Abstract
Stem and progenitor cells derived from human tissues are being developed as cell sources for cell-based assays and therapies. However, tissue-derived stem and progenitor cells are heterogeneous. Differences in observed clones of stem cells likely reflect important aspects of the underlying state of the source cells, as well as future potency for cell therapies. This paper describes a colony analysis and picking device that provides quantitative analysis of heterogeneous cell populations and precise tools for cell picking for research or biomanufacturing applications. We describe an integrated robotic system that enables image acquisition and automated image analysis to be coupled with rapid automated selection of individual colonies in adherent cell cultures. Other automated systems have demonstrated feasibility with picking from semisolid media or off feeder layers. We demonstrate the capability to pick adherent bone-derived stem cells from tissue culture plastic. Cells are efficiently picked from a target site and transferred to a recipient well plate. Cells demonstrate viability and adherence and maintain biologic potential for surface markers CD73 and CD90 based on phase contrast and fluorescence imaging 6 days after transfer. Methods developed here can be applied to the study of other stem cell types and automated culture of cells.
Introduction
Human tissue is the source of cells for virtually all cell-based assays and cell therapies. Unless colony selection is performed, the culture expanded cells of any tissue source represent the progeny of a heterogeneous population of tissue-resident progenitors. In vitro, each progenitor gives rise to a clone of progeny, and can therefore be defined as a “colony forming unit” (CFU). The prevalence and biological performance of individual CFUs vary widely from clone to clone from a single donor, and from donor to donor. These differences are likely to reflect important aspects of the underlying health or state of the source tissue.
Methods for quantitative measurement of colony formation and biological performance have been developed using automated image analysis 1 and contributed to the development of an ASTM standard that will help advance this field across many CFU subtypes. 2 While image analysis is a valuable tool for assessing heterogeneity, there is an opportunity in basic science research to better characterize heterogeneity by selecting different clones for subculture and analysis. This also presents an opportunity in biomanufacturing to make objectively informed choices about which CFU subtypes or clones should be included or excluded from a specific cell product or cell-based assay.
Clonal analysis can be used to characterize heterogeneity of CFUs or as a tool for clonal selection for specific applications. A number of methods can be used to perform clonal analysis on stem cell populations, but each is faced with its own limitations. 3 Fluorescence-activated cell sorting (FACS) requires a large number of analyzable cells. FACS can also damage cells and therefore change the proliferative and differentiation potential of a cell population. 4 Limited dilution methods to isolate cell clones are well characterized but time-consuming and imprecise with respect to the founding cell. 5 Defining an optimal plating density for the limited dilution approach is also challenged by variation in stem cell prevalence between clinical patient samples and tissues. 6 Cloning rings 7 or cell scraping 8 can be used to manually isolate clones of interest. However, this is a manual process that is not conducive for high-throughput applications. Manual processes introduce variability due to technician technique, the subjective nature of visual inspection, and limited documentation to demonstrate reproducibility. 9
To improve objective analysis, control, precision, reproducibility, and documentation, we developed an integrated robotic system that enables image acquisition and automated image analysis that can be coupled with rapid, automated selection of individual colonies in adherent cell cultures. Automation has been recognized as an effective approach in cell culture to reduce technician variability and increase throughput of cell selection. 10 A number of automated systems exist for different selection applications, including antibody production, induced pluripotent stem cells, and embryonic stem cells.11–13 These applications involve picking cells from semisolid media or off of feeder layers, which are specific to certain cell types. The adhesion environment and methods required for cell picking in these applications are different when compared with cells cultured on tissue culture plastic. In this paper, we present methods to address the unique quantitative imaging and selective cell transfer needs of cells on tissue culture plastic. Human bone–derived progenitors cultured in vitro are used to demonstrate this method. In this process, cells are efficiently harvested, maintain viability, and maintain biological potential based on cell marker expression.
Methods
We have developed a robotic platform that enables automated imaging, image analysis, and cell selection, as shown in Figure 1A . The system is capable of performing multichannel phase contrast, brightfield, and fluorescence imaging at multiple magnifications. High-precision motion control enables fast and accurate image acquisition and montaging. Image processing and analysis algorithms enable identification of cells and their location in a culture vessel. Automated motion controls enable precise position of a pipette tip above the culture surface to select desired cells and transfer to another culture vessel for expansion or biomolecular analysis. Methods and software have been developed for component integration, image acquisition, image montaging, pipette tip management, culture surface detection, and resolving stage and imaging coordinates.
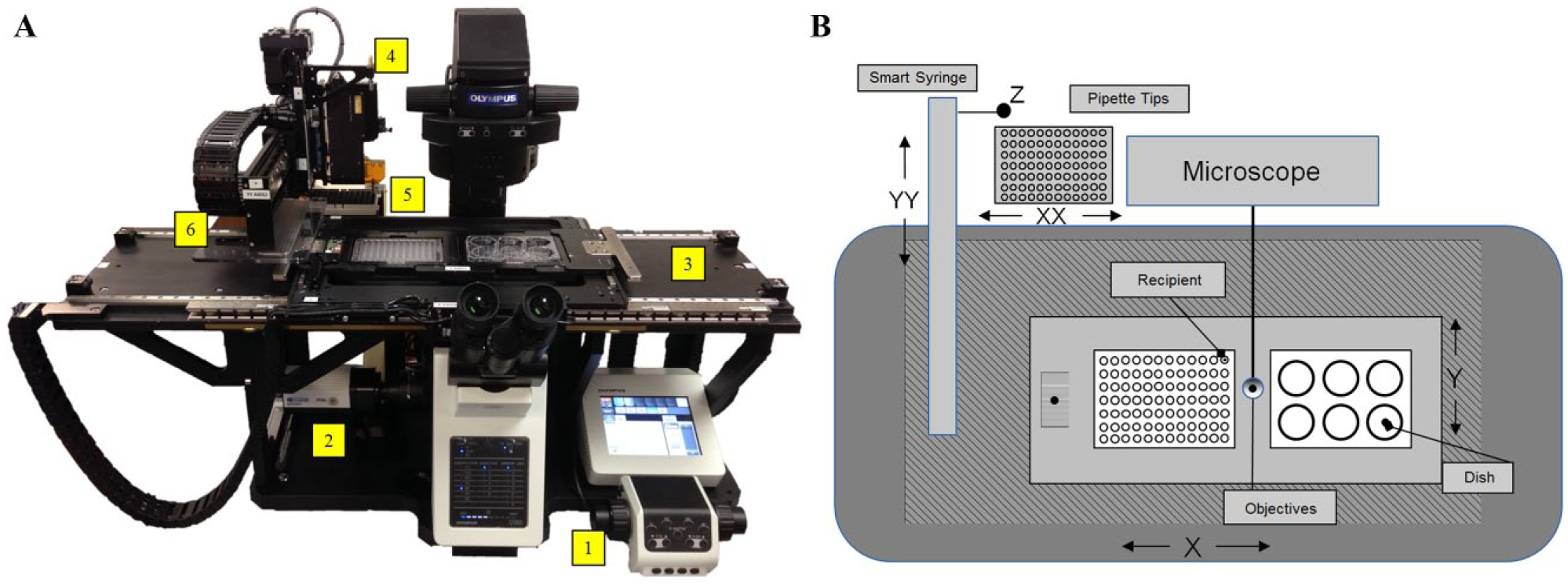
Overview of integrated system. (
Hardware
The system comprises a Parker Hannifin Smart Syringe pump (Parker Hannifin, Mayfield Heights, OH), Olympus IX83 microscope (Olympus, Tokyo, Japan), QImaging Retiga 2000R camera (QImaging, Surrey, British Columbia), XCite Fluorescence 120PC Q Lamp (Excelitas Technologies, Waltham, MA), and NE-9000 pump (New Era Pump Systems, Farmingdale, NY). Qiagen 250 µL (Thermo Fisher Scientific, Waltham, MA) filtered disposable pipette tips were used in conjunction with the Smart Syringe pump to perform cell selection. Precision motion control for imaging and positioning of microfluidics is enabled by Parker Hannifin mSR, 401XE, and LCR22 positioners. Motion control is integrated into a single ACR9000 controller (Parker Hannifin).
Structure
A custom anodized aluminum frame was designed around the microscope to accommodate a stage for positioning culture vessels in the X and Y axes ( Fig. 1B ). A cross axis (YY) for positioning of fluidics and an axis for positioning of a disposable tip rack (XX) serve as secondary axis systems. The fluidics and pipette tip used for picking are raised and lowered by the Z axis. Two mSR linear positioners are used to control the stage. The linear positioners’ repeatability of ±0.1 microns is critical to accurately image cells and define their position on the stage. The stage is designed with two holders capable of accommodating various culture vessels, including well plates, culture dishes, and slides. A cross axis holding a third mSR series linear positioner was used to position the Smart Syringe over the cells for selection and/or fluid dispenser for management of culture media. These are mounted on two separate 401XE ballscrew positioners.
The Smart Syringe pump axis contains an LCM201 (OMEGA Engineering, Stanford, CT) force sensor whose signal is processed by a DMD-465WB (OMEGA Engineering) bridge sensor and provides feedback to the controller when the Smart Syringe tip is in contact with an object. This feedback is necessary for the correct seating of a disposable pipette tip on the Smart Syringe. It is also used as a sensing mechanism to detect the cell culture surface. Precise surface detection is necessary for control of the pipette tip height during cell selection. An LCR22 positioner carries Qiagen disposable tips (Qiagen, Valencia, CA) for the Smart Syringe.
A tip sensor was developed to enable detection of the x and y positions of each disposable pipette tip. Disposable tips were connected to the Smart Syringe pump by a press fit adapter. It is necessary to detect the position of the tip in the x and y positions due to variation between tips and how the tips were connected to the adapter. Detection was enabled by integration of two optical sensors and a printed circuit board to provide a digital output when the tip was detected by the sensors.
Imaging Hardware
The IX83 microscope provides robust automation capabilities, including automated shutter, objective, fluorescence filters, and condenser control. Objectives of 2×, 4×, 10×, and 20× were selected to image cells. Fluorescence filters (excitation/emission wavelengths) for green fluorescence protein (470/525 nm), Texas Red (560/630 nm), and Cy5 (620/700 nm) were selected. These wavelengths provide sufficient spectral separation and avoid the detrimental side effects of ultraviolet excitation wavelengths on cell viability. 14 A QImaging Retiga 2000R camera provides a large charge-coupled device (CCD) array, 7.4 µm pixel size, and high sensitivity. An XCite 120 PC Q lamp provides a metal halide fluorescence source.
Software
Custom software was developed in Python and GTK to integrate control of the multiple hardware components into a single graphical user interface (GUI) ( Fig. 2 ). Software development kits from Parker Hannifin, Olympus, QImaging, and Excelitas were provided from manufacturers. These development kits do not use the same computer language. Olympus, QImaging, and Excelitas use C++, while Parker Hannifin uses C#. To simplify the integration of these disparate devices, individual drivers were developed for each component. Each driver accepts high-level commands through a transmission control protocol (TCP) port from the GUI to control individual hardware components. The GUI enables acquisition of large field-of-view (FOV) images by acquiring high-resolution tagged image file format (TIFF) images across the imaging area and montaging them into one single image. Software was designed to enable saving of image information in the TIFF header in JSON or XML format to provide compatibility with the Open Microscopy Environment TIFF format. 15
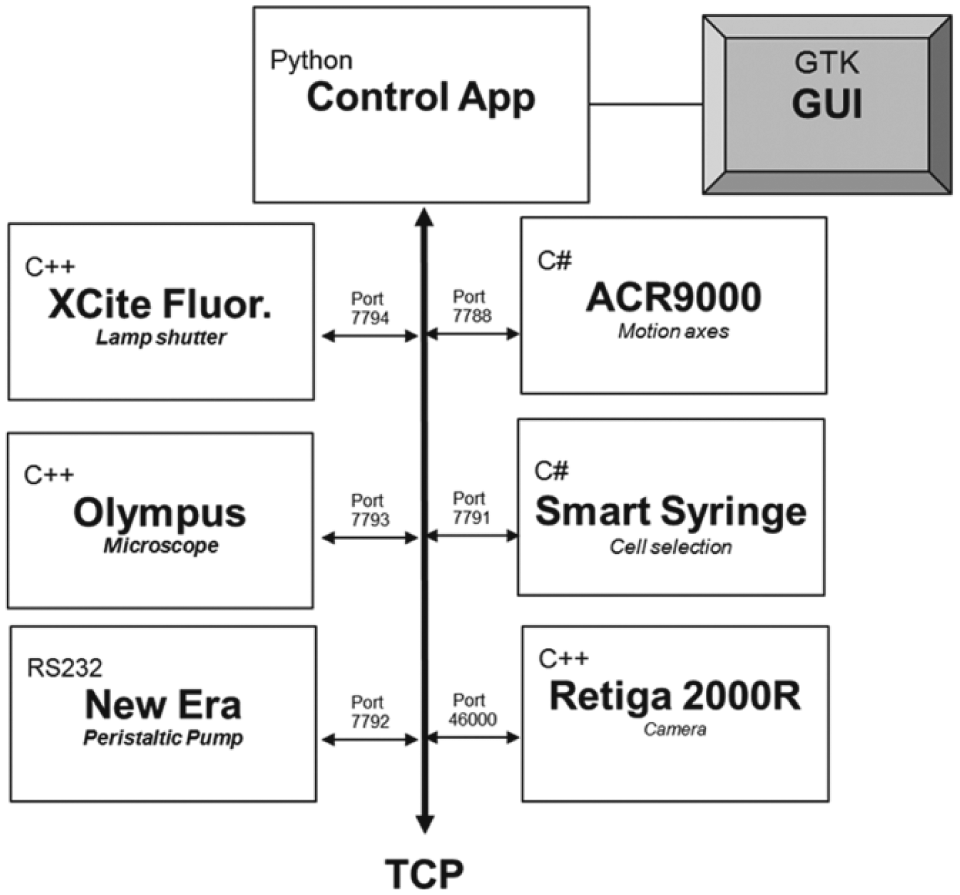
Software overview of multicomponent integration, communication, and languages into a single GUI.
Analysis of this image is used to identify cells or cell populations of interest for cell picking. Automated image processing and analysis algorithms for spatially locating specific cells or cell colonies are implemented separately in MATLAB and Fiji, adapted from previously described methods. 16
Cell Culture
Discarded bone core samples from the proximal femur were received from three patients undergoing hip arthroplasty procedure. Discarded samples were de-identified, human subject research exempt, and acquired under a protocol approved by the Cleveland Clinic Central Biorepository. Samples were sterilely minced in alpha minimum essential medium (α-MEM) (Life Technologies, Grand Island, NY) into 1–2 mm fragments using forceps and a sharp osteotome. Fragments were mechanically agitated in α-MEM medium to dislodge and remove marrow-derived cells. Marrow-derived cells were discarded. To isolate cells that were tightly adherent to the trabecular bone, medium containing 100 U/mL collagenase type I (Worthington Biochemical, Lakewood, NJ) in Hank’s buffered salt solution was added to the fragments of trabecular bone for 1.5 h at 37 °C. Collagenase activity was stopped using complete α-MEM media with 10% fetal bovine serum (FBS; Atlanta Biologics, Lawrenceville, GA) at a 1:1 ratio. Cells were passed through a 70 µm cell strainer (Fisher Scientific, Hampton, NH) to filter out bone fragments and large debris. Cells were plated in a six-well plate (Corning, Kennebunk, ME) at a plating density of 125,000 cells/cm2. Cells were cultured to allow colony formation in α-MEM media, 10% FBS, and 1% penicillin/streptomycin (Sigma-Aldrich, St. Louis, MO).
Cell Imaging
Cells were plated in a six-well plate and cultured for 14 days. Cells were labeled using directly conjugated antibodies PE-CF594-CD73 (BD Bioscience, San Jose, CA), and Alexa647-CD90 (BioLegend, San Diego, CA) by adding fluorescent antibodies at a 1:150 ratio directly to the culture media. This technique has been used in several imaging applications for live, surface marker detection.17,18 To prepare for picking, cells were trypsinized using 0.005% trypsin-EDTA (Sigma Aldrich) in phosphate-buffered saline for 7 min. Complete media was added to deactivate the trypsin and prevent complete dissociation of the cells from the culture surface.
Before imaging the cells, media and trypsin were removed from the well. Live Cell Imaging Solution (Thermo Fisher Scientific) was replaced in the well to reduce background autofluorescence. Wells were scanned at 4× magnification in a 12 × 16 grid pattern to image the entire well. After acquisition, software performed 2 × 2 binning to achieve an improved signal-to-noise ratio for fluorescence imaging and a final resolution of 3.71 microns/pixel. Wells were scanned at a stage velocity of 10 mm/s, acceleration of 50 mm/s2, and deceleration of 50 mm/s2. These parameters were utilized to prevent the induction of waves in the media that interfere with phase contrast imaging ( Fig. 3 ).
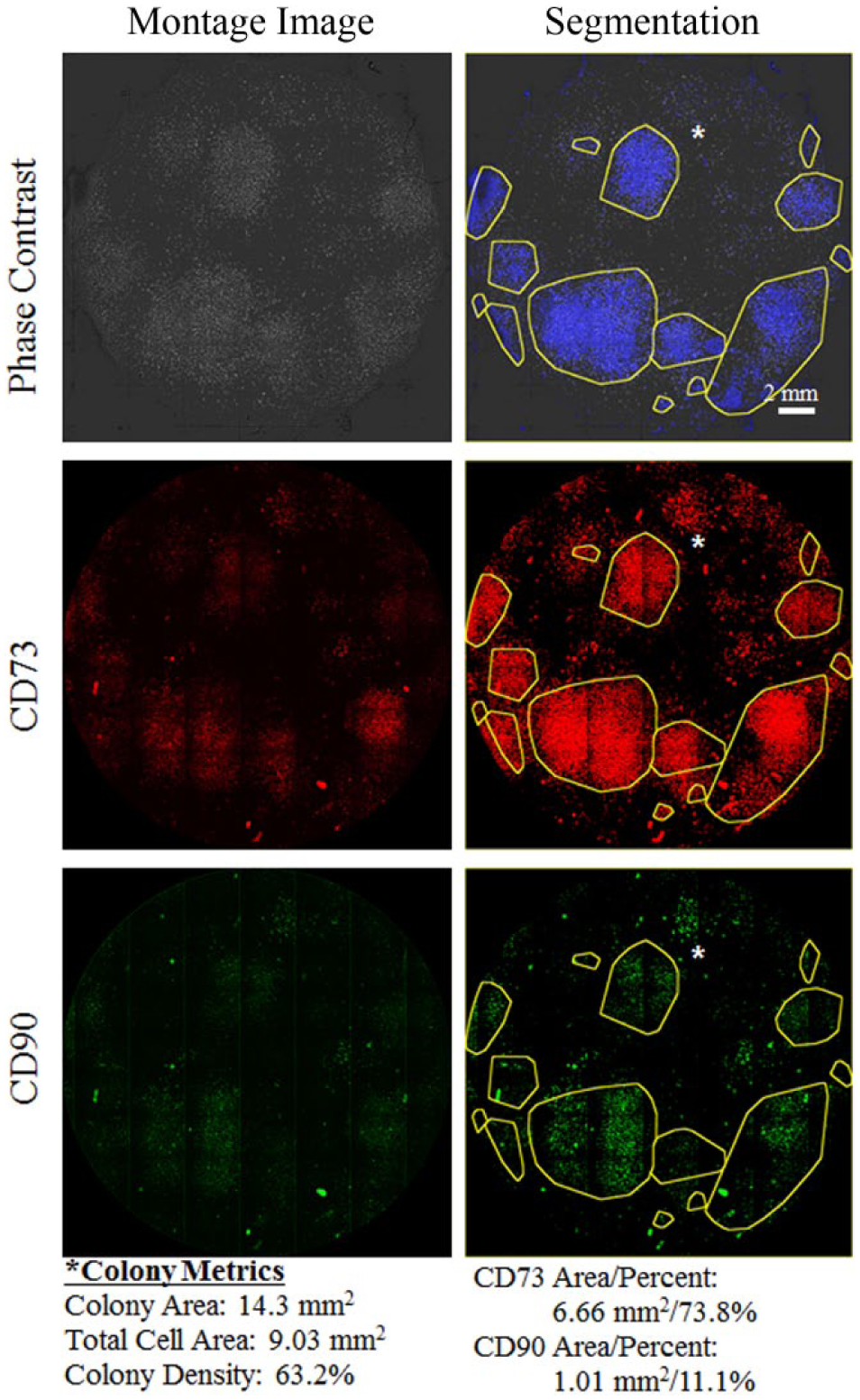
Phase contrast and fluorescence images of bone-derived stem cells after 14 days in culture. Cells are segmented from phase contrast images (blue). Colonies (yellow) are identified based on cell segmentation. Original phase and fluorescence images shown in the first column. Segmented images shown in the second column. Colony metrics shown for colony marked with asterisk.
Cell Picking
Cells were selected from segmented cells by positioning the pipette tip at a height of 20 µm off the surface of the plate and aspirated at a flow rate of 250 µL/s. Harvested cells were transferred to individual wells on a 96-well plate. Images were taken before and after selection to document the region of cell removal, the characteristics of the selected cells, and the efficiency of cell harvest at the target location. If removal efficiency after the first pick was poor, the tip was repositioned 200 microns from the original picking position and cells were removed again. Cells were imaged and restained 2 and 6 days after selection to characterize the viability and cell marker expression following cell selection.
Results
Cells and colonies were segmented in day 14 samples ( Fig. 4A ). Cells were selected from regions identified as colonies in the segmented images. Cells were removed from target locations as documented by images acquired before and after selection ( Fig. 4A , B ). These before and after images were segmented to identify the area of cells that were removed ( Fig. 4E ). While the tip used for picking had an inner diameter of 533 microns, cells were removed in a region that was 1000 microns in diameter. Based on cell area data from 16 pick locations, cells were harvested at an 88.9% ± 6.9% efficiency (mean ± standard deviation). All cells that were transferred to wells in a 96-well plate remained viable and capable of expressing CD73 and CD90 2 and 6 days after selection.
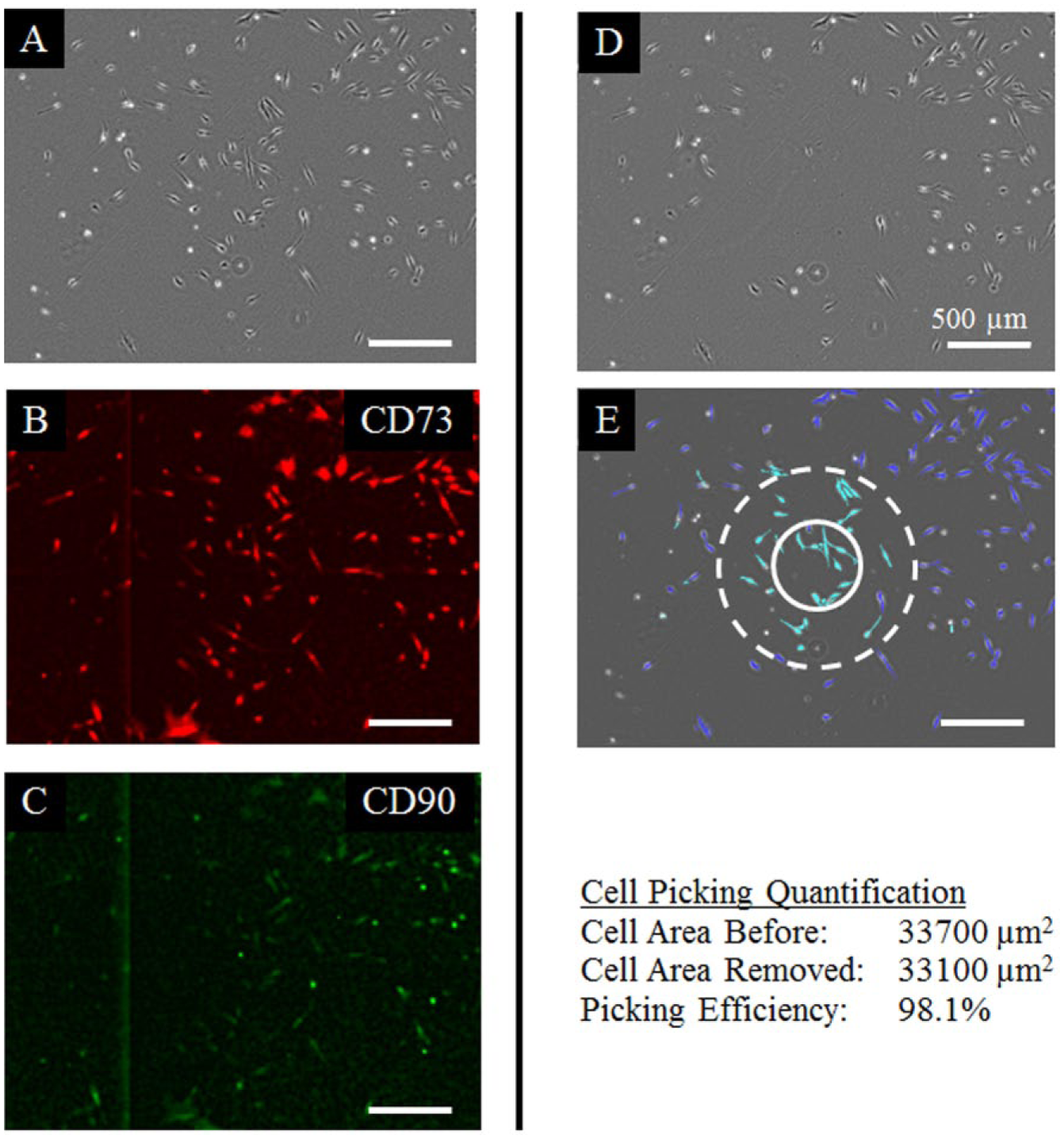
Cell picking evalution. (
Discussion
The device design and data presented here demonstrate the capability to reproducibly image cells, automatically segment cells and colonies, efficiently remove selected cells, confirm viability after cell transfer, and document the entire process. By using phase contrast, we present a noninvasive method to identify cells and colonies. These methods can be augmented using live cell labeling for quantitative area expression of surface markers ( Fig. 4B , C ). The device and methods shown here are novel developments that demonstrate the picking of cells from tissue culture plastic. We expect these methods to be directly applicable to picking adherent tissue-derived cells and stem cell types. Future work will define cell-specific assay and selection protocols for a variety of cell types beyond bone-derived stem cells, which have different morphology and adhesion strength to the culture surface. 19 While we use customized image processing and analysis algorithms in this study, the software was designed to capture images that allow users to use or develop alternative image analysis algorithms to quantitatively assess or guide cell selection.
The work to date has focused on short-term process steps required for the viable transfer of cells. Future work includes examining biologic potential in both shorter time frames and long-term culture across multiple passages. In this study, cells were picked from colonies cultured for 14 days following cell harvest. Picking cells before day 14 can elucidate different information about cell heterogeneity than picking at day 14 ( Fig. 5 ).
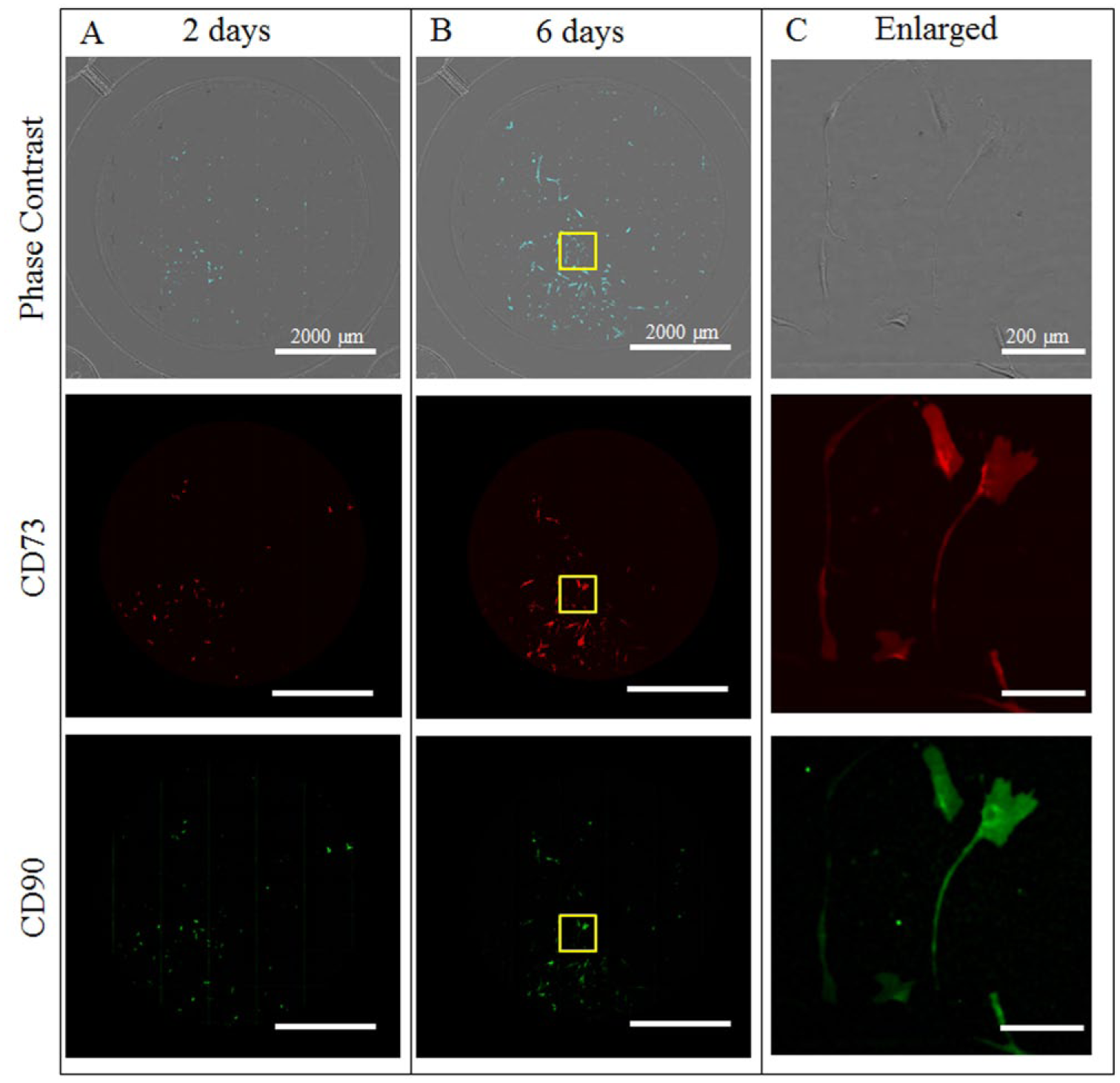
Cells are imaged in wells of 96-well plate. (
While we expect our cell picking methods to impact long-term biologic potential of cell populations no differently than current manual picking methods, future works include examining biological potential, including gene expression or differentiation potential, across multiple passages. This includes time-lapse studies to pick cell-based dynamic characteristics, such as proliferation rate, and confirm maintenance of dynamic characteristics. Manual application of similar methods using low-concentration trypsin and pipetting has been shown to be less detrimental to cell viability than current clonal isolation methods involving cell scraping. 20 Alternatively, these picking methods can be used as a form of negative selection where nondesired cells are removed from a culture vessel such that a purified population of desired cells with a given biological potential remains unaltered in a culture vessel.
The methods developed here advance the field of stem/progenitor cell biology by enabling investigators to better characterize and reproducibly select specific stem cell populations from heterogeneous populations of cells. The device described here was designed to be placed into a standard biosafety cabinet for use under sterile conditions ( Fig. 6 ). This enables the device to be readily incorporated into existing facilities, a recognized need in biomanufacturing. 21 Both biomanufacturing and basic science can benefit from use of this device by enabling investigators to quantitatively define and document the characteristics of the cells that they are working with while they are in culture and precisely sample, move, or remove selected cells.

Current device configuration inside laminar flow hood.
Footnotes
Acknowledgements
This work was supported by funding from the Cleveland Clinic Product Development Fund and from the Parker Hannifin Corporation.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
