Abstract
Background:
Currently, biomarkers do not have a role in diagnosis or outcome prediction models for patients with recurrent anterior shoulder instability.
Purpose/Hypothesis:
The purpose of this study was to compare gene expression differences in the blood and tissue of patients with anterior shoulder instability with and without significant glenoid bone loss (GBL). The hypothesis was that the severity of GBL would be associated with the expression level of genes in the blood and capsular tissue of patients with anterior shoulder instability.
Study Design:
Descriptive laboratory study.
Methods:
Consecutive patients with anterior shoulder instability undergoing arthroscopic and open shoulder stabilization were prospectively enrolled. Blood and anterior capsular tissue specimens obtained during surgery were compared between patients with significant GBL (≥10%) and patients without significant GBL (<10%). RNA was extracted, and a panel of 277 inflammatory genes was utilized to quantify gene expression at the RNA level using a probe-based RNA quantification platform. Differential expression analysis was performed to identify genes expressed at different levels between patients with and without significant GBL. The expression levels of the subset of genes identified were used to generate a ridge regression model to predict the severity of GBL. Quantitative polymerase chain reaction was performed to confirm probe-based RNA findings.
Results:
A total of 17 patients were included, with a mean age of 26 years. Overall, 7 patients had <10% GBL (mean, 2.3%), and 10 patients had ≥10% GBL (mean, 16.4%). There were 9 genes that were identified as significantly differentially expressed in the blood, and 5 of these (IFIT1, IFIT3, IFI44, PRKCB, and OAS2 with P values of 1 × 10−5, 1 × 10−4, 1 × 10−4, 1 × 10−4, and 6 × 10−4, respectively) were confirmed using nonparametric tests. The authors developed a model using the 5 genes to predict the severity of GBL, and this had an accuracy of 88% (95% CI, 64%-99%). By quantitative polymerase chain reaction, IFIT3 was confirmed to be significantly differentially expressed (P = .012).
Conclusion:
There were significant gene expression differences in the blood of patients with (≥10%) and without (<10%) significant GBL. The differential expression of 5 genes allowed for the development of an accurate predictive model and transcriptomic biomarker to predict the severity of GBL.
Clinical Relevance:
The addition of a blood biomarker to current outcome prediction models may provide increased accuracy in identifying those at risk of failure from arthroscopic Bankart repair.
Anterior shoulder instability is common in young athletes, with an incidence in the United States population of 0.08 per 1000 person-years.21,29 The outcomes of surgically managed anterior shoulder labral tears with instability are favorable in this population.2,9,10,23 However, delayed or nonoperative management carries the risk of developing chronic labral tears and attritional bone loss of the glenoid and humerus secondary to repeat instability events.11,25 The mechanical factors associated with recurrent anterior shoulder instability are well known and include glenoid bone loss (GBL), humeral bone loss, glenoid version, morphological variations of glenoid concavity, capsular hyperlaxity, and anterior labral morphology. In contrast, little is known about the pathobiology of anterior shoulder instability and attritional GBL.12,14,18,27,30 The identification of novel serum biomarkers may expand our understanding of this abnormality and improve the diagnosis and tracking of injury severity and GBL progression. Furthermore, biomarkers may provide the potential to improve prognostic forecasts for existing treatment options and to develop novel treatment modalities for predicted failure.
Previous studies have investigated the association of protein biomarkers with the development of shoulder instability. Owens et al19,20 conducted 2 prospective studies of 1050 young military athletes and found that increased serum relaxin concentrations and lower baseline collagen type II cleavage levels were associated with the likelihood of acute shoulder instability. In a study evaluating whether preoperative serum protein biomarkers of cartilage turnover and inflammation are associated with injury severity in patients with shoulder instability, Yu et al 31 found that inflammatory biomarkers including high-sensitivity C-reactive protein (hs-CRP), interleukin-8 (IL-8), and macrophage inflammatory protein–1β (MIP-1β) were not associated with specific shoulder lesions. However, cartilage oligomeric matrix protein (COMP) was elevated in Hill-Sachs lesions. 31 Recently, Galvin et al 15 identified synovial periostin, a proinflammatory catabolic protein, as a significant target in anterior shoulder instability and found that it moderately correlated with the size of the Hill-Sachs lesion.
Currently, blood biomarkers do not have a role in the diagnosis or monitoring of injury severity in young patients with recurrent anterior shoulder instability. The identification of a biomarker associated with the severity of an injury and the extent of GBL in anterior shoulder instability may provide novel information to track injury progression in patients who choose nonoperative management. Additionally, current prognostic models that predict failure rates after arthroscopic Bankart repair rely solely on demographic and radiographic variables.5,13 The addition of a patient-specific biological signature may improve the accuracy of outcome prediction models through a precision medicine approach.
The purpose of this study was to compare gene expression differences in the peripheral blood and tissue of young patients with recurrent anterior shoulder instability with and without significant GBL. Additionally, we sought to identify a peripheral blood transcriptomic biomarker for the reliable delineation of the severity of GBL in patients with anterior shoulder instability.
Methods
Study Design and Patients
This study is a prospective evaluation of all patients undergoing surgery for symptomatic recurrent anterior shoulder instability at a single academic institution. Study approval was obtained from our institutional review board (No. 222038), and all participants provided written informed consent before biological specimen collection. The investigators have adhered to the policies for the protection of human participants as prescribed in 45 CFR 46.
All patients aged 18 to 45 years undergoing arthroscopic and open shoulder surgery for recurrent anterior shoulder instability were included. We excluded patients with collagen disorders, posterior or multidirectional shoulder instability, functional shoulder instability, adhesive capsulitis, or diagnoses other than symptomatic unidirectional anterior shoulder instability. A total of 17 patients were prospectively enrolled. Patients were indicated for arthroscopic or open shoulder stabilization if they had history, physical examination, and advanced imaging (magnetic resonance imaging [MRI], magnetic resonance arthrography [MRA], or computed tomography [CT]) findings consistent with an anterior inferior glenoid labral tear and symptomatic anterior shoulder instability. The following procedures were performed: arthroscopic Bankart repair (n = 6), arthroscopic Bankart repair with Hill-Sachs remplissage (n = 6), and open Latarjet procedure (n = 5).
Data Collection
All patients underwent preoperative standard MRI/MRA and/or CT with 3-dimensional (3D) reconstructions. The glenoid defect area was calculated according to the validated Pico surface area–based method 4 on standardized en face views of a CT scan with 3D reconstruction while viewed using a Synapse picture archiving and communication system (Version 5; FUJIFILM). For patients without a CT scan with 3D reconstruction (n = 8), the glenoid defect area was measured on sagittal oblique sequences of MRI/MRA with a 3.0-T MRI scanner. There were 2 independent observers who performed the GBL measurements, and the interobserver reliability was excellent, with an intraclass correlation coefficient of 0.95. Significant GBL was defined as ≥10%. Therefore, 2 groups of patients were formed: GBL <10% and GBL ≥10%. The Hill-Sachs lesion length was measured on axial sequences of MRI with a line from the articular insertion of the rotator cuff to the medial margin of the Hill-Sachs lesion. 25 The same 2 independent observers measured the Hill-Sachs lesion on axial sequences of MRI, and the intraclass correlation coefficient was good (0.85).
Collection of Blood and Tissue Specimens
Arthroscopic shoulder stabilization procedures were performed in the lateral decubitus (n = 11) and beach-chair (n = 1) positions. All open Latarjet procedures (n = 5) were performed in the beach-chair position. After an examination under anesthesia, patients underwent sterile preparation and draping. For arthroscopic shoulder stabilization cases, the arthroscope was first inserted through a standard posterior viewing portal, and a rotator interval portal was established under direct visualization. Redundant anterior inferior capsulolabral tissue that was going to be debrided as part of the standard of care for that procedure was collected before surgical reconstruction. For open Latarjet procedures, resected anterior inferior capsulolabral tissue was sent as a specimen, as we do not perform capsulolabral repair as part of the open Latarjet procedure. The tissue specimens were cut into pieces small enough for insertion into cryovials, snap frozen in liquid nitrogen, and stored at −80°C.
After tissue specimen collection, 3 mL of peripheral blood was collected and placed in a DNA/RNA Shield Blood Collection Tube (Catalog No. R1150, Lot No. PNG430299; Zymo Research). It was gently inverted 10 times, placed on ice, and transported within 15 minutes to the hospital's research laboratory, where it was aliquoted into multiple 1000-µL cryovials and stored at −80°C.
Total RNA Isolation
RNA from snap-frozen tissue was extracted by first pulverizing tissue in a Bio-Pulverizer device (Thomas Scientific) supercooled with liquid nitrogen in the presence of TRIzol (Thermo Fisher Scientific). Next, pulverized tissue was funneled into centrifuge tubes, allowed to thaw, and then homogenized with Pro200 (PRO Scientific) for 1 minute. This mixture was incubated at room temperature for 45 minutes and inverted every 15 minutes. After incubation, chloroform was added and the tube shaken. After a final 15-minute incubation, the tubes were centrifuged and extraction performed using the RNeasy Lipid Tissue Kit (Qiagen) per the manufacturer's instructions. RNA was extracted from blood preserved in DNA/RNA Shield (Zymo Research) using the Quick-RNA Whole Blood Kit (Zymo Research). Briefly, 400 µL of thawed blood was processed per the manufacturer's instructions, which included deoxyribonuclease treatment. The elution of RNA was performed with 33 µL of 1 mM Tris (pH 8). RNA quality and quantity were assessed using the NanoDrop One spectrophotometer (Thermo Fisher Scientific), Qubit 1X dsDNA High Sensitivity Assay Kit (Thermo Fisher Scientific), and TapeStation system (Agilent), enabling the calculation of an RNA integrity number (RIN). Aliquots of RNA were placed in LoBind tubes (Eppendorf) and frozen at −80°C until use.
Quantitative Polymerase Chain Reaction (qPCR)
To prepare complementary DNA, total RNA was reverse transcribed using the SuperScript VILO cDNA Synthesis Kit (Thermo Fisher Scientific) on a thin-walled 96-well PCR plate with the QIAamplifier (Qiagen). qPCR was performed using TaqMan predesigned assays (IFIT1: Hs03027069; OAS2: Hs00942643; PRKCB: Hs00176998; IFIT3: Hs01922752; IFI44: Hs00197427; GAPDH: Hs02786624; GUSB: Hs00939627; HPRT1: Hs02800695; PGK1: Hs00943178). The default thermocycling program was selected using TaqMan Fast Advanced Master Mix (Thermo Fisher Scientific), and samples were run in triplicate or quadruplicate. A “no template” control was included on each PCR plate.
Processing of Samples
Gene expression was assayed on an nCounter SPRINT Profiler (NanoString) using the “Human Inflammation Panel” customized to include additional probes relevant to the question of interest, such as COMP. See Appendix 1A and 1B for an entire list of genes included. For each nCounter run, 12 RNA samples were normalized to 10 ng/µL. Then, 5 µL of each was added to 10 µL of Reporter CodeSet (NanoString) and Reporter Plus Master Mix (NanoString). Capture CodeSet (NanoString) and Capture Plus Master Mix (NanoString) were prepared, and 3 µL was added to the samples. These were hybridized for 16 hours at 65°C under a heated lid (at 70°C). The SPRINT cartridge (NanoString) was loaded with the 12 samples and run on the nCounter SPRINT Profiler.
Statistical Analysis
Transcript count data generated on the nCounter underwent the manufacturer's standard quality control assessment in nSolver software (NanoString) before being exported to CSV files and imported into R using the NanoStringDiff package. 28 The same package was used to perform differential expression analysis to determine whether there were differences in gene expression between the <10% GBL and ≥10% GBL groups. In addition, the Wilcoxon rank-sum test was performed on the differentially expressed genes to identify a subset of genes with an especially high confidence of being differentially expressed. P values were adjusted to account for multiple comparisons and the false discovery rate. This adjusted P value is referred to as the Q value. In these analyses, the raw gene expression data were normalized by dividing the gene expression by the geometric mean of the expression of the housekeeping genes. The subset of normalized and differentially expressed genes was subjected to principal component analysis. Ridge regression was performed to predict the severity of GBL based on gene expression levels. The genes included in this model were the differentially expressed genes identified in the above analysis (IFIT1, IFIT3, IFI44, PRKCB, OAS2) after normalization against housekeeping genes (CLTC, GAPDH, GUSB, HPRT1, PGK1, TUBB) as described above. Statistical analyses were performed in R (Version 4.0.2).
Results
A total of 17 patients (16 male, 1 female) undergoing surgery for recurrent anterior shoulder instability were prospectively enrolled with a mean age of 26 years (range, 20-41 years). Overall, 7 patients had <10% GBL with a mean GBL of 2.3% (range, 0%-8%), and 10 patients had ≥10% GBL with a mean GBL of 16.4% (range, 10%-25%). All 7 patients in the <10% GBL group had on-track Hill-Sachs lesions, whereas only 1 of 10 patients in the ≥10% GBL group had an on-track lesion. Demographic characteristics of the cohort are included in Table 1. All RIN values were ≥7, which signifies that we had high RNA quality for analysis: 3 were between 7 and 8, 11 were between 8 and 9, and 3 were >9. RIN values were compared between GBL groups, and no differences were found (P = .1).
Patient Characteristics a

Data are shown as No. or mean (range). GBL, glenoid bone loss.
Blood and Tissue Specimens
No genes in the anterior capsular tissue specimens were differentially expressed (Table 2). There were 9 genes that were identified as significantly differentially expressed in the peripheral blood: IFIT1, IFIT3, IFI44, PRKCB, OAS2, CCL3, FGFR2, CXCL10, and NOD1 (Table 3 and Figure 1). Of these, 5 (IFIT1, IFIT3, IFI44, PRKCB, and OAS2 with P values of 1 × 10−5, 1 × 10−4, 1 × 10−4, 1 × 10−4, and 6 × 10−4, respectively) were confirmed using nonparametric tests; this helped to ensure that the assumptions of the statistical methods used in the analysis of blood specimens did not unduly influence the results (Table 4 and Figure 2).
Differential Expression Analysis of Tissue Specimens

Differential Expression Analysis of Blood Specimens


Differential expression analysis of the blood specimens identified 9 differentially expressed genes, as depicted in the volcano plot. Q value = P value adjusted for multiple comparisons.
Nonparametric Test Confirmation for Differential Analysis of Blood Specimens


Gene expression of IFI44, IFIT3, PRKCB, IFIT1, and OAS2 between patients with <10% glenoid bone loss (GBL) and patients with ≥10% GBL.
Exploratory Data Analysis With Unsupervised Clustering
Principal component analysis was performed to visualize differences in the gene expression profiles. Significant differences were observed across principal components 1 and 2 between the gene expression profiles of patients in the low and high GBL groups (Figure 3A), with the first 2 components explaining approximately 95% of the variation in the data (Figure 3B). The genes IFI44, IFIT1, IFIT3, and OAS2 explained between 22% and 25% of the variability in principal component 1, while PRKCB explained <4%. In contrast, PRKCB explained 90% of the variability observed in principal component 2, while IFIT1 explained 7%, and the other genes explained <2%.

(A) Principal component analysis depicting differences in the gene expression profiles observed in the blood of patients with low and high glenoid bone loss (GBL). (B) Principal component analysis of the gene expression profiles of the blood specimens revealed that approximately 95% of the variation in the data was explained by principal components 1 and 2.
Transcriptomic Classifier to Predict Severity of GBL
We developed a predictive model (ridge regression) using the 5 differentially expressed genes in the blood to accurately predict the severity of GBL (<10% and ≥10%). The predictive model had an accuracy of 88% (95% CI, 64%-99%), a sensitivity of 0.90, a specificity of 0.86, and an area under the receiver operating characteristic curve of 0.99 (95% CI, 0.95-1.00) (Figure 4).
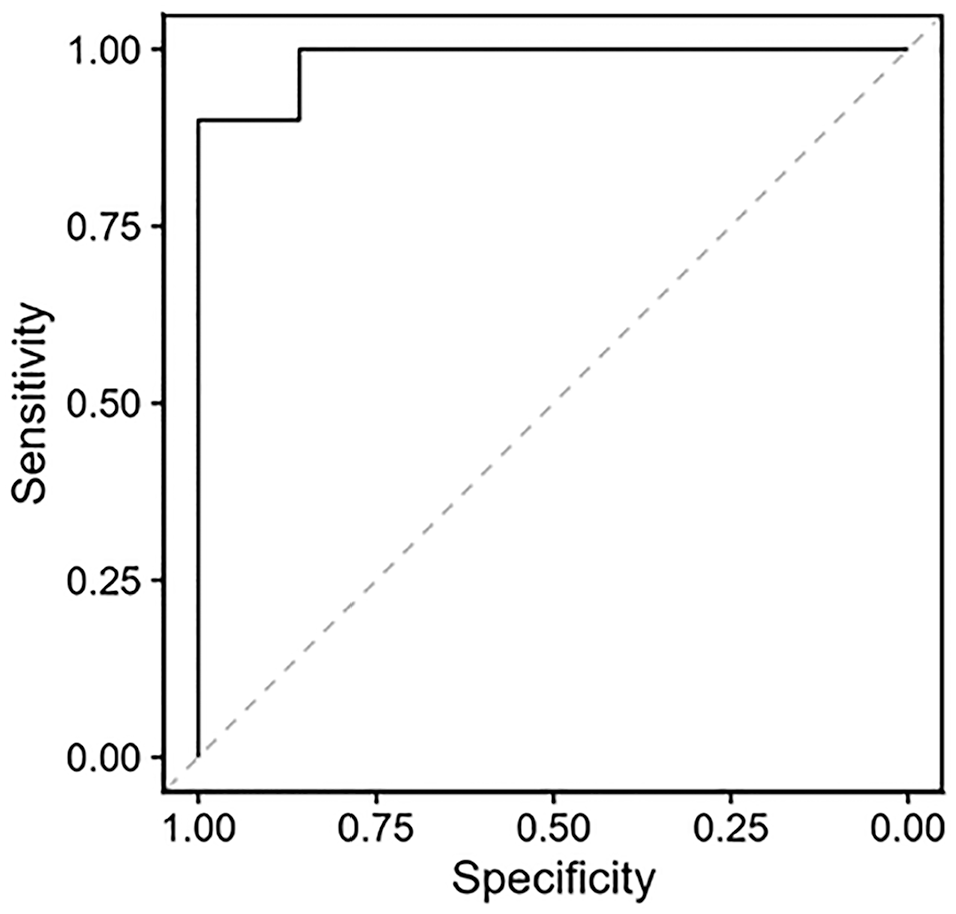
Receiver operating characteristic curve depicting the sensitivity and specificity of the model differentiating between low and high glenoid bone loss (GBL) based on gene expression.
Quantitative Polymerase Chain Reaction
Although nCounter is generally considered to be more accurate than qPCR because there is no need for complementary DNA generation and no concern regarding primer amplification bias, we also evaluated the expression of the 5 genes that we identified via qPCR. We used off-the-shelf primers, which did not target the same sequence as the nCounter probes. Only one gene, IFIT3, exhibited statistically significant gene expression differences between the GBL groups (P = .012). P values for the other genes did not reach statistical significance (IFI44: P = .70; IFIT1: P = .42; OAS2: P = .69; PRKCB: P = .59), suggesting that specific gene isoforms captured by nCounter but not qPCR could be involved.
Discussion
The primary findings of this study are that there were significant gene expression differences in the peripheral blood of patients with anterior shoulder instability with (≥10%) and without (<10%) significant GBL. There were 5 inflammatory genes that were differentially expressed between the 2 groups: IFIT1, IFIT3, IFI44, PRKCB, and OAS2 (P values of 1 × 10−5, 1 × 10−4, 1 × 10−4, 1 × 10−4, and 6 × 10−4, respectively). We used a ridge regression model and the 5 differentially expressed genes to develop a blood transcriptomic biomarker to accurately predict the severity of GBL in patients undergoing surgery for recurrent anterior shoulder instability. The predictive model had an accuracy of 88% (95% CI, 64%-99%), a sensitivity of 0.90, a specificity of 0.86, and an area under the receiver operating characteristic curve of 0.99 (95% CI, 0.95–1.00). We utilized qPCR to confirm our nCounter data. Of the 5 genes that were found to be significantly different between the low and high GBL groups, IFIT3 was the only gene significantly differentially expressed by qPCR. The reason for the difference in results between these 2 methods is unclear and may be secondary to the methodology of the techniques. nCounter has been shown to be a more accurate method for low abundance genes because of its direct measurements of mRNA fragments with capture and reporter probes, whereas qPCR is accurate for highly abundant expressed genes with its indirect enzymatic methodology.8,22,32 In addition, the oligonucleotide sequences used in nCounter and the qPCR primers differ. This raises the possibility that the discrepancy between our nCounter and qPCR results could be derived from differences in the expression levels of specific isoforms of the genes. Future prospective studies are needed with larger samples sizes. These novel peripheral blood transcriptomic data may assist in tracking GBL progression and injury severity in young patients with recurrent anterior shoulder instability; however, these findings need to be replicated in larger prospective studies.
Few studies have examined the role of biomarkers in the diagnosis, prognosis, and management of patients with shoulder instability. Yu et al 31 conducted a study on 33 patients undergoing surgical shoulder stabilization to determine whether preoperative biomarkers of cartilage turnover and inflammation were associated with specific lesions in shoulder instability. The protein biomarkers of COMP, hs-CRP, IL-8, and MIP-1β were collected at the time of surgery. Patients with Hill-Sachs lesions had a 31% increase in COMP plasma levels (P = .046). No other significant differences were observed for any of the protein biomarkers including Hill-Sachs lesions, capsular injuries, bony Bankart fractures, and SLAP (superior labrum anterior and posterior) lesions. COMP, CRP, IL-8, and MIP-1β (CCL4) genes were all queried in our customized gene panel, and they were not found to be differentially expressed between the low and high GBL groups.
Belangero et al 7 sampled the injured anterior inferior glenohumeral capsule and compared this to the unaffected anterior superior capsular region in 18 patients with traumatic anterior shoulder instability. The authors found the reduced expression of one of the collagen genes (COL5A1). In a subsequent study, the same research group evaluated the tissue samples of patients with recurrent anterior shoulder instability compared with controls and found that recurrent shoulder dislocations were associated with the upregulation of TGFβR1 in the anteroinferior capsule, which is a gene involved in the regulation of collagen cross-linking. 6 Although these data are interesting and provide improved understanding of pathobiology, they are not helpful to clinicians in the management of patients. We included the following collagen genes (COL1A1, COL1A2, COL3A1, COL4A2, COL5A1, COL5A2, COL6A3) in our commercially available customized gene expression panel (nCounter) of 277 genes and did not find any differential expression in the capsular tissue or blood between the low and high GBL groups.
Recently, Aleem et al 1 examined the gene expression of glenoid cartilage tissue samples among patients with acute instability (<3 dislocations), chronic instability (≥3 dislocations), and glenohumeral osteoarthritis. They found that glenoid articular cartilage in the osteoarthritis group displayed higher expression levels of CCL3, CHST11, GPR22, PRKAR2B, and PTGS2 than cartilage in the group with acute or chronic shoulder instability. We also identified CCL3 as a gene of interest. In our analysis, CCL3 was not significantly different in the anterior capsular tissue specimens (P = .018; Q = .65). Although the P value was significant, the Q value (an adjusted P value that corrects for inflation of the type I error rate during multiple hypothesis testing) was not significant. It was significantly different in the blood (P = .00013; Q = .008); however, with nonparametric confirmation testing, the result was not significant (P = .07). In the blood, the analysis demonstrated that the CCL3 gene was downregulated in the high GBL group (log fold change = −2.32). Given the recent findings of Aleem et al, 1 as well as our analysis, CCL3 is a gene of significant interest and should be investigated in larger studies as a target for the potential prevention of glenohumeral osteoarthritis.
IFIT1, IFIT3, and IFI44 were all significantly differentially expressed genes between patients with and without significant GBL. All of these genes were overexpressed in patients with <10% GBL. The mechanism for these findings is unclear. Previous literature has found that IFIT1 regulates Wnt/β-catenin signaling. 16 This pathway comprises a family of proteins that play critical roles in tissue homeostasis. Specifically, the canonical Wnt signaling pathway affects the proliferation and differentiation of mesenchymal stem cells and osteoblast progenitor cells in addition to osteoclast function. This pathway is critical to the maintenance of bone homeostasis. 17 On qPCR validation, IFIT3 was the only gene that was found to be significantly differentially expressed. Further investigation is needed in larger sample sizes to validate and potentially replicate these findings, to better understand the mechanism, and to confirm the initial nCounter results demonstrating the significant differential expression of all 3 genes (IFIT1, IFIT3, IFI44).
The PRKCB gene was significantly overexpressed in the patients with ≥10% GBL. This gene has been well examined, and interestingly, it is significantly overexpressed in Ewing sarcoma, the malignant pediatric bone tumor. Surdez et al 26 found that the transcriptional activation of PRKCB was directly regulated by the fusion oncogene EWSR1-FLI1, which drives the growth of Ewing sarcoma. Additionally, the authors discovered that PRKCB loss induced apoptosis in vitro and prevented tumor growth in vivo. These findings provide a proof of concept for potential targeting of PRKCB as a therapeutic gene in Ewing sarcoma treatment. The mechanism of PRKCB overexpression in patients with increasing GBL is unclear; however, both shoulder instability with GBL and Ewing sarcoma feature rapid bone turnover, which could account for the similar expression. 33
Previous studies have attempted to identify diagnostic and prognostic biomarkers for use in clinical practice for shoulder instability and have been unsuccessful.19,20,31 Further, studies have identified tissue-level significant differences in gene expression6,7; however, these tissue transcriptomic data are less useful in the clinical setting. These novel peripheral blood transcriptomic findings provide initial data that may lead to a relevant biomarker that can be added to current prognostic outcome prediction models.
Limitations of the study include the small sample size and the lack of inclusion of patients without shoulder instability. Additionally, not all patients had CT scans with 3D reconstructions. Overall, 8 patients without CT scans had the glenoid defect area measured on sagittal oblique sequences of MRI/MRA with a 3.0-T MRI scanner. While CT is considered the gold standard for measurements of GBL, MRI has been validated as a reliable alternative.3,24 Also, we controlled for the potential variability in GBL measurements with excellent interobserver reliability. Additionally, despite the Q values (adjusted P values) being statistically significant, the log fold change was relatively small for the 5 differentially expressed genes. Therefore, future larger studies are needed to confirm these findings. Furthermore, on qPCR validation, only IFIT3 was significantly differentially expressed. nCounter holds several advantages and distinct differences over qPCR. It directly measures mRNA with reporter and capture probes and produces a digital readout without the need for multiple indirect enzymatic reactions of reverse transcription and PCR DNA amplification, which introduces the possibility of amplification bias.8,22,32 qPCR does have several advantages, including its low cost and accuracy for highly abundant genes. We are confident of our results using the nCounter because of our strict quality control measures, which included ensuring appropriate RIN values, the complete absence of quality control failures in our nCounter runs, and the significant adjusted P values. In future work, we hope to customize qPCR primers to match the oligonucleotides used in nCounter, ensuring that the same isoforms are assayed in both experimental modalities. Lastly, these transcriptomic data are based on 2 groups categorized by the severity of GBL. It is well known that bone loss in anterior shoulder instability is bipolar, with loss on the glenoid and humeral sides. 12 Future investigation is needed to examine the influence of variations of bipolar bone loss on transcriptomics.
Strengths of the study include the prospective enrollment of patients with the collection of blood and tissue biological specimens. Additionally, rigorous scientific and statistical methods were followed with the confirmation of RNA quality with high RIN values, confirmation of gene expression results with nonparametric methods, highly significant P values (.000013, .00011, .0001, .00019, and .0006), and strict quality control measures for the nCounter. Additionally, another strength is the development of a novel accurate statistical model to identify a transcriptomic biomarker for the reliable delineation of the severity of GBL. Finally, we thought that it was critical to include the qPCR findings, as any future transcriptomic biomarkers have to be reproduced on platforms that are available for use in laboratories and accessible to clinicians at a low cost.
Conclusion
There were significant gene expression differences in the blood of patients with anterior shoulder instability with (≥10%) and without (<10%) significant GBL. The differential expression of 5 genes allowed for the development of an accurate predictive model and transcriptomic biomarker to predict the severity of GBL. These novel blood transcriptomic data may assist in tracking GBL and injury progression in patients with recurrent anterior shoulder instability.
Supplemental Material
sj-xlsx-1-ojs-10.1177_23259671251336394 – Supplemental material for Transcriptomics of Anterior Shoulder Instability: Differences in Gene Expression in the Blood of Patients With and Without Significant Glenoid Bone Loss
Supplemental material, sj-xlsx-1-ojs-10.1177_23259671251336394 for Transcriptomics of Anterior Shoulder Instability: Differences in Gene Expression in the Blood of Patients With and Without Significant Glenoid Bone Loss by Joseph W. Galvin, Patrick Rooney, Alec Egan, John M. Tokish, Jason A. Grassbaugh, Brendan Masini, Katherine E. Free, Marit K. Bastian, Laurel H. Gillette and Zachary T. Colburn in Orthopaedic Journal of Sports Medicine
Supplemental Material
sj-xlsx-2-ojs-10.1177_23259671251336394 – Supplemental material for Transcriptomics of Anterior Shoulder Instability: Differences in Gene Expression in the Blood of Patients With and Without Significant Glenoid Bone Loss
Supplemental material, sj-xlsx-2-ojs-10.1177_23259671251336394 for Transcriptomics of Anterior Shoulder Instability: Differences in Gene Expression in the Blood of Patients With and Without Significant Glenoid Bone Loss by Joseph W. Galvin, Patrick Rooney, Alec Egan, John M. Tokish, Jason A. Grassbaugh, Brendan Masini, Katherine E. Free, Marit K. Bastian, Laurel H. Gillette and Zachary T. Colburn in Orthopaedic Journal of Sports Medicine
Footnotes
Final revision submitted October 29, 2024; accepted December 6, 2024.
One or more of the authors has declared the following potential conflict of interest or source of funding: This study was funded through the Musculoskeletal Injury Rehabilitation Research for Operational Readiness program administered by the Geneva Foundation (award No. HU00011920011). The Defense Health Agency and the United States Army filed a Patent Cooperation Treaty patent for the transcriptomic biomarker that is reported in this article. The views expressed in this article are those of the authors and do not reflect the official policy of the Department of the Army, the Department of Defense, or the United States Government. The investigators have adhered to the policies for protection of human subjects as prescribed in 45 CFR 46. J.W.G. has received support for education from Smith & Nephew, Kairos Surgical, ImpactOrtho, and Summit Surgical; consulting fees from FH Orthopedics; and hospitality payments from Encore Medical and Wright Medical. P.R. has received support for education from Summit Surgical. A.E. has received support for education from Summit Surgical. J.M.T. has received consulting fees, royalties, and honoraria from Arthrex; consulting fees from DePuy Synthes, ImpactOrtho, and Medical Device Business Services; and support for education from Paladin Technology Solutions, Supreme Orthopedic Systems, Goode Surgical, and ImpactOrtho. J.A.G. has received support for education from Summit Surgical and hospitality payments from Stryker and Arthrex. B.M. has received support for education from Summit Surgical and hospitality payments from Gemini Mountain Medical. AOSSM checks author disclosures against the Open Payments Database (OPD). AOSSM has not conducted an independent investigation on the OPD and disclaims any liability or responsibility relating thereto.
Ethical approval for this study was obtained from Madigan Army Medical Center (222038).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
