Abstract
Purpose
Interleukin-1 is the main proinflammatory cytokine in osteoarthritis (OA). Several single-nucleotide polymorphisms (SNPs) within the IL-1 gene cluster (IL-1β, IL-1R1, and IL-1RN) have been determined, but their associations with knee OA remain poorly understood. The present study aimed to identify the associations between IL-1 SNPs and knee osteoarthritis.
Methods
This meta-analysis and systematic review included all comparative studies published in the MEDLINE/PubMed, Embase, Google Scholar, and Cochrane Library databases. We performed a systematic search to identify relevant studies on the evaluation of the correlation between the IL-1 gene and knee OA published up to February 2020 that met the eligibility criteria. Nine studies on a total of 2256 knees with OA and 3527 healthy knees met the eligibility criteria. Results associated with IL-1A, IL-1B, IL-1R1, and IL-1RN SNPs were extracted and compared between knees with OA and healthy knees. Methodological quality was assessed using the Newcastle–Ottawa scale (NOS). All studies with fair or good quality were included.
Results
The meta-analysis showed that the risk of knee OA is decreased by the IL-1RN*1 and IL-1RN*1/*1 genotypes and increased by the IL-1RN*2 and I-L1RN*1/*2 genotypes. The systematic review revealed only two studies associating the IL-1RN allele, none associating the IL-1B polymorphism, and only one study associating IL-1A and IL-1R1 polymorphisms with knee OA.
Conclusions
Several IL-1RN alleles and genotypes play a role in knee OA but other genetic variations in the IL-1 region were still conflicting in its association with knee OA.
Introduction
The pathogenesis of osteoarthritis (OA) is not yet fully understood. It was originally thought to result from the failed repair of damaged cartilage. 1 This theory has been expanded to include contributions from the bone and synovial tissue as well as inflammatory factors.1–3 Interleukin-1 (IL-1) is the regulator of inflammation by controlling the innate immune process. IL-1 itself is expressed in a wide range of tissues and cells. 4 IL-1 is synthesized due to stress response. By binding to its receptors, it will activate other immune response and other hematological changes. 5 IL-1 also associated with the pathogenesis of knee OA due to the fact that an increased expression of this cytokine correlates with worsening symptoms and more progressive radiological features of knee OA.6–8 Interleukin-1 is regarded as a major proinflammatory cytokine in addition to Tumor Necrosis Factor (TNF) and IL-6, and its biological activity is mediated through specific cell surface receptor known as the IL-1 receptor type I (IL-1R1). 9 Another natural inhibitor of IL-1 activity, IL-1 receptor antagonist (IL-1RA), is produced by several types of cell including chondrocytes and synovial fibroblasts, and functions as a competitive antagonist in the process of osteoarthritis.8,10 Another mechanism is through the binding of IL-1β to the IL-1 receptor (IL-1R1), which induces a pro-inflammatory reaction resulting in cartilage destruction. Progressive cartilage destruction might be followed by subchondral destruction manifesting as progressive radiological grading. Pro-inflammatory reactions also cause synovial inflammation, that might manifest as pain. 11
The latest concept of OA pathogenesis involves a genetic component that might influence inflammation (cytokines), anti-inflammation, and receptor binding processes. Several types of OA are associated with a genetic component,12–14 but controversies are also found among different ethnicities. 15 Variations in cytokine levels among individuals become a plausible explanation for differences in disease susceptibility and severity, 15 and this association is particularly true for cytokine gene polymorphisms and OA. The most consistent association involves a broad region on human chromosome 2q13–32, which represents the IL-1 gene cluster. 16 Several DNA variants within the IL-1 gene cluster have been reported, including single-nucleotide polymorphisms (SNPs) in IL-1β, IL-1R1, and IL-1RN. However, specific genetic factors and polymorphisms associated with these conditions remain poorly understood. Several studies have generated conflicting findings in terms of this association with various OA phenotypes.
The present meta-analysis and systematic review aimed to clarify an association between genetic polymorphisms in the IL-1 gene cluster and the development of knee OA. We hypothesize that there is a correlation between genetic polymorphisms in the IL-1 gene cluster and the development of knee OA.
Material and methods
Search strategy
Based on the Preferred Reporting Items for Systematic Reviews and Meta Analysis (PRISMA) guidelines, we performed a systematic review using the MEDLINE/PubMed, Embase, Google Scholar, and Cochrane Library databases to identify relevant studies evaluating the correlation between IL-1 gene and knee OA published up to February 2020 that met the eligibility criteria. We conducted a systematic search using the following medical subject headings (MeSH) and key terms: ((((Knee osteoarthritis) AND Single Nucleotide Polymorphism) OR SNP) OR Gene) AND Interleukin-1) OR IL-1))). We limit our search to studies conducted on humans and published in English. Two authors (NCB, ILG) separately reviewed all abstracts for eligibility. We manually-searched the references from the included articles and reviews to avoid missing any relevant studies.
Inclusion and exclusion
The inclusion criteria were (1) studies comparing knee OA patients and normal patients; (2) studies investigating the relationship between the IL-1 gene or polymorphisms and knee OA; (3) studies conducted on human study; (4) studies published in English; (5) studies providing the number subjects for their case and control groups; and (6) studies providing the genotype and/or allele frequency for their case and control groups. The initial search yielded 18,693 references. After the studies were compared against the inclusion criteria, 18,684 studies were excluded due to the following reasons: irrelevant topic (not correlated with IL-1 in knee OA), full text not available, not a comparative study (no control group), no comparison with a healthy control group, incomplete data, and being meta-analyses or systematic review articles. After exclusions and evaluation by two authors, 9 studies met the inclusion criteria. We found 1 cohort study and 8 case-control studies. Using the Newcastle-Ottawa scale (NOS) for quality assessment, we reviewed the 9 articles (Figure 1). All fair and good-quality studies were included. In each article, we reviewed the comparison between the knee OA group and the healthy control group in terms of the frequency of IL-1 gene polymorphism. Flowchart shows the numbers of articles initially identified as well as the exclusion and inclusion steps.
The quality of all articles was assessed by using the Newcastle-Ottawa scale (NOS). Each study is interpreted as having good, fair, and poor quality. The eight case-control and one cohort studies are of good quality. All studies adequately reported a succinct study aim and an appropriate outcome measure. Due to the limitations and wide variation in terms study quality, we were only able to perform meta-analysis for the IL1RN polymorphism while a systematic review was performed for the other IL1 polymorphisms.
Data extraction
In each article, we collected the demographic characteristics, sample size, phenotype information, the number of genotypes in cases and controls, and the results of the study.
Statistical analysis
The distribution of genotypes and alleles is summarized in a data table. The distribution of the genotypes in the control population was assessed using Hardy–Weinberg equilibrium (HWE). p values of <0.05 indicated that the genotype distribution deviated from HWE. General data were evaluated using meta-analysis. Heterogeneity was assessed using chi-squared tests. Random and fixed-effect models were applied when p < 0.05 or p > 0.05, respectively. Sources of heterogeneity were identified using a meta-regression analysis. Publication bias was assessed using funnel plots, and Begg and Egger tests helped to reach a clear conclusion. After determining the effect model, the strength of the association between the IL-1-RN Variable Number Tandem Repeat (VNTR) polymorphism and the risk of OA was assessed using odds ratios (OR) and confidence intervals (CI). Two authors independently completed the analysis and obtained the same results. All data were statistically analyzed using Review Manager (RevMan) [Computer program] Version 5.3. Copenhagen: The Nordic Cochrane Centre, The Cochrane Collaboration, 2014. p values of <0.05 were considered significant.
Results
Newcastle–Ottawa scale (NOS) quality assessment of case-control studies.

Quantitative synthesis
Studies comparing IL-1 gene polymorphisms between knees with OA and healthy knees.


Forest plot of IL1RN VNTR allele.
The meta-analysis revealed that some IL-1RN VNTR genotype polymorphisms were also significantly associated with knee OA. The IL-1RN*1/*1 polymorphism decreased the risk of knee OA (OR, 0.59; 95% CI, 0.37–0.95; I2 = 82, p = 0.03). Interleukin-1RN*1/*1 and IL-1RN*1/*2 polymorphisms remained significantly associated with knee OA in Caucasians (OR, 0.50; 95% CI, 0.28–0.89; I2 = 84, p = 0.02 and OR, 1.77; 95% CI 1.03–3.06, I2 = 86, p = 0.04, respectively). Subgroup analysis revealed a significant association between IL-1RN*1/*2 genotype polymorphism and knee OA in Caucasians (OR, 2.20; 95% CI, 1.10–4.43, I2 = 88, p = 0.03). Other IL-1RN VNTR (rs419598) alleles or genotypes polymorphisms were not significantly associated with knee OA (Figure 3). Forest plot of IL1RN VNTR genotype.
Heterogeneity tests
The allele contrast model analysis revealed significant heterogeneity (I2 = 69; p < 0.0001); therefore, a random effects model was adopted. Subgroup heterogeneity was also high (I2 = 68.4; p = 0.02). The genotype model analysis also indicated significant heterogeneity (I2 = 67; p < 0.0001) and the subgroup heterogeneity was I2 = 47.1; p = 0.08. Subgroup analysis based on rand type of study did not identify any source of heterogeneity.
Publication bias
Publication bias was evaluated with funnel plots using the IL-1RN*1 allele as a sample. The results showed asymmetry for the IL-1RN*2, IL-1RN*1/*1, and IL-1RN*1/*2 alleles, but not for any other IL-1RN alleles or genotypes (Figure 4). Funnel plot IL1RN*1 allele shows asymmetric funnel plot in IL1RN*1.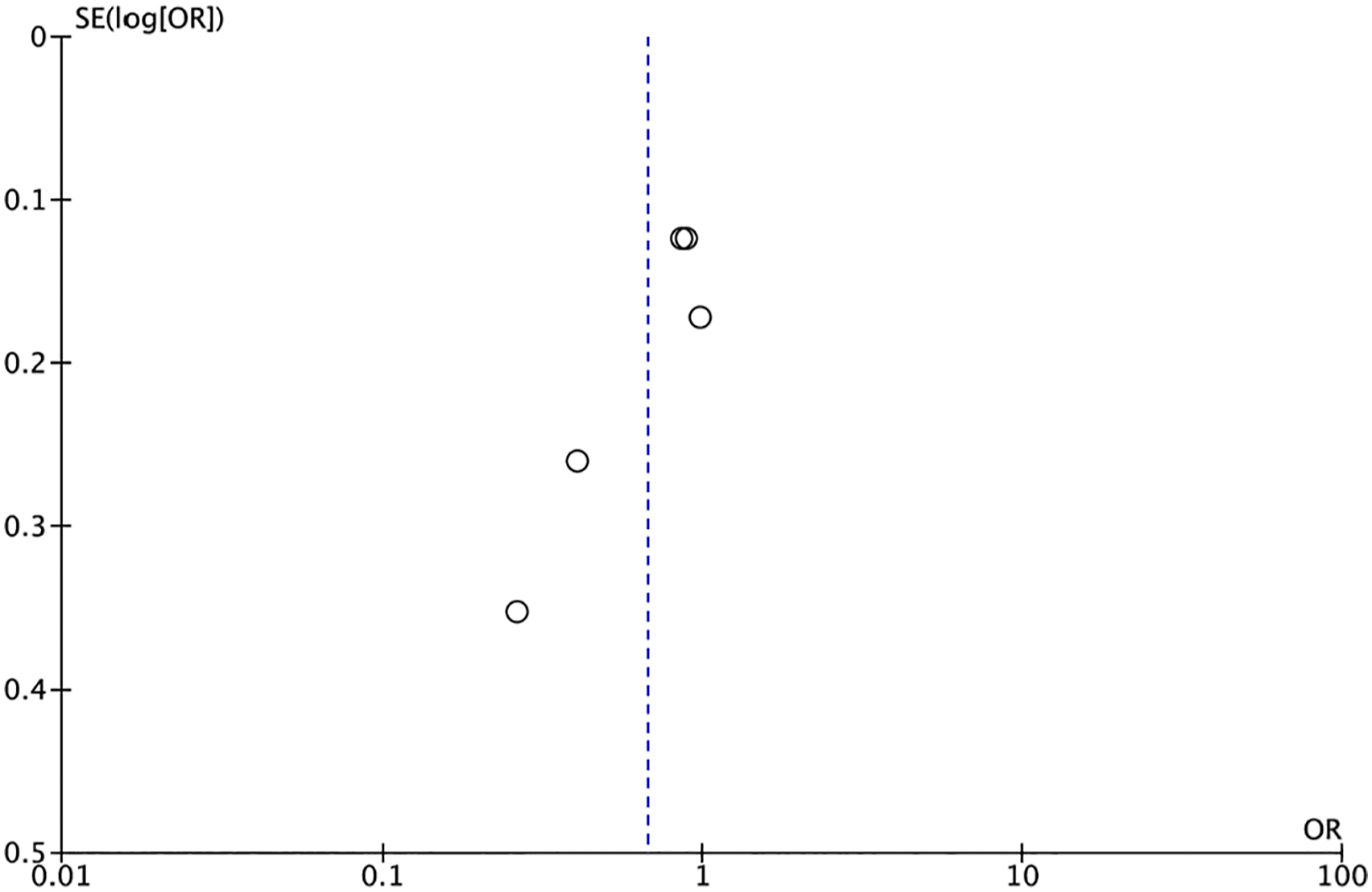
Systematic review
The IL-1RN allele was evaluated in seven studies, among which, two found an association between knee OA and IL-1RN*1 allele SNP rs419598, and the IL-1RN*2 SNP rs419598. Interleukin-1RN*1 SNP rs419598 decreased the risk, while the IL-1RN*2 polymorphism increased the risk of knee OA in both studies. One study evaluated different IL-1RN SNP, (11,100 known as rs315952), and found no association between these IL-1RN alleles and knee OA. Six studies investigated associations between IL-1RN genotypes and knee OA. Two of these studies associated a decreased and an increased risk for knee OA with IL-1RN*1/*1 SNP rs419598 and IL-1RN*1/*2 SNP rs419598, respectively (Table 2).
Among the nine studies, five that evaluated associations between IL-1B polymorphisms and knee OA found no associations for the IL-1B allele and the IL-1B genotypes, −511C/T (SNP rs16944), +3954 (SNP rs1143634), 5810 (SNP rs1143633), or in +3953. Three studies found an association between IL-1A and knee OA. However, one study found an association between the IL-1A-899 allele (SNP rs1800587) and knee OA but we found no matching association based on gender. Another study found no association between the IL-1A-899 allele with knee OA but associated the IL-1A-899 genotypes CC and CT with knee OA. One study found no association between IL-1A+4845 SNP allele and knee OA.
One of the nine studies evaluated IL-1R1 polymorphisms SNP rs10490571, rs12712127, rs956730, rs3917225, and rs3917318. Among these five SNPs, only two were associated with knee OA: rs956730 decreased the risk (OR, 0.74; 95% CI, 0.57–0.97, p = 0.028) while rs3917225 increased the risk of knee OA (OR, 1.38; 95% CI, 1.09–1.75, p = 0.007). Interleukin-1R1 rs956730 SNP was associated with a decreased risk of knee OA in an additive model (OR, 0.73; 95% CI, 0.56–0.96; p = 0.026); but when the age and sex were matched, no association was found between knee OA and IL-1RN SNP rs956730 regardless of whether it was dominant, recessive, or additive. Interleukin-1RN SNP rs3917225 genotypes GG, dominant GG+GA, recessive GG, and an additive model were associated with an increased risk of knee OA. The correlation persisted in age- and sex-matched dominant (adjusted OR, 1.47; 95% CI, 1.04–2.07, p = 0.030), recessive (adjusted OR, 1.75, 95% CI, 1.08–2.85, p = 0.023), and additive models (adjusted OR, 1.40; 95% CI, 1.09–1.79; p = 0.007).
One study evaluated the associations between the IL-1RN polymorphism and matrix metalloproteinase-9 (MMP-9), tissue inhibitor of matrix metalloproteinases-1 (TIMP-1), as well as hyaluronic acid (HA) and pentosidine (PEN), which are cartilage oligomeric matrix proteins (COMP) and markers of synovitis and cartilage destruction. They found no associations between IL-1RN and any of these proteins. Some studies also investigated the haplotype of IL-1 family polymorphisms. Interleukin-1B -511C/T-IL-1RN VNTR was not associated with knee OA, but the IL-1A(889)-IL-1B(3954)-IL-1B(511)-IL-1RN VNTR haplotype 1-2-1-1 (C-T-G-4) decreased the risk of knee OA (OR, 0.31; 95% CI, 0.11–0.84, p = 0.011). Age- and sex-matched models also found an association between this haplotype and knee OA in women (OR = 0.14, 95% CI = 0.04–0.47, p < 0.001). Interleukin-1A(889)-IL-1B(3954)-IL-1B(511)-IL-1RN VNTR haplotype 1-1-1-1/1-1-2-1 (C-C-G-4/C-C-A-4) also decreased the risk of knee OA (OR, 0.46; 95% CI, 0.22–0.94, p = 0.021) as did haplotype 1-2-1-1/all (C-T-G-4/all) (OR, 0.30; 95% CI, 0.10–0.81; p = 0.008).
Correlation between IL-1 gene polymorphisms and PROM and laboratory findings in knee OA.

WPS = WOMAC pain subscale; WPFS = WOMAC physical function subscale; WSS = WOMAC stiffness subscale.
Correlation between IL-1 gene polymorphism and ROA grading in knee OA.

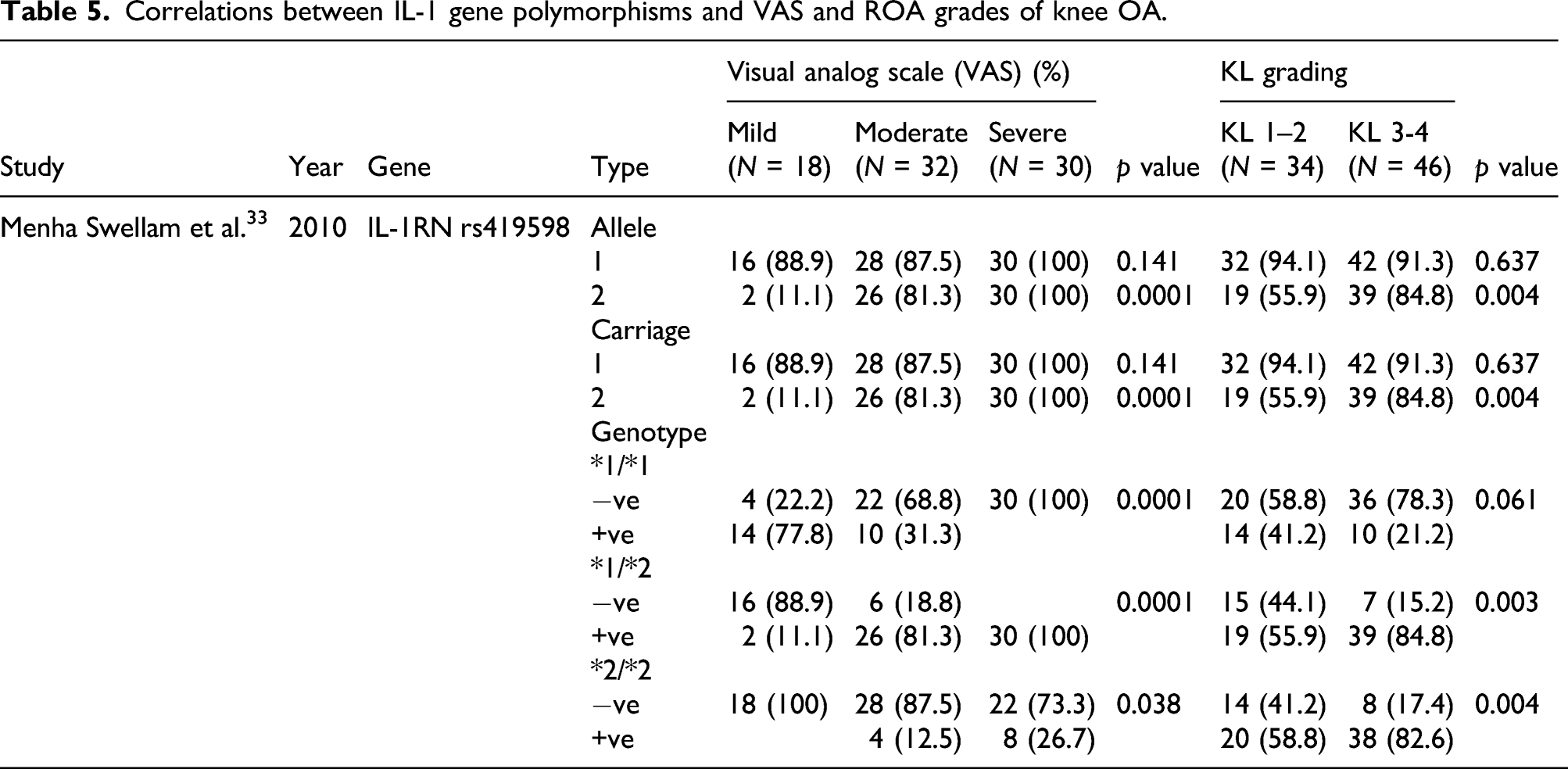
Correlations between IL-1 gene polymorphisms and VAS and ROA grades of knee OA.
Discussion
We found that IL-1RN*2 allele and IL-1RN*1/*2 genotype rs419598 increased the risk of knee OA, whereas the IL-1RN*1/*1 allele decreased the risk of knee OA. Furthermore, IL-1RN VNTR was correlated with knee OA in Caucasians. We were unable to analyze this correlation in Asians because only one study has evaluated this in Asian population. Since only a handful of studies have compared IL-1 gene polymorphisms between knees with OA and healthy knees, only the IL-1RN VNTR could be assessed in a meta-analysis.
Genetic factors are among the many causes of OA, including that of the knee. Although several studies and meta-analyses have assessed various sites of OA, the results have remained inconclusive.17–21 The conflicting results among studies were probably due to factors other than genetics, such as the environment, age, gender, ethnicity, and BMI. Combinations of factors and the contributions of individual factors that influence knee OA remain unknown. 22–24 In addition, the sites of knee OA, whether medial or lateral, affect the risk for severity or progression.25,26
Cytokines play key roles in the pathogenesis of synovitis and cartilage destruction in OA. 27 Various differentially expressed cytokines due to polymorphisms in the genes encoding cytokines such as IL-1 also play important roles in cartilage and bone destruction in OA. Interleukin-1α, IL-1β, and their naturally occurring antagonist (IL-1RA) are encoded by genes located within the 430-kb region, known as the IL-1 gene family. The IL-1A, IL-1B, and IL-1RN genes produce IL-1α, IL-1β, and IL-1Ra, respectively. Interleukin-1Ra counteracts the IL-1 action by binding to the IL-1 receptor without activating it. 28
The results of the correlations between IL-1RN VNTR and ROA were inconsistent. They did not significantly differ based on KL grading, but did so when evaluated with dichotomous data; the frequency of IL-1RN VNTR was lower for severe knee OA. Wu et al. 29 also found that IL-1RN rs380092 and rs315952 decreased the risk of knee OA, while other specific IL-1RN and IL-1R1 increased the risk of knee OA. Attur et al. 30 also found that the IL-1RN SNP rs419598, rs9005, and the haplotype rs419598/rs315952/rs9005 function as protective factors in knee OA. The differences in the results of these studies were probably due to the OA grading on the medial or lateral compartment as revealed by Hunter et al., 31 who found that OA in the lateral and medial compartments differs based on risk factors for disease severity and progression. Attur et al. 8 compared the lateral and medial compartment OA and found a difference in plasma levels of IL-1Ra.
This systematic review showed that IL-1B polymorphisms were not associated with knee OA in Asian and Caucasian populations. These findings differed from those of Kanoh et al., 20 who evaluated 303 normal knees and 56 ROAs. They found that the rate of ROA was significantly higher for the T/T, than the C/C or CT genotypes. Others have also associated IL-1B SNP with knee OA.18,21 Barakat et al. found that the same concentrations of IL-1B in different compartments of the same knee result in different region of cartilage damage. 32 Knee joint chondrocytes are more sensitive to stimulation with IL-1B than other joints, 33 but the present systematic review did not found an association between IL-1B polymorphisms and knee OA. Whether IL-1B polymorphisms are associated with knee OA remains inconclusive.
Moxley et al. found no association between IL-1A SNP, IL-1B SNP, IL-1RN, and knee OA. Evaluations of these three genes using extended haplotypes found no association between genetic variations in the IL-1 region and knee OA. 17 In contrast, others have found that an extended IL-1 region haplotype is associated with knee OA.34,35
This meta-analysis has some limitations. Firstly, the IL-RN VNTR polymorphisms studies were heterogeneous, although subgroup analysis did not identify any potential sources. This might have resulted in misunderstanding the meta-analysis results. Secondly, the total sample size of the eligible studies was insufficient to reach a convincing conclusion. Thirdly, all the samples in this study were taken from peripheral blood which can result in bias. As we know, some SNP of IL-1β also correlates with heart disease or outcome of other disease. Lastly, our conclusion of the meta-analysis was based on unadjusted estimations. Many factors influence OA, including gender, ethnicity, BMI, lifestyle, and hormones. Since complete data were unavailable, we could not conduct meta-regression analyses, despite the studies included in the analysis being of good qualities.
Conclusion
We concluded that several IL-1RN alleles and genotypes play a role in its severity, but other genetic variations in the IL-1 region were still conflicting in its association with knee OA. Further study is needed to evaluate the association between genetic variations in the IL-1 region and knee OA especially across different ethnicities.
Supplemental Material
sj-pdf-1-osj-10.1177_23094990221076652 – Supplemental Material for Interleukin-1 genetic polymorphisms in knee osteoarthritis: What do we know? A meta-analysis and systematic review
Supplemental Material, sj-pdf-1-osj-10.1177_23094990221076652 for Interleukin-1 genetic polymorphisms in knee osteoarthritis: What do we know? A meta-analysis and systematic review by Nicolaas C Budhiparama, Imelda Lumban-Gaol, Herawati Sudoyo, Rahadyan Magetsari and Tri Wibawa in Journal of Orthopaedic Surgery
Footnotes
Declaration of conflicting interests
NCB reports personal fees, non-financial support and other from DePuy Synthes, other from Zimmer Biomet, other from Gruppo Bioimpianti, outside the submitted work; and Editorial board/Reviewer: VJSM, CORR, J of Arthroplasty, AJSM, JOS, KSSTA, JISAKOS, BJJ, BJO, Knee Journal, OJSM, JIMR, JSHS, KSRR, CiOS, ANZ Journal of Surgery, JOINTS, Arthroplasty, Arthropaedia, Hip & Knee Journal. The rest of the authors declare no conflict of interest.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Registration
This study was registered in OSF with DOI 10.17605/OSF.IO/W2CY3.
Data availability statement
The data that supports the findings of this study are available within the article [and its supplementary material]
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
