Abstract
Assessment of a tumor’s molecular makeup using biofluid samples, known as liquid biopsy, is a prominent research topic in precision medicine for cancer, due to its noninvasive property allowing repeat sampling for monitoring molecular changes of tumors over time. Circulating exosomes recently have been recognized as promising tumor surrogates because they deliver enriched biomarkers, such as proteins, RNAs, and DNA. However, purification and characterization of these exosomes are technically challenging. Microfluidic lab-on-a-chip technology effectively addresses these challenges owing to its inherent advantages in integration and automation of multiple functional modules, enhancing sensing performance, and expediting analysis processes. In this article, we review the state-of-the-art development of microfluidic technologies for exosome isolation and molecular characterization with emphasis on their applications toward liquid biopsy–based analysis of cancer. Finally, we share our perspectives on current challenges and future directions of microfluidic exosome analysis.
Introduction
Tissue biopsy is often required for cancer diagnosis and prognosis. However, tissue biopsy is highly invasive for most primary tumors and metastatic diseases,1,2 especially brain cancer, lung cancer, and ovarian cancer, which require difficult surgeries. 3 The obtained tissue quality and quantity highly determine the diagnostic precision at the molecular level, including mutation characterization. Tumor tissues are heterogeneous and evolve over time.2,4 Sampling of entire tissue with dynamic representatives is not possible. Therefore, assessment of the molecular makeup of tumors from a biofluid sample is of great research interest. 2 Although technology challenging, noninvasive blood-based liquid biopsy allows repeat monitoring for clinical oncologists to gain a broader molecular understanding of tumors without the need for a tissue biopsy.5,6 Indeed, several blood-based biomarker tests that have been developed have been around for decades and are still debatable, including prostate-specific antigen (PSA) screening for prostate cancer, carcinoembryonic antigen (CEA) for colorectal cancer, CA19-9 for pancreatic cancer, and CA125 for ovarian cancer, due to the lack of reliability and specificity.7,8 Recent technological developments being applied to liquid biopsies are capable of reproducibly detecting mutations at very low allelic frequencies. 9 However, lack of confidence in blood-based biomarkers still prevents widespread utilization of liquid biopsy for cancers.
Recent research of extracellular vesicles (EVs) and exosomes has revealed that early-stage tumors constitutively release vesicles carrying various tumor markers.10–12 Exosomes, which are membrane vesicles of endocytic origin (30–150 nm diameter, Fig. 1 ), are systemically detectable in the blood of various cancer patients and have been shown to correlate well with tumor progression, immune response suppression, angiogenesis, and metastasis.13–15 Exosomes are stable carriers of enriched genetic material and proteins from their cell of origin, thereby holding great promise for identifying early-stage tumors.16,17 Compared to well-studied circulating tumor cells (1–10 circulating tumor cells [CTCs]/mL of blood), exosome release from tumor cells is an active process with concentrations of ≥109 vesicles/mL in blood.11,18 As illustrated in Figure 2 , exosomes sensitively reflect tumor status; therefore, substantial investigations have focused on the essential physiologic and pathophysiologic functions of circulating exosomes as a surrogate for tumor liquid biopsy.14,19–22
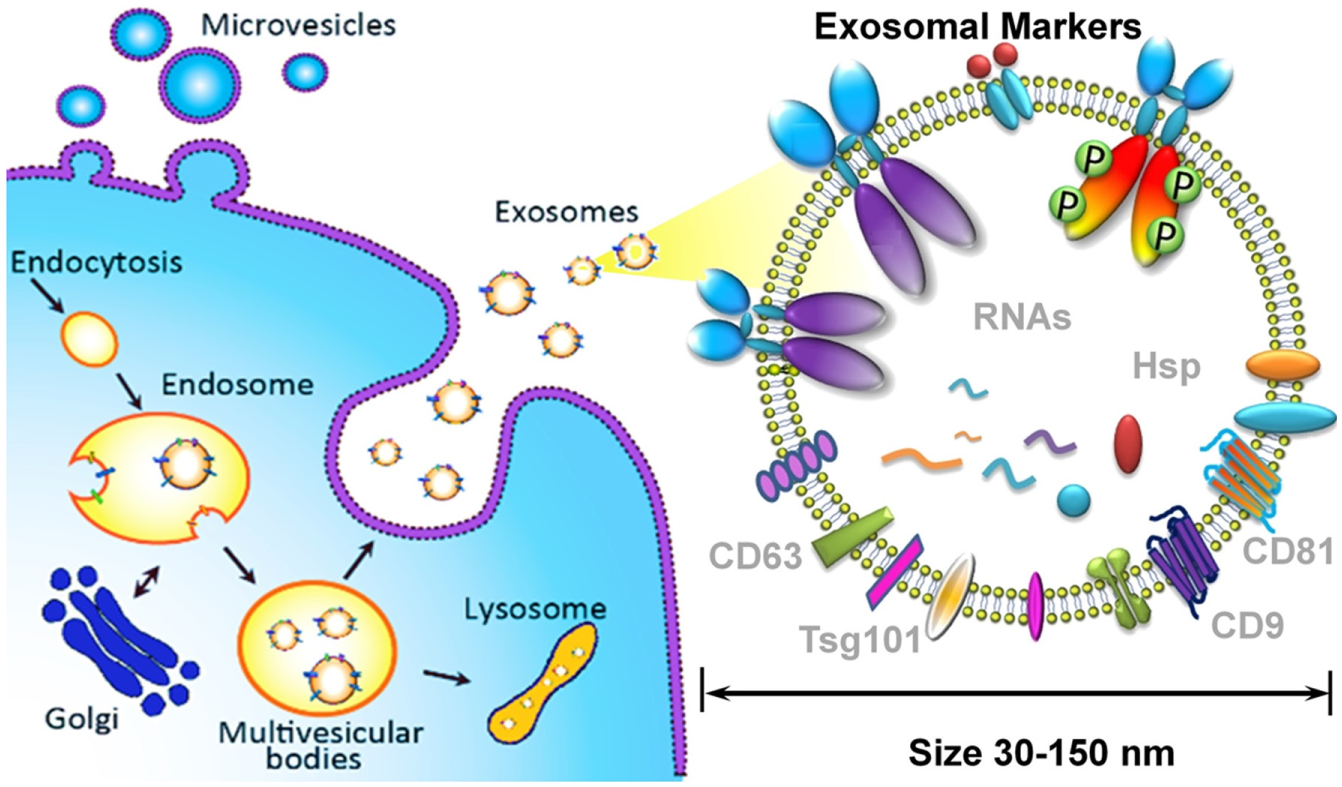
Exosome biogenesis, properties, and molecular composition.
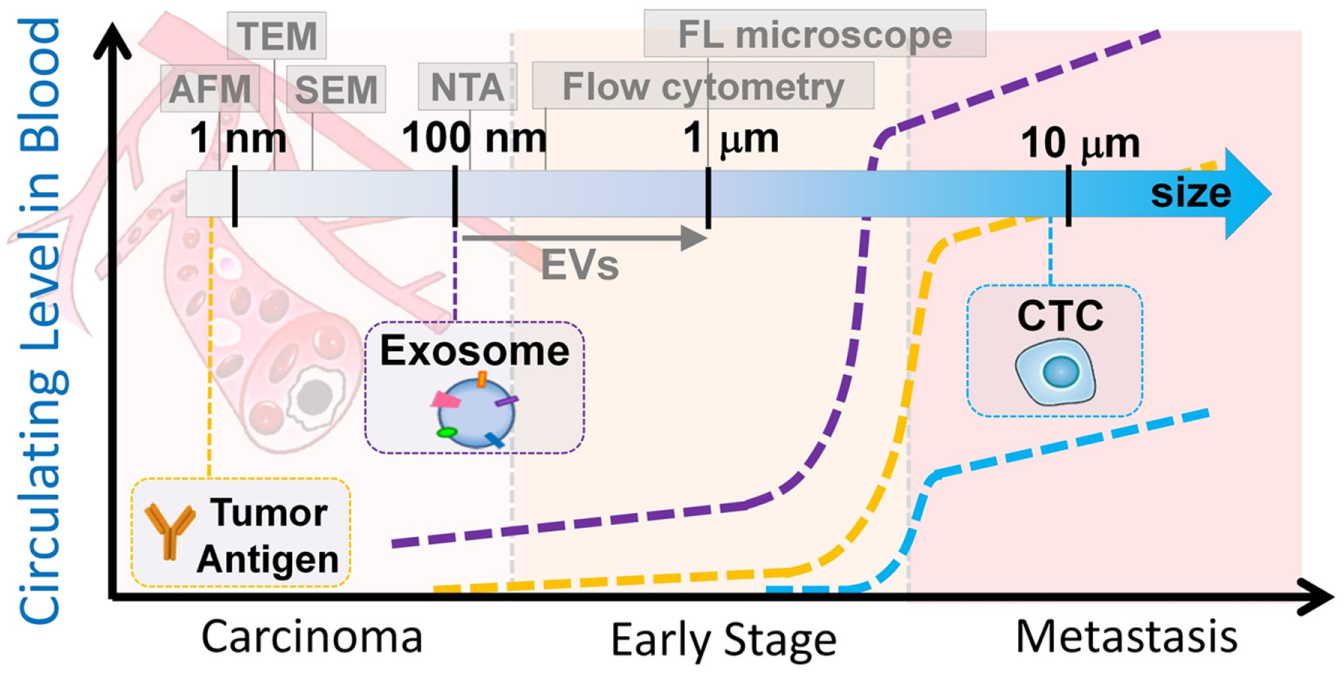
Circulating tumor antigens, exosomes, and CTCs associated with tumor status (not to scale). Scale bar shows the detection resolution of various benchtop instruments.
However, isolation and molecular analysis of such diverse nanoscale exosome vesicles for clinical utilization are technically challenging.21,23–26 Size overlap complicates exosome definition due to the presence of other membrane-derived subcellular structures, such as apoptotic vesicles, exosome-like vesicles, membrane particles, and ectosomes.27,28 Current exosome purification methods, including ultrafiltration and sucrose gradient ultracentrifugation, are tedious and time-consuming (>10 h) and cannot completely discriminate exosomes from other EVs.29–31 Although conventional filtration isolates microvesicles (MVs) with a uniform size of less than 150 nm, forced filtration and shearing force may cause membrane fusion and loss of integrity. 32 Molecular analysis of isolated MVs is primarily performed using Western blot, enzyme-linked immunosorbent assay (ELISA), or mass spectrometry, which require lengthy processes and concentrated exosome samples. 33 Nanoparticle tracking analysis (NTA) and flow cytometry have demonstrated limited reliability for detecting particles smaller than 200 nm.33,34 Transmission electron microscopy (TEM) and atomic force microscopy (AFM) have been utilized often to investigate exosome morphology and size distribution ( Fig. 2 ), 35 but microscopic protocols do not inherently exhibit high-throughput and rapid measurement. Microfluidic lab-on-a-chip technology has recently been spotlighted as a promising approach for exosome isolation and molecular analysis, owing to its low-volume consumption, 36 high capability of functional module integration, 37 quick analysis time, and high sensitivity, 38 as well as a sample-to-answer format.39–41 Several microfluidic approaches, such as isolation, quantification, and molecular profiling, have been previously developed for exosome study. 42 We review the state-of-the-art microfluidic technologies for isolating exosomes and their applications for exosome-based liquid biopsy analysis of cancer. Other emerging microfluidic approaches developed for fabrication and production of therapeutic exosomes 43 are not within the scope of this article.
Microfluidic Exosome Isolation
Since 2012, research efforts have increased dramatically to develop microfluidic platforms in order to isolate exosomes. Compared to benchtop methods, microfluidic technology offers fast isolation speed, high yield and efficiency, automation, and functional integration for streamlined exosome molecular analysis. 42 Vesicle size (30–150 nm) and surface markers (immunoaffinity) are mandatory in order to identify a specific population of circulating MVs that primarily consists of exosomes. 10 Therefore, current reported microfluidic platforms can be classified as immunoaffinity-based exosome isolation and size-based exosome isolation.
Immunoaffinity-based exosome isolation can be implemented into microfluidic devices by manipulating affinity particles/magnetic beads, or modifying microchannel surfaces with antibodies. In 2010, Chen et al. reported the first microfluidic exosome isolation platform, which used an anti-CD63 functionalized surface for immunocapture of exosomes from human sera. 44 Kanwar et al. developed a similar platform called ExoChip, which utilized a surface-functionalized (anti-CD63) circular microchamber to capture exosomes, followed by fluorescent carbocyanine dye (DiO) staining for quantitation. 45 Although various microstructures were configured to enhance mixing and capture efficiency, capture capacity was still limited by available surface area and antibody immobilization density. Therefore, spherical particles or immunomagnetic beads were introduced into microfluidic devices to enhance capture capacity. We recently reported two microfluidic platforms46,47 for large-scale exosome isolation and molecular profiling of both surface and intravesicular markers by manipulating immunomagnetic capture beads in a microfluidic, multistage circuit. Dudani et al. introduced 20 µm polystyrene beads conjugated with biotinylated anti-human CD63 into a microfluidic device that utilized inertial lift forces at a finite Reynolds number, in order to position microparticles and exchange solutions for rapid purification. 48 This approach provided high-flow-rate isolation of exosomes, which greatly increased throughput and processing volume compared to other methods. However, premixing and incubation of capture beads with samples were needed.
Although immunoaffinity-based isolation generates specific exosome populations and reflects molecular expression levels, size-based isolation has advantages of size uniformity without sample bias. Wang et al. fabricated a microfluidic device consisting of an array of porous silicon nanowire-on-micropillar. 49 The inter-nanowire spacing was tuned within a range of 30–200 nm to create a high density of interstitial sites, which allowed physical trap of exosomes. Davies et al. in situ prepared nanoporous membranes in a microfluidic filtration system to isolate vesicles from whole blood with tunable size cutoff (<500 nm). 50 However, these physical trapping approaches were restricted by exosome saturation limit and recovery rate. A continuous flow design, without capacity limitation, is ideal for on-chip high-throughput processing and integration of downstream sample preparation. Lee et al. developed an acoustic nanofilter chip that can fractionate exosomes (diameter < 200 nm) from cell culture media and blood in a continuous flow manner. 51 Santana et al. built a microfluidic obstacle array based on the principle of deterministic lateral displacement and fractionated MVs with size cutoff of 250 nm. 52 Currently, isolation with greater size resolution below 200 nm still needs to be improved.
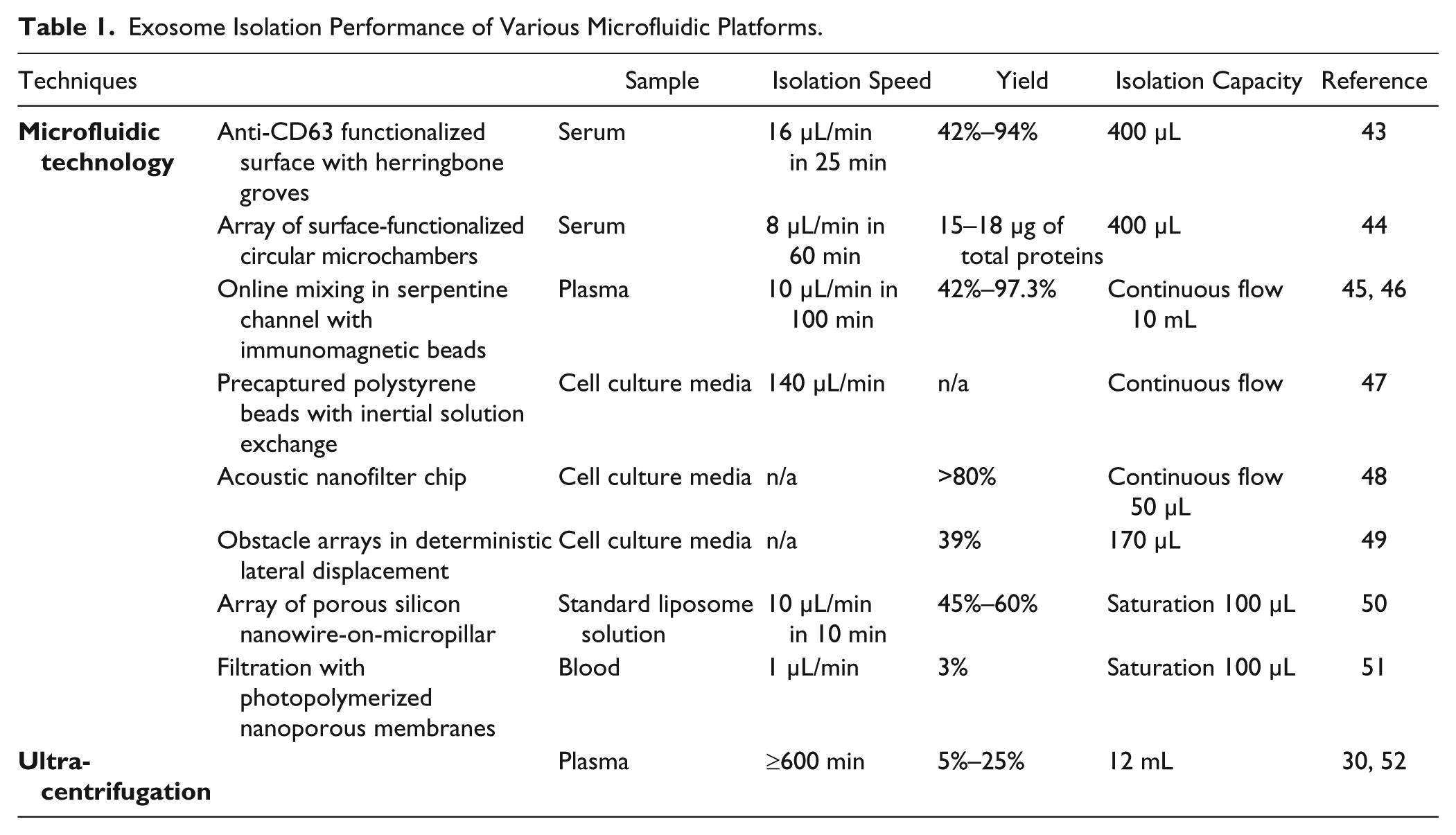
Table 1 summarizes exosome isolation performance from various microfluidic platforms in terms of yield, capacity, and efficiency. As shown in the table, microfluidic technology as an advanced approach substantially increased exosome isolation speed and efficiency compared to a classic benchtop ultracentrifugation approach. In addition, it has been reported that diverse subpopulations of vesicles secreted by various intracellular mechanisms were present in exosome preparations obtained by differential ultracentrifugation. 31 Although size is the most acceptable criterion for exosome identification (smaller than 150 nm), it is not a strict feature of exosomes. Several current preparations invariably contain varying proportions of other membranous vesicles that co-purified with exosomes, such as shed MVs and apoptotic blebs. Tauro et al. observed that EpCAM immunoaffinity-captured exosomes contain at least double the amount of enriched exosome markers and exosome-associated proteins, compared to centrifugation and density-based separations, although these MVs all range from 40 to 100 nm. 53 Therefore, it is crucial in the future to design novel microfluidic methods that combine size and marker features for purifying and characterizing vesicle types and allowing precise analysis of respective exosome functions.
Exosome Isolation Performance of Various Microfluidic Platforms.
Microfluidic Exosome Sensing toward Liquid Biopsy
Liquid biopsy analysis of tumors has beneficially impacted clinical care, especially regarding treatment decision guidance; exosomes are significant contributors to such progress. 54 Several advanced microfluidic exosome measurement and sensing technologies have been developed to characterize exosome physical, biological, and molecular properties. Akagi et al. developed a microfluidic immunoelectrophoresis approach to detect bound antibodies to exosome surface markers without need of fluorescent labeling. 55 Immunobinding of exosomes with antibodies has resulted in zeta potential changes, thus exhibiting different migration mobilities under electrophoresis depending on marker expression levels. Equipped with a laser dark-field microscope, this platform can measure size, zeta potential, and surface markers from exosomes. 55 In contrast to flow cytometry, this method does not require fluorescent labeling for detecting bound antibodies. Addition-ally, the shift in the zeta potential is not associated with the size of vesicles, as long as the density of bound antibodies is not changed. However, the sensitivity or resolution of the zeta potential change in terms of exosome particle number has not been discussed, and the equipment setup is relatively expensive and complicate. Vaidyanathan et al. introduced a tunable alternating current electrohydrodynamic flow (nanoshearing) in a microfluidic channel that enhances the specificity of capture for sensitive detection of exosomes. 56 The multiplexed device allows simultaneous detection of multiple exosome markers on-chip using colorimetric readout visible to the naked eye. The detection method is simple and straightforward. Quantitative detection of markers HER2, PSA, and CD9 was demonstrated for breast cancer diagnosis. More efforts have been made to extract intravesicular exosomal biomarkers (microRNA) for seeking interconnections with cancer disease study and diagnostic potential. Taller et al. substantially improved exosome lysing and the RNA detection process in ~1.5 h by integrating a surface acoustic wave with an ion exchange nanomembrane sensor. 57 Note that conventional benchtop methods require at least ~24 h of processing time. Wei et al. employed a nonuniform electrical field to disrupt exosomes and release harbored exosomal RNA/protein biomarkers for on-site monitoring through immune recognition.58,59 The hCD63-GFP expressing exosomes from lung cancer cell line H460, which was stably transfected with hCD63-GFP, were found in vivo to transport to saliva, in addition to serum.
In order to profile both surface and intravesicular exosomal markers, we have developed a cascading chip to integrate exosome isolation, enrichment, chemical lysis, protein immunoprecipitation, and sandwich immunoassay assisted by chemifluorescence detection 47 ( Fig. 3a , b ). Analysis of both type 1 insulin growth factor receptor (IGF-1R) and its intravesicular phosphorylation status from non-small-cell lung cancer patient plasma-derived exosomes was demonstrated. Significant overexpression of exosomal IGF-1R was observed in lung cancer patients compared to healthy individuals. In addition, we employed an ExoSearch chip for blood-based diagnosis of ovarian cancer using multiplexed measurement of three exosomal tumor markers (CA125, EpCAM, CD24) from the same population of CD9 positive exosomes 46 ( Fig. 3c ). Results showed significant diagnostic power (a.u.c. = 1.0, p = 0.001) through a training set of ovarian cancer patient plasma ( Fig. 3d , e ). In contrast, the diagnostic accuracy of using exosomal particle concentrations measured by NTA was poor, with an a.u.c. of only 0.67 ( Fig. 3f ), due to the relative large uncertainty in size and concentration measured by NTA. Note that counting exosomes alone has been found insufficient for cancer diagnosis. The results from receiver operating characteristic (ROC) analysis suggested that ExoSearch chip enables sensitive multiplexed exosomal marker detection for blood-based diagnosis of ovarian cancer with significant predictive power. In order to further improve the exosome detection sensitivity and boost the limit for early detection of cancer, we recently introduced a novel graphene oxide/polydopamine (GO/PDA) nanointerface. 60 The GO-induced formation of 3D nanoporous PDA surface coating enabled ultrasensitive on-chip ELISA assay of ovarian cancer plasma-derived exosomes at 50 exosomes/μL.
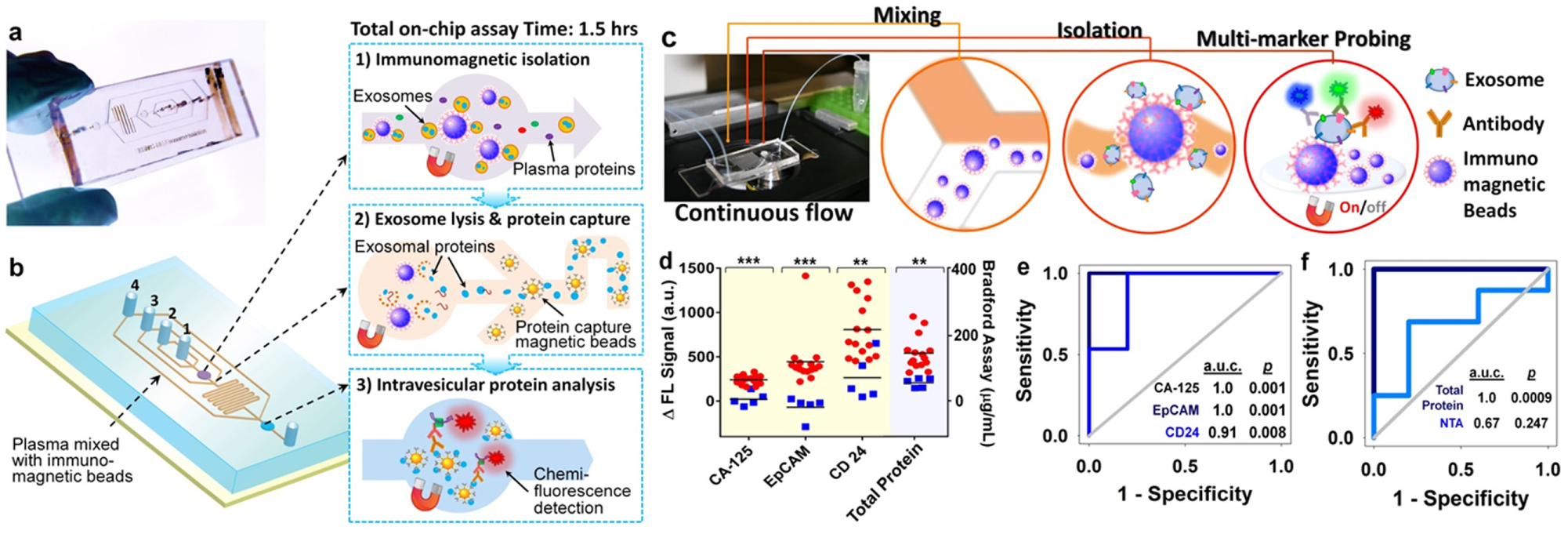
Microfluidic analysis of exosomal protein markers for liquid biopsy of cancer. (
Dr. Hakho Lee’s research group pioneered a study of diagnostic and prognostic roles of circulating exosomes using microfluidic sensing technology. In 2012, Shao et al. devised a microfluidic nuclear magnetic resonance system (µNMR) in which MVs bound with marker-specific magnetic nanoparticles resulted in faster decay of the NMR signal depending on protein expression levels (
Fig. 4a
). The significance of using blood exosomes in diagnosis and monitoring treatment responses for patients with glioblastoma was demonstrated by probing nine exosomal markers.
61
Efficacy of drug treatment, including temozolomide (TMZ) and geldanamycin, has been evaluated using µNMR, as MV numbers decreased in proportion to drug concentration. This platform has better detection sensitivity than standard ELISA and flow cytometry analyses, and further enhancement of sensitivity is under development. Rho et al. refined the exosome sample preparation step in integration with µNMR. A filter membrane cartridge was incorporated to directly isolate and enrich exosomes from blood before labeling with target-specific magnetic nanoparticles for µNMR sensing.
62
Im et al. introduced a nanoplasmonic exosome (nPLEX) assay platform that consists of arrays of lattice nanoholes patterned in a gold film within parallel microfluidic channels for label-free, surface plasmon resonance (SPR) sensing (
Fig. 4b
).
63
Ascites samples from ovarian cancer patients and 10 ovarian cancer cell lines were studied by probing 71 exosomal protein markers. Levels of EpCAM and CD24 markers showed significant responses for improving ovarian cancer diagnosis with an accuracy of 97%. The nPLEX platform provided high-throughput screening capability with improved sensitivity, compared to µNMR platform. However, the system complexity of both nPLEX and µNMR requires nanofabrication and skilled operations, which limits the application in routine clinical tests. In 2015, Shao et al. expanded the work by investigating exosomal mRNA in glioblastoma patient blood during treatment, since serial rebiopsy of primary tumors is difficult.
64
Real-time PCR analysis of RNA contents released from immunomagnetically captured exosomes was integrated into a microfluidic chip (iMER) for studying key exosomal mRNA markers that could probe epigenetic status of a primary tumor and potentially predict TMZ drug resistance. Results from the above several novel platforms supported the use of exosomes as tumor cellular surrogates. In parallel with CTC research, exosomes were shown to be abundant and stable in circulation, providing significant practical value as surrogates of tumors for liquid biopsy (
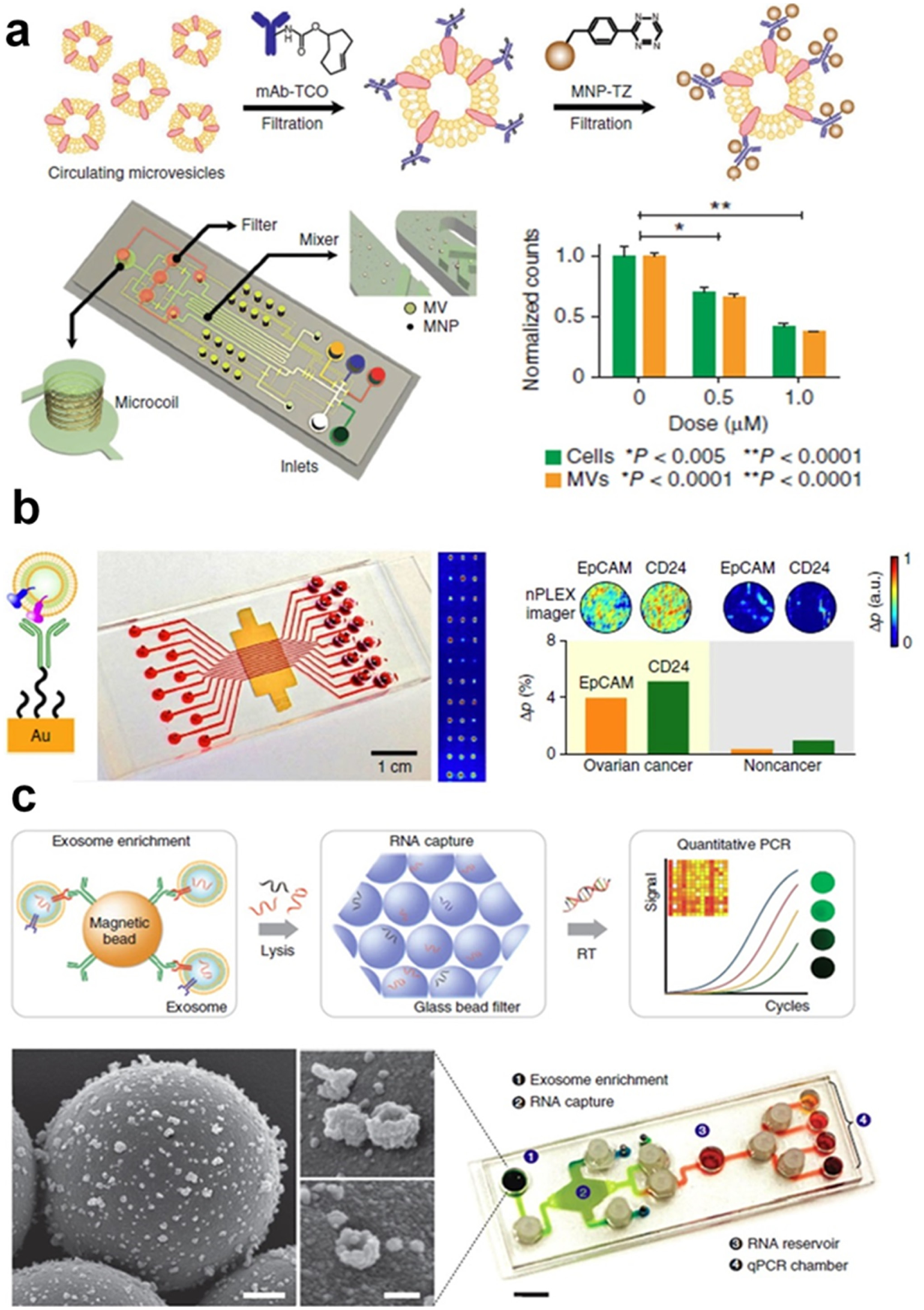
Microfluidic exosome molecular profiling for monitoring of disease and treatment. (
Discussion
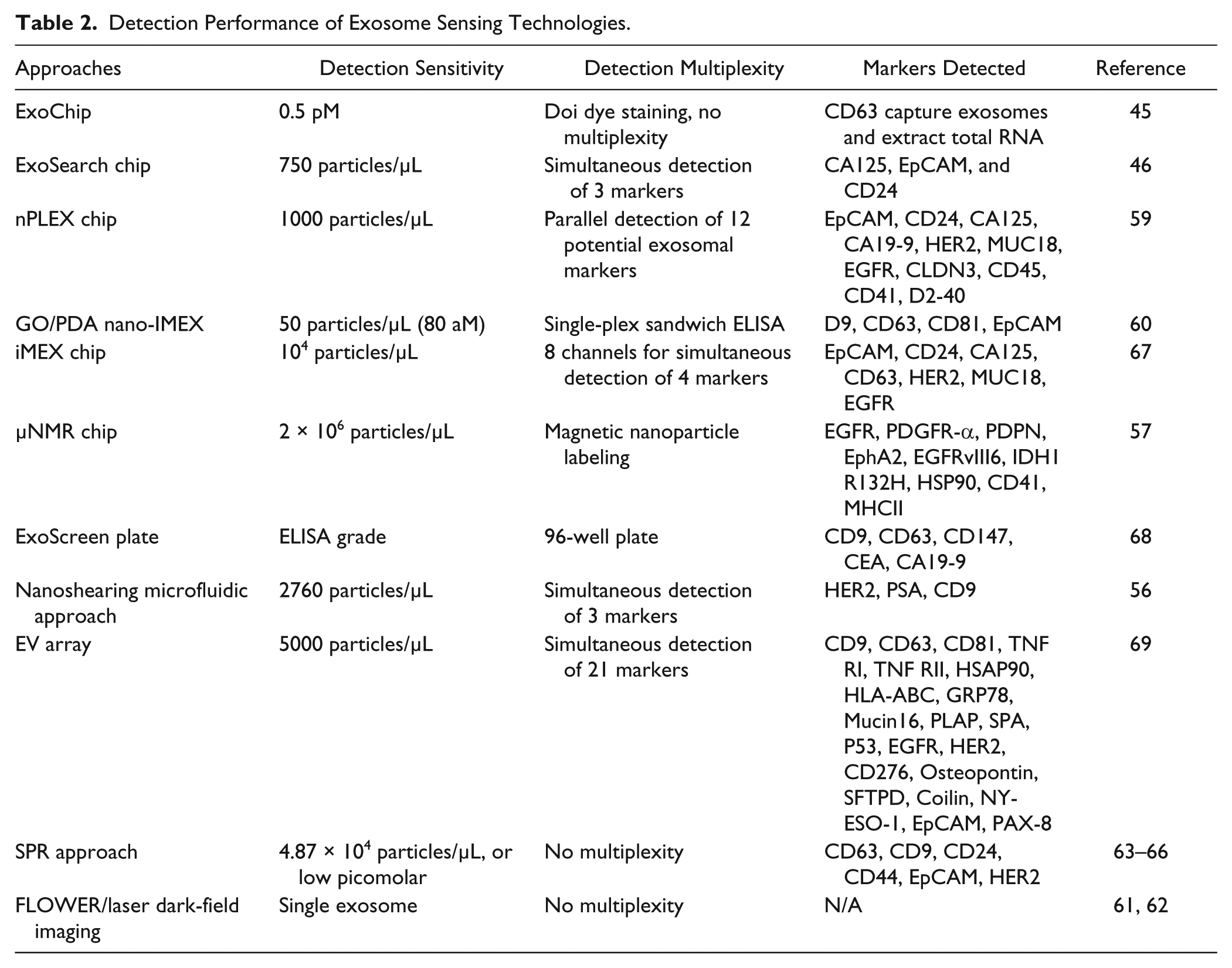
Tumor-derived circulating exosomes have attracted increasing interest for noninvasive cancer diagnosis and monitoring of treatment response, as a promising alternative to liquid biopsy. In addition to microfluidic technology, several other emerging approaches have been implemented to study exosomes and their molecular compositions, such as single exosome detection,65,66 SPR and imaging,67–70 magnetoelectrochemical sensor, 71 ExoScreen 96-well plate, 72 and EV array.73,74 We summarized detection sensitivity and sensing capability in Table 2 , which showed superior performance compared to conventional Western blotting analysis and chemiluminescence ELISA. Such improvements in terms of analytical sensitivity and specificity will greatly address current hurdles in liquid biopsy of cancers. We foresee that a growing number of investigators across disciplines will devote more efforts to advancing exosome research in the next 5 years and beyond.
Detection Performance of Exosome Sensing Technologies.
Exosome secretion is a dynamic process, producing diverse populations with 5-fold differences in size and 104-fold differences in concentration; 27 as a result, precise measurement and analysis of exosomes is challenging. To date, standardized isolation protocols are still lacking, and the field of exosome research lags behind CTC research because the definition and characterization of exosome types are not yet firmly established. Specific exosomal markers for quantitative evaluation of exosomes with cell origins are urgently needed. In order to increase understanding of exosomes and quantitatively decipher exosomal components, more novel technologies are needed for comprehensive characterization of surface and intravesicular compositions. Interconnections between exosomal RNA, surface protein topography, and posttranslational modification could aid identification of exosome associated with cancer phenotypes. It is worth mentioning that increased MV counts with prolonged storage of blood have been observed, but the protein profile per vesicle in blood displayed negligible changes during blood aging based on Rho et al.’s study. 62 These findings point out that sample preparation protocols are critical and need to be standardized. Using highly precise fluid control and automation, microfluidic technology has the capability to address the above issues in sample preparation and isolation, with more power for molecular characterization and sensitive detection. However, in order to achieve clinical utilities in liquid biopsy for cancer, tremendous efforts are still needed to improve the adaptability of the microfluidic technologies to clinical settings and promote the commercialization of the systems. Some of the microfluidic platforms described above require off-chip sample preparation steps, which introduce extensive manual interventions. Microfluidic technologies are still largely developed for operation under the research settings and require expensive setup and well-trained professional to run tests. Simple colorimetric or mobile device–based readout from blood drops is garnering great interest. Other essential research efforts include improving exosome processing throughput and enhancing assay reliability. Overall, in addition to providing new abilities to better elucidate the biology and clinical relevance of exosomes, robustness and ease of use will be the key considerations in microfluidic research to drive the benchtop-to-bedside transition of both technology innovation and scientific advances in exosome biology. Cancer is a complicated and dynamic disease. For accomplishing personalized cancer medicine, the development of a reliable, novel liquid biopsy platform will have tremendous benefit. We anticipate that microfluidic technology will play a game-changing role in exosome analysis and liquid biopsy of cancer.
Footnotes
Acknowledgements
We would like to acknowledge our collaborator, Dr. Andrew Godwin at the University of Kansas Cancer Center, and fund donor Harvey McCarter for supporting our research.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the K-INBRE Developmental Research Project Award from NIH/NIGMS (P20GM103418) and the Innovative Research Award from the Terry C. Johnson Cancer Research Center to M.H., and the J. R. and Inez Jay Award from the University of Kansas, the COBRE Research Project Award under P20GM103638 (NIGMS), and NIH/NCI grant R21CA186846 to Y.Z.
