Abstract
A microplate technology based on a Compact Cell Culture Disc (CCCD) is described. This new tool employs a multitude of miniature cell culture chambers set into a planar compact disc suitable for computer-aided microscopic or spectroscopic analysis of single cells or cell clones in large numbers. The aim is to perform automated, accurate and reproducible studies of morphologic and physical-chemical parameters, in rapid succession under physiologically and optically ideal, uniform and reproducible conditions.
Keywords
INTRODUCTION
The CCCD prototype here described is essentially a “sandwich” consisting of two 10 cm silicon wafers microstructured by chemical micromachining techniques. In properly designed laboratory settings, this new device will permit Computer-Aided cell SEeding, Microcloning, visual or instrumental Analysis, Telemetric Evaluation (“CASE-MATE”) and eventual storage (in vitro or after fixation) of hundreds to thousands of separately confined biological specimens, in the form of single living cells or monoclonal cellular aggregates. The silicon-based CCCD is primarily intended for “predictive assays” (Table 1) based on tumour biopsy specimens from individual patients. Presently, applications are prepared in support of a research program aiming at the evaluation of new cancer treatment modalities based on the “new radiations” in radiotherapy (protons and other light ions, neutrons, monochromatic X-rays, alpha- or beta-emitting radionuclides, visible or near-visible light). The CCCD design concept appears to be of general interest, however, in the context of cellular biotechnology, because it appears to be applicable to many other clinical or industrial R&D activities in need of computer-aided multiparameter assays (or high-throughput single-parameter screening) of monoclonal cell specimens in large numbers.
Tumour biopsy parameters using predictive assays
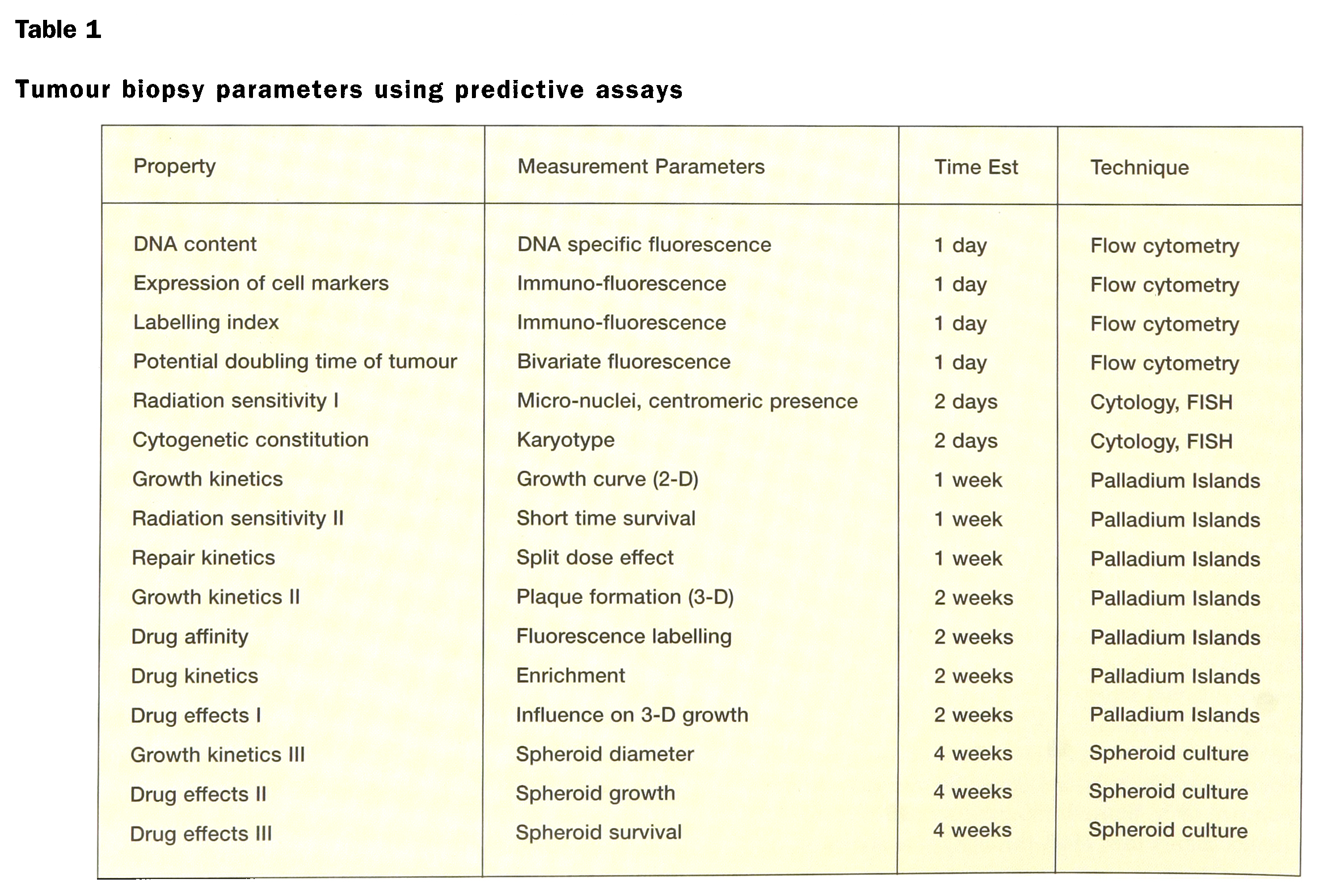
The microchambers offer advantageous conditions for culture of daughter cell populations (up to 1000–10,000 cells per microchamber) from single cells (cell cloning). The large number of cell clones produced on one single CCCD could be studied individually, in rapid succession. The uniform and reproducible culture conditions provided are of significant advantage, especially for bioassays in tumour diagnosis and for tumour therapy evaluation (cytological studies and investigation of tumour response to treatment). To this end, single dispersed cells from the tumour can be guided into individual CCCD micropits by computer-aided cell seeding. (Alternatively, one can use randomized seeding from a diluted tumour cell suspension, a technique that allows implantation and cloning of 1 or 0 cells per micropit.) A tumour biopsy containing typically some 100,000 to 1,000,000 cells could then be conveniently analysed as several thousand individually isolated tumour and stroma cell clones in primary cell culture.
MATERIALS AND METHODS
The CCCD — a 10 cm silicon-based compact cell culture disc: The CCCD is designed to permit the implantation of single cells individually, each one in its own separate micropit. The individual micropits consist of shallow (typically 50–300 micrometer deep), cylindrical depressions defined by holes (800 or 1600 micrometer in diameter) in an “upper wafer” that faces geometrically conform optically transparent quartz windows in the “lower wafer”. If desired, the micropit could be provided with a coated spot offering cellular anchorage, for example a “palladium island” (1), centrally deposited on the agarose-covered quartz window, to permit optimal conditions for microscopic observation of attached single cells or cell clones (Fig. 1).
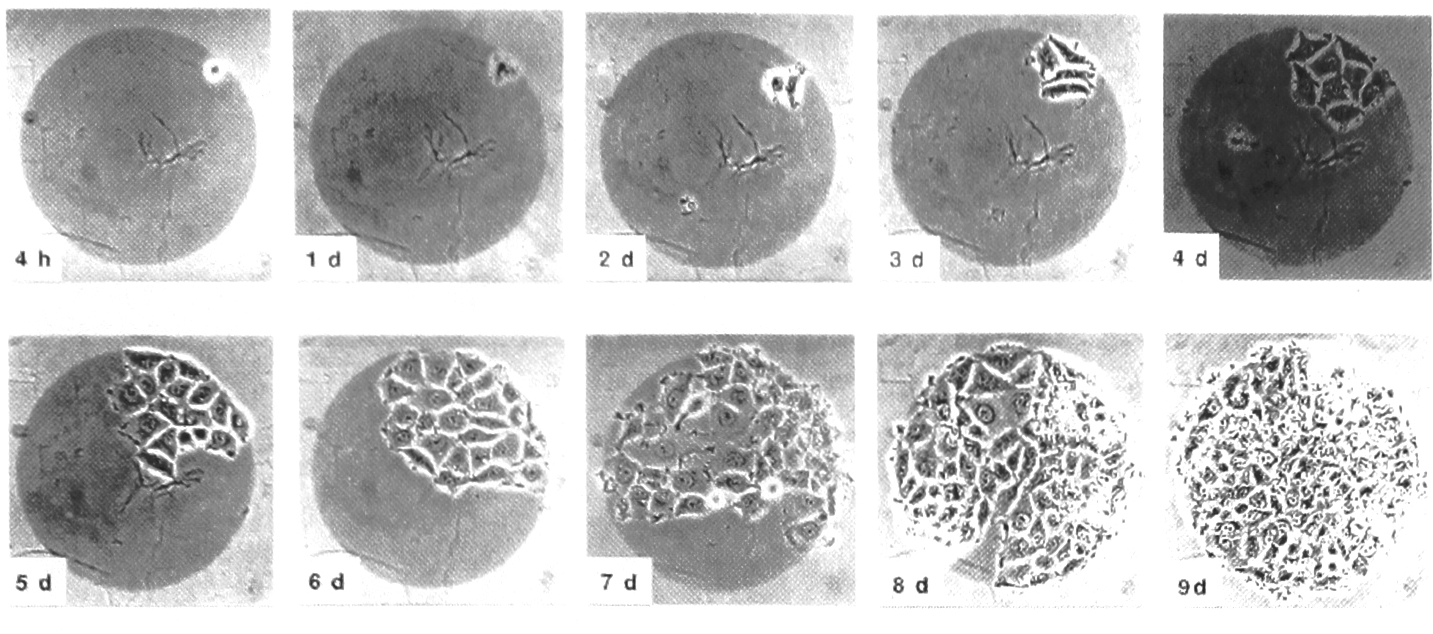
Single adenocarcinoma cell seeded on Pd island to produce cell clone. From ref.1 (courtesy Elsiver, Amsterdam)
After implantation (“seeding”) the micropits can be covered by a thin plastic membrane forming a semipermeable cover foil for the CCCD. Each single pit then forms a “microchamber”, in the form of a sterile microscopic compartment inaccessable to virus or bacteria. In this way it is also possible to achieve optimized and uniform supplies of respiratory gasses, nutrients, antibiotics, reagents etc. to all the CCCD microchambers (Fig. 2a).
The patterns of holes and windows are geometrically conform in their layout on the two wafers, respectively (Fig. 2b).
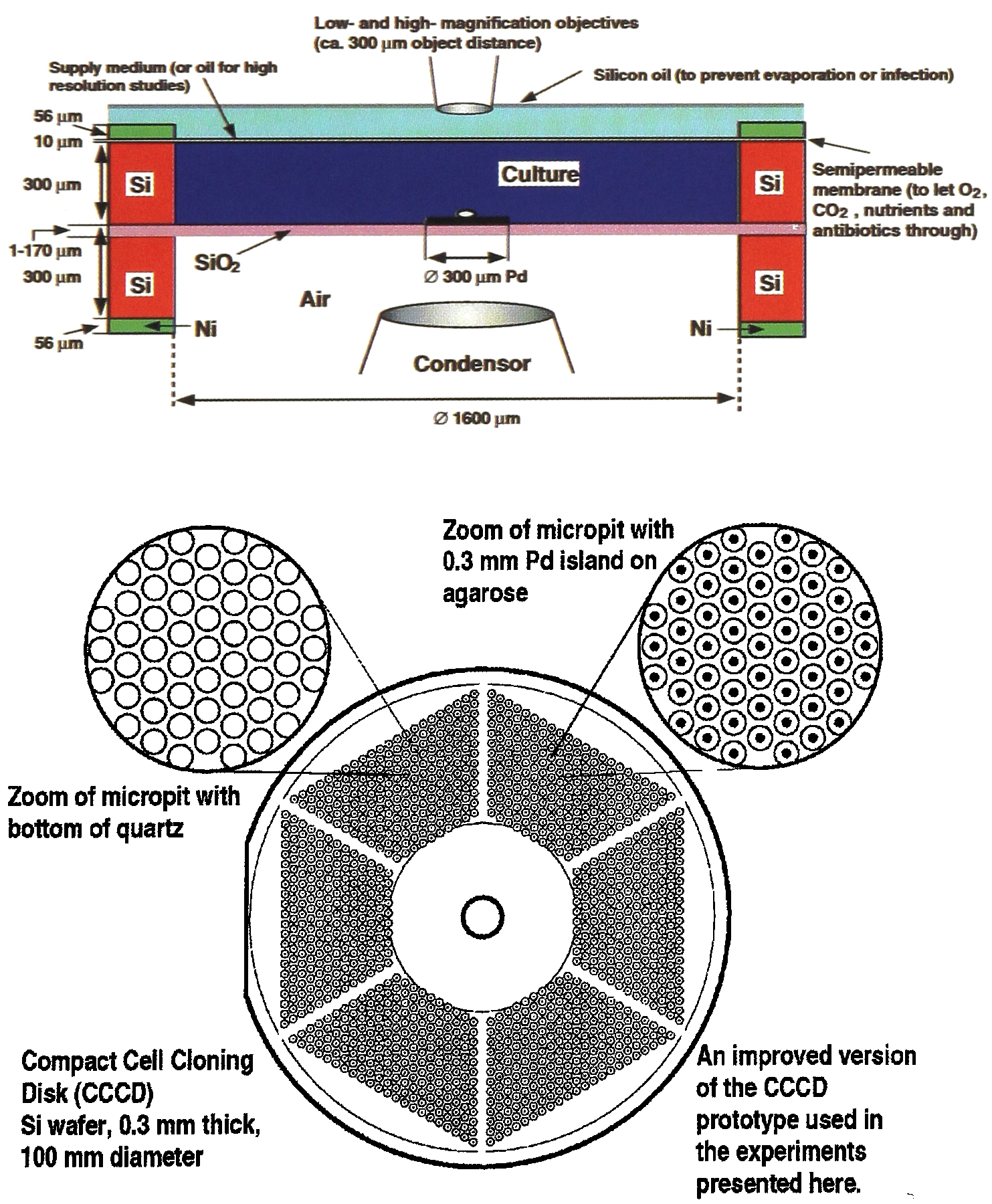
Design features of a 10 cm CCCD (pit diameter 1.6 mm):
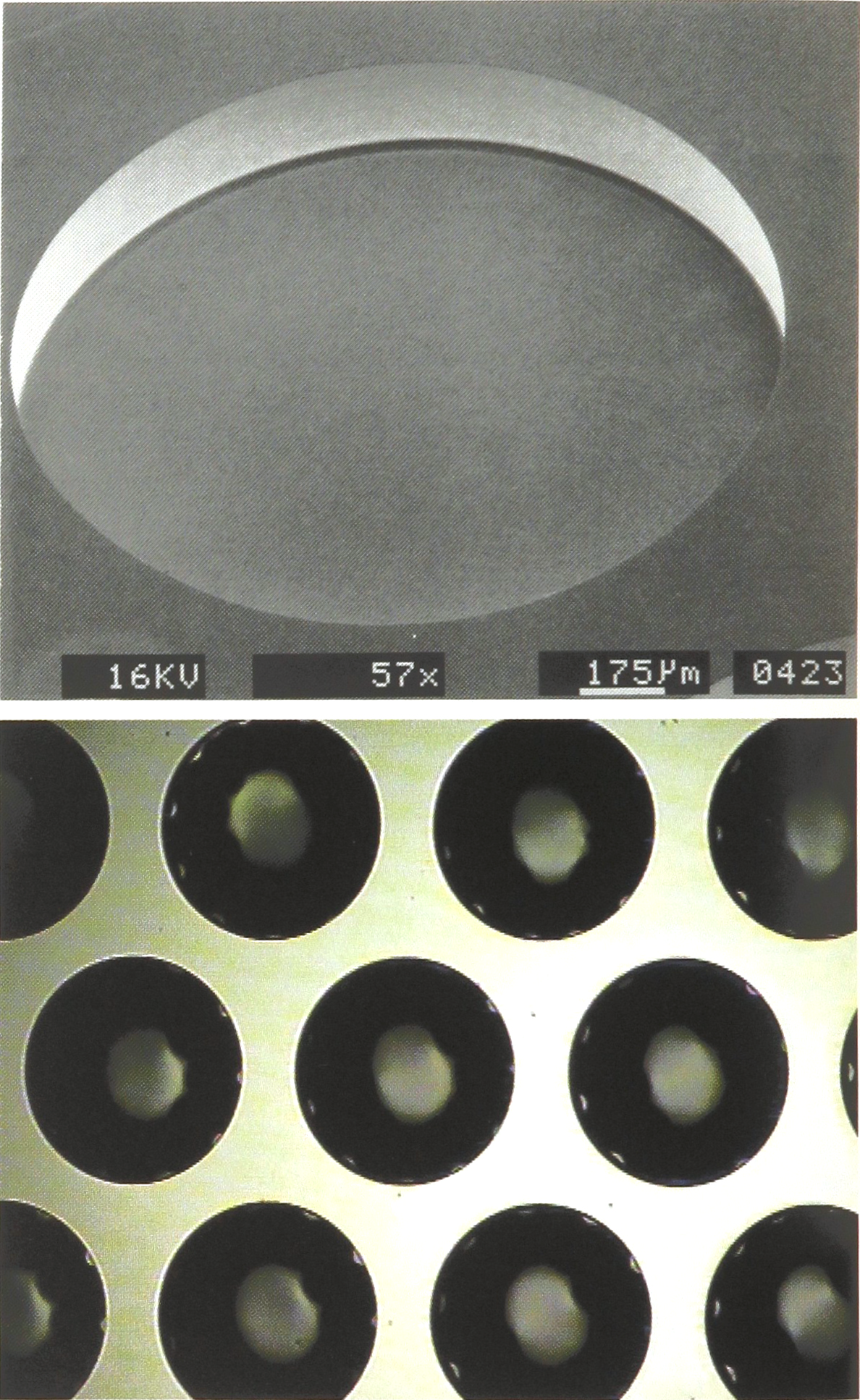
Micrographs showing details of a CCCD pit-array in accord with Fig. 2, manufactured by “Chemical micromachining”. (a) SEM of a single 1.6 mm pit with a ca. 2 micrometer thick quartz window. (b) Optical micrograph showing part of the CCCD pit-array.
a) in KOH following standard procedures (2). Pits produced by similar methods have been described before (3).
b) by dry etching with a high-rate, anisotropic etching process developed by the STS Corporation. Fig. 3a shows a SEM of a 1.6 mm diameter micropit produced in such a way. The advantage of this method is that a higher density of pits can be realised per unit area because of the almost vertical sidewalls produced by this process.
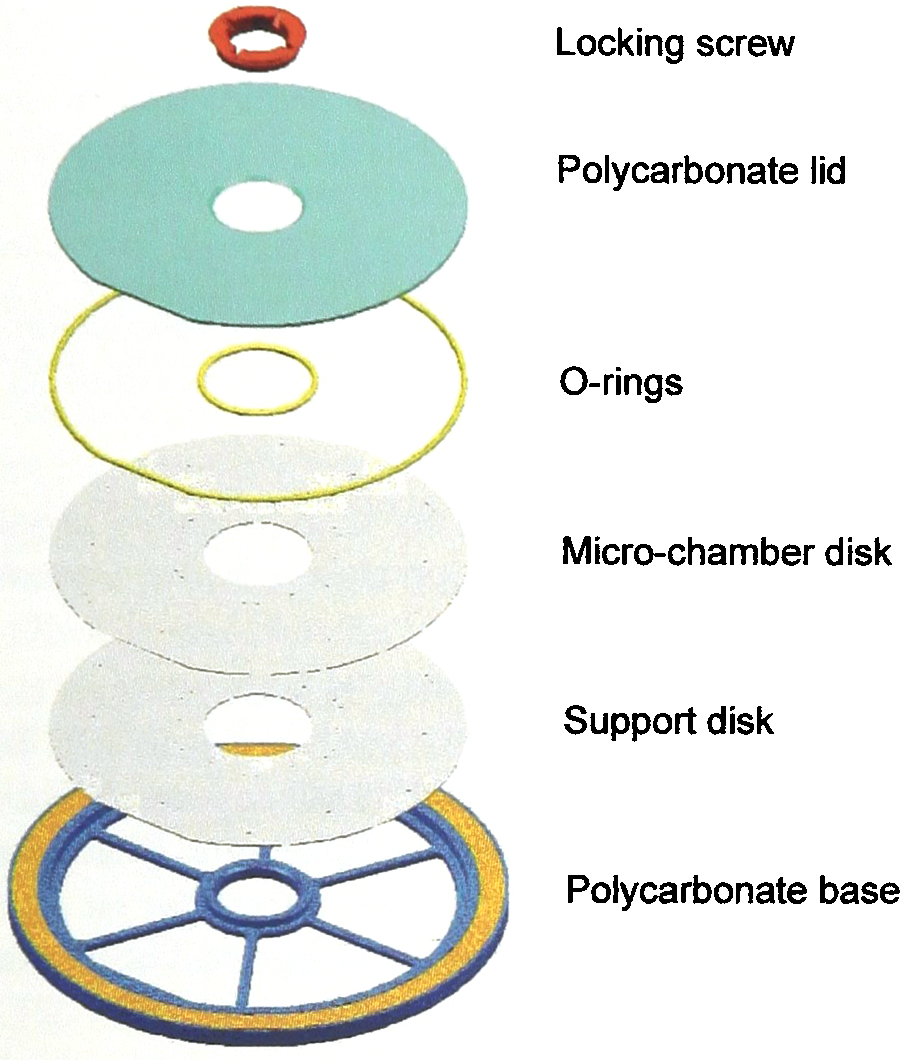
Embedment of a 10 cm CCCD in a 12 cm CD structure provides a common source of low-molecular weight medium components for the microchambers, mechanical protection and a convenient structure for opto-mechanical data storage. The yellow band represents space reserved for a megabyte file containing specimen data (in the form of a virtual bar code) and instructions for CCCD coordinate control and settings choosen for computer-aided seeding, microscopic or spectroscopic analysis, telemetric data evaluation, etc.
COMPUTER-AIDED SEEDING, MICROCLONING, ANALYSIS AND TELEMETRIC EVALUATION (CASE-MATE)
As an application example now under development, we employ confocal micros copy-based measurements (Figs. 5 and 6) permitting evaluation of the prerequisites for targeted cancer therapies in individual patients, through studies of pharmacokinetics and biological response in cell populations isolated from biopsy or surgical specimens. This new means for improved tumour therapy evaluation and treatment planning will serve a clinical community interconnected through an IT network similar to the networks for macroscopic treatment planning that are now implemented to serve the Swiss and Italian hadrontherapy communities.
An important advance the CASE-MATE system brings to oncology research in general, is the ability to measure biological parameters and simultaneously take tumour heterogenety into account (6). Major challenges to targeted therapy are to bring sufficient amounts of a toxic principle, e.g. boron in the context of neutron capture therapy, selectively to the cancer cells and to reduce background therapy dose as much as possible. Pretreatment evaluation of boron distribution, through neutron capture microradiography, tumour-genetic characteristics, and also normal-tissue radiosensitivity should employ parameters that can all be addressed using CASE-MATE system. Furthermore, evaluations of tumour clonogens and cell-death fractions using microcloning and cytotoxicity assays are possible, and the role of genetic instability in clinical tumour eradication and tumour promotion as a secondary consequence of the therapy can be studied. To date the design and preliminary testing of the prototype CASE-MATE system has been performed. Successful primary cultivation and microscopic analyses of single cells and clones isolated from clinical biopsy specimens have also been achieved.
DISCUSSION AND CONCLUSIONS
In the fields of molecular and cellular genetics, microbiology, oncology, radiobiology, pharmacology, etc. there is a need to culture individuals cells, cell clones and even multicellular microorganisms in order to study their growth, morphology and molecular-genetic, biochemical or biophysical characteristics. The system described permits the isolation, continued culture, and investigation of single cells, clones, and other biological microspecimens in large numbers. It was especially designed for computer-aided investigations of growth, and the structural and functional characteristics displayed during the development of monoclonal cultures.
During the preparations for these investigations, different anchorage-dependent cell lines were cultured on palladium islands (Fig. 1). A fundamental idea behind the CCCD design is the placement of such cell-attracting “coated spots” within a honeycomb of micropits which can be closed and sealed with a common cover. To the selected cells in any given micropit, specific nutrients and agents can be given in order to investigate the effects on cell populatoin growth. A pair of superimposed, circular 0.3 mm thick silicon wafers forms the basis of the CCCD. Its diameter can be varied from 5 to 20 cm depending on the number of micropits required. This structure can best be compared with a standard compact disc filled with a honeycomb of micropits.
The individual micropits of the CCCD permit the study of individual cells and cell clones, in both small and large numbers, i.e. from single cells up to a few thousand cells, under optically optimized conditions. The covered micropit (the “microchamber”) with its defined medium and growth conditions forms a sterile microlaboratory providing conditions for control and examination of the individual cells and their growth under favourable optical conditions. Control of both the chemical and physical milieu of the cells is possible. Autofluorescence (Fig. 5) or the fluorescence from chemical markers (Fig. 6) provide important measurement parameters that can preferably be analysed by confocal laser scanning microscopy adapted to the optical features of the CCCD microchambers. Both the whole CCCD and the individual growth chambers can be maintained under sterile or non-sterile conditions; for short term, long term, and permanent experimentation, transport and storage.
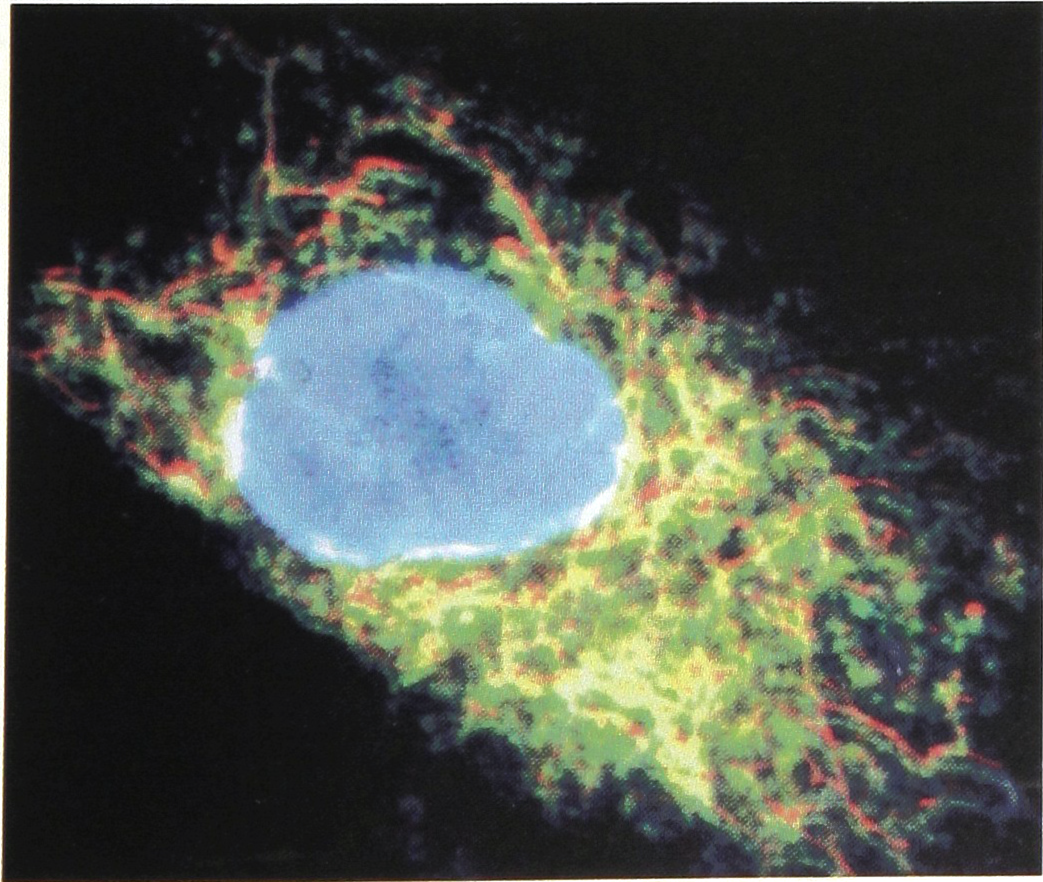
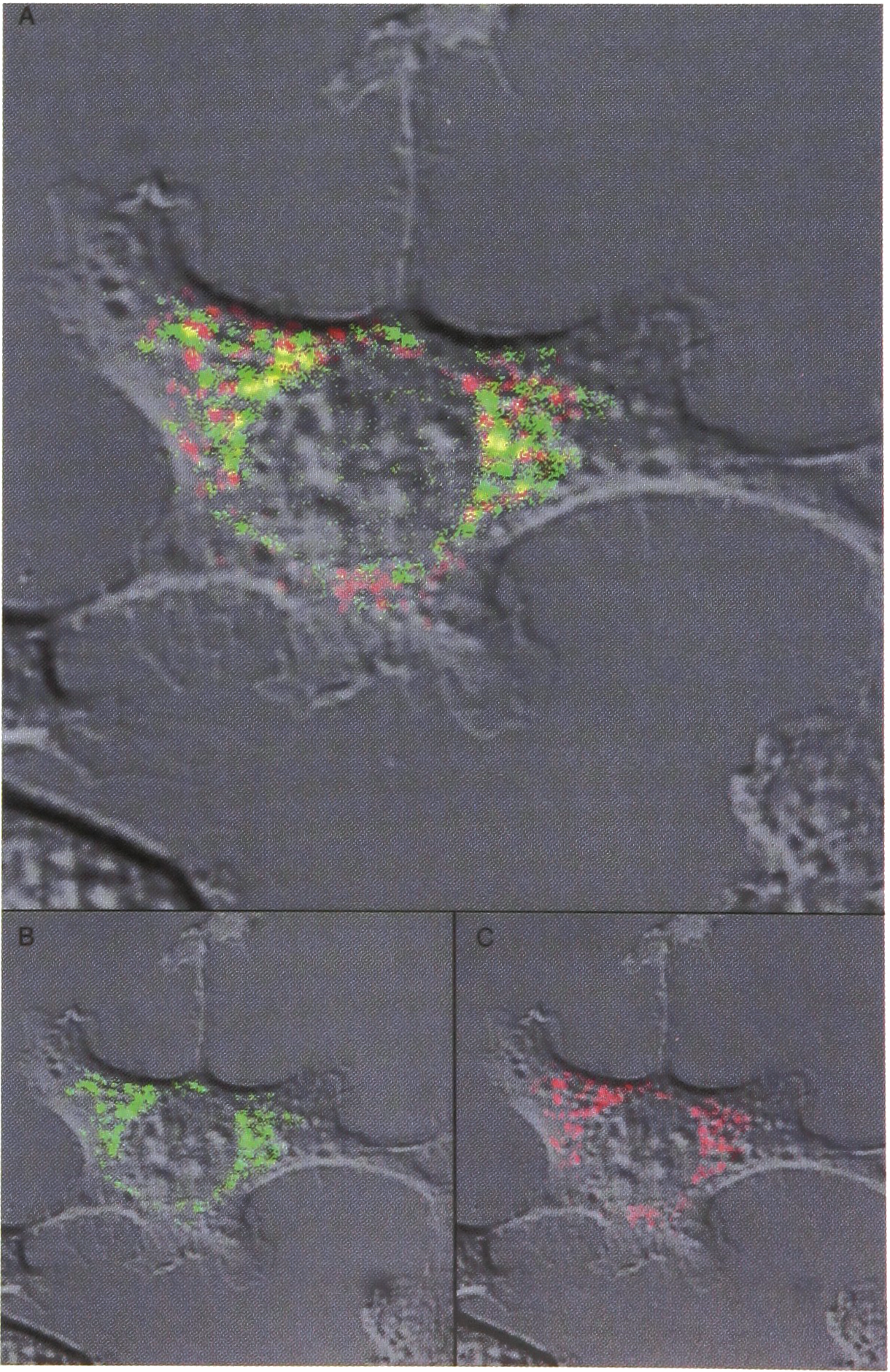
To illustrate the use of autofluorescence, as an alternative to observation through intravital staining (cf. Fig. 6) for outlining the position and contour of cell nuclei by computer-aided image analysis of cell growth: (b) Laser scanning microradiograph (LSM) of auto-fluorescence (green) in fibroblasts, and (c) LSM of the corresponding rhodamine-fluorescence (red). Autofluorescence is mainly produced by NADH in mitochondria as demonstrated by the yellow colour in (a) that represents colocalization of the green and red fluorescence. From Ref. 4. (courtesy J. Microscopy)
LSM visualization of the Hoechst substance affinity to cellular DNA. From ref. 5 (courtesy J. Marelius, Uppsala University, Sweden)
Footnotes
ACKNOWLEDGEMENTS
We are indebted to R. Widmer of IBM Research, Zurich (CH), for etching some wafers with the STS process and to Microcloning CCCD AB, Jönköping (S) for financial support.
