frequency of ELF3 mutation. LumU tumors have high cell cycle activity and APOBEC signatures. Stroma-rich tumors show significant infiltration of smooth muscle and fibroblasts. Ba/Sq tumors have a high rate of mutations in TP53 and RB1, with infiltration by CD8+ T cells and NK cells. NE-like tumors also show frequent TP53 and RB1 mutations and have the worst prognosis.32,33
Investigation of molecular subtypes in the UROMOL2021 cohort using scRNA-seq and ST revealed the coexistence of multiple molecular subtypes in individual tumors. 34 In a cohort of 48 cases of NMIBC and MIBC, the scRNA-seq-defined subtypes corresponded to the bulk RNA-seq-defined subtypes in 79% of NMIBC tumors and 67% of MIBC tumors. 34 These are high concordance rates, but several subtypes are present in each tumor. Therefore, the intratumor subtype heterogeneity needs to be considered.
NMIBC
NMIBC is diagnosed via TURBT. According to current guidelines, high-risk NMIBC should be managed with a combination of a second TURBT and intravesical Bacillus Calmette-Guerin (BCG) therapy. 35 However, some patients treated with BCG experience recurrence or progression to MIBC. 6 Tumor heterogeneity and poor response to BCG therapy in patients with tumors of various grades and those with recurrent disease have been investigated using scRNA-seq.
In one study that evaluated BCG resistance, the transcriptomes before and after treatment with intravesical pembrolizumab plus BCG were compared by ST. 36 After treatment, the tumor area of interest (AOI) showed enrichment of pathways related to immune modulation and cytokine signaling. In the TME, there were significant increases in the numbers of NK cells, CD8+ T cells, and Tregs. The early recurrent TME had a high stromal score, enriched angiogenesis, and increased expression of exhaustion markers before and after treatment. BCG-resistant tumors enhance lymphocyte exhaustion, so they respond better to intravenous administration of drugs. 37
Localized MIBC
Current guidelines recommend radical cystectomy as a treatment option for cases of repeatedly recurrence after BCG therapy or that have progressed to MIBC. 6 Neoadjuvant chemotherapy (NAC) is recommended for this procedure, 38 and the treatment response to NAC has been analyzed by ST.39–41 ST has been used to compare patients with NAC who had a pathological complete response (pCR) or not (non-pCR) after surgery. In pCR patients, the TME was rich in B cells and CD3+ T cells. By contrast, non-pCR patients had increased numbers of ARG1+ cells and cells related to immune checkpoints (such as LAG3, ICOS, and STING), which are characteristics of an immunosuppressive microenvironment. 41
scRNA-seq and ST have been used to identify subpopulations of epithelial cells, including a subset with high CDH12 expression, representing undifferentiated epithelial cells in patients with a poor prognosis. A subset of epithelial cells expressing CDH12, β-catenin, and other epithelial markers has been identified. CDH12-rich bladder cancers show elevated PD-L1/L2 expression and are surrounded by exhausted T cells and other immune cells. These cancers are associated with a poor response to NAC but high sensitivity to immunotherapy. 39
In a cohort of bladder tumors treated with pembrolizumab preoperatively (PURE01), integrated bulk RNA-seq and ST analysis identified five subtypes. In the luminal-excluded subtype, which had a poor response rate, FGFR3 mutations were frequent and KDM5B was identified as a suppressor of tumor immune signaling. The inhibition of KDM5B enhanced the immunogenicity of FGFR3-mutant bladder cancer cells, indicating the usefulness of combination therapies. 40
Metastatic MIBC
Systemic therapies such as chemotherapy and ICIs are options for metastatic bladder cancer. However, such cancer shows tumor heterogeneity and a poor response to systemic therapies, resulting in a low 5-year survival rate. Therefore, new therapeutic targets are under investigation.
In a study of MIBC that used scRNA-seq, the inhibition of CD39 in mice led to the proliferation of immune cells such as CD8+ T cells, NK cells, and cDC1 dendritic cells, resulting in suppression of the growth of bladder cancer. CD39, a transmembrane protein encoded by ENTPD1, converted ATP and ADP into adenosine (ADO). The inhibition of CD39 had a synergistic effect with cisplatin, indicating its potential as a therapeutic target. 42
scRNA-seq has been used to identify IL-6 as a correlate of poor response to anti-PD-L1 therapy in bladder cancer. IL-6-STAT signaling inhibits the differentiation of cytotoxic T lymphocytes, and in mouse models, combined inhibition of PD-L1 and IL-6 results in a greater reduction in tumor size than anti-PD-L1 monotherapy. This suggests that IL-6 has potential as a therapeutic target. 43
scRNA-seq has also identified LRFN2 as a novel immune-suppressive target in MIBC. LRFN2 reduces the recruitment of CD8+ T cells by decreasing the levels of proinflammatory cytokines and chemokines, leading to spatial exclusion of tumor and immune cells. Knockout of LRFN2 in a mouse model increased the effectiveness of ICIs therapy, making LRFN2 a potential target for combination therapy with ICIs. 44
Cancer-associated fibroblasts (CAFs)
Based on scRNA-seq of the expression of RGS5 and PDGFRA, cancer-associated fibroblasts (CAFs) have been broadly classified as myofibroblastic CAFs (mCAFs) and inflammatory CAFs (iCAFs). iCAFs are characterized by a high expression of chemokines and are enriched for genes involved in ECM degradation, cell-migration regulation, and angiogenesis. By contrast, mCAFs show high expression levels of genes related to ECM and regulation of focal adhesions. 45
CAFs in NMIBC
ICAM1+ iCAFs have been identified in recurrent bladder cancer via scRNA-seq. ICAM1+ iCAFs secrete CCL2, which binds to CCR2 on myeloid-derived suppressor cells (MDSCs) enriched in THBS1+ monocytes. 46
Cancer stem cells (CSCs) may contribute to the recurrence and treatment resistance of cancer. In bladder cancer, CSCs express markers such as CD44, ALDH1A1, and SOX4. 47 In recurrent bladder cancer, ICAM1+ iCAFs interact with the CSC ligand CD44 via the FGF2 receptor. 46 Another subtype of iCAFs, ITGA11+ CAFs, has been identified by scRNA-seq and ST. These cells interact with the surface receptor SELE on lymphatic endothelial cells via ITGA11, promoting lymphangiogenesis. Furthermore, ITGA11+ CAFs express CHI3L1, which is implicated in remodeling the extracellular matrix (ECM), thereby increasing its permeability to cancer cells and facilitating lymphovascular invasion and lymph node metastasis. In a study on a mouse model, the combined inhibition of ITGA11 and CHI3L1 resulted in a significant reduction in lymph node metastasis. 48
CAFs in MIBC
The following reports on CAFs in MIBC using scRNA-seq are available. iCAFs are involved in the production of CXCL12, which interacts with CXCR4 on, and thereby recruits, tumor-associated macrophages (TAMs). 45 Among other CAF subsets, interferon-regulated CAFs (irCAFs) have been found between the differentiation trajectories of iCAFs and mCAFs, and they express the urea transporter SLC14A1 on their surface. By secreting WNT5A, irCAFs promote the adoption of a stem cell phenotype by tumor cells. Activation of the cGAS-STING pathway contributes to the synthesis of IFNs, which induces SLC14A1+ CAF formation. High expression levels of irCAFs are associated with poor responses to NAC and immunotherapy, as well as poor outcomes. 49
COL11A1+ fibroblasts are a CAF subset present in tumors but not in normal fibroblasts. These cells express LRC15, ITGA11, SPHK1, and FAP, thereby promoting ECM remodeling and inhibiting antitumor immune responses by upregulating MIF. MIF induces the secretion of OPN/SPP1 by macrophages, promoting tumor progression. High levels of COL11A1+ CAFs are associated with poor prognoses of bladder and lung adenocarcinomas. 50 An analysis of CAF-related genes identified LRP1, ANXA5, SERPINE2, ECM1, RBP1, GJA1, and FKBP10 as prognostic markers. Patients in a high-risk group with elevated expression levels of these genes exhibited poor responses to immunotherapy. 51 ST has shown that recurrent MIBC tumors have higher tumor heterogeneity and strong interactions between epithelial and immune cells. Spatial analysis has revealed the proximity of CD8+ T cells to plasma cells at the tumor boundary, with minimal presence of monocytes, dendritic cells, and mast cells. In recurrent tumors, COL4A1–SDC1 interactions are increased between CAFs and malignant or immune cells. 52 In summary, CAFs are involved in the progression and treatment resistance of bladder cancer, and certain subtypes may be biomarkers for the prognosis and treatment response.
Immune cells
Regarding immune cells, TAMs, cytotoxic T cells, B cells, and dendritic cells have been implicated in the formation of the TME. 53 scRNA-seq has been used to investigate the composition and functions of immune cells in the TME.
TAMs typically consist of M2 macrophages, which are implicated in the transition to immunosuppression in a manner involving surface CD276. Inhibiting CD276 stimulates T-cell activation, and so combination with anti-PD-1 therapies may enhance the efficacy of CD276 inhibitors. 54 Additionally, a high mRNA level of macrophage-derived MMP12 is associated with decreased overall survival in many types of cancer. In bladder cancer, higher MMP12 expression is correlated with increased malignancy. 55 TAM-related genes such as TBXAS1, GYPC, HPGDS, GAB3, ADORA3, and FOLR2 are specifically expressed by M2 macrophages and are suggested to be involved in TAM polarization. 56 The M1/M2 ratio is positively associated with better prognosis in bladder cancer, while EMP1 promotes tumor invasion and metastasis through EMT and focal adhesion (FA). EMP1 may interact with macrophages and tumor cells through the FN1-SDC1 pathway. 57 In subsets of myeloid phagocytes different from macrophages, the Msc2IR score, which predicts the response to immunotherapy based on the ratio of immune response gene signatures to tumorigenic inflammatory signatures, has been inferred from scRNA-seq data. 58
The use of scRNA-seq and TCR-seq in combination has identified not only cytotoxic CD8+ T cells but also clonally expanded cytotoxic CD4+ T cells and tumor-specific Tregs. CD4+ T cells kill autologous tumor cells but are inhibited by Tregs. 59 The abundance of several subsets of T cells, including exhausted T cells, exhausted NKT cells, Ki67+ T cells, and B cell-like T cells, is high in bladder tumors, and their presence is associated with a poor prognosis. 60 Potential biomarkers for exhausted T cells include IKZF3 and TRGC2. PD1hi CD200hi CD4+ and PD1hi CD200low CD4+ subpopulations have been identified among CD4+ exhausted T cells. PD1hi CD200hi cells are resistant to immunotherapy and promote the EMT and angiogenesis. Malignant cells recruit PD1hi CD200hi CD4+ T cells with which they communicate via the m6A-mediated GAS6-AXL axis. 61
In B-cell MIBC subtypes, IFN-stimulated B-cell subtypes (B-ISG15) and germinal center-related B-cell subtypes (B-LMO2, B-STMN1) are significantly increased. Tertiary lymphoid structures (TLSs) are follicular structures characteristic of germinal centers and have CD19+ B cells in the central region surrounded by T cells. Their presence is associated with an improved survival rate in patients with MIBC. 62 Furthermore, high B-cell infiltration is associated with better prognosis; in one study, a B-cell gene signature constructed from bladder cancer scRNA-seq data predicted favorable outcomes in 21 of 33 types of cancers. 63
Plasma cells (PCs) are associated with tumor cells, and the ligand-receptor pairs LAMB3/CD44 and ANGPTL4/SDC1 are linked to a poor OS and non-response to immunotherapy in ST analysis. Plasma cells can be divided into IgG1 and IgA1 subtypes; IgG1 PCs are associated with immune responses and IgA1 PCs are related to cell cycle pathways. High-grade bladder cancers have greater infiltration of IgA1 PCs and enrichment of the MK and MIF signaling pathways. 64 ST has been used to construct a tumor-specific T-cell signature (TstcSig) based on related genes to predict the outcomes of bladder cancer immunotherapy. Tumor-specific CD4+ T cells show increased expression of CXCL13 and T-cell exhaustion markers, whereas CD8+ T cells have high expression levels of cytotoxic-related genes and immunoregulatory factors. The expression levels of VAMP5, TIGIT, LCK, CD27, and CACYBP are positively associated with T-cell infiltration. 65
Comparisons of tumor, immune infiltration, and stromal regions by ST have revealed distinct gene signatures and enriched pathways in each region. The tumor ROI is enriched for genes involved in translation elongation and EIF2 signaling, whereas the stromal ROI is rich in collagen-remodeling genes and ECM gene sets. Immune-infiltration ROI is enriched for neutrophil degranulation genes (such as CXCL8), MHC class I processing, and the HER2 signaling pathway. 66
In summary, the immune microenvironment of bladder cancer is made up of a variety of immune cell types, which are involved in tumor progression and treatment responses. Certain immune cell subtypes and gene signatures may be useful biomarkers for predicting prognosis and planning treatment.
Gender differences in bladder cancer
Bladder cancer is more common in men, with a male-to-female ratio of approximately 4:1. Women are 40 times more likely to develop urinary tract infections than men 67 and may be more likely to have locally advanced bladder tumors at the time of diagnosis. Smoking is a risk factor, and may influence these statistics, but women also tend to have higher tumor grades, implicating molecular biological factors. Although the influence of sex hormones is known, 68 the molecular mechanisms underlying gender differences in bladder cancer are unclear. A study that used scRNA-seq and ST explored the influence of the TME on tumor tissues in both genders. 69 scRNA-seq found that, in females, genes related to responses to bacterial stimuli and intrinsic apoptotic signaling were upregulated. In males, genes associated with ribosome biogenesis were upregulated. Urothelial cells in the S and G2/M phases were more prevalent in males (55%) than in females (35.2%), and the transcription factor MYC was more active in the urothelial cells of males. This suggests that the higher cell proliferation activity in males protects the urothelial barrier against bacterial invasion. Increased ribosome biogenesis may reduce the binding of ribosomal proteins to MDM2, increasing the binding of the latter to p53, which promotes p53 degradation and may increase the risk of tumor formation in males. Additionally, fibroblasts in females had higher expression levels of collagen genes.
In a mouse study of bladder cancer using scRNA-seq, metabolic pathways such as glycolysis and gluconeogenesis, as well as PKA signaling, were more active in males. By contrast, thrombin, endocannabinoid tumor suppression, and oxytocin signaling pathways were more active in females. In male CAFs, PI3 K/AKT signaling was upregulated, and cell–cell signaling mediated by PTN was abundant, with TLR4 being more active upstream. In females, the tumor-suppressor factors EGR2 and EGR3 were more active, potentially explaining the lower incidence of bladder cancer in women. 70 In another study, scRNA-seq indicated that CD163+ macrophages and B cells were more prevalent in female patients with NMIBC. Costimulation of M2-like macrophages by bacterial lipopolysaccharide and IL-10 induced the production of CXCL13, a chemokine critical for B-cell recruitment. 71 Additionally, the expression of HAVCR1, which is linked to IL-10 production, is increased in both genders. Larger numbers of CD163+ macrophages and B cells are associated with worse recurrence-free survival in males and females. 72
About 10–40% of male mice show the loss of the Y chromosome (LOY), and ST has revealed that LOY is associated with poor prognosis in patients with bladder cancer. Y− tumors show greater proliferation than Y+ tumors. In immunocompromised mice, Y− tumors proliferate in a T-cell-dependent manner. Integration of scRNA-seq and ST has revealed increased expression of exhausted T-cell markers such as TOX and TIM3 in Y− tumors, with improved responsiveness to ICIs. 73
Regarding gender differences, scRNA-seq and ST have shown that males with bladder cancer have higher expression levels of genes related to tumor formation and cell proliferation. By contrast, females have increased expression levels of tumor suppressor factors, suggesting a difference in incidence. Additionally, LOY is associated with a poor prognosis and is related to the effectiveness of ICIs.
Future perspectives and challenges
Bladder cancer is a highly prevalent disease characterized by considerable cellular heterogeneity. Here we reviewed the latest approaches to the analysis of bladder diseases, focusing on the combination of ST and scRNA-seq.
The development of ST from scRNA-seq has enabled visualization of the spatial organization of tissues and cells. Spatially localizing tumors is crucial for assessing tumor formation and disease progression, which may provide insight into the characteristics of bladder cancer. Technologies such as Visium and GeoMx DSP are frequently used in ST studies of bladder diseases, and these techniques enable the visualization of major spatial structures. However, the spatial resolution provided by these technologies is limited, as the analyzed spots often contain mixed cell populations. Therefore, they are typically used in conjunction with scRNA-seq to enhance analytical precision. Integration scRNA-seq and ST can address these limitations, and several tools have been developed for this purpose. Tools like RCTD and CARD enable us to predict the types and proportions of cells present in mixed populations derived from ST through deconvolution.74,75 However, tools for integrating other omics data remain limited, presenting a challenge for future development.
The software provided with Visium and DSP enables data analysis and visualization, allowing spatial mapping. However, for other ST tools, the computational methods for data analysis are highly complex, requiring advanced knowledge of statistics and mathematical algorithms, highlighting the need for standardized and practical software solutions. To enhance the accuracy and reproducibility of tissue management and analysis, it is essential to establish clear and standardized protocols. Another challenge is the high cost of reagents and equipment per sample. Additionally, a limitation of scRNA-seq and ST is the small sample size, as increasing the number of samples would require higher costs. Furthermore, due to the difficulty in collecting sequential samples, the lack of untreated cases makes it impossible to distinguish between treatment effects and the natural progression of the disease. Considering that genome and transcriptome analyses, which were previously very expensive, have become more affordable with technological advancements, the development of cost-effective techniques and protocols for scRNA-seq and ST is anticipated to further improve accessibility.
Results obtained from scRNA-seq and ST have the potential to be translated into clinical applications, including classifying bladder cancer subtypes, predicting treatment responses, and identifying novel biomarkers for personalized therapy. Insights from exploratory research using scRNA-seq and ST could be simplified for practical use by employing immunohistochemistry to classify subtypes of bladder cancer more conveniently. Furthermore, if molecular targets related to treatment resistance and liquid biopsy markers essential for personalized therapy are identified, the need to perform scRNA-seq and ST on all clinical specimens could be significantly reduced. In this context, scRNA-seq and ST serve as powerful tools in preclinical research, paving the way for groundbreaking discoveries that are expected to advance the treatment of bladder cancer. Notably, a clinical trial (NCT06373055) is currently investigating the use of ST to assess the effects of NAC in MIBC. This trial aims to explore whether there are distinct microenvironmental characteristics in MIBC cases that respond exceptionally well or poorly to NAC. Looking ahead, it is anticipated that further clinical applications of these techniques will be realized.
Conclusions
We summarize the advancements in scRNA-seq and ST technologies related to bladder diseases. The continuous development of scRNA-seq and scRNA-seq-based ST from bulk RNA-seq has enabled the spatial visualization of tissues and cells. These techniques enable the identification of genes associated with the pathogenesis of TME-mediated bladder diseases, thereby providing mechanistic insight and clinically meaningful information. However, cost and accessibility, as well as the statistical and mathematical complexity of ST data, are challenges that must be overcome to facilitate the use of these techniques in clinical practice.
Footnotes
Acknowledgments
This study was supported by a Translational Research Grant for Urological Oncology.
Authors’ contributions
KY conducted the review of the literature and wrote the first draft of the manuscript. All authors contributed to the conception of the study and the interpretation of data. All authors approved the final version of the manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by JUA Science Award.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
