Abstract
Epidermal growth factor receptor (EGFR) mutations are the most common carcinogenic driver mutations in non-small-cell lung cancer (NSCLC) patients, while invasive tissue biopsy has certain inherent defects. PubMed, Ovid Medline, Embase, and the Cochrane Library were systematically searched on January 4, 2020, using the keywords “liquid biopsy,” “EGFR,” and “NSCLC.” The pooled sensitivity and specificity of EGFR mutations in paired tissue and blood were calculated. The accuracy was assessed by receiver operating characteristic curve. The meta-regression of the subgroup was performed to analyze the heterogeneity. Hazard ratio (HR) and 95% confidence interval (CI) were combined for evaluating the impact of EGFR mutation in tissue and liquid blood biopsy. A total of 40 studies with 5,995 patients were involved in the study. The pooled sensitivity was 68% (95% CI = 60–75%), and the specificity was 98% (95% CI = 95–99%). The diagnostic odds ratio was 88 (95% CI = 40–195), and the area under the curve was 0.91 (95% CI = 0.88–0.93). In the meta-regression, the sensitivity and specificity remain lower in the Asian studies than non-Asian studies (sensitivity: 66% vs. 73%, P = 0.04; specificity: 96% vs. 97%, P = 0.03, respectively). The EGFR mutation was associated with a better progression-free survival than wild type in both tissue (HR = 0.54, 95% CI = 0.34–0.85, P = 0.007) and blood (HR = 0.81, 95% CI = 0.71–0.92, P = 0.001) detection. Peripheral blood liquid biopsy had a better specificity for detecting EGFR mutation in NSCLC patients, while tissue biopsy still needs to be undertaken for negative blood biopsy patients due to its lower sensitivity.
Introduction
Lung cancer is one of the most common cancers and the leading cause of cancer-related deaths worldwide.1,2 More than 1.8 million new lung cancer cases (13% of total diagnosed cancer cases) and 1.6 million lung cancer–related deaths (19.4% of the total) are diagnosed globally every year. 3 The incidence of non-small-cell lung cancer (NSCLC) accounts for approximately 85–90% of all new-onset lung cancers. For the treatment of NSCLC according to the diagnosis and treatment guidelines, patients who are in earlier stages than IIIA strive for surgical resection, and advanced patients mainly adopt medical treatment. 4 Although improvement has been made in the treatment of lung cancer in recent years, the 5-year relative survival rate of NSCLC is still 18%, and more than 50% of cases are diagnosed as advanced. 5 Epidermal growth factor receptor (EGFR) mutations are the most common carcinogenic driver mutations, with a positive rate of 30–50% in Southeast Asian populations. Notably, in those lung adenocarcinoma patients who did not have a history of smoking, the mutation rate is as high as 50–60%. 6 As studies reported previously, EGFR mutation status can not only predict the prognosis of patients but also predict the efficacy of EGFR tyrosine kinase inhibitors (EGFR-TKIs).7,8
For the detection of oncogene mutations, tissue samples obtained from surgical specimens or tissue biopsies are the “gold standards.” 9 Tissue biopsy, however, has some inherent defects in clinical practice. First, the invasive approach may lead to complications during the procedure.9,10 Second, compared with metastatic tumor tissues, the mutations found in different metastatic clones of genetic mutations can be significantly different. A small tissue sample will lead to bias in genetic test results. 11 Lastly, with the use of chemotherapy and targeted drugs, patients’ genetic mutations will also be changed. 12 Because of this, alternative methods are urgently needed to obtain relevant molecular biological information. In recent years, “liquid biopsy” based on free nucleic acid has been receiving more and more attention.
Circulating tumor DNA (ctDNA) is produced by apoptotic or necrotic tumor cells, circulating tumor cells, or metastatic tumors. In other words, ctDNA is considered to be extracellularly produced by tumors. 9 ctDNA originates from somatic cells and is normally relatively stable in quantity. 13 ctDNA fragments carrying tumor-specific genes can be extracted from the blood and furtherly examined.14,15 This non-invasive “liquid biopsy” test can be easily and repeatedly performed, and at the same time is able to reflect the mutation of tumor genes in patients. 16 Compared with traditional biopsy methods, ctDNA has the potential effect of reflecting all heterogeneous mutations in real time during the treatment of patients. For this reason, ctDNA offers new possibilities for diagnosis, assessment of prognosis, and patient follow-up compared to traditional tissue biopsy. The diagnostic accuracy reported in different studies, however, remained controversial. Thus, we designed the systematic review and meta-analysis to evaluate the diagnostic accuracy of ctDNA in peripheral blood “liquid biopsy” for detection of EGFR mutation in NSCLC, and to assess the prognostic effect of EGFR mutation in association with the treatment response and survival outcome.
Method
This study was designed by the Preferred Reporting Items for Systematic Review and Meta-Analysis (PRISMA) guidelines. 17
Search Strategy
This systematic review and meta-analysis were designed to compare the sensitivity and specificity between tissue and blood in detecting EGFR-mutated NSCLC in the re-biopsy process. PubMed, Ovid Medline, Embase, and the Cochrane Library were systematically searched on January 4, 2020. Moreover, related websites and Google Scholar were searched for the gray literatures. The keywords were designed by an experienced librarian. Briefly, the keywords were “re-biopsy” or “repeat biopsy” or “liquid biopsy” or “plasma biopsy” or “circulating tumor DNA” or “circulating cell-free DNA,” combined with “EGFR” and “NSCLC.” All the studies containing titles and abstracts were imported into Endnote for duplicate deletion and literature screening.
Criteria for Inclusion and Exclusion
All the studies comparing sensitivity and specificity between tissue biopsy and blood biopsy in EGFR-mutated NSCLC were included in the meta-analysis. The inclusion criteria were as follows: (1) The tissue biopsy and blood biopsy should be paired in the same patient; (2) the number of true positives (TP), true negatives (TN), false positives (FP), and false negatives (FN) should be reported; (3) all the patients should be diagnosed with NSCLC; and (4) the EGFR driver gene should be detected. Studies were limited in prospective or retrospective cohort studies. Other meta-analyses, reviews, letters, comments, and conference reviews were evaluated for the further inclusion of the studies.
The exclusion criteria were as follows: (1) The involved sample size was fewer than 10, (2) the study did not contain a distinct group of patients with NSCLC, (3) the study did not contain the pairs between tissue and blood biopsy, (4) the study had no full data to extract the sensitivity and specificity, and (5) the study was not written in English.
Literature Screening, Data Extraction, and Quality Evaluation
Two investigators (Naiqun Wang and Xiaolian Zhang) independently screened the titles and abstracts according to the inclusion and exclusion criteria. The full text was further evaluated if the abstracts could not be determined. The third investigator (Wei Lu) was adapted for discussion if any disagreement existed.
A standard Excel spreadsheet was designed for data extraction, and the following information was collected from the original studies: the study characteristics (author, publish year, country, institution, recruitment period, study design, etc.); patient characteristics (EGFR mutation, age, gender, smoking status, tumor stage); the biopsy method in tissue and blood; the cases of TP, FP, FN, and TN in comparison to the sensitivity and specificity between tissue and blood; and the concordance between two biopsy methods. The sensitivity equals TP / (TP + FN), while specificity equals TN / (TN + FP). The tissue biopsy was regarded as the non–reference standard. If the studies reported the association between EGFR mutation and short- and long-term outcome of the NSCLC patients, the treatment response, including overall response rate, disease control rate, complete response, partial response (PR), stable disease (SD), and progression of disease (PD), was assessed after the Response Evaluation Criteria in Solid Tumors (RECIST) was recorded. 18 Moreover, the hazard ratio (HR) with a 95% confidence interval (CI) was extracted from the regression or survival plot for progression-free survival (PFS) and overall survival (OS).
Two researchers (Weihua Yin and Shaorong Deng) independently assessed the quality of the included papers. The quality assessment was followed by the Quality Assessment for Studies of Diagnostic Accuracy Score-2 (QUADAS-2) tool, with four questions to assess a high, unclear, or low risk of bias. 19
Statistical Analysis
All calculations were performed by Stata 15.0 software (Stata Corporation, College Station, TX). In terms of diagnostic meta-analysis, the MIDAS module was used. For each study, the sensitivity and specificity were calculated, and the analysis was based on a summary receiver-operator characteristics (SROC) curve.20,21 The binary variables were regressed to calculate pooled sensitivity and specificity. 22 The covariates included study institution (multiple or single), country (Asian or not), median recruitment year (later than 2010 or not), sample size (more than 40 or not), blood samples (plasma or serum), and detection method applied in peripheral blood and tumor tissue (same or not). The treatment response in EGFR mutation and wild type in either tissue or blood detection was compared and combined using relative risk (RR). The survival analysis was combined using HR with 95% CI. If the HR was not described explicitly in the univariate or multivariate analysis, we summarized the time-to-event data through the survival curve based on Tierney’s method. 23 The χ2 test and I2 statistics were used for heterogeneity assessment (I2 ≥ 50% indicating the presence of heterogeneity). When the heterogeneity was detected, the Spearman correlation coefficient was calculated to judge whether the threshold effect existed. A P-value less than 0.05 was set as a significant difference.
Results
Literature Selection
A total of 5,143 studies were found based on the search strategy. A flowchart of the literature selection is shown in Figure 1 . After screening the titles and abstracts, 243 studies were scanned in full text. After exclusion of studies, a total of 40 studies were finally included in our systematic review and meta-analysis.24–63
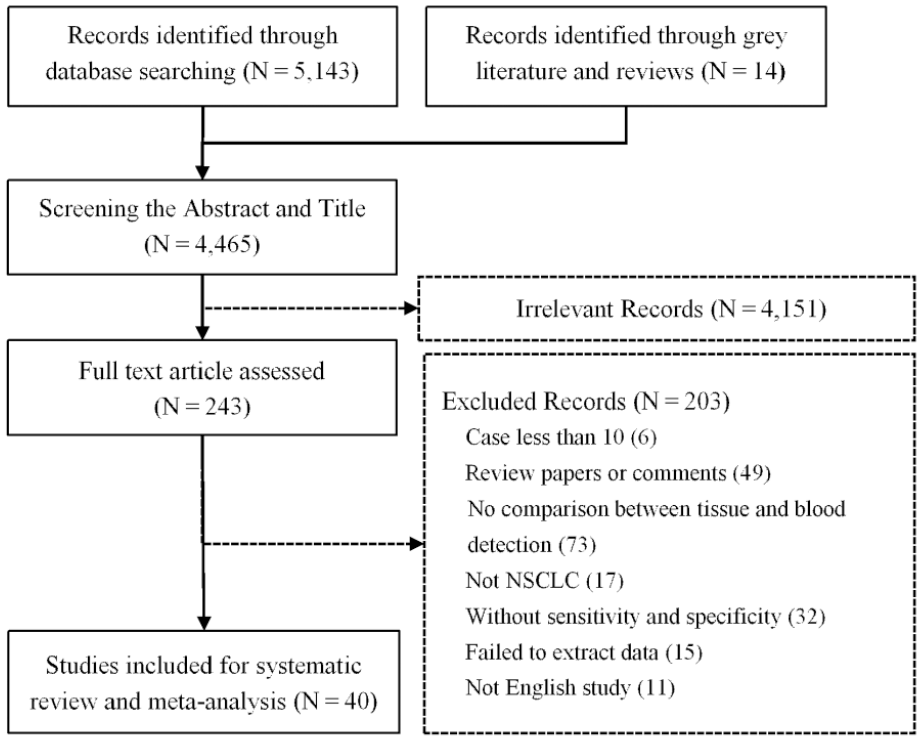
Flowchart of the literature screening.
Characteristics of Included Studies
The characteristics of the included studies were listed in Table 1 . A total of 5,995 patients were detected for EGFR mutation. The publishing year was between 2006 and 2019, with a recruitment year ranging from 1992 to 2018. Three studies were multicenter analysis, while single-center studies included China (18 studies), Japan (6 studies), United States (5 studies), Korea (3 studies), Australia (2 studies), India (1 study), Turkey (1 study), and Denmark (1 study). The median age was 61.6 years. 55% of patients were male, and 42% of patients did not have a history of smoking. The advanced-stage patients (tumor stage IIIB–IV) represented 95% of the patients (ranging from 47% to 100%). The quality of included studies was assessed by QUADAS-2 guidelines ( Suppl. Figs. 1 and 2 ). A total of four domains were evaluated for risk of bias assessment, and three factors were used for assessing the applicability concerns. The majority of studies are combined with a low risk of bias or unclear bias in patient selection (92.5%), index test (80%), reference standard (95%), and flow and timing (85%).
The Characteristics of NSCLC Patients in the Included Studies.
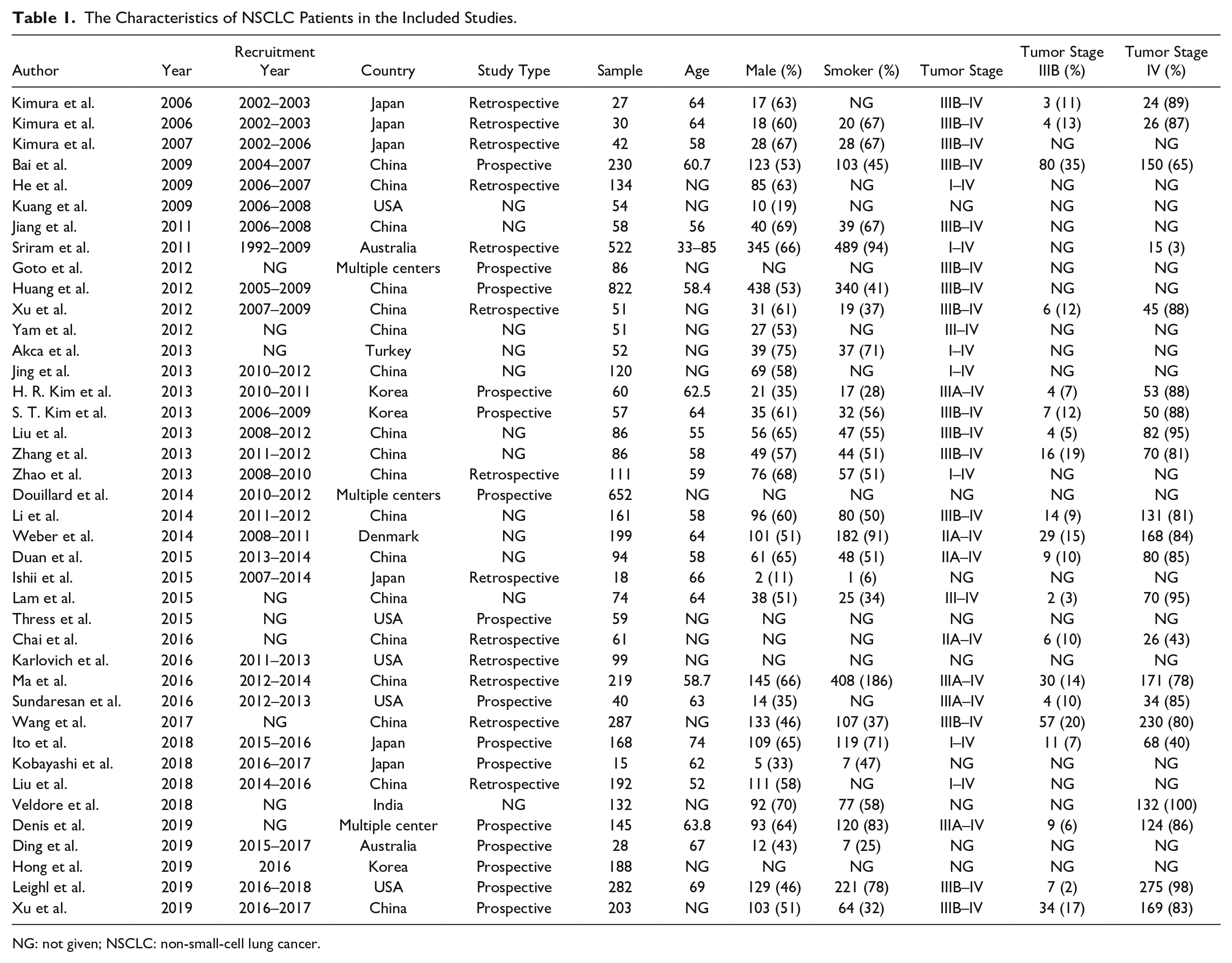
NG: not given; NSCLC: non-small-cell lung cancer.
Comparison of Detection Rates of EGFR Mutations in Paired Blood and NSCLC Tissue
As shown in Table 2 and Figure 2 , all studies provided the data for calculating the sensitivity and specificity in evaluating the effectiveness of blood biopsy on the detection of EGFR mutations. Nine studies used PCR to detect the EGFR mutation in the tissue, the pooled specificity of which was 97%, while the sensitivity was only 51%.28,31,38,42,46,47,53,60,61 Thirteen studies used the amplification refractory mutation system (ARMS) or scorpion ARMS to detect the EGFR mutation in blood, the pooled specificity of which was 98%, while the sensitivity was 66%.29,32,34,40,43,44,49,50,52,54,57,59,63 In the six studies using gene sequencing to detect the EGFR mutation, the pooled specificity was 98%, and the sensitivity was 79%.24–26,30,58,62 Thirty-eight studies used the same detecting methods between tissue and blood biopsy. The pooled sensitivity was 68% (95% CI = 60–75%), and the specificity was 98% (95% CI = 95–99%). Besides, the results showed a positive likelihood ratio of 28.7 (95% CI = 13.4–61.5) and a negative likelihood ratio of 0.33 (95% CI = 0.26–0.41). The diagnostic odds ratio (DOR) was 88 (95% CI = 40–195), and the area under the curve was 0.91 (95% CI = 0.88–0.93), showing that the peripheral blood test had a high diagnostic performance in detecting EGFR mutation ( Fig. 3A ). The results, however, showed a high heterogeneity with I2 of 85.56% and 90.58%. Thus, after assessing the potential bias of the included studies, we used the random-effect model to weigh those studies. The Spearman correlation coefficient was calculated to assess the threshold effect. The Spearman correlation coefficient was −0.31, and the P-value was 0.10 (>0.05), confirming that the threshold effect was not significant. Besides, we assessed the Deek’s funnel plot test to assess the publication bias of the included study; the P-value was 0.71 (>0.05), suggesting no evidence of publication bias ( Fig. 3B ).
The Methodology of Detection and the Data for EGFR Mutations in Paired Tumor Tissue and Blood of Patients.
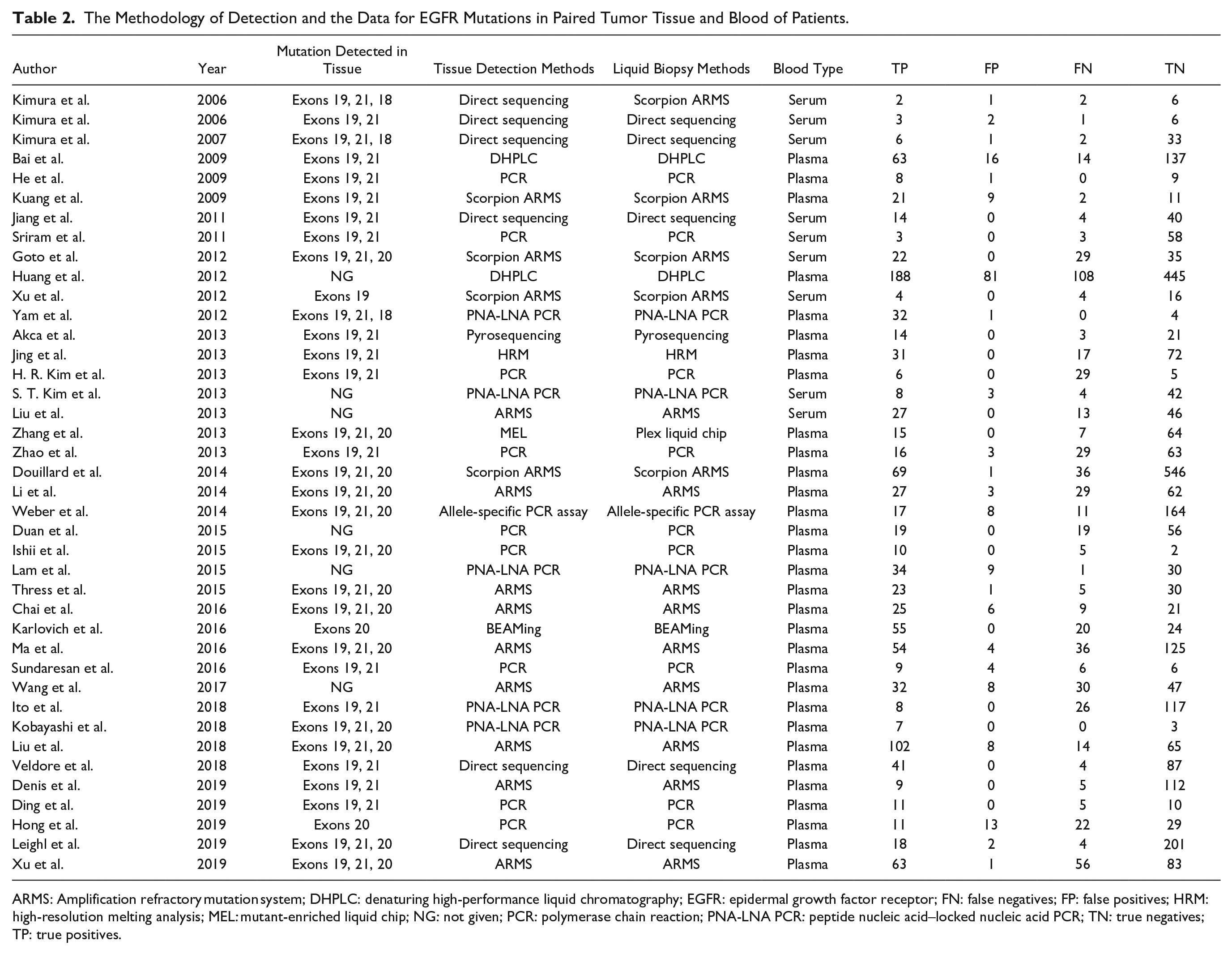
ARMS: Amplification refractory mutation system; DHPLC: denaturing high-performance liquid chromatography; EGFR: epidermal growth factor receptor; FN: false negatives; FP: false positives; HRM: high-resolution melting analysis; MEL: mutant-enriched liquid chip; NG: not given; PCR: polymerase chain reaction; PNA-LNA PCR: peptide nucleic acid–locked nucleic acid PCR; TN: true negatives; TP: true positives.
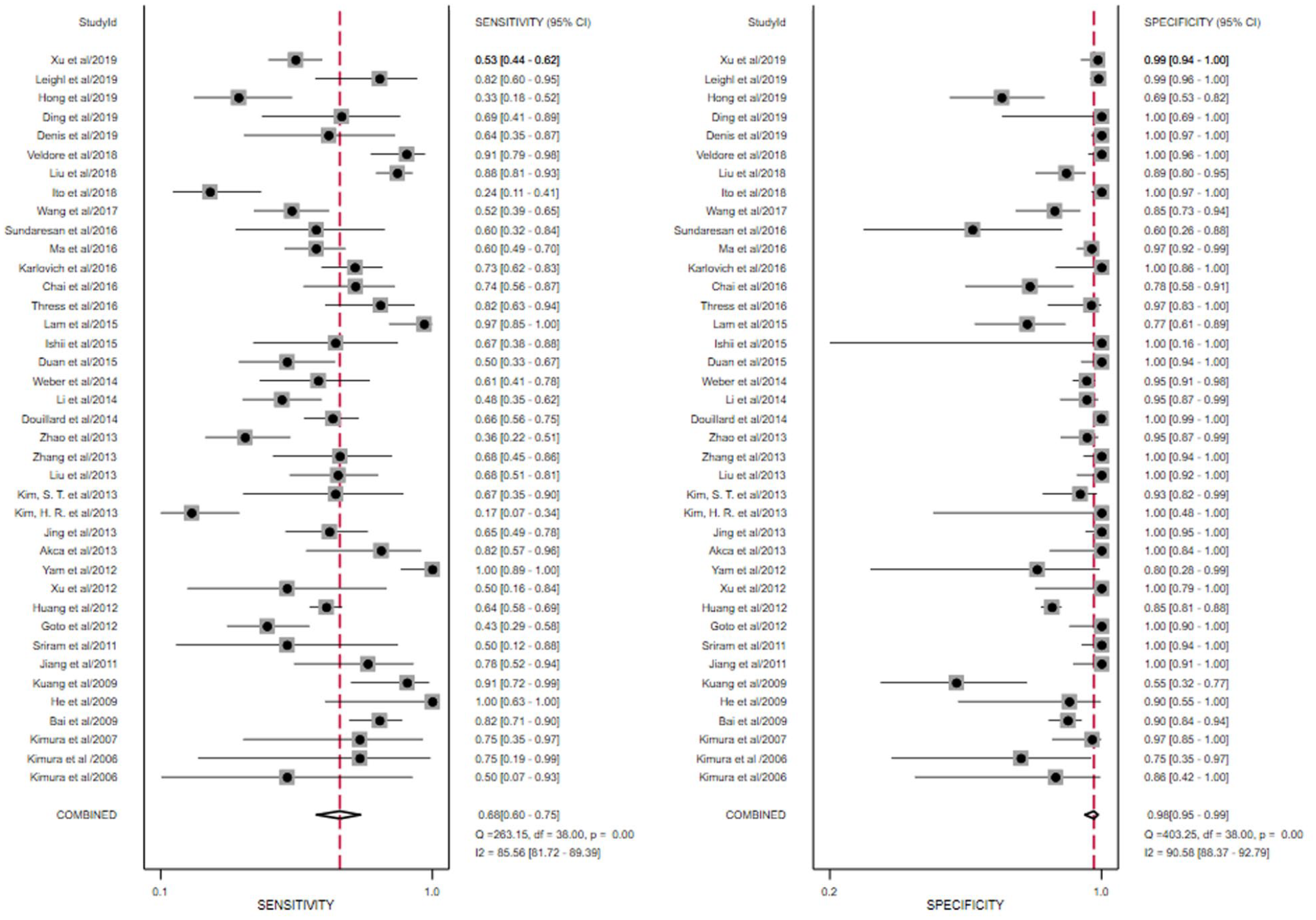
The forest plot of the pooled sensitivity and specificity of liquid biopsy in detecting epidermal growth factor receptor (EGFR) mutation in non-small-cell lung cancer (NSCLC) patients.
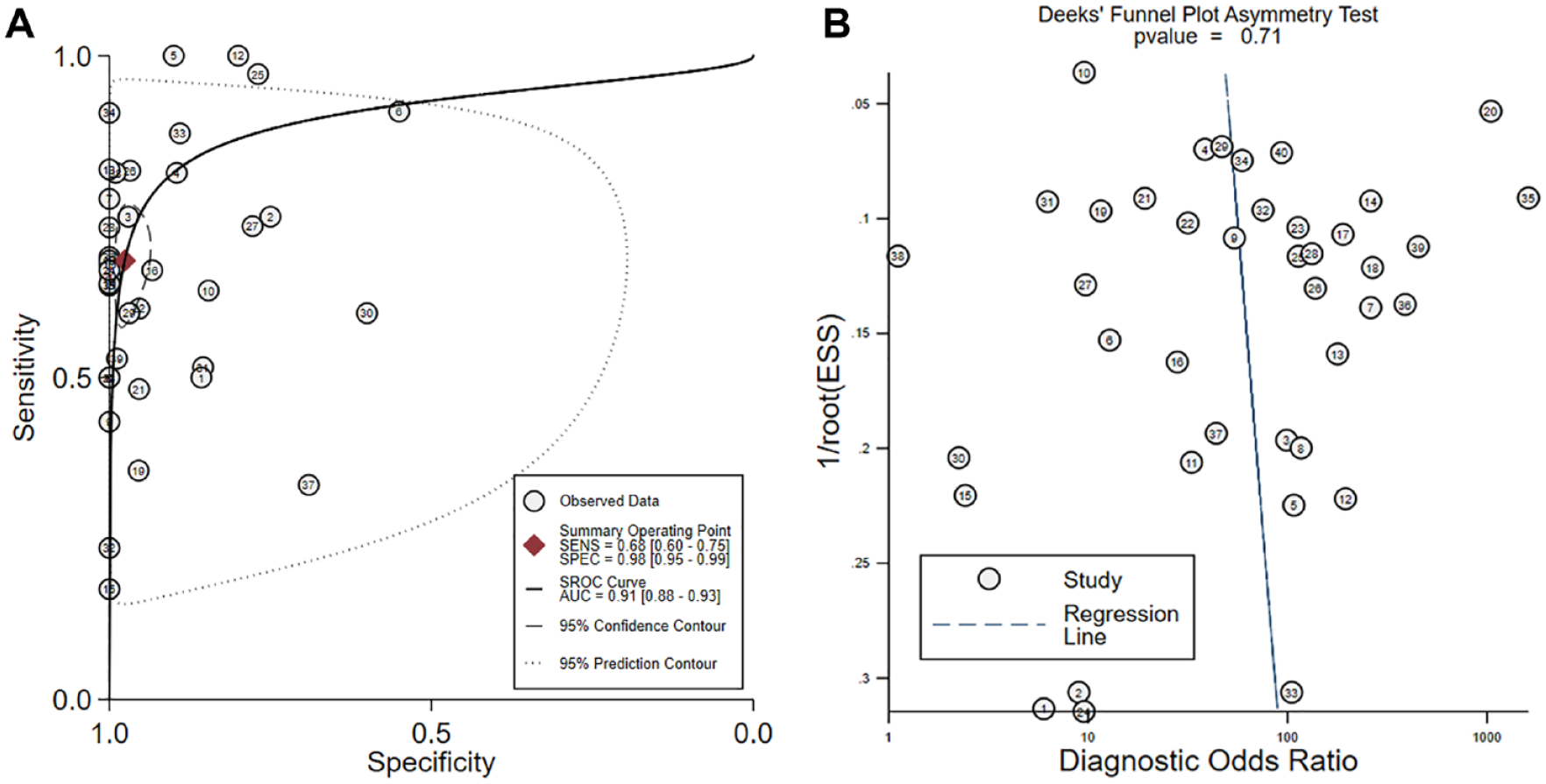
(A) Summary receiver operating characteristics (SROC) plots showed a good accuracy of liquid biopsy in detecting epidermal growth factor receptor (EGFR) mutation in non-small-cell lung cancer (NSCLC) patients. (B) Deek’s funnel plot showed no significant publication bias.
Meta-Regression and Subgroup Analysis
The meta-regression and subgroup analysis were shown in Figure 4 . The multicenter studies32,43,59 pooled a higher specificity (100%) than the single-center studies (97%)24–31,33–42,44–58,60–63 (P < 0.001). Besides, the similar detection methods24–40,42–59,61–63 between tissue and blood pooled a lower specificity (97%) than the two studies41,60 without similar methods (100%) (P < 0.001). Due to the small sample size of studies in those two subgroups, however, the specificity showed the statistical significance, while the difference of specificity did not remain significant. Moreover, the sensitivity and specificity remain lower in the studies published by Asian countries than those published in other areas (sensitivity: 66% vs. 73%, P = 0.04; specificity: 96% vs. 97%, P = 0.03, respectively). No statistical significance was found in the median recruitment year (after 2010 or not), sample size (more than 40 or not), or blood sample (plasma or serum) subgroups.
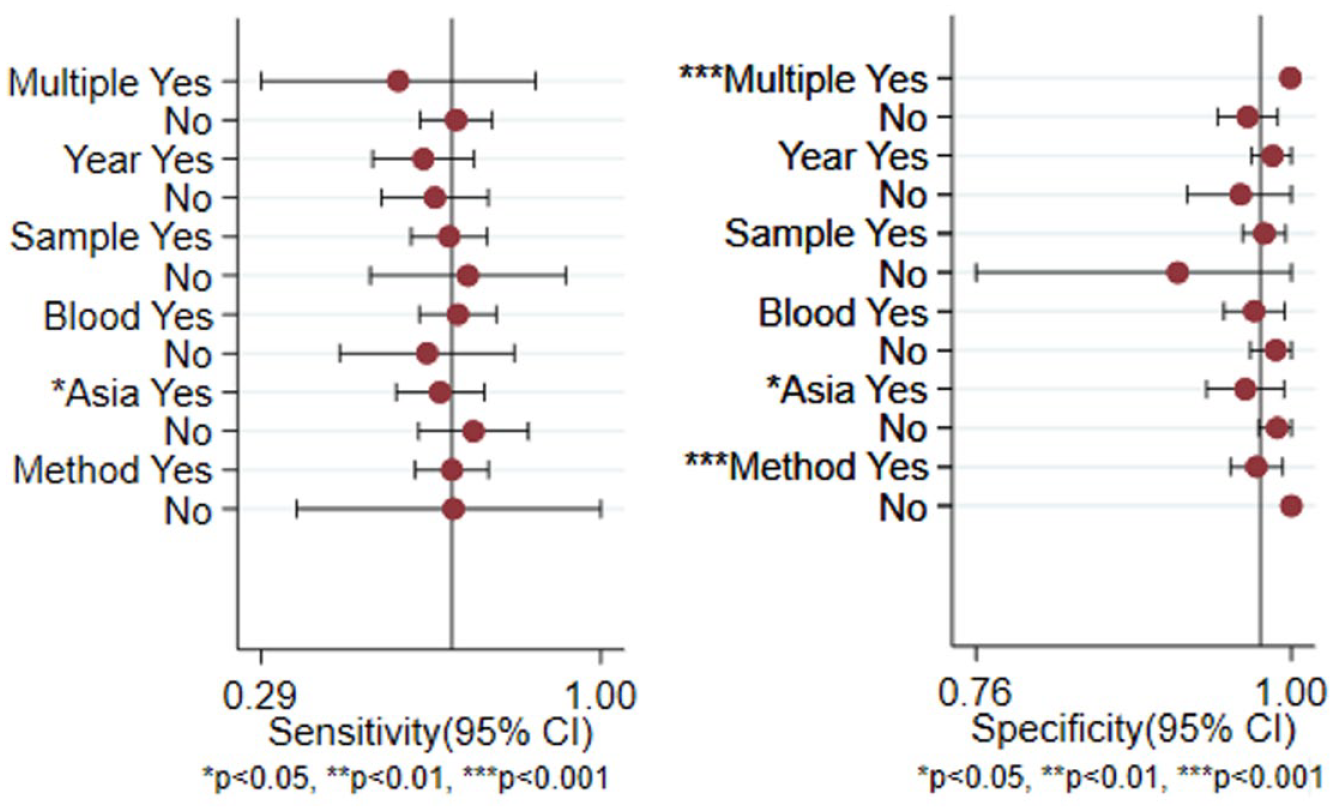
Forest plots of subgroup meta-regression for sensitivity and specificity. The covariates included multiple center (yes or no), country (Asia or not), median recruitment year after 2010 (yes or no), sample size more than 40 (yes or no), blood sample from plasma (yes or no), and similar method applied in peripheral blood and tumor tissue (yes or no).
The Predictive Value of EGFR Mutation in Treatment Response
The association of EGFR mutation and treatment response is shown in Figure 5A (mutation detected in tissue) and Figure 5B (mutation detected in blood). For the samples detected in tissue, the PR rate was 56% in EGFR mutation NSCLC compared to 12% in wild-type NSCLC (RR = 0.30, 95% CI = 0.19–0.47, P < 0.001). Besides, the SD/PD rate was 26% in EGFR mutation NSCLC compared to 55% in wild-type NSCLC (RR = 1.59, 95% CI = 1.18–2.16, P < 0.001). Similarly, for the samples detected in blood, the PR rate was 54% in EGFR mutation NSCLC compared to 20% in wild-type NSCLC (RR = 0.53, 95% CI = 0.38–0.72, P < 0.001). Also, the SD/PD rate was 28% in EGFR mutation NSCLC compared to 54% in wild-type NSCLC (RR = 1.64, 95% CI = 1.19–2.25, P < 0.001).
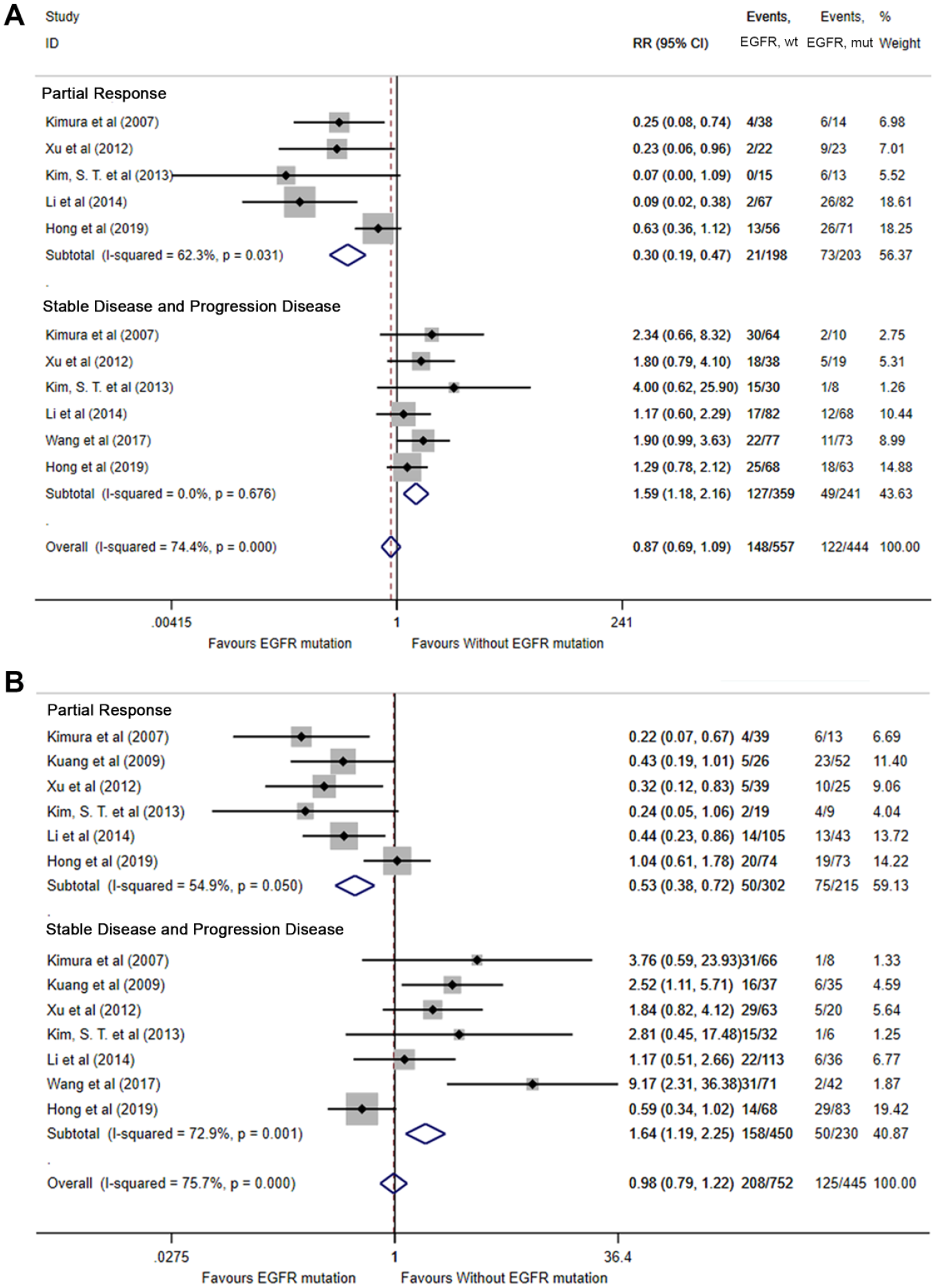
The predictive value of epidermal growth factor receptor (EGFR) mutation in association with treatment response in (A) tissue biopsy and (B) liquid biopsy.
The Predictive Value of EGFR Mutation in Long-Term Survival
The combination of HRs in evaluating the predictive value of EGFR mutation in survival was shown in Figure 6A (PFS) and Figure 6B (OS). Generally, in terms of PFS, the EGFR mutation was associated with a better PFS than wild type in both tissue detection (HR = 0.54, 95% CI = 0.34–0.85, P = 0.007) and blood detection (HR = 0.81, 95% CI = 0.71–0.92, P = 0.001). In terms of OS, however, no significant difference was found that EGFR mutation was associated with a better OS in both tissue detection and blood detection (all P > 0.05).
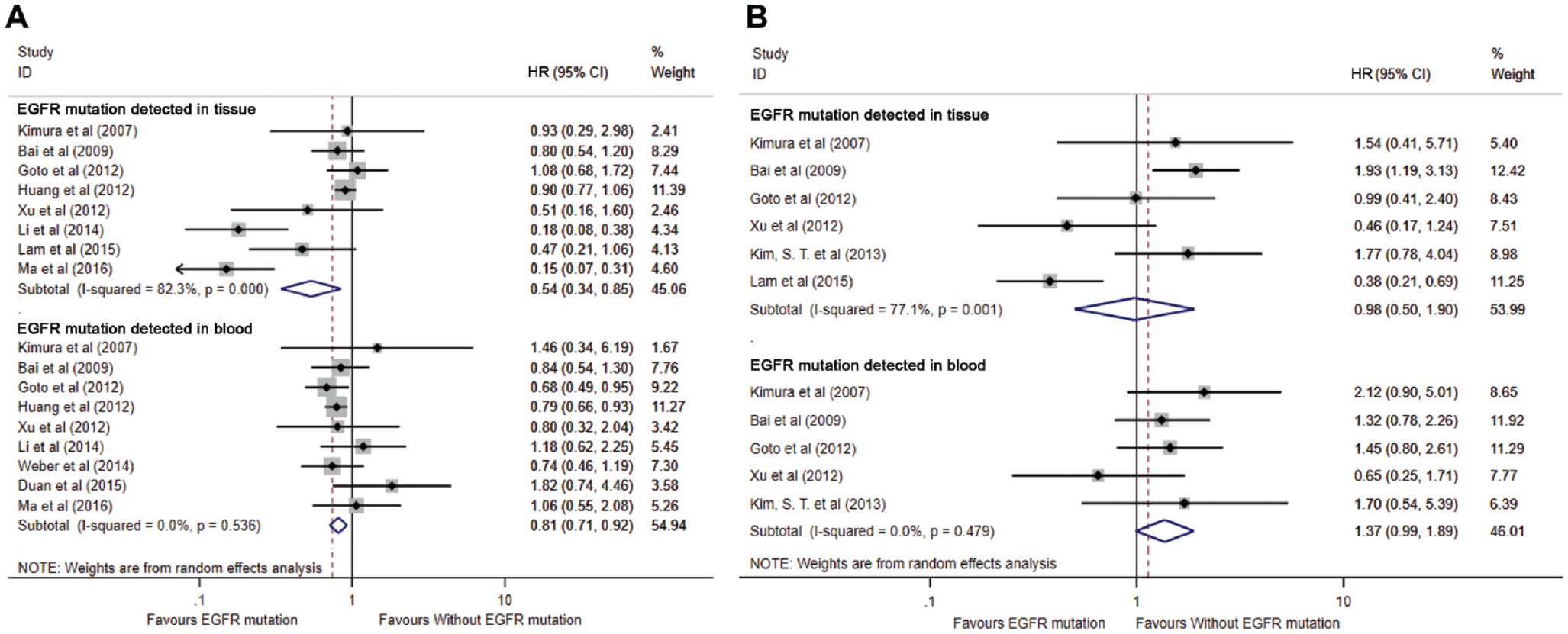
The predictive value of epidermal growth factor receptor (EGFR) mutation in association with (A) progression-free survival and (B) overall survival.
Discussion
This is a large-scale meta-analysis, including 40 studies (5,995 patients) investigating the diagnostic accuracy of peripheral blood ctDNA-based “liquid biopsy” in the detection of EGFR mutation in patients with NSCLC. In our study, the peripheral blood biopsy approach is characterized by a pooled sensitivity of 0.68 and a pooled specificity of 0.98, which were higher compared to the previous meta-analysis with a sensitivity of 61% and specificity of 90%. 64 The heterogeneity still existed in the meta-analysis, which was similar to the previous study report. 65 Through the Spearman correlation coefficient test, Deek’s funnel plot test, and meta-regression, we suggested that the study centers (multiple and single center), consistency of detecting methods between tissue and blood, and human races may be the factors associated with the heterogeneity. Due to the small sample size of the multiple-center analysis and inconsistency between the two biopsy methods, however, a small difference between specificity could not affect the usefulness of liquid biopsy in clinical application, although a statistical significance was found. Our study suggested that the specificity and sensitivity remained lower in the studies published by Asian countries than in other areas. For one reason, this may be because we included 27 Asian studies with the recruiting period between 2006 and 2019 and compared those to 13 non-Asian studies with the recruiting period between 2009 and 2019. The non-Asian studies may have more ARMS and sequencing instead of PCR. For another reason, the proportion of EGFR mutation NSCLC patients was different in Asian and other human races, and thus resulted in the limited difference between Asian studies and non-Asian studies. The application of peripheral blood in the different human races still needs to be demonstrated further with multicenter analysis. Moreover, our results also demonstrated that EGFR mutation was combined with better treatment response and PFS, which might be associated with the therapeutic efficacy of second-generation or third-generation EGFR-TKI. 66 In contrast, patients without EGFR mutation may not benefit from EGFR-TKI, thus resulting in a higher rate of SD and PD. There was no significant difference in OS, however, which may be affected by the short follow-up period in some studies.
Current research suggests that patients with EGFR mutations may detect EGFR mutation status when treating with EGFR-TKI and finding the early stage of EGFR-TKI resistance.7,12 Compared with the difficulty in obtaining tissue samples from patients with advanced lung cancer and the heterogeneity existing in the tissue samples, ctDNA gene detection has the advantages of being less invasive and avoiding the heterogeneity of tumors existing in different locations of the lung. 67 ctDNA is derived from tumor cells in the blood circulation of tumor patients. There are four main hypotheses about how tumor cells release free DNA into the blood circulation: (1) lysis of circulating tumor cells, (2) apoptosis of tumor cells, (3) necrosis of tumor cells, and (4) spontaneous release of tumor cells (exocrine). 68 Based on the ctDNA carrying part of the genetic information of tumor cells, ctDNA can be used for molecular diagnosis of tumors, especially for EGFR mutation detection. At present, several guidelines and institutions have demonstrated the efficacy of blood biopsy in detecting EGFR mutation in NSCLC patients.69,70 Moreover, the key advantage of ctDNA is its high specificity, and its misdiagnosis rate is extremely low. When tissue biopsy is difficult to perform, and the result of liquid biopsy is positive, EGFR-TKI treatment can be attempted. As the results showed in our study, however, peripheral blood gene detection is less sensitive. The previous study showed that sensitivity also depends on the state of the tumor, such as tumor size, whether there is lymph node metastasis, and tumor burden. Also, it is considered to be related to the lower abundance of ctDNA mutations. Oellerich et al. and Fiala et al. also stated that the FN results can occur in particular with small tumor size (e.g., <1 cm), due to the insufficient mutant DNA fraction (<0.01%). 71 Besides, different detection methods had a different lower abundance of ctDNA. Yang et al. found that digital droplet PCR (ddPCR) can detect 0.04% mutations. The sensitivity of ctDNA detection is limited by the abundance of EGFR mutations. 72 Besides, the lower limit of detection of the ARMS is 1% mutations. 73 Moreover, EGFR mutations are abundant at 20% to 30% before they can be detected by direct sequencing. 74 Currently, commonly used detection methods include ddPCR, BEAMing, and so on, making it possible to detect trace amounts of DNA quantitatively. In advanced tumors, digital detection has obvious advantages. Mutations in tumor tissues are completely consistent with mutations in ctDNA, which has extremely high sensitivity and specificity. At present, next-generation sequencing (NGS) technology based on PCR is more widely used for the detection of ctDNA. 75 The advantage of NGS technology is that it can detect multiple genes and mutation sites on the same sample, and in addition to point mutations, it also can detect a variety of mutations such as gene amplification, rearrangement, and chromosomal mutations. 76 In our study, we suggested that the mutation detection by sequencing has the highest sensitivity comparing to ARMS and normal PCR, which could achieve up to 79% in liquid biopsy.
Except for detecting the EGFR mutations in advanced-stage NSCLC patients, the liquid biopsy based on ctDNA was used as a monitoring method in recent years, especially with the development of NGS and other sequencing technology. Besides, some studies suggested that liquid biopsy could be used for identifying hidden residual diseases, thereafter providing a new strategy for clinical treatment decisions.77,78 More recently, Aaron Newman et al. have invented a highly sensitive ctDNA library construction and detection method, CAPP-Seq, in patients with stage I NSCLC. The sensitivity of this method is 50% in patients with stage I and 100% in patients with stage II–IV. The specificity reaches 96% with a mutant gene ratio of 0.02%. The team has recently improved the CAPP-Seq technology platform so that when the proportion of mutant genes is as low as 0.004%, the sensitivity reaches 90%. This is by far the lowest ctDNA mutation concentration that can be detected on all technology platforms. 79 There is still much to be explored and improved in the detection technology of ctDNA.
There were some limitations in our meta-analysis. We detected heterogeneity among the included studies. Although a meta-regression analysis was performed, only the human race was associated with the heterogeneity in sensitivity, but this may not be the only factor associated with heterogeneity. Second, apart from the factors analyzed, the included studies differed in many aspects, such as the percentage of different types of EGFR mutation, treatment methods, blood collection time, methodological quality, and the relationship between blood collection and treatment. These undocumented differences may be a potential source of heterogeneity. Moreover, the number of factors included in the meta-regression analysis is relatively small. Therefore, the results are likely to be biased. Although the detection threshold for mutant DNA in liquid biopsies will decrease as technology improves, it is not easy to determine the clinical correlation between low-level mutant DNA in plasma and the probability of responding to EGFR-TKIs. Further efforts still need to be undertaken in future clinical research and practice.
Conclusion
Peripheral blood liquid biopsy had a better specificity for detecting EGFR mutation in NSCLC patients, while tissue biopsy still needs to be undertaken for negative blood biopsy patients due to its lower sensitivity. EGFR mutation detected in either tissue biopsy or liquid biopsy was related to a better PFS compared to wild-type NSCLC patients.
Supplemental Material
Supplement_file – Supplemental material for The Diagnostic Accuracy of Liquid Biopsy in EGFR-Mutated NSCLC: A Systematic Review and Meta-Analysis of 40 Studies
Supplemental material, Supplement_file for The Diagnostic Accuracy of Liquid Biopsy in EGFR-Mutated NSCLC: A Systematic Review and Meta-Analysis of 40 Studies by Naiqun Wang, Xiaolian Zhang, Feilong Wang, Min Zhang, Bo Sun, Weihua Yin, Shaorong Deng, Ying Wan and Wei Lu in SLAS Technology
Footnotes
Supplemental material is available online with this article.
Author’s Contribution
Design of the meta-analysis: Feilong Wang, Min Zhang, Bo Sun, and Wei Lu. Literature screening: Naiqun Wang and Xiaolian Zhang. Quality assessment: Weihua Yin and Shaorong Deng. Statistics analysis: Naiqun Wang, Wei Lu, and Ying Wan. Writing and revision: Naiqun Wang, Xiaolian Zhang, Feilong Wang, Min Zhang, Bo Sun, Weihua Yin, Shaorong Deng, Ying Wan, and Wei Lu.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
