Abstract
Background:
Epigenetic mechanisms play vital roles in the activation, differentiation, and effector function of immune cells. The breast and kidney-expressed chemokine (CXCL14) mainly contributes to the regulation of immune cells. However, its role in shaping the tumor immune microenvironment (TIME) is yet to be elucidated in renal cell carcinoma (RCC).
Objectives:
This study aimed to elucidate the role of CXCL14 in predicting the efficacy of immunotherapy in patients with RCC.
Methods:
CXCL14 expression and RNA-sequencing, single-cell RNA-sequencing (scRNA-seq), and survival datasets of RCC from public databases were analyzed, and survival was compared between different CXCL14 levels. The correlation between CXCL14 and immune infiltration and human leukocyte antigen (HLA) gene expression was analyzed with TIMER2.0 and gene expression profiling interactive analysis. Institutional scRNA-seq and immunohistochemical staining analyses were used to verify the relationship between CXCL14 expression level and the efficacy of immunotherapy.
Results:
CXCL14 was expressed in fibroblast and malignant cells in RCC, and higher expression was associated with better survival. Enrichment analysis revealed that CXCL14 is involved in immune activation, primarily in antigen procession, antigen presentation, and major histocompatibility complex assemble. CXCL14 expression was positively correlated with T-cell infiltration as well as HLA-related gene expression. Among the RCC cohort receiving nivolumab in Checkmate 025, the patients with CXCL14 high expression had better overall survival than those with CXCL14 low expression after immunotherapy. scRNA-seq revealed a cluster of CXCL14+ fibroblast in immunotherapy responders. Immunohistochemistry analysis verified that the patients with high CXCL14 expression had an increased proportion of high CD8 expression simultaneously. The expression level of CXCL14 was associated with CXCR4 expression in RCC.
Conclusion:
CXCL14 expression is associated with immunotherapy response in RCC. It is a promising biomarker for immunotherapy response prediction and may be an effective epigenetic modulator in combination with immunotherapy approaches.
Plain language summary
Kidney-expressed chemokine (CXCL14) regulates immune cells. We studied how it affects the body’s immune response to kidney cancer based on public and private database and staining. We found that higher levels of CXCL14 in kidney cancer were linked to better patient survival. CXCL14 seems to help activate the immune system. When patients with high CXCL14 levels received immunotherapy, they tended to survive longer than those with low levels. Fibroblasts with CXCL14 were present in patients responding to immunotherapy. Further tests confirmed that high CXCL14 levels were related to more immune cells. CXCR4 may be its receptor in kidney cancer. This suggests that measuring CXCL14 levels could help predict how well a patient might respond to immunotherapy for kidney cancer.
Keywords
Introduction
Each year, there are more than 400,000 new cases of kidney cancer, and renal cell carcinoma (RCC) is the most prevalent type. The survival of metastatic RCC is poor, with only 10% of patients living 5 years after diagnosis.1,2 In recent years, the advent of immunotherapy targeting immune checkpoints has largely reshaped the treatment paradigm of metastatic RCC and remarkably prolonged the survival of this fatal disease. However, heterogeneous responses to immunotherapy are common, and the prognosis of non-responders is generally poor.3–5 Although emerging biomarkers such as sCD27 and the H&E score of tumor-infiltrating immune cells have shown promising efficacy,6,7 no well-established biomarkers could reliably predict response to immunotherapy in RCC. Therefore, it is imperative to identify novel biomarkers to assist in potential patient selection.
Emerging evidence indicates that the tumor immune microenvironment (TIME) is a key factor in determining the efficacy of immunotherapy in tumor treatment, 8 and epigenetic mechanisms play important roles in immune cells within the TIME. CXC and CC chemokines play significant roles in the tumor microenvironment, influencing various aspects of solid tumor progression, including angiogenesis, immune cell infiltration, metastasis, tumor microenvironment, and therapeutic targets. 9 CXCL14, which is also called breast and kidney-expressed chemokine, belongs to the CXC subfamily of chemokines and is expressed widely and highly in normal tissue.10–12 It is related to infiltration of immune cells, cell mobilization, maturation of dendritic cells, and increased expression of the major histocompatibility complex I (MHC-I) molecules.13–15 Although the prognostic value of CXCL14 is inconsistent among different cancers,16–20 growing evidence demonstrates its place in the regulation of TIME. In glioma, the expression of CXCL14 by tumor cells has been shown to enhance the chemotaxis of cytotoxic T cells, boost CD8+ T-cell response, and improve overall survival (OS). 21 Another report revealed that CXCL14 could influence DC maturation through NF-κB signaling activation in head and neck squamous cell carcinoma. 22
In RCC, a previous study revealed that CXCL14 could induce apoptosis. 23 Given the interplay between CXCL14 and TIME, this chemokine has the potential to serve as a biomarker for assessing the response to immunotherapy in the treatment of RCC. However, the actual distribution of CXCL14 in tissue and its association with treatment response to immunotherapy in RCC remain unclear, due to the lack of further analysis. Nowadays, single-cell RNA sequencing (scRNA-seq) is an effective technique for uncovering the complex molecular features of the TIME at the level of a single cell.
Therefore, in this study, we aimed to explore the interplay between CXCL14 expression and TIME, as well as its predictive value for immunotherapy in metastatic RCC.
Methods
RNA analysis of CXCL14 using public datasets
We obtained data on CXCL14 mRNA expression by analyzing the RNA-seq gene expression matrix of clear cell renal cell carcinoma (ccRCC) and papillary renal cell carcinoma (pRCC) from The Cancer Genome Atlas (TCGA). Patients with incomplete clinicopathological data were excluded. CXCL14 expression levels in normal tissue and tumor tissue in TCGA ccRCC and pRCC were analyzed using the GEPIA2 web tool (http://tisch.comp-genomics.org/home/). Raw counts of RNA sequencing were log2-transformed into log2 transcripts per million (TPM) values. We analyzed disease-specific survival (DSS) and OS data from the TCGA RCC dataset to evaluate the predictive significance of CXCL14. The R software package ‘survminer’ was used to produce survival curves for CXCL14 mRNA expression with optimal cutoffs, as well as to evaluate OS and DSS. The patients were classified into two groups based on CXCL14 mRNA expression levels, using the quartile method. To explore the value of CXCL14 for immunotherapy, the OS of CXCL14 high and low groups was compared among RCC patients receiving nivolumab in the Checkmate 025 trial. 24 Kaplan–Meier survival curves were constructed to illustrate the disparity in survival duration.
Immune infiltration analysis
To explore the potential impact of CXCL14 on immune infiltration, we conducted an analysis of the correlation between CXCL14 expression levels and T-cell infiltration using the TIMER2.0 database. The immunedeconv R package, which integrates advanced algorithms (xCell, MCP-counter, TIMER, EPIC, CIBERSORT, and quanTIseq), is utilized by the TIMER2.0 database. 25 For the correlation analysis, we considered a Spearman’s r value greater than 0 with a p value <0.05 as indicative of a positive correlation, while a Spearman’s r value less than 0 with a p value <0.05 was considered indicative of a negative correlation, after adjusting for tumor purity.
Gene correlation analysis in GEPIA
We utilized the online platform gene expression profiling interactive analysis (GEPIA) (http://gepia.cancer-pku.cn/index.html) to identify the genes significantly correlated with our target gene. GEPIA 26 is an online tool for RNA expression analysis, which integrates data from TCGA and the GTEx projects and includes a vast collection of tumor and normal samples. This study employed GEPIA to create survival curves for OS and DSS, using gene expression levels, and statistical analysis was performed using the log-rank test. We conducted gene expression correlation analysis using the given sets of TCGA expression data and investigated the association between CXCL14 expression levels and human leukocyte antigen class I (HLA-I) class (HLA-A, HLA-B, and HLA-C) and HLA-II class (HLA-DPA1, HLA-DQA1, HLA-DQB1, HLA-DRA, HLA-DRB1, and HLA-DRB2) gene expression levels using the GEPIA web tool. The correlation between CXCL14 and HLA RNA-Seq (expressed as TPM expression value) was analyzed with Spearman’s correlation coefficient.
Differentially expressed genes and function pathway analysis
Differentially expressed genes (DEGs) were detected through the ‘limma’ package in R, utilizing criteria of an adjusted p value <0.05, baseMean >50, and |logFC| >0.5. We defined CXCL14 high and low expression groups with the quartile method according to CXCL14 TPM value. DEGs were analyzed with R package DESeq2. After calculating DEGs, we got a list of upregulated genes in the CXCL14 high expression group. To identify disparities in biological functions between the CXCL14-high and CXCL14-low groups, Kyoto Encyclopedia of Genes and Genomes (KEGG), Gene Set Enrichment Analysis (GSEA), and Gene Ontology (GO) enrichment analysis were performed. For conducting GO analysis, we employed the enrichGO function of the R package ‘clusterProfiler’, and we derived GO annotations from genome-wide annotation packages (org.Hs.eg.db). 27 Biological processes with a p value <0.05 were selected for further analysis. By performing GSEA to the gene list, we ranked the genes according to a moderated t-statistic, which compared patients with high and low CXCL14 expression. normalized enrichment score (NES) represents the normalized enrichment score.
Single-cell analysis
To conduct scRNA-seq, we obtained raw data for GSE111360 and GSE171306 from the GEO database and processed the data using the Seurat package in R with R Studio. 28 We utilized the Tumor Immune Single-cell Hub database to investigate the correlation between CXCL14 expression levels and the infiltration of immune cells. 29 Furthermore, we conducted validation by performing scRNA-seq on four samples of RCC from the Sun Yat-sen University Cancer Center (SYSUCC). Fresh tissue samples were obtained from four patients with non-ccRCC (nccRCC) after immune checkpoint inhibitors (ICI)-based treatment. Three of the samples were obtained from lymph nodes, while the remaining sample was taken from a site on the abdominal wall. The raw sequencing fastq files were aligned to the GRCh38 human genome reference genome using cellranger V6.0.2 software, resulting in a gene expression matrix. Cells were selected under the following conditions: (1) cells not expressed more than 2500 genes or less than 200 genes; (2) mitochondrial count ratio in a cell less than 0.2; and (3) belong to non-doublet cells. R package Seurat was used for analysis in the next steps. The gene expression count matrix was normalized by the ‘log’ method. The matrix is scaled with the ScaleData function in the Seurat R package (Default Parameters). Defining features of highly variable genes in this matrix was performed with the Seurat FindVariableGenes function. The RunPCA function was utilized to conduct principal component analysis. Nearest-neighbor graph construction was performed on the FindNeighbors function (dims.use = 1:15). Cell clusters were formed by the FindClusters function (resolution = 0.5). The R package Rtsne was used for t-Distributed Stochastic Neighbor Embedding (tSNE) dimension reduction analysis of all single-cell transcriptome data. We defined each cell cluster according to known marker genes.
Immunotherapy response prediction
To investigate the potential predictive value of CXCL14 for immunotherapy, we analyzed 120 patients with ccRCC who had both treatment response information and transcriptomic data from the Checkmate 025 cohort. Patients were stratified into two cohorts based on CXCL14 mRNA expression levels using the quartile method. In addition, a validation dataset of 22 RCC patients from SYSUCC was used to verify the predictive value of CXCL14 on the efficacy of immunotherapy. All patients were identified using pathology reports, and experienced uropathologists reviewed all cases. The treatment responses were assessed based on the Response Evaluation Criteria in Solid Tumors version 1.1. 30 The response to ICI treatment was defined as complete response (CR), partial response (PR), or stable disease (SD). Non-response was defined as the progression disease (PD) after ICI treatment. The disease control rate (DCR) was calculated as the total proportion of patients who achieved either CR, PR, or SD following ICI treatment.
Tissue specimens and the immunohistochemistry staining
The collection of tissue samples for this study was conducted in accordance with the ethical standards of the Helsinki Declaration. All patients who participated in this study were informed about the purpose, procedures, and potential risks and benefits of the research, and provided their written informed consent before any study-related procedures were conducted. The samples were evaluated independently by two uropathologists. The uropathologists conducted a secondary review of cases with interobserver disagreements, and any remaining discrepancies were resolved through mutual consensus. The samples collected were immediately frozen in liquid nitrogen and stored in a −80°C freezer for further analysis. Follow-up was conducted every 3 months through phone calls or visits by our team to inquire about the patient’s survival status until either death or the end of the follow-up period. The formalin-fixed paraffin-embedded tissue blocks were cut into 5-mm-thick sections, which were then subjected to immunohistochemical (IHC) staining. CXCL14 antibody (Abcam, Shanghai, China, 1:500 dilution), CXCR4 antibody (Bioss, Beijing, China, 1:500 dilution), and CD8 antibody (Ascend Biology, China, 1:100 dilution) were used to stain these tissues. Normal human kidney tissue and human placenta tissue were used for primary antibodies as positive controls. Human kidney cancer tissues with secondary antibodies alone were used as negative controls. IHC visualization was performed using a Jiangfeng automatic slide scanner (KF-PRO-020, Ningbo Jiangfeng Bioinformation Technology Co., Ltd., Ningbo, China). The cutoff point for CXCL14, CXCR4, and CD8 expression was determined concerning previous literature. 31 The staining intensity was graded on a scale of 0–3 (0: negative staining, 1: weak staining, 2: moderate staining, and 3: strong staining). 32 Percentage scores were assigned on a scale of 1–4, based on the extent of staining: 1 (1–25%), 2 (26–50%), 3 (51–75%), and 4 (76–100%). Percentage and intensity scores were multiplied for final scores. Low expression was defined as scores ranging from 0 to 6, while high expression was defined as scores ranging from 7 to 12. 33
Statistical analysis
Comparisons were conducted using Fisher’s two-tail test. The generation of dot plots for correlation analysis was carried out using the ggplot2 package, and the correlation analysis was conducted using Spearman’s rank correlation test. This investigation employed the R packages survival and survminer in the R environment to conduct survival analysis. Survival curves were constructed using the Kaplan–Meier method and were compared using the log-rank test. All statistical analyses were conducted with R (version 4.1.0, http://www.r-project.org/) and SPSS software (version 25.0, SPSS Inc., Chicago, IL, USA). The level of statistical significance was set at p < 0.05.
Results
CXCL14 expressed in fibroblast and malignant cells in RCC
To explore the distribution of CXCL14 in RCC, we conducted an analysis of bulk mRNA expression from tumor and normal tissues in the TCGA database using the GEPIA2 web tool. We observed that the expression level was similar between tumor and normal tissue in ccRCC and pRCC [Figure 1(a)]. Meanwhile, there was no significant difference between the CXCL14 expression level and the clinical stage of RCC [Figure 1(b)]. In the analysis of two ccRCC single-cell public datasets (GSE171306, GSE111360), we found the expression of CXCL14 was most prominent in fibroblast and malignant cells. Other immune cells, such as T cells, B cells, macrophages, and NK cells, hardly expressed CXCL14 [Figure 1(c) and (d)].

(a) Relative CXCL14 mRNA expression in normal and tumor tissues from clear cell renal cell carcinoma (ccRCC) and papillary renal cell carcinoma (pRCC) data. (b) Relative CXCL14 mRNA expression in different clinical stages of RCC. (c) The UMAP shows the annotation and color codes for cell types in the GSE171306 dataset. (d) The UMAP shows the annotation and color codes for cell types in the GSE111360 dataset.
High CXCL14 expression is associated with a favorable prognosis in RCC
The role of CXCL14 in cancer is intriguingly paradoxical. It could exert either supportive or inhibitory functions on tumors depending on the cancer type. Using the TCGA database, we conducted a Kaplan–Meier survival analysis to evaluate the prognostic significance of CXCL14 in RCC. Our analysis focused on ccRCC and pRCC data and revealed that patients with high CXCL14 expression in ccRCC had significantly better DSS and OS compared to those with low CXCL14 expression [5-year DSS 74.8% versus 60.5%, p = 0.015; 5-year OS 67.6% versus 52.3%, p = 0.0063; Figure 2(a)]. Similarly, in pRCC, we could observe a trend of better DSS (85% versus 76%, p = 0.19) and OS (85% versus 69%, p = 0.15) in the CXCL14 high group [Figure 2(b)]. These findings suggest that CXCL14 might exert an inhibitory effect on RCC development.

Kaplan–Meier analysis according to the mRNA expression of CXCL14 from TCGA (a) ccRCC data (b) pRCC data.
CXCL14 expression level is related to immune activation and cell metabolism
Given the tumor-inhibitory function of CXCL14 in RCC, we performed GO and KEGG enrichment analysis of up-regulated genes in the CXCL14 high group to investigate the potential mechanisms involved. Results from GO enrichment analysis [Figure 3(a)] indicated that the genes were predominantly enriched in two biological processes, immune activation and cell metabolism. The immune activation process included antigen processing and presentation, MHC protein assembly, and T-cell activation. It implied the potential role of CXCL14 in immune regulation and activation in TIME. Another related process was cell metabolism, such as monocarboxylic acid metabolic process, small molecule biosynthetic process, and fatty acid metabolic process. In KEGG enrichment analysis, CXCL14 was related to metabolic pathways like peroxisomes and glycolysis/gluconeogenesis pathways [Figure 3(b)].
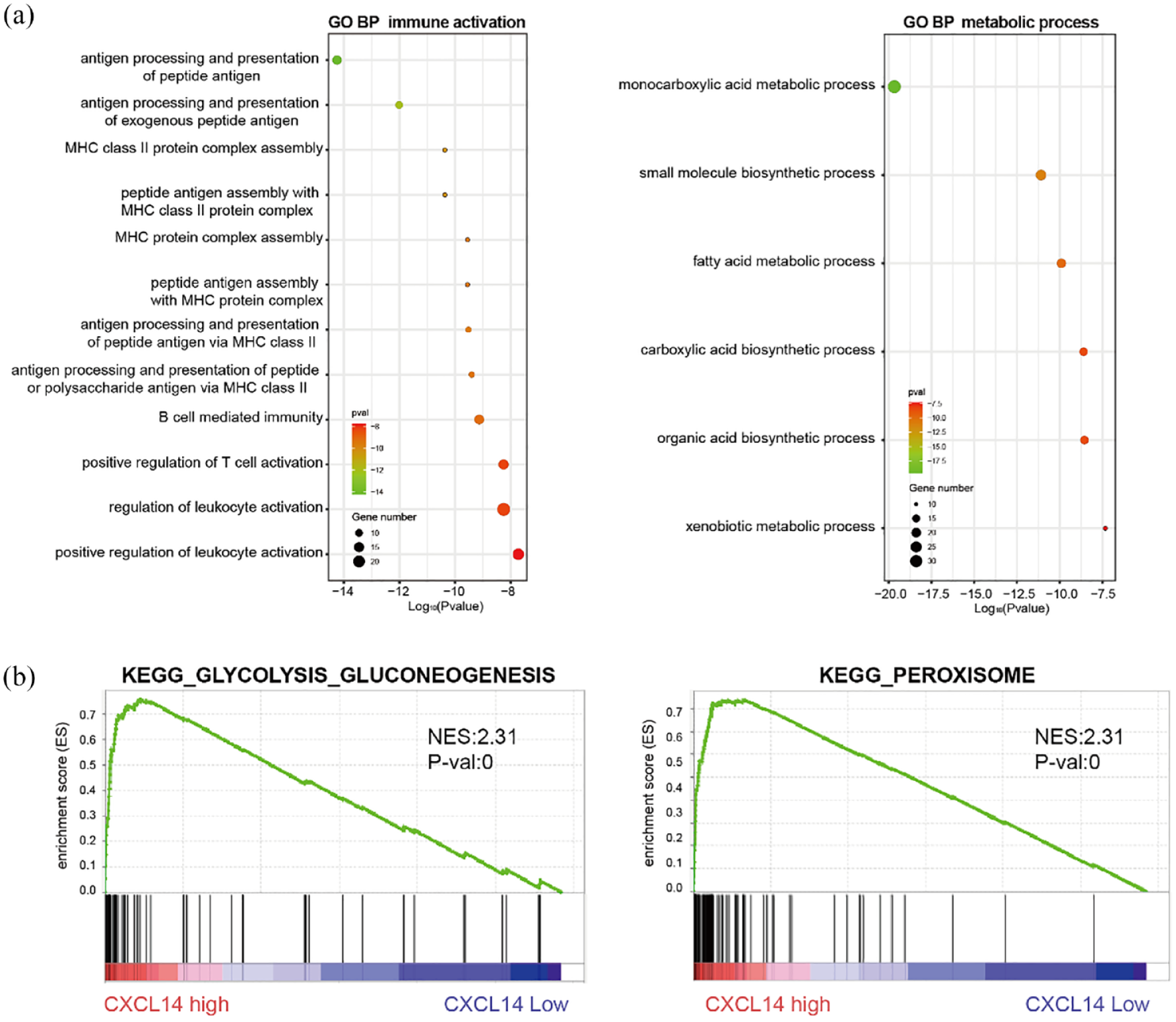
(a) Functional enrichment analysis of CXCL14 genes for Gene Ontology biological process in immune activation and metabolic process. (b) Gene Set Enrichment Analysis plots show that CXCL14 expression correlates positively with GLYCOLYSIS_GLUCONEOGENESIS and PEROXISOME in the published The Cancer Genome Atlas renal cell carcinoma gene expression profiles.
CXCL14 expression level is positively correlated with the degree of immune infiltration
Since our analysis indicated the association between CXCL14 and immune activation, we further explore its interplay with TIME. Previous studies reported that T-cell activation may be a mechanism for CXCL14-dependent tumor regression. 22 Hence, we examined the possible correlation between CXCL14 expression levels and the infiltration of T cells in RCC. The results [Figure 4(a)] showed that CXCL14 expression has significant positive correlations with infiltrating levels of CD8+ T cells (CIBERSORT, r = 0.215, p = 3.11e−06; EPIC, r = 0.137, p = 3.13e−03; XCELL, r = 0.171, p = 2.16e−04), CD8+ effector memory T cell (XCELL, r = 0.203, p = 1.09e−05), CD8+ central memory T cell (XCELL, r = 0.198, p = 1.88e−05), CD4+ Th1 cell (XCELL, r = 0.14, p = 2.60e−03), CD4+ effector memory T cell (XCELL, r = 0.262, p = 1.16e−08), follicular helper T cell (CIBERSORT, r = 0.118, p = 1.16e−02; CIBERSORT-ABS, r = 0.12, p = 1.02e−02), and gamma delta T cell (CIBERSORT, r = 0.184, p = 6.91e−05; CIBERSORT-ABS, r = 0.174, p = 1.75e−04). As the final executors of antitumor immune responses, CD8 T cells are recognized as the most important antitumor immune cells. 34 Hence, we conducted an IHC analysis to investigate the expression of CD8 and CXCL14 in tumor tissues. The results revealed a significant positive correlation between high CD8 expression and high CXCL14 expression [p = 0.025, high group: 13, low group: 17; Figure 4(b)–(d)], which was consistent with our previous results. These findings suggest that CXCL14 plays an important role in T-cell infiltration in the RCC microenvironment.

(a) The correlation of CXCL14 expression level with T-cell infiltration level in TCGA ccRCC dataset. (b) The histogram shows the relationship between CXCL14 expression level and CD8 expression level in 30 SYSUCC RCC cohorts. (c, d) The representative immunohistochemical pictures of CD8 in high and low CXCL14 groups in the SYSUCC RCC cohort (high group: 13, low group: 17).
CXCL14 expression level is positively correlated with HLA expression level
HLA-I genes, including HLA-A, HLA-B, and HLA-C, encode MHC class I molecules. MHC class I molecules present antigenic peptides to cytotoxic CD8+ T cells, which enables them to eliminate tumor cells. HLA-II genes, including the HLA-D gene family (such as HLA-DP, HLA-DQ, and HLA-DR), encode MHC class II molecules. MHC class II molecules are mainly distributed on the surface of antigen-presenting cells, assisting in helper T-cell activation. 32 We analyzed TCGA ccRCC gene expression level relationship between CXCL14 and HLA-I [Figure 5(a)] and HLA-II-related genes [Figure 5(b)] using GEPIA2 web tool. We found that CXCL14 expression has significant positive correlations with the expressions of HLA-I genes, including HLA-A (r = 0.375, p = 3.18e−19), HLA-B (r = 0.295, p = 3.82e−12), and HLA-C (r = 0.27, p = 2.27e−10). Among HLA-II-related genes, a positive correlation has also been observed in HLA-DPA1 (r = 0.238, p = 2.7e−08), HLA-DQA1 (r = 0.219, p = 3.32e−07), HLA-DQB1 (r = 0.21, p = 9.86e−07), HLA-DRA (r = 0.199, p = 3.65e−06), HLA-DRB1 (r = 0.263, p = 6.79e−10), and HLA-DQB2 (r = 0.249, p = 5.46e−09). These findings suggest that CXCL14 may enhance antigen presentation in RCC.

(a) The correlation of CXCL14 with HLA-I genes expression level in TCGA ccRCC dataset. (b) The correlation of CXCL14 with HLA-II genes expression level in TCGA ccRCC dataset.
High CXCL14 expression is correlated with good immunotherapy response
Given the close correlation between CXCL14 and immune response, we analyzed the single-cell profile of the tumor tissues from four nccRCC patients treated with immunotherapy. A total of approximately 32,479 individual cells were classified into 19 principal cell clusters. We identified the following clusters: epithelial cells, T cells (CD4+ naive/memory T cells, Treg, cytotoxicity T cells, exhausted T cells, proliferation T cells, RSAD2+ T cells, and tissue-resident memory T cells); NK cells; Macrophage cells (C1QA+ Macrophage and S100A8+ Macrophage); B cells (naive/memory B cells, plasma cells, and RSAD2+ B cells); mast cells; fibroblasts; endothelial cells; and DC cell [Figure 6(a)]. We observed a cluster of fibroblasts that exhibited high expression levels of the CXCL14 gene [Figure 6(b)]. Interestingly, these CXCL14+ fibroblasts were prominent only in responding patients [Figure 6(c) and (d)]. In addition, data from the Checkmate 025 cohort demonstrated superior survival for high CXCL14 mRNA expression patients after immunotherapy [5-year OS, 45.8% versus 10.0%, p = 0.011, Figure 6(e)]. Meanwhile, CXCL14 mRNA expression failed to show any association with survival in the group of patients treated with everolimus, supporting its potential predictive value for immunotherapy (Supplemental Figure S1). Hence, we postulated that elevated CXCL14 expression in fibroblasts could potentially be linked with an improved response to immunotherapy.

(a) The tSNE plot shows the single-cell profile of 4 SYSUCC nccRCC patients. (b) The tSNE plot shows CXCL14 expression distribution in the SYSUCC nccRCC single-cell cohort. (c) The tSNE plot shows the source of each cell according to immunotherapy response in the SYSUCC nccRCC single-cell cohort. (d) The histogram shows the distribution of immunotherapy response about CXCL14+ fibroblast in the SYSUCC nccRCC single-cell cohort. (e) Kaplan–Meier analysis according to the mRNA expression of CXCL14 in Checkmate 025. (f) The histogram shows the relationship between CXCL14 expression level and immunotherapy response in 22 SYSUCC nccRCC cohorts. (g) Immunohistochemical (CXCL14) analyses of tumor tissue sections from progression disease and partial response patients. Scale bar: 50 μm. (h) Representative images of a PR patient and a patient after receiving ICI-based treatment.
To testify this hypothesis, we explored the relationship of CXCL14 with treatment response at the protein expression level. We performed IHC staining on the samples of 22 RCC patients treated with immunotherapy. Baseline characteristics were similar between the CXCL14 high and the CXCL14 low group (Supplemental Table S1). In addition, no significant difference in DCR was found between different ages (p = 0.670), gender (p = 0.221), International mRCC Database Consortium (IMDC) risk group (p = 0.999), stage (p = 0.074), and number of metastasis groups (p = 0.415). Patients with CXCL14 high expression showed significantly higher DCR ICI-based treatment than those with CXCL14 low expression [76.9% versus 22.2%, p = 0.027, Figure 6(f) and (g)]. Patients with high CXCL14 expression generally had better responses after immunotherapy. Representative imaging of the responder and non-responder and the corresponding CXCL14 expression by IHC is shown in Figure 6(g) and (h).
CXCR4 may be the receptor of CXCL14 in RCC microenvironment
The receptor of CXCL14 remains controversial, and ACKR2, CXCR4, and GPR85 have all been proposed as potential receptors for CXCL14.35–38 To identify the potential receptor of CXCL14 in RCC, we analyzed the level of these three receptors in the single-cell datasets. The levels of ACKR2 and GPR85 were exceedingly low in all single-cell sequencing datasets. Meanwhile, CXCR4 expression was prominent in all datasets. In dataset GSE171306, CXCR4 expression was low in CXCL14+ fibroblasts and malignant cells, in contrast with high expression in other cells [Figure 7(a)]. In dataset GSE111360 and scRNA-seq from SYSUCC, the CXCR4 expression was higher in other cells compared with CXCL14+ fibroblasts [Figure 7(b) and (c)]. Next, we investigated the expression levels of CXCR4 and CXCL14 in tumor tissues through IHC analysis. Our findings revealed a significant positive correlation between CXCR4 expression and CXCL14 expression [p = 0.03, high CXCL14 expression group: 10, low CXCL14 expression group: 12; Figure 7(d)–(f)]. These results suggest that CXCR4 is the most likely potential receptor for CXCL14 in RCC. Since a positive correlation was observed between CXCL14 and CD8+ T cells, we conducted multiple immunofluorescence staining to examine their potential interaction. We found that CXCR4 receptors were present in CD8-positive T cells (Supplemental Figure S2).
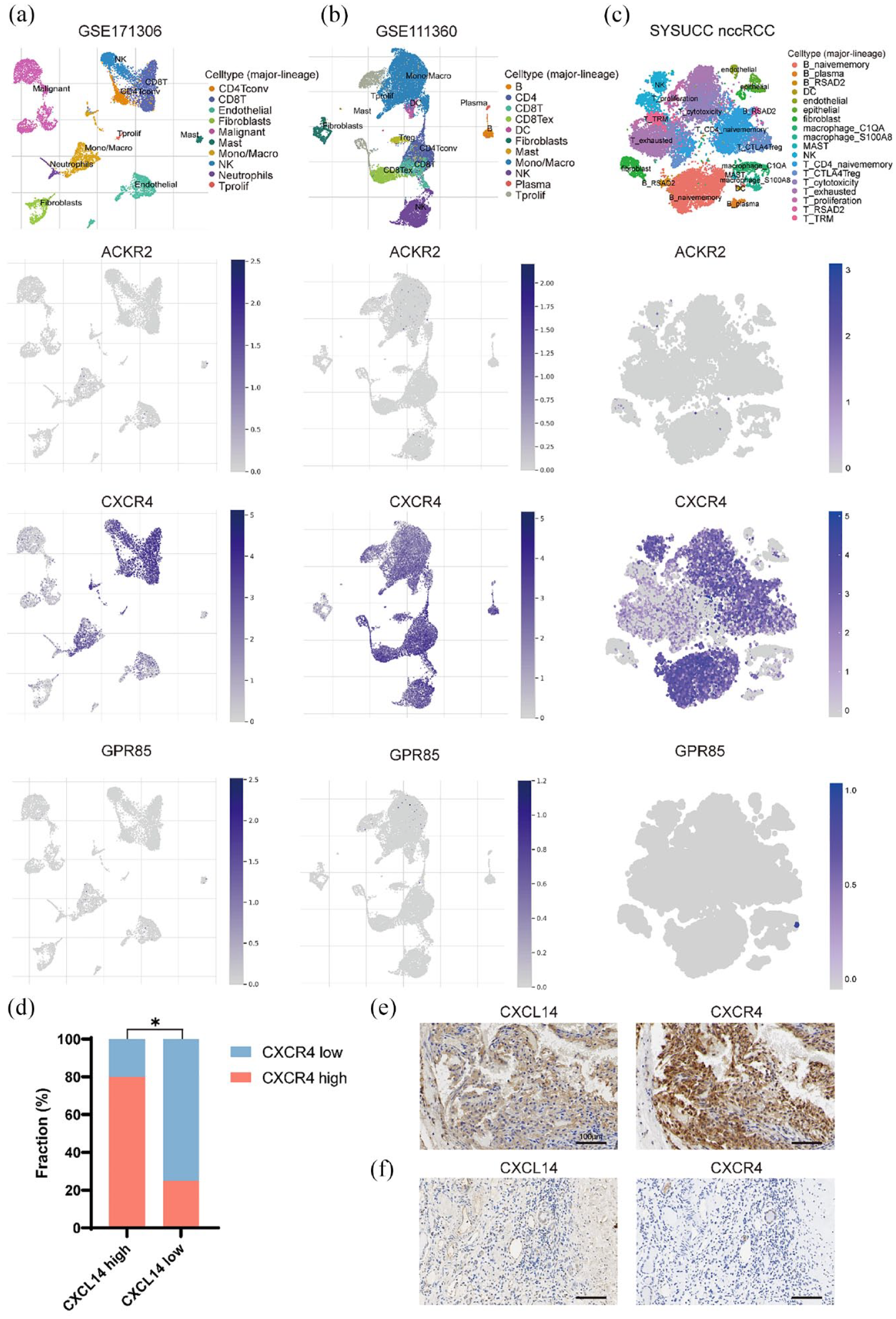
The expression profile of ACKR2, CXCR4, and GPR85 in GSE111360 (a), GSE171306 (b), and Sun Yat-sen University Cancer Center (SYSUCC) non-clear cell renal cell carcinoma (nccRCC) (c) single-cell datasets. (d) The histogram shows the relationship between CXCL14 and CXCR4 expression levels in the SYSUCC RCC cohort. (e, f) The representative immunohistochemical pictures of CXCR4 in high and low CXCL14 groups in the SYSUCC RCC cohort (high CXCL14 expression group: 10, low CXCL14 expression group: 12).
Discussion
CXCL14 is a relatively newly identified chemokine with complicated functions, and its role in RCC is poorly understood. In the present investigation, our findings revealed that increased levels of CXCL14 expression were significantly associated with improved survival. CXCL14 was associated with immune activation, supported by its correlation with T-cell infiltration and HLA-related gene expression. Single-cell dissection further revealed the link between response to immunotherapy and a group of CLXC14+ fibroblasts. In clinical observation, higher CXCL14 was associated with better response to treatment and favorable survival in the immunotherapy cohort. These results suggest that higher CLCX14 may be involved in immune activation in TIME to improve response to immunotherapy in RCC.
The role of CXCL14 in cancer is conflicting since it could exert supportive or inhibitory function depending on the cancer type. High expression of CXCL14 was associated with increased aggressiveness in lung, breast, colorectal, and thyroid papillary cancers. Meanwhile, its elevated expression was related to tumor suppression in breast and head and neck cancer.22,39 Kumar et al. 21 discovered that in glioma, CXCL14 had no direct impact on tumor cell proliferation in vitro. However, CXCL14 overexpression was associated with slower tumor growth in vivo, 40 and the survival of mice harboring CXCL14+ tumor was significantly longer. 21 One of the potential reasons for its antitumor function lies in its interaction with TIME. Indeed, CXCL14 has been reported to participate in mediating immune profiles in breast cancer, glioma, and sarcoma.21,41,42 In glioma, CXCL14 promoted directional migration and chemotaxis of activated CD8+ T cells in a dose-dependent way in vitro. In vivo, CXCL14 was associated with prolonged survival in a CD8+ T-cell-dependent manner. 21 CD8-positive lymphocytes played a complex role in immunotherapy efficacy and prognosis in RCC. Several researchers have reported that CD8+ T-cell infiltration in RCC is not necessarily associated with a favorable prognosis or a robust response to immunotherapy.43–46 Giraldo et al. 43 found that the expression of immune checkpoints and the localization of DC cells in the tumor microenvironment regulate the clinical impact of CD8+ T cells in ccRCC. Meanwhile, several studies have reported that CD8+ T-cell infiltration is associated with better response to immunotherapy.47,48 These conflicting results underscore that the degree of CD8+ T-cell infiltration is insufficient to assess its clinical significance. Its actual clinical impact may be modulated by the presence of immunosuppressive elements, tumor heterogeneity, and specific immunoevasive mechanisms. Furthermore, CXCL14 could also induce dendritic cell maturation and monocyte differentiation into macrophages. Studies in head and neck cancer and oral cavity cancer both indicated that it could increase tumor-infiltrating lymphocytes through restoration of MHC-I expression on tumor cells.22,40 Our results also supported that CXCL14 is primarily involved in antigen processing and presentation in RCC, probably owing to its regulation of HLA-related gene expression. Collectively, these results suggest that CXCL14 may promote T-cell infiltration in TIME through increasing antigen presentation, realized by inducing dendritic cell and macrophage maturation and enhancing HLA-related gene expression in RCC. This immune-active microenvironment may explain the superior prognosis of patients with CXCL14 high expression in RCC.
The complicated effect of CXCL14 in tumors may result from its secreting cell type and its receptor.11,40 Our study found a positive correlation between CXCR4 and CXCL14. Kumar et al. 21 discovered that CXCR4 expression was significantly higher in CXCL14 high cell lines and that inhibition of CXCR4 led to blockade of CXCL14-mediated chemotaxis of activated CD8+ T cells in glioma. These suggest that among the three potential receptors, CXCR4 seemed to be the most sensible receptor for CXCL14 in RCC. Notably, CXCR4 was found to be most abundantly expressed by immune cells, both in our study and in previous reports.21,49 CXCR4 also serves as receptor for CXCL12, an immunosuppressive cytokine. CXCL12/CXCR4 interaction plays a critical role in immune cell recruitment and tumor progression. Previous studies observed conflicting influences of CXCL14 on the CXCL12/CXCR4 pathway. It can present as inhibition, synergistic activation or function during its interaction with CXCL14 and CXCR4.36,50,51 This indicates that CXCL14 may exhibit different behavior in different tumor and tumor microenvironments. In our scRNA-seq analysis, CXCL14 was most predominantly expressed in fibroblasts in RCC. These results suggest a potential cross-talk between fibroblasts and immune cells. Hornburg et al. 52 have identified three main clusters of cancer-associated fibroblast (CAF) in ovarian cancer. One CAF subpopulation characterized by cytokine/chemokine signaling markers, such as CXCL14, CCL2, and SOCS3, exhibits a trend toward higher tumor immune infiltration. In addition, this CAF subpopulation may assist in CD8+ T-cell recruitment, which is crucial for CXCL14-mediated tumor suppression.22,52 Therefore, the presence of fibroblasts expressing CXCL14 in RCC suggests a potentially immune-friendly microenvironment, which is important for immunotherapy application.
ICI has largely shifted the paradigm of RCC treatment, and TIME has been recognized as a vital factor in its success. In particular, several studies have dissected TIME in ccRCC at the single-cell level and found tissue-resident T cells as an important element in response and resistance to ICI. Klein et al. 42 reported that the abundance and functionality of tumor-infiltrating T cells were elevated in the CXCL14 high group compared to the CXCL14 low group in sarcoma. In our nccRCC single-cell profile of patients receiving ICI-based treatment, CAF with high CXCL14 expression was only observed in responding patients. Further analysis at transcription and protein expression levels revealed superior responding rate and OS after ICI-based treatment in the CXCL14 high group. These data verify our hypothesis that CXCL14 expressing CAF may create an immune-active microenvironment by increasing T-cell recruitment and activation, which facilitates the antitumor effect of ICI. CXCL14 may have both predictive and prognostic value for RCC patients receiving ICI-based treatment.
Although the results are promising, this study has some limitations that need to be addressed. First, the predictive and prognostic significance of CXCL14 at the protein level needs to be further investigated with a larger sample size. Second, although this study has uncovered a potential relationship between CXCL14 and TIME, the underlying mechanism of immune regulation by CXCL14 in RCC is still in mist. We will further dissect these findings in future studies. Lastly, our study did not elucidate the reaction of CXCL14 after ICI exposure in RCC, which shall be further investigated.
Conclusion
In conclusion, elevated expression of CXCL14 in RCC is associated with superior response to ICI treatment via its modulation on TIME, primarily through the enhancement of T-cell infiltration and HLA-related gene expression. CXCL14 holds promise as a potential biomarker and epigenetic modulator for personalized prediction and aiding in the identification of suitable patients who may benefit from ICI treatment in RCC.
Supplemental Material
sj-docx-1-tam-10.1177_17588359231217966 – Supplemental material for CXCL14 as a potential marker for immunotherapy response prediction in renal cell carcinoma
Supplemental material, sj-docx-1-tam-10.1177_17588359231217966 for CXCL14 as a potential marker for immunotherapy response prediction in renal cell carcinoma by Qiwen Pan, Ruiqi Liu, Xinyue Zhang, Lingling Cai, Yilin Li, Pei Dong, Jianming Gao, Yang Liu and Liru He in Therapeutic Advances in Medical Oncology
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
