Abstract

Over the past two decades methodologies used to analyze accurately nucleic acids and determine the sequence or expression of a gene have improved continuously Today, automated DNA-based assays and test systems are already available to a limited research market. However, before DNA-based tests will find broad applications in the healthcare and other industries, the development of technologies to lower substantially costs and speed up DNA analyses and data interpretation will be required.
The latest technological advances paving the way to integrated and automated DNA-based diagnostic tests were presented during IBC's recently held conference on “Biochip Arrays and Integrated Devices for Clinical Diagnostics”, March 5–6, 1997, in San Diego. The meeting was opened by Dr. Stanley Abramowitz, Program Manager of the Advanced Technology Program (ATP) at the National Institute of Standards and Technology (Washington, DC). ATP's mission is to stimulate US economic growth by supporting the development of high-risk, high-payoff technologies through industry-driven, cost-sharing partnerships. In the genomics area, the scope of a specific ATP program that focuses on “Tools for DNA Diagnostics” is the development of DNA-based technology platforms that make use of the data generated from the human genome project. Such platforms will form the basis to develop commercially viable applications in specific industries, such as the healthcare industry (e.g., drug discovery and disease diagnosis), forensics, agriculture, veterinary medicine, environmental monitoring, and the food industry.
According to Dr. Abramowitz, commercialization of DNA-based diagnostic tests faces great technological challenges in the areas of sample preparation, sample analysis (assay and detection capabilities), and data management. For example, the trend towards miniaturization of sample sizes and assay volumes requires manipulating DNA in nanoliter volumes and increasingly more sensitive detection systems. The solutions to these challenges will require expertise from a broad range of disciplines, such as molecular biology, surface chemistry, computer science, and engineering. The key to commercial success will result from successful integration of miniaturization, parallelization and automation of all steps involved in DNA-based testing, ranging from sample preparation to analytical procedures and data management.
Currently, promising approaches to analyze DNA use electrophoresis, hybridization, primer extension or amplification-based assays, often in conjunction with one another. In the area of electrophoresis, the technology has moved from ultra-thin slab gels to capillaries with microfabricated channels to electrophoretic chips. For example, Dr. Gregory Went at CuraGen Corporation (Branford, CT) presented the company's high-throughput microchannel array system that allows for the analysis of differential gene expression in diseased and normal tissue at the rate of 500 genes a minute. With this rate, the bottleneck for using differential expression profiles in advanced disease diagnosis has shifted from assay capacity to sample preparation.
Successful integration of a microfabricated electrophoretic chip with a silicon-based PCR chip has been presented by Dr. Phil Belgrader from the Lawrence Livermore National Laboratory (Livermore, CA). Using micro-electromechanical system technology, the laboratory has developed a portable, inexpensive, and high-throughput DNA chip-based instrument, the Miniature Analytical Thermal Cycler Instrument. The successful development of this instrument has demonstrated that commercially viable DNA-based instrumentation will become a reality. As an extension to electrophoresis-based analytical systems, Dr. Joseph Monforte at GeneTrace Systems, Inc. (Henlo Park, CA), described the development of an ultrahigh-throughput sequencing system using mass spectrometry. GeneTrace's spectrometric machine analyzes DNA fragments as they fly through a vacuum rather than traveling along a gel. Mass spectrometry enables DNA sequence analysis with a 300 fold higher resolution, at a 100 fold greater speed, and at one fiftieth of the cost. Using their spectrometric technology, the company proposes the feasibility of a benchtop DNA sequencer with the capability of resequencing over one million bases a day to screen for mutations associated with cancer and genetic diseases.
In the area of DNA analysis, commonly used approaches such as hybridization-based assays have seen the technology shift from hybridization using soluble DNA probes to DNA arrays immobilized on a chip. During the two-day meeting, the development of a variety of array chips was discussed, an example of which is illustrated in Figure 1. Affymetrix's integrated gene chip system contains an automated fluidics station, a photolithographically constructed oligonucleotide array chip (see figure 2) with up to 100,000 probes per 1.5cm2, a laser-based scanner, and software for data analysis. Dr. Glenn McGall from Affymetrix, Inc. (Santa Carla, CA), presented the development of three types of gene chips, the resequencing chip designed to detect gene mutations using short DNA probes, the genotyping chip to detect polymorphism in larger disease genes and the expression chip to determine gene expression profiles. In collaboration with several pharmaceutical and biotechnology companies, Affymetrix's gene chip systems are under development for many clinical applications, such as detection of infectious, genetic and neoplastic diseases, testing drug resistance of infectious agents, and predicting drug reactions in individual patients.
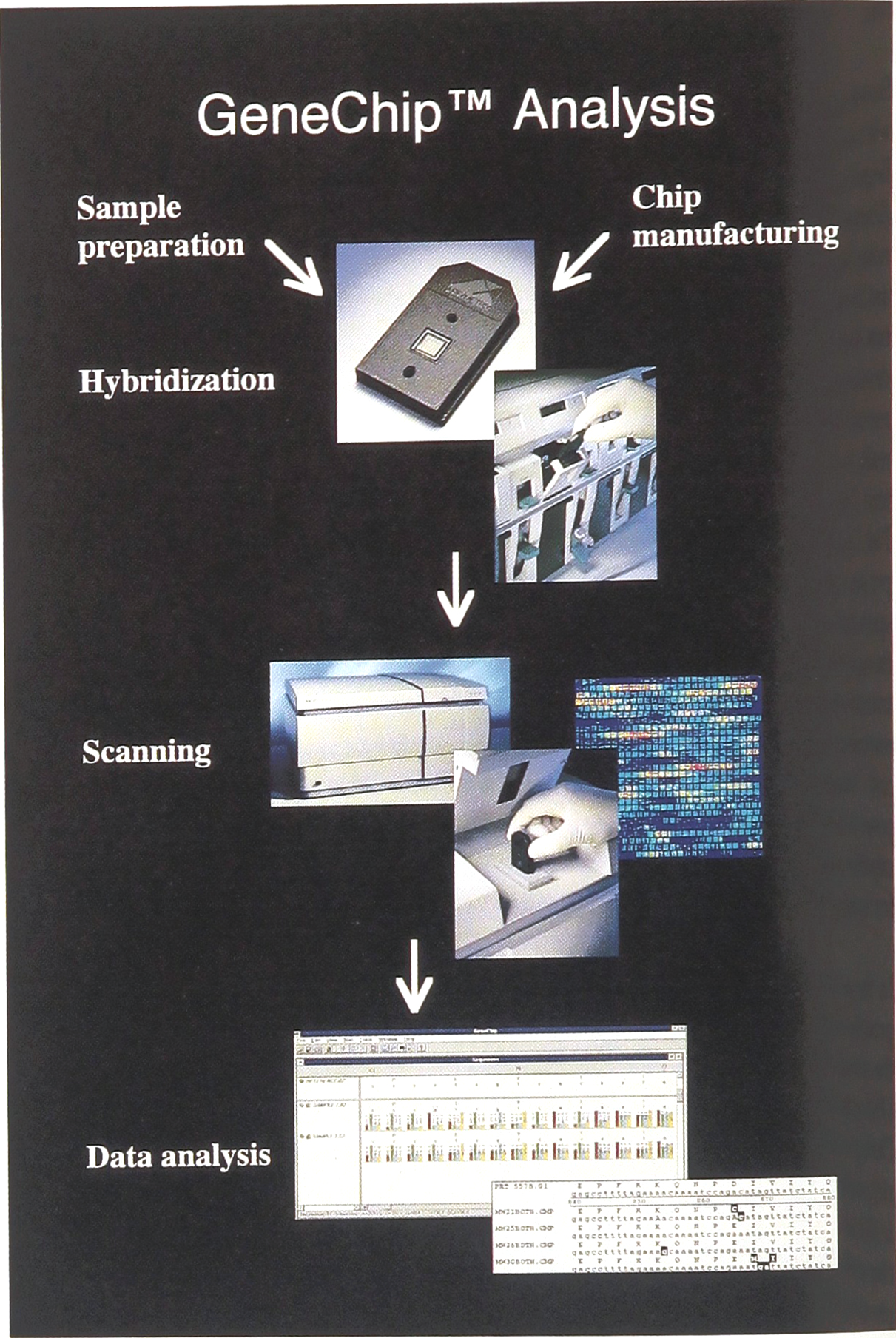
During the two-day meeting, the development of a variety of array chips was discussed
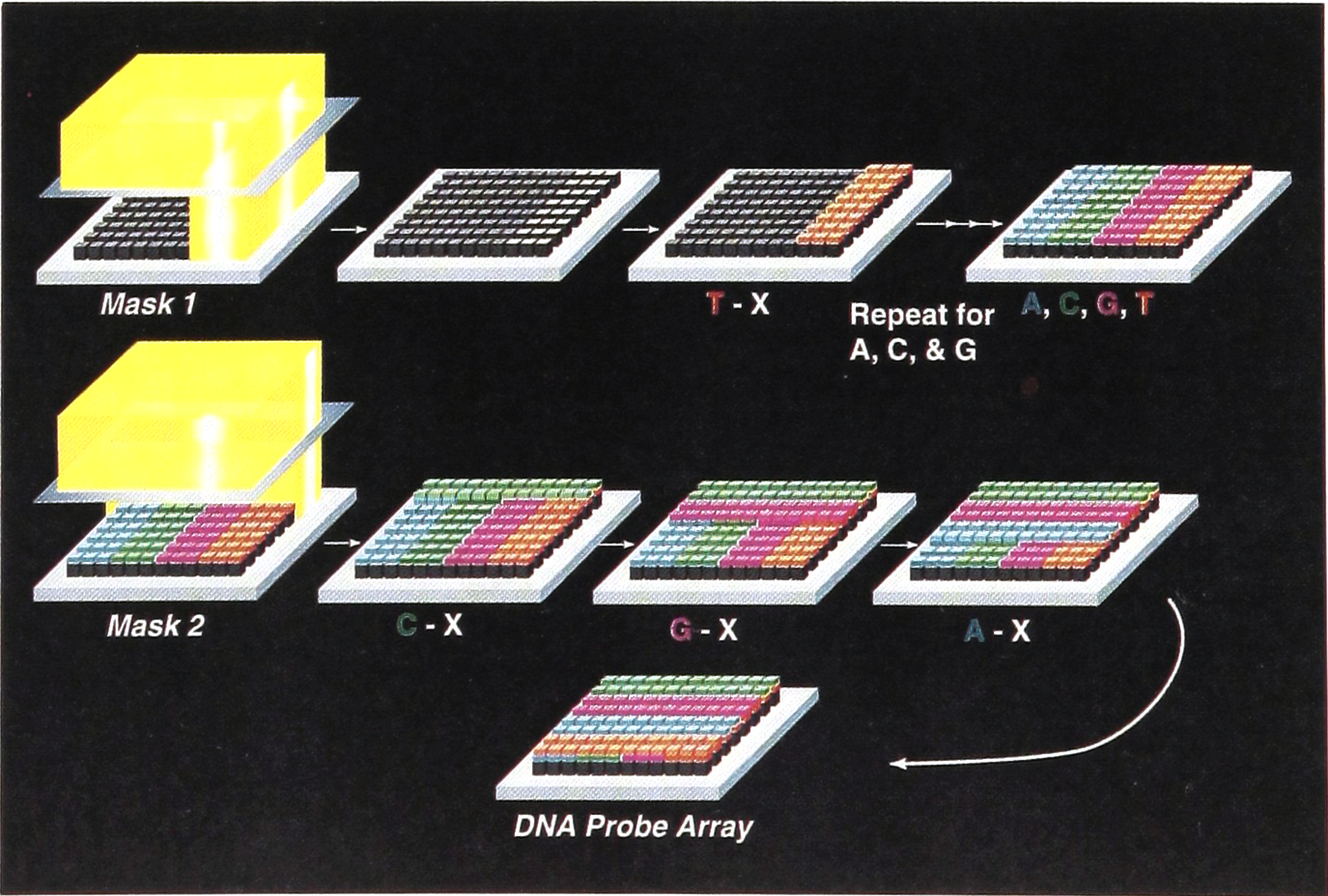
Affymetrix's integrated gene chip system contains an automated fluidics station, a photolithographically constructed oligonucleotide array chip
Other examples of integrated hybridization-based assay systems have been presented by Dr. Michael Heller at Nanogen Inc. (San Diego, CA), Dr. Mich Eggers at Genometrix Inc. (The Woodlands, TX), and Dr. Robert Matson at Beckman Instruments Inc. (Fullerton, CA), to name just a few. The electronic device developed by Nanogen is a fully integrated microlaboratory with a multiplex gene chip at its core. Each chip contains 25 independent test locations where samples can be analyzed using multiple sequence sites in the probe and in the sample. Using their device, matches and mismatches between probe and sample sequences can be detected within seconds. The chip's multiplex analytical capability makes Nanogen's prototype device ideal for diagnosis of infectious diseases in a point-of-care type of clinical setting. In contrast, Genometrix's and Beckman's microchips, with their high-density oligonucleotide arrays, have been designed for use in high-throughput test laboratories screening thousands of samples quickly and inexpensively (less than one dollar a test). Beckman's integrated genosensor is currently being tested for its usefulness in early detection of cancer based on the presence of mutations in the ras oncogene.
Advantages of primer extension over hybridization-based assays have been discussed by Dr. Tony Murray at Pharmacia Biotech, Inc. (Piscataway, NJ), and Dr. Steve Head at Molecular Tool, Inc. (Baltimore, MD). While hybridization-based diagnostic tests are limited to screening for mutations at known sites, array primer extension (APEX) technology in addition allows for detection of unanticipated polymorphism and mutations. The ability to uncover unknown mutations has important clinical applications, e.g. in the diagnosis of cancer using a p53 assay or identification of drug resistant TB strains. Pharmacia's APEX technology involves ex situ DNA sequencing followed by surface applications of the synthesized oligonucleotides. This approach has the advantage of great accuracy, thereby reducing quality control issues, but limits the test volume due to low array density on the chip.
Finally, alternative analytical approaches using novel enzymes were presented toward the end of the meeting. Dr. Victor Lyamichev at Third Wave Technologies, Inc., (Madison, WI), discussed the use of a novel DNA exonuclease (CFLP) to detect mutations. CFLP generates highly specific, easily quantifiable patterns of DNA fragments due to the enzyme's specificity for DNA secondary structures. CFLP assays can detect mutations even in a heterogeneous sample allowing tumor cell detection from biopsies or identification of antibiotic resistant infectious agents. In the area of DNA amplification assays, Dr. Terry Walker at Becton Dickinson Research Center (RTP, NC) presented the use of two enzymes in strand displacement amplification (SDA), an isothermal and fast alternative to PCR-based amplification methods. The rapidity and accuracy of SDA-based assays is clinically useful for the diagnosis of contagious diseases where appropriate treatments need to be initiated quickly (e.g., in tuberculosis patients).
The message from IBC's meeting on “Biochip Arrays” can be summarized by a quote from Dr. Robert Matson at Beckman Instruments, the meeting's chairman of the second day: “yesterday we learned … that you now can perform PCR in the car en route to the golf course”. In other words, as we enter the next century, we will witness the introduction of quick, cheap, convenient, and reliable DNA-based diagnostics with broad clinical applications.
Proceedings from the Biochip Arrays conference are currently available. Please call IBC at 508-804-5441 for information.
