Abstract
In recent years, various label-free biosensing technologies have been developed for studying the real-time kinetics of diverse biomolecular interactions. These biosensors partially take the place of fluorescence-based methods by providing a comparable sensitivity as well as retaining the conformational and functional integrality of biomolecules to be investigated. However, to completely eliminate the need of fluorescence, throughput is the next big consideration. Microarrays provide a high-throughput platform for screening tens of thousands of biomolecular interactions simultaneously, and many compatible fluorescent scanners have been commercially available. The combination of microarrays and label-free biosensors will be of great interest to researchers in related fields. Microarrays are fabricated by spotting, imprinting, or directly synthesizing biomolecules on solid supports such as glasses, silicon wafers, and other functionalized substrates, and they have been applied to detect DNAs, proteins, toxins, and so on in surface plasmon resonance (SPR) imaging systems and oblique-incidence reflectivity difference (OI-RD) microscopes. Current challenges include increasing sensitivity, reducing sampling time, improving surface chemistry, identifying captured molecules, and minimizing reagent consumption. Future research directions are to improve the instruments themselves, modify the microarray surface for more efficient analyte capture, and combine the systems with mass spectrometry and microfluidics.
Keywords
Introduction
Biosensors are important tools in characterizing biomolecular interactions, which have great applications in proteomics, drug screenings, and even medical diagnosis. In the past few years, various methods have been developed and commonly used to detect interactions between two molecules. Traditional measurements such as chromatography and immunoprecipitation are widely used for detecting protein–protein interactions. Often, fluorescence-based methods are used because of their high sensitivity and usability. For example, fluorescence labeling is used to detect both real-time binding kinetics1,2 and endpoint images (images taken at certain time points). 3 Usually, one or more biomolecules are tagged chemically with fluorophores (Cy3-, Cy5-dye or quantum-dot), and with sophisticated scanning and detecting components, these methods provide very high throughput and dynamic range. However, some problems regarding these “labeled” methods have been pointed out.4–7 First, labeling biomolecules (especially proteins) can change their conformational and functional properties. Those extrinsic tags can have significant influence on both binding affinity and isothermal equilibrium. Second, the heterogeneity of biomolecules makes quantitative measurements difficult because the efficiency of labeling may vary from one molecule to another. Third, labeling biomolecules is very laborious and expensive.
As a result, label-free biosensors based on optical,8–11 mechanical,12,13 and electrical techniques14–17 were developed to resolve these problems. Among these, optical biosensors have been the leading research fields because they have many advantages over others such as good sensitivity, high throughput, flexible system integration, and various measurable physical properties. In label-free optical biosensors, target biomolecules are often immobilized on solid substrates, and the optical properties change in response to the bindings of probe molecules. These properties can be refractive index (RI),18–20 absorption,21–24 emission,25–28 or scattering. 29
Early in the 1940s, optical methods based on light interference and ellipsometry were developed for studying biomolecular interactions. 30 Recently, RI-based detections have been applied in designing and constructing various label-free optical biosensors. For example, the surface plasmon resonance (SPR) technique works by measuring the change in coupling conditions, in which surface plasmon polaritons are excited in a very thin metal film, in response to the change in RIs of surface-bound molecules above the film. 31 Resonant mirror (RM), a leaky waveguide-based technique, uses the evanescent field to probe changes in the RI at the sensing surface. 32 Reflectometric interference spectroscopy (RIFS), which determines the change in thickness and RI of a thin deposited layer by measuring the interference of a light source caused by partial reflection at the interface, works well in affinity detection of low-molecular-weight analytes. 33 Elliposometry measures the differential changes in both phase and magnitude of the reflection coefficients for p-polarized and s-polarized components of a monochromatic light, 34 and it has been applied to both real-time and endpoint detections.35,36
In the past few years, the oblique-incidence reflectivity difference (OI-RD) technique, a particular and the most sensitive form of optical elliposometry, has been developed as an emerging label-free biosensor.34,37 When the light has an oblique angle of incidence and is reflected from a surface, the reflection coefficients of its s- and p-polarized components will change disproportionately to the physical and chemical properties of this surface, such as thickness, mass density, and optical relative permittivity. By polarization-modulated nulling, OI-RD directly measures the difference in reflection coefficient changes (both magnitude and phase) between p- and s-polarized components of the laser beam.
There are two key points commonly used to evaluate the performance of a label-free optical biosensor: sensitivity and throughput. In terms of surface mass density, the detection sensitivities are about 0.78 pg protein/mm2 for SPR, 7.36 pg protein/mm2 for RM, and 1.50 pg protein/mm2 for RIFS. 38 In SPR-based techniques, gold or magnetic nanoparticles are sometimes conjugated to probes to decrease the detection limit. For example, Mousavi et al. reported using functionalized magnetic nanoparticles (MNPs) for signal enhancement in conjunction with SPR on gold nanoslits. 39 It was claimed that this method was capable of measuring down to 30 fM of the probe molecule in a 7-µl sample (corresponding to 1.26 × 105 molecules).
On the other hand, to increase the efficiency of parallel detection, throughput is a main consideration in designing label-free biosensors. To maximize the number of simultaneous measurements, it is crucial to integrate a high-throughput platform into the sampling and/or the detecting system. Microarray is an optimal choice because it can accommodate hundreds to tens of thousands of biomolecules for reactions with one desired probe. A microarray is composed of submillimeter-sized spots immobilized on solid substrates, which is compatible with surface-based sensing techniques such as SPR and OI-RD. In this review, we first introduce the fabrication and application of microarrays. Then, principles and recent developments in combining microarrays with SPR imaging (SPRi) and OI-RD microscopes for high-throughput and label-free biosensing are presented. This information provides a broad and deep horizon in developing the next-generation biosensors.
Microarrays
Introduction
A microarray consists of surface-immobilized, regularly arranged biomolecules such as DNAs, proteins, small molecules, carbohydrates, and even cells or tissues. They are spotted, imprinted, or directly synthesized on solid supports such as glasses, silicon wafers, and other functionalized substrates. In 1991, Fodor reported the first DNA microarrays fabricated on glass chips by combining the photolithographic method with a directed synthesis of oligonucleotides. 40 Two years later, he co-started Affymetrix (Santa Clara, CA), which released the first commercial DNA microarray, GeneChip. Concurrently, a group at Stanford University developed a robot for mechanically printing and immobilizing myriad PCR-derived cDNA on glass slides. 41 DNA microarrays have since been used widely for high-throughput analysis of nucleic acids at the transcriptional level. In 2000, MacBeath and Schreiber fabricated the first protein microarray with more than 10,000 spots of protein G immobilized on aldehyde-functionalized glass slides. 42 One year later, Zhu and colleagues reported the first “proteome-on-a-chip” by cloning, overexpressing, purifying, and arraying 5800 yeast open reading frames on a functionalized glass slide. 3 Successes in protein microarrays have enabled characterization of human proteins in the postgenomic era. Advances in arraying hardware, surface immobilizing chemistry, detection methods, and data acquisition and analysis have made microarrays a versatile platform for high-throughput screening and characterization.
Microarray Fabrication
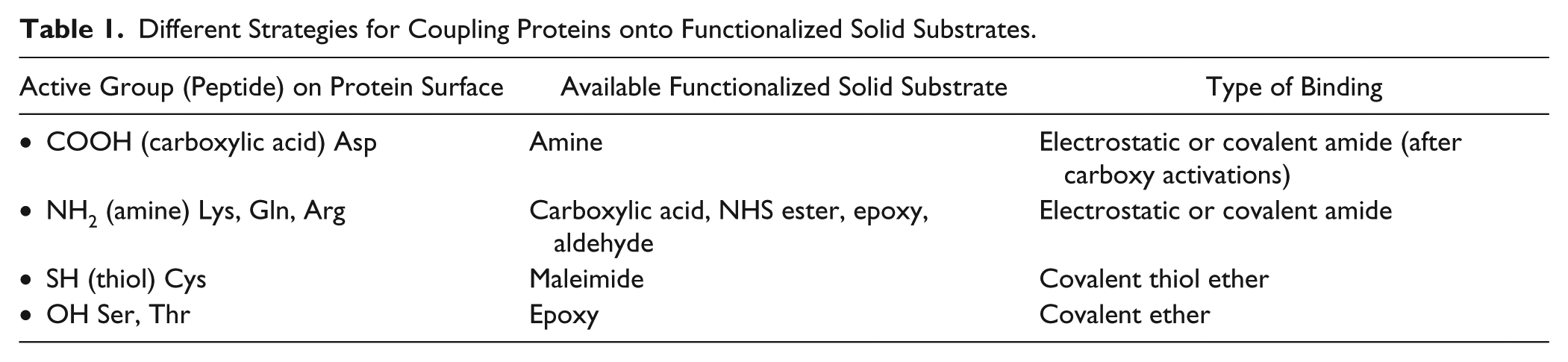
Surface chemistry plays a crucial role in microarray fabrication. Solid substrates coated with specific chemical groups are commercially available for capturing various types of targets.43-45 For example, Table 1 lists different strategies for coupling proteins onto functionalized solid substrates. A robotic arrayer accommodating target-loaded microplates and functionalized substrates is used for programmable and automated microarray fabrication. In fabrication and processing, issues such as spot heterogeneity, nonspecific interactions, cross-talks, and denaturation of biomolecules could occur. 46 Spot-to-spot variation in location, size, and target density sometimes makes it difficult to quantify the measurement. This is mostly due to liquid transfer and uneven surface chemical or morphological variation of a glass slide. Nonspecific interactions between analytes and unprinted yet functionalized surfaces reduce the detection sensitivity and specificity. Suitable blocking steps can avoid this phenomenon.47,48 Cross-talks between separated spots, mainly due to insufficient washing steps during printing, give false readouts when one is looking for possible hits. In surface-based detection techniques using microarrays, preserving the intrinsic properties and functions of immobilized targets is important. This is particularly crucial in the case of protein targets, in which functional groups on their surfaces may be blocked after immobilization. To overcome this deficiency, linkers are typically inserted between targets and the solid surface so that the key chemical properties of the targets are kept more or less intact.49–51
Different Strategies for Coupling Proteins onto Functionalized Solid Substrates.

Types of Microarrays
The precursor of microarrays was the use of immunoassays aiming at the analysis of protein mixtures deriving from recombinant DNA technologies. 52 The access to genome sequencing stimulated the need for multigene screening and emerging technologies in DNA microarrays. To elucidate gene function in posttranslational modifications, protein microarrays composed of membrane proteins, antibodies, or enzymes are now widely used for many different purposes. Biologists take advantage of these state-of-the-art technologies and use them in all kinds of applications, such as screening for protein ligands, drug discovery, and even better understanding in glycome and metabolome.
DNA microarrays can be composed of short oligonucleotides (10~20 nucleotides), long oligonucleotides (50~100 nucleotides), or polymerase chain reaction (PCR)-amplified complementary DNA (cDNA; 100~3000 base pairs). Short oligonucleotide targets (fewer than 20 bases) can be in situ synthesized on a solid support, whereas longer oligonucleotide and cDNA targets need to be spotted directly onto glass substrates. DNA microarrays can be applied in binding assays to screen against proteins, cells, or other oligonucleotides with different sequences for desired purposes and applications.
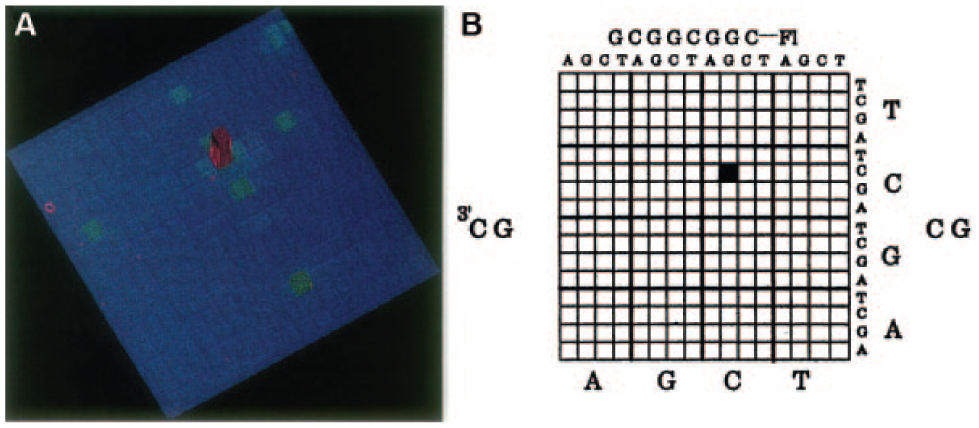
The first DNA microarray came out in 1991, when Fodor and colleagues fabricated an array of dinucleotides using a photolithographic light-directed synthesis method. 40 Soon after this, the same group exploited an identical strategy to construct an array of 256 different octanucleotides mainly for fast and high-throughput analysis of complementary DNA sequences. 53 Figure 1 shows the hybridization to a matrix of this 256-octanucleotide array. 53 In 1995, Brown et al. developed an automatic robot for mechanically printing and immobilizing myriad PCR-derived cDNAs on glass slides. 41 In the following decade, DNA microarrays, some incorporating more than tens of thousands of different oligonucleotides, have been demonstrated as powerful tools for gene expression profiling,54–60 drug validation,61–64 and DNA sequencing.65–69 In a gene-expression-profiling experiment, the expression levels of thousands of genes are simultaneously monitored to study the effects of certain treatments, diseases, and developmental stages on gene expression.
(
The human genome consists of around 25,000 genes, which are responsible for producing as many unmodified proteins. Posttranscriptional modification of these genes and of the proteins from the original genes expands the number of proteins far beyond 25,000. Therefore, it is necessary to directly array proteins to study their interactions with other molecules, such as small molecules, peptides, DNA fragments, and carbohydrates.
One major challenge in fabricating protein microarrays is to immobilize a diverse range of proteins while maintaining their properties. Proteins can be hydrophilic or hydrophobic, can be negatively or positively charged, and are more prone to changes in buffers, temperature, and humidity than nucleic acids. Using a high-precision pin-type arrayer designed for DNA microarrays, MacBeath and Schreiber successfully arrayed protein targets on aldehyde-functionalized glass slides. 42 They printed more than 10,000 spots of protein G onto half of a slide (1600 spots per square centimeter) and were able to use this microarray for studying ligand-binding reactions. Zhu and Snyder subsequently fabricated the first “proteome microarray” by immobilizing 5800 yeast proteins from the entire yeast genome. 3 The proteins were tagged with gluthathione-S-transferase-polyhistidine (GST-[His]6) and printed on aldehyde-functionalized or nickel-coated slides. This microarray was used to screen for phospholipid-interacting as well as calmodulin-binding proteins, and it has since been commercialized (Yeast ProtoArray, Invitrogen, Carlsbad, CA) for a variety of applications. In 2002, the first commercially available functional protein microarray was released by Sense Proteomic (Maidenhead, UK). Since then, antibodies, 70 membrane proteins, 71 enzymes, 72 and many other different proteins can now be directly immobilized on solid substrates through various chemical or physical methods.73,74 Different types of protein microarrays and their applications are illustrated in Figure 2 . 43
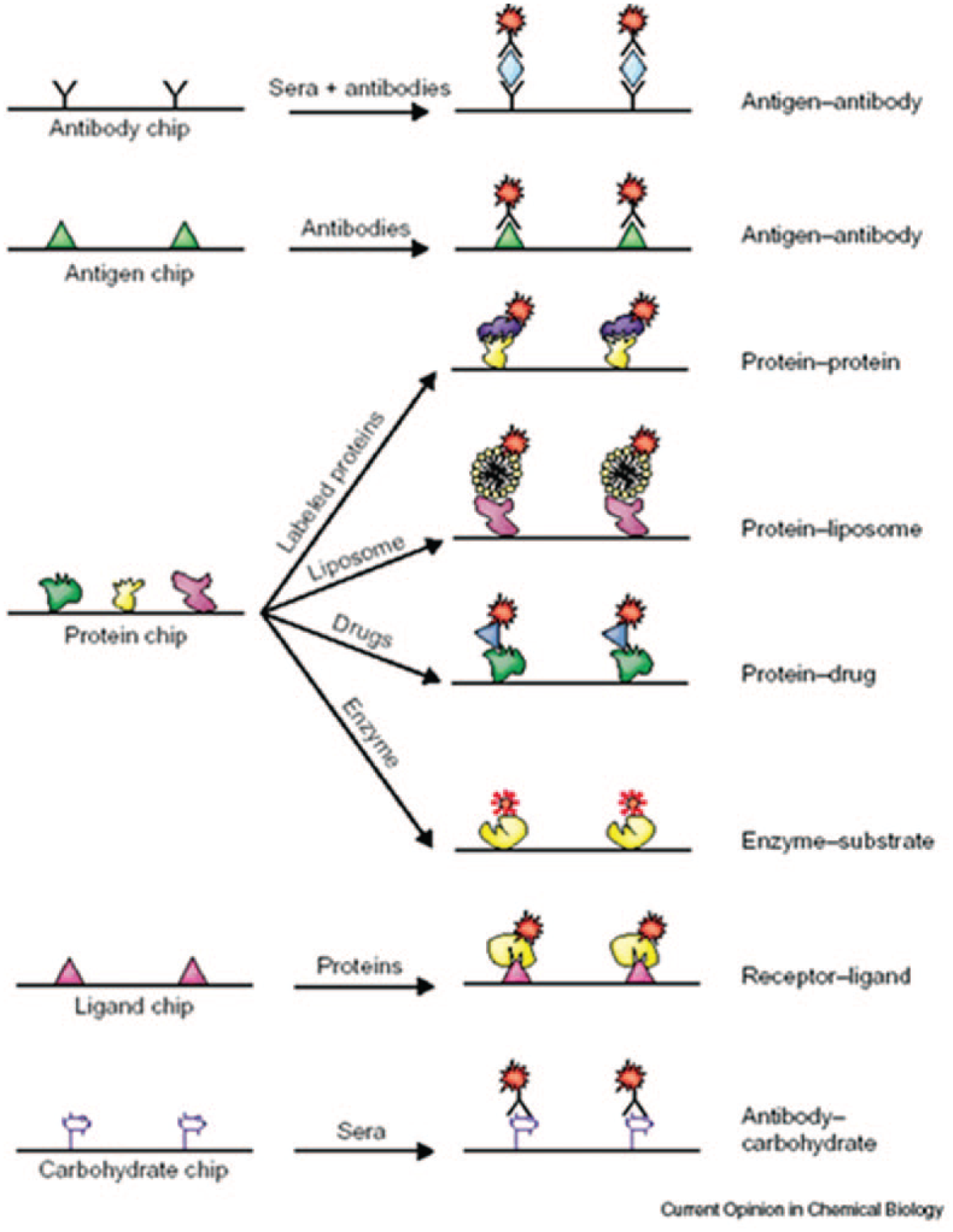
Different types of protein microarrays and their applications. Reprinted from Current Opinion in Chemical Biology Vol. 7, pp. 55, Protein chip technology. 43
Peptides are functionally stable, and they are capable of retaining their activities after immobilization on substrates. Similar to DNA immobilization, peptide targets can be either in-situ synthesized step by step on a solid or membrane support, or directly printed onto glass slides using a robotic arrayer. In the latter case, presynthetic peptides are easily functionalized to accommodate any necessary chemistry for immobilization. Peptide microarrays can serve as a facile and robust platform for protein studies in a parallel and high-throughput manner.
In the early 1980s, Geysen and colleagues described a procedure for rapid synthesis of hundreds of peptides on solid supports, and they subsequently used them to map for B-cell epitopes. 75 Several years later, Frank developed the SPOT synthesis technique, which allows simultaneous, parallel syntheses of multiple peptides at distinct positions on a membrane support. 76 In 1991, Fodor et al. fabricated a peptide microarray of 1024 peptides on a 1.6 cm2 glass substrate using the light-directed, spatially addressable parallel chemical synthesis method. 40 As shown in Figure 3 , Lam et al. developed the one-bead one-compound (OBOC) combinatorial peptide library method for synthesizing a spatially separable, but not addressable, peptide array. 77 This array was screened by an enzyme-linked calorimetric method, and then desired beads were picked and identified by peptide microsequencing. Peptide microarrays have been widely used to study protein recognition and functions. Takahashi et al. constructed a novel protein detection system using a designed peptide library with fluorescent labels, and they found that various proteins with recognition patterns corresponding to “protein fingerprints” could be detected using this immobilized peptide library. 78 Usui et al. reported designing peptide arrays with designed secondary structures (e.g., loop, alpha helix, or beta strand) for protein characterization using fluorescent fingerprint patterns. 79 Recently, to characterize the substrate specificity of an enzyme and look for its potential inhibitors, peptide microarrays have been applied to probe enzymatic activity in vitro. Shigaki et al. developed a peptide microarray, by chemoselectively immobilizing all substrate peptides on amino-coated glass slides, to detect protein kinase activity in cell lysate. 80 Other groups also reported using peptide microarrays for kinase functionality and inhibition study.81,82 Moreover, peptides binding to cell surfaces are useful in studying cell–cell communications and targeting for cancer cells. Falsey and colleagues demonstrated that peptide microarrays can be used to determine the binding specificity of peptides against different cell lines, as well as the functional cell signaling of attached cells using immunofluorescence techniques. 83
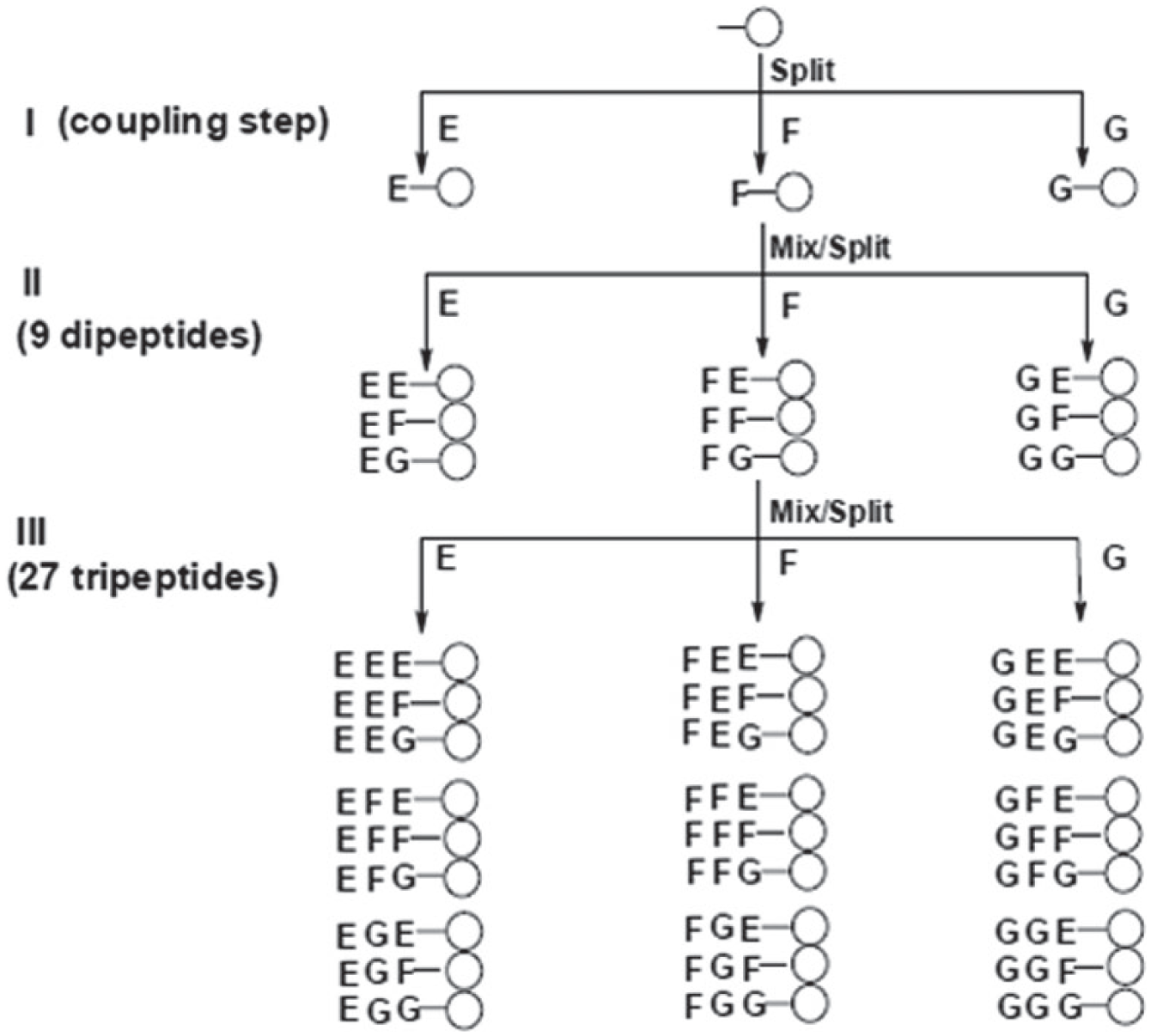
An example of creating a tripeptide library using the one-bead one-compound (OBOC) method. In step I, a pool of resin beads is distributed (split) into three separate reaction vessels, each with a single amino acid (E, F, and G). After the coupling reactions are complete, all beads in a single vessel are distributed (split) into another set of separate reaction vessels, each with a single amino acid (step II). This cycle is repeated again to extend the peptide chain to three amino acids (step III). Reproduced from Nature, Vol. 354, pp. 82, A new type of synthetic peptide library for identifying ligand-binding activity. 77
Small chemical molecules are known to be important regulators of protein activities. They provide effective handles in selective and specific perturbations of protein functions. This property makes small molecules viable drug candidates, and therefore characterizing protein–small molecule interactions is basically the first step in drug discovery. Small molecule microarrays (SMMs) composed of small molecules such as drugs, natural products, peptides, and combinatorial libraries provide a high-throughput platform for studying protein functions and finding potential protein ligands.
One of the major challenges in SMM application, however, is to efficiently immobilize a large variety of small molecule compounds with diverse structures on a singly or even multiply functionalized solid support while maintain the innate binding properties of the molecules. SMMs were first reported by MacBeath et al. in 1999. 84 They distributed one synthesized bead into each well of a microtiter plate, released compounds from the beads, and dissolved them in a small volume of suitable solvents. Then, a high-precision robot was used to array these compounds on maleimide-functionalized glass slides. This microarray was screened against cyanine 3 (Cy3)- or cyanine 5 (Cy5)-labeled proteins, and reaction constants of some protein–small molecule interactions were estimated. A few years later, the same group introduced the diversity-oriented synthesis (DOS) approach to generate a library of 3780 encoded small molecules, 85 in which compounds were synthesized using the OBOC technique originally designed for peptide libraries. 86 Since then, many efforts have been made in the fabrication of SMMs, including use of ketone-modified macromolecular scaffolds, 87 immobilization of hydrazide-linked compounds on epoxy-coated slides, 88 nonspecific isocyanate-mediated surface immobilization,89,90 and photoactivated capture on gold surfaces for SPR imaging. 91 By screening SMMs against proteins or other biomolecules, researchers were able to identify small molecule ligands and potential drug candidates.92–98
Carbohydrates play important roles in a wide range of biological events, particularly those involving cell surfaces, through interactions with proteins. These carbohydrate-binding proteins, the lectins, have very high specificity to their binding moieties. 99 Specific interactions between lectins and carbohydrates are essential in many biological processes, such as regulation of cell adhesion,100,101 metastasis, 102 immune response,103,104 and pathogen–host infections. 105 Carbohydrate microarrays composed of many distinct glycans immobilized on solid substrates provide ways to simultaneously detect multiple bindings of proteins, viruses, and cells to every glycan under identical conditions. There are a number of methods to efficiently immobilize various carbohydrates, including mono-, di-, oligo-, and polysaccharides, on functionalized solid supports while maintaining their innate functionalities. 106 Unmodified carbohydrates can be directly immobilized either covalently on hydrazide-coated glass slides 107 or noncovalently on nitrocellulose-coated glass slides. 108 Amine- and maleimide-derivatized carbohydrates can be covalently attached onto N-hydroxysuccinimide (NHS)- and thiol-activated glass substrates, respectively.109,110 Moreover, biotinylated carbohydrates are immobilized on NeutrAvidin-coated surfaces, providing a stronger and more orientation-specific attachment. 111
To date, many research groups have reported using carbohydrate microarrays for protein characterization and profiling.112,113 Blixt et al. constructed a carbohydrate microarray of 200 natural and synthesized glycan sequences representing the structures of major glycoproteins and glysolipids. They used this microarray to analyze specific glycan-binding proteins, including C-type lectins, siglecs, galectins, anticarbohydrate antibodies, lectins from plants and microbes, and intact viruses.114,115 Carbohydrate microarrays have also been used to profile the specificity of influenza virus receptors116,117 and human serum antibodies118,119 for the purpose of developing new carbohydrate-based vaccines and drugs.
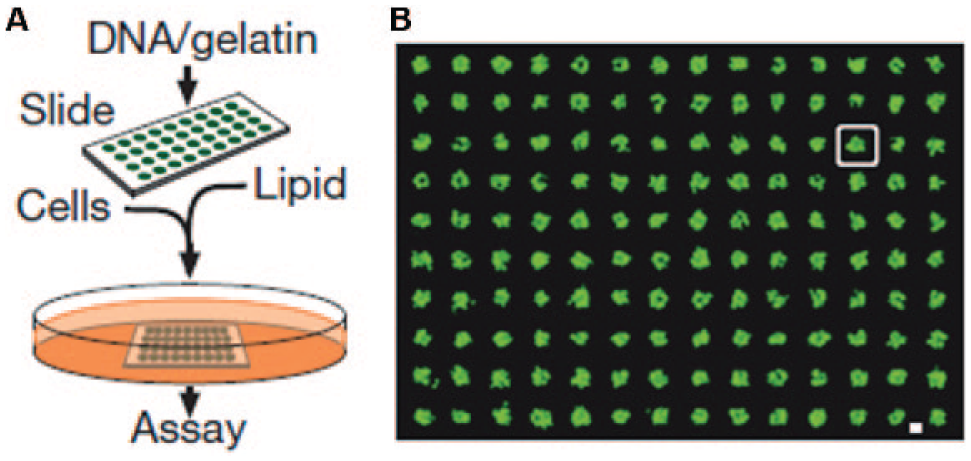
Living organisms are composed of all kinds of cells with various capabilities. These cells perform specific functions, interact with one another, and respond to environmental stimuli. Cell microarrays provide a platform for potential in vivo screening and characterization of cellular interactions in a high-throughput manner. In 2001, Ziauddin et al. developed the reverse-transfected cell microarrays, in which mammalian cells were cultured on a glass slide with different DNA–gelatin mixtures printed in defined locations. 120 As shown in Figure 4 , cells growing above DNA spots take up the nucleotides underneath and become transfected. Ziauddin et al. applied a cell microarray expressed from 192 different cDNAs in screening for proteins involved in tyrosine kinase signaling. These reverse-transfected cell microarrays can also be used to identify potential drug candidates and evaluate their binding specificities. 121 Schwenk et al. reported using a robotic arrayer to print cells from different cell lines onto poly-L-lysine-coated slides via electrostatic interactions for rapid screening of cell-surface-specific antibodies. 122 Recently, cell microarrays were used in drug discovery, 123 stem cell fate studies, 124 and evaluation of DNA double-strand breaks and DNA repair inhibitors. 125 In addition to cells, tissue specimens can be assembled as tissue microarrays, providing a high-throughput platform for studying anatomic pathology such as multitumor analysis and cancer progression analysis.126–130
(
Use of Microarrays in SPRi Microscopy
Introduction
Surface plasmon resonance (SPR), as it is commonly known in the field of life sciences, refers to the consequence of exciting a surface-bound electromagnetic wave at the interface between a metal and a transparent material. A SPR is excited optically with an electromagnetic wave (angular frequency, or ω), and it has a wave number along the interface as
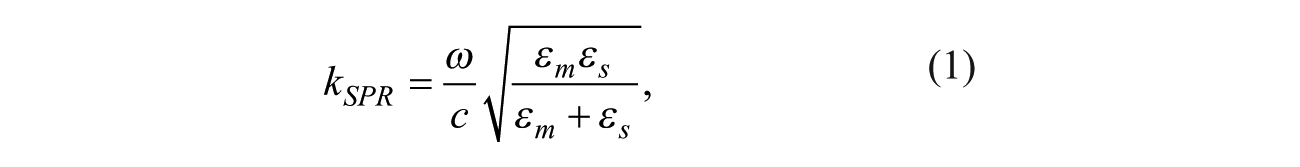
where εm is the optical relative permittivity of the metal and εs is the optical relative permittivity of the transparent material. εm is a complex quantity with
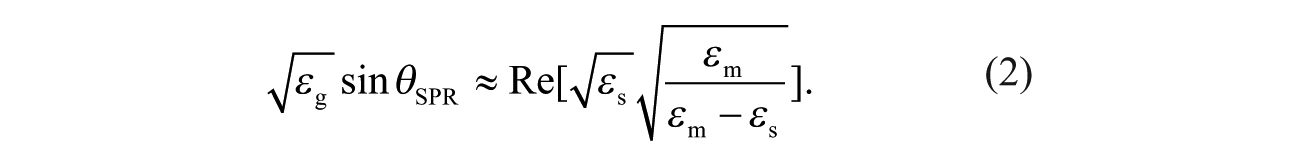
As a result of
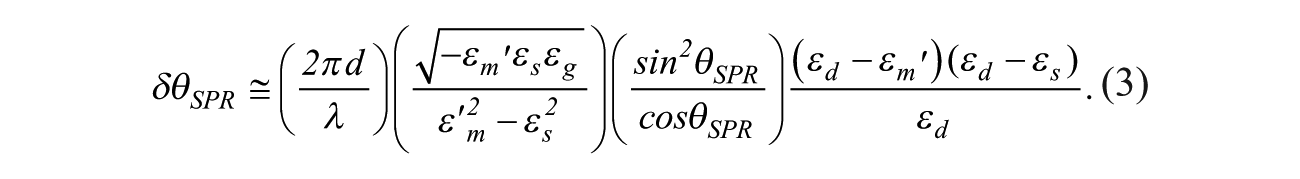
By measuring δθSPR, one can obtain the physical properties (d and εd) of the ultrathin film or the biomolecular layer. For example, for the gold–water interface where εs = 1.77 and
In the SPR spectroscopy mentioned in this article, usually the incidence angle is scanned at a fixed wavelength (or the wavelength is scanned at a fixed incidence angle) to locate the resonance angle. Changes in the resonance angle throughout time are displayed in a graph called a sensorgram. The real-time sensorgram is a direct representation of the interactions between biomolecules on the sensor surface. In contrast, the SPR imaging, or the SPR microscopy, measures the reflection coefficient of a monochromatic incident light at a fixed angle and a fixed wavelength. An expanded and collimated light is shone on the entire area to be monitored, and a sensor array (typically, a closed-circuit display (CCD) camera) is used to “image” the reflection coefficient at different locations. When a biomolecular binding occurs, the intensity of the reflected light in the corresponding CCD pixel changes proportionally. In addition to the temporal resolution given by the sensorgram, SPRi also provides a spatial resolution by simultaneously measuring the reflected signals from numerous spots. Therefore, combining SPRi with microarrays allows for high-throughput, parallel detections of biomolecular interactions.
SPRi Instrumentation
In most cases, SPRi microscopes use the Kretschmann configuration, in which a prism is used in the total internal reflection (TIR) geometry. As shown in Figure 5A , the incident light is reflected from the backside of the substrate with immobilized biomolecules. The intensities of the reflections from spots with reactions and spots without reactions are different, and they are imaged by a sensor array ( Fig. 5B ). Simultaneously, these signals are monitored throughout time to acquire the sensorgram ( Fig. 5C ). A number of SPRi instruments have been designed and constructed based on this Kretschmann configuration.137–142 In the SPRi microscope built by Shumaker-Parry and Campbell, 143 a He–Ne laser beam was p-polarized, expanded, and collimated before passing through a hemi-cylindrical prism and illuminating on the gold-coated glass slide. A CCD camera was used to collect the focused reflected light and create an image. For real-time and in situ measurements, a fluidic system, consisting of a flow chamber (volume ~ 15 µL), a syringe pump, two switching valves, and laminar flow tubes, was integrated and computer controlled. This microscope was reported to have a detection limit of 0.072% change in absolute reflection coefficient when simultaneously measuring all 200 μm × 200 µm areas with the 24 mm2 beam size and 1-s time averaging. This corresponded to a refractive index of 1.8×10−5 and a surface mass density of 1.2 ng protein/cm2. A representative SPR image and corresponding real-time curves are shown in Figure 6 . 143 In another work by Chen et al., 144 the beam from a wavelength-tunable monochromator was collimated, polarized, and shone on a prism-coupled SPRi chip. A cooled CCD camera was used to collect the reflected light, and the “smart image-processing software” with filter processing was used to improve the signal-to-noise ratio and correct the distribution of images. In this SPRi microscope, the detection limit in terms of probe concentration was claimed to be as low as 10 fg/µL.
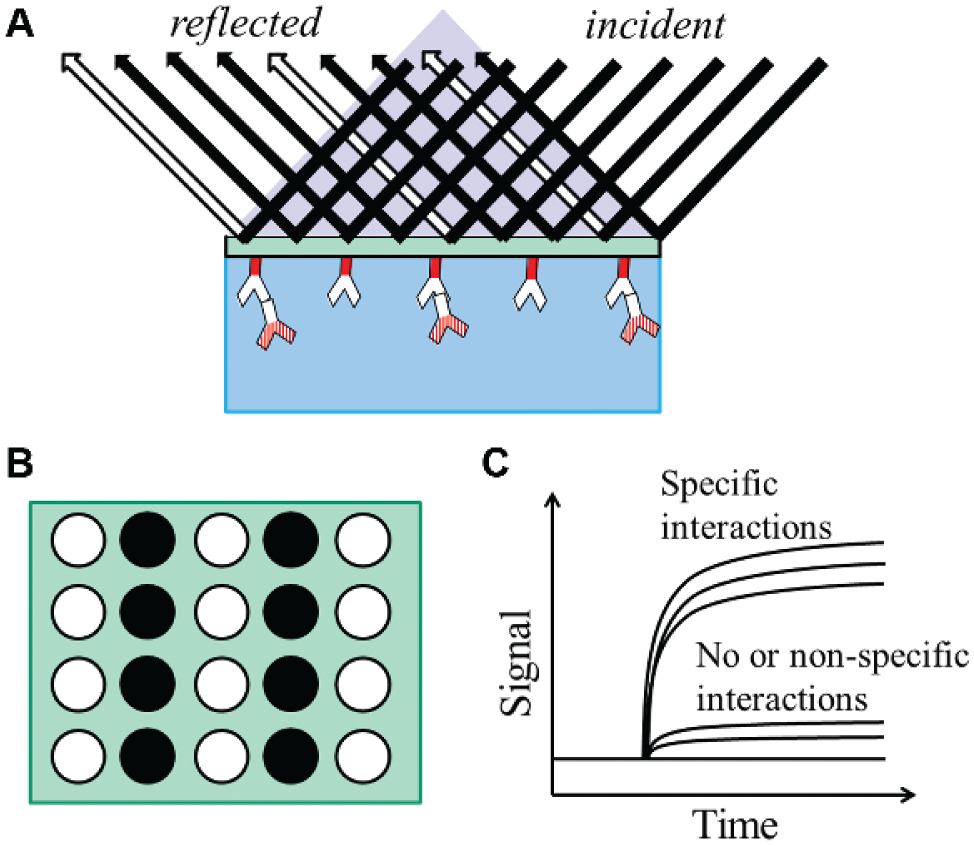
Surface plasmon resonance imaging (SPRi) microscopy. (
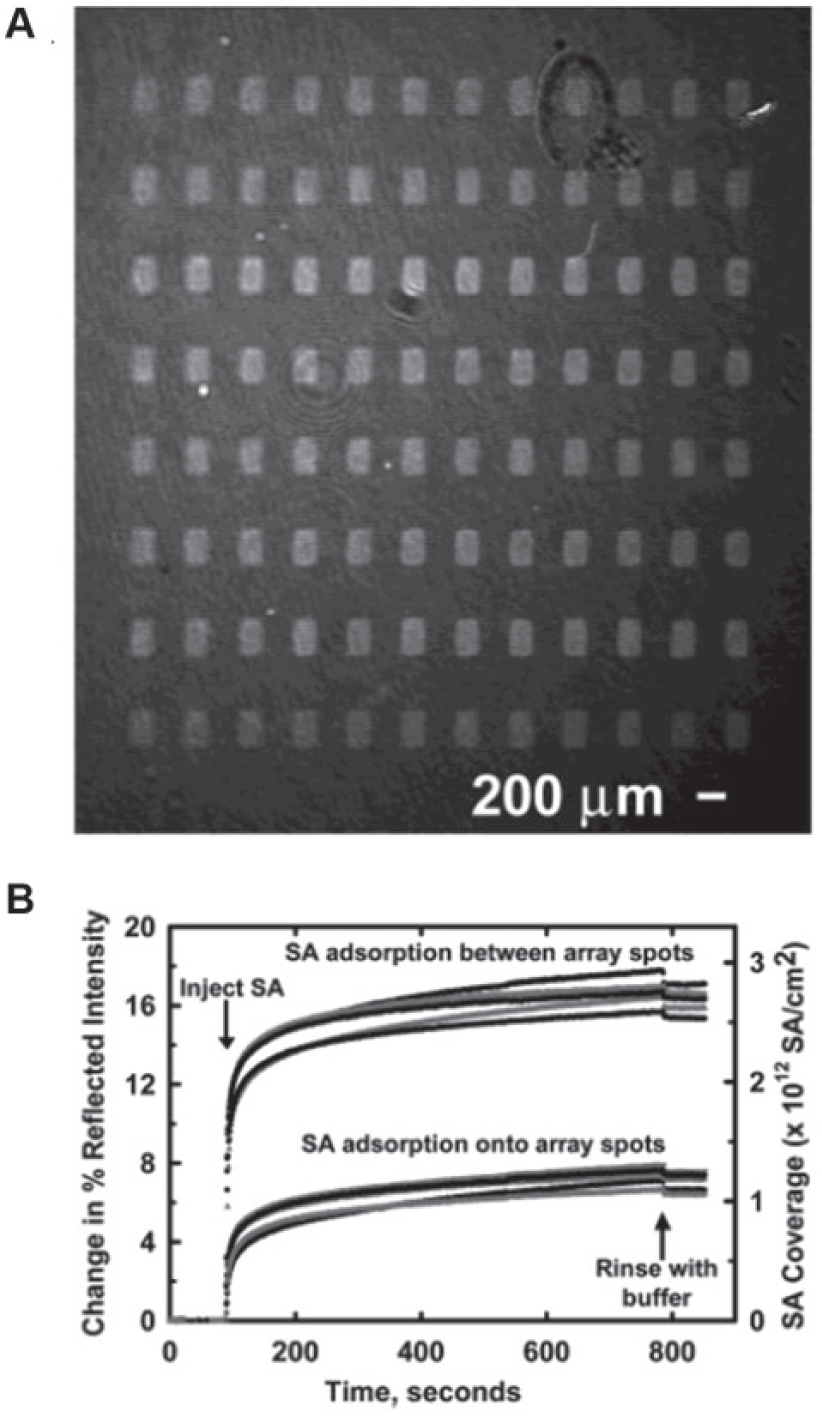
(
An alternative scheme is that of the grating-coupled SPR system, in which the metal surface is periodically corrugated. The light diffracted by the grating has different orders, with at least one inducing SPR. Without the need of prisms, this setup is cheaper and more usable. Lindquist et al. reported using nanohole arrays in thin gold film to attain submicron resolution in SPRi pixels. 145 In this setup, each pixel of a 3×3 nanohole array was surrounded by plasmonic Bragg mirrors to block interference between adjacent sensing pixels. Yao et al. described a new plasmonic crystal optics that provided excellent analytical sensitivity at visible wavelengths. 146 Cylindrical nanowells were fabricated on the surface of UV-curable polyurethane, which provided a large area with spatially uniform arrays for SPRi applications. This system was capable of detecting biomolecular interactions with a probe mass difference of as low as 25 dalton. Linman et al. fabricated gold-coated glass array substrates for SPRi analysis with very good image contrast and sensitivity. 147 A simulation showed that the etched structure enhanced the SPR intensity by threefold compared to standard planar gold substrates. SPR images of an etched glass array (left column) and gold island array (right column) as water fills the flow cell are shown in Figure 7 . 147 There are several commercially available SPRi microscopes reported to provide simultaneous measurements of more than 1300 binding reactions with a detection limit of ~0.3 ng protein/cm2 and a time resolution of 1 s.141,148,149 The SPRimger II from GWC Technologies (Madison, WI, USA), the IBIS-MX96 from IBIS Technologies (Hengelo, The Netherlands), and the OpenPlex from HORIBA Scientific (Kyoto, Japan) use the Kretschmann configuration. The Biacore Flexchip from GE Healthcare (Uppsala, Swedem) uses the grating-coupled SPR. The IBIS-MX96 operates in a scanning-angle mode, in which the SPR angle shifts are directly monitored in response to probe-target bindings. This provides the instrument with a 10-fold larger dynamic range compared to other systems. 142
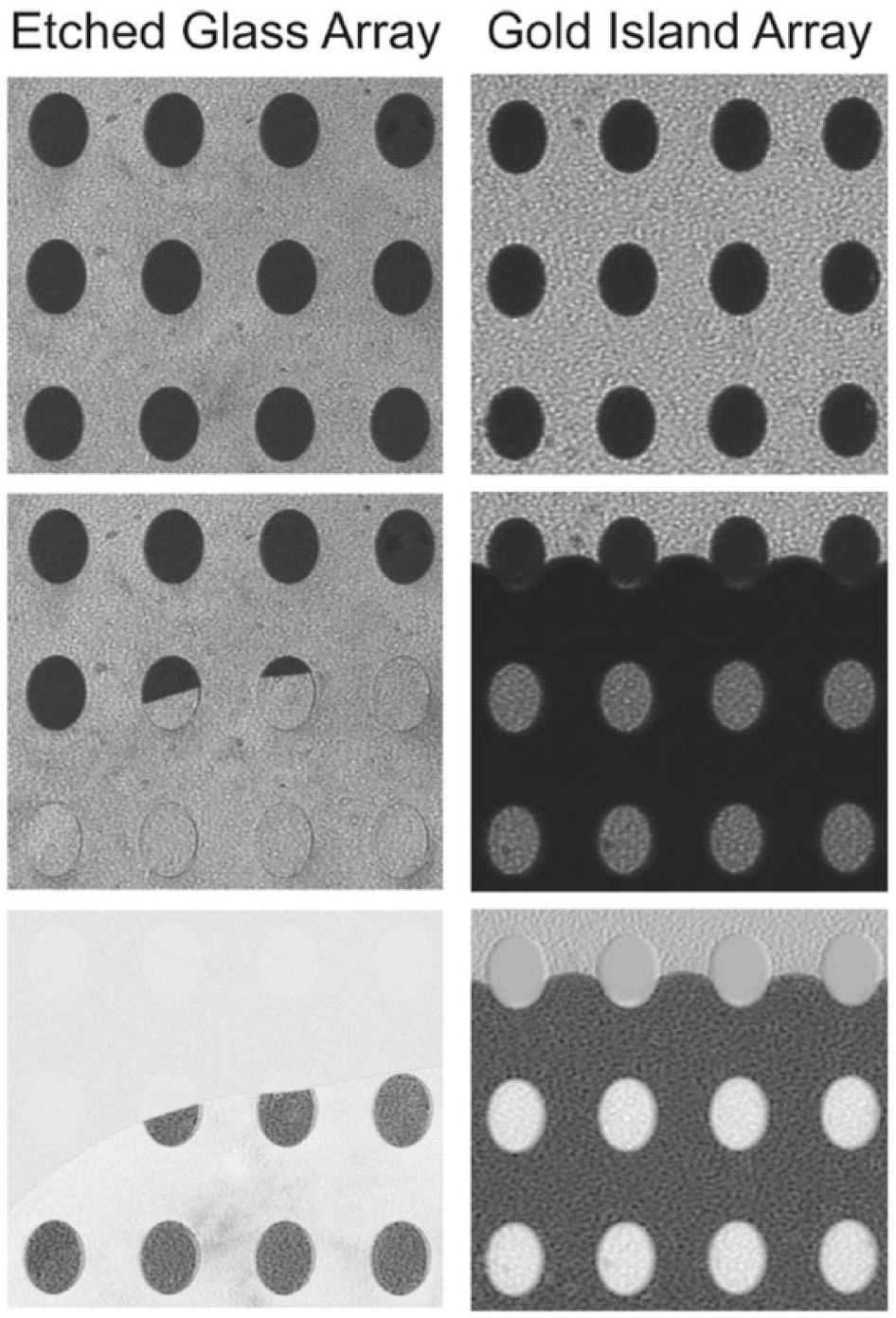

Surface plasmon resonance (SPR) images of an etched glass array (left column) and gold island array (right column) as water fills the flow cell. For each column, from top to bottom: initial SPR image taken in air, water covering the substrate surface, and the resulting difference image. Reprinted from Analytical Chemistry, Vol. 83, No. 15, pp. 5936, Etched Glass Microarrays with Differential Resonance for Enhanced Contrast and Sensitivity of Surface Plasmon Resonance Imaging Analysis. 147 Copyright © 2011 American Chemical Society.
Examples
When SPRi microscopes were first developed, DNA microarrays were combined with the systems for simultaneously detecting biomolecular interactions. By immobilizing DNA molecules on SPR-active surfaces, Corn’s group studied various DNA–DNA, RNA–DNA, and protein–DNA interactions.150–161 For example, they reported a 4×3 SPRi microarray, in which each element had an individual microfluidic channel. The microarray consisted of single-stranded DNAs and RNAs fabricated by chemical and/or enzymatic reactions in these microchannels, and it was used to detect DNAs and proteins at the femtomole scale. 153 Campbell’s group also worked on combining DNA microarrays with a SPRi microscope.162,163 A 10×12 double-stranded DNA (dsDNA) microarray was fabricated on planer gold surfaces using a robotic microspotting system. Using the prism-based SPR microscopy, they successfully detected the bindings between yeast transcription factor Gal4 and 120 dsDNA molecules with a sensitivity of around 0.5 pg of bound proteins (i.e., 2×107 molecules) in each array spot of 200 µm diameter. 162
Corn’s group also reported a simple method for fabricating high-density, multiplexed antibody microarrays for detecting protein biomarkers with SPRi microscopy. 164 Antibodies were immobilized on alkanethiol-modified gold substrates via a carbonyldiimidazole (CDI) coupling reaction, and then an antibody microarray with spot sizes of 200~750 µm was created, again using the CDI chemistry. A SPRi microscope could detect two biomarkers (molecular weights, 11.8 kDa and 13.4 kDa) binding specifically to this antibody array. In addition, protein microarrays could be fabricated from dsDNA microarrays by cell-free enzymatic synthesis in a microfluidic format, and SPRi microscopy was used to verify the formation of these arrays. 151 Campbell et al. demonstrated combining a SPRi microscope with 96-spot protein microarrays for high-throughput detection of antibody–antigen reactions. 141 They have shown that this system was capable of monitoring 300 protein spots simultaneously with a time resolution of 1 s and a detection limit of ~0.4 ng protein/cm2. In 2006, a commercial SPRi microscope, the Lumera Corporation’s Proteomic Processor (Washington, USA), was developed and reported to offer parallel, high-throughput analysis of biomolecular interactions on high-density microarrays (>1000 spots in 1.0 cm2). 165 As a proof of principle, monoclonal antibodies (MAbs) to an oncofetal glycoprotein [carcinoembryonic antigen (CEA)] were printed in triplicate spots at a density of 1140 spots/1.0 cm2. CEA antigens were brought into contact with the MAbs microarray to obtain both the binding specificity and the kinetic constants.
In 2003, Smith et al. first reported fabricating carbohydrate microarrays on gold films to study carbohydrate–protein interactions with SPRi microscopy. 166 Disulfide bonds on gold surfaces were used to capture thiol-modified carbohydrates. The equilibrium dissociation constants (KD) of concanavalin A (ConA) binding to mannose and jacalin binding to galactose were measured to be 200±50 µM and 16±5 µM, respectively. Karamanska et al. printed a number of N-biotinylated carbohydrates on neutravidin-coated gold substrates for detecting their bindings to a lectin Ricinus communis agglutinin (RCA). 167 A 40-spot microarray was imaged to obtain the affinity binding data with as little as 10~20 µg of lectin per experiment. Tyagi et al. described a photochemical strategy to generate carbohydrate microarrays on SPRi surfaces. 168 Their results showed a detection limit of 0.18~3.1 nM for various carbohydrates binding to ConA. Linman’s group also reported the fabrication and characterization of a new carbohydrate microarray with synthetic sialosides for SPRi analysis of lectin–carbohydrate interactions.113,169 Biotinylated sialosides were contact-printed on neutravidin-functionalized substrates, and Sambucus nigra agglutinin (SNA) was reacted with the 400-element microarray. A relative standard deviation (RSD) less than 5% from element to element was obtained, which assured the good quality of microarrays.
In addition to the above-mentioned examples, SPRi microscopy has also been applied to detect other types of microarrays. By combining SPRi microscopy with photo-crosslinked small molecule microarrays, Kanoh et al. studied the interactions between small molecules (estrogenic and androgenic substances) on gold surfaces and proteins (estrogen receptor alpha) in solutions. 170 Neumann et al. described the design and immobilization of fragment libraries in generating chemical microarrays for SPRi. 171 Simultaneous measurements of protein probes binding to more than 9000 fragments were performed to obtain the affinity date. They also demonstrated the successful identification of inhibitors with low molecular weights for applications in drug screening. Peterson et al. reported using SPRi contrast to measure cell–substrate interactions and mass changes at the substrate interface. 172 By monitoring changes in surface protein mass and area, SPRi allowed the quantification of cell-secreted and -deposited materials. The authors concluded that this sensitive technique is capable of bridging the gap between molecular (e.g., DNA–protein and protein–protein interactions) and cellular (e.g., cell–cell and cell–substrate interactions) measurements.
Use of Microarrays in OI-RD Microscopy
Introduction
OI-RD, a particular form of optical ellipsometry, measures the differential changes in both phase and magnitude of the reflection coefficients for p- and s-polarized components of a monochromatic light in response to small changes in physical and chemical properties of deposited thin films. With an incidence angle of θ, a monochromatic light is reflected from a substrate [optical relative permittivity (εs)]-ambient [optical relative permittivity (εamb)] interface. Let
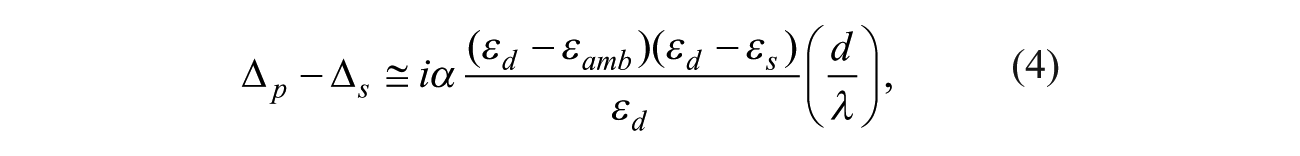
where
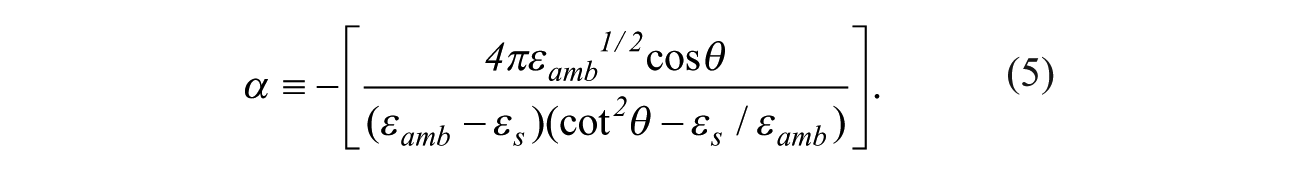
For real optical relative permittivity constants εamb, εs, and εd, only the imaginary part of
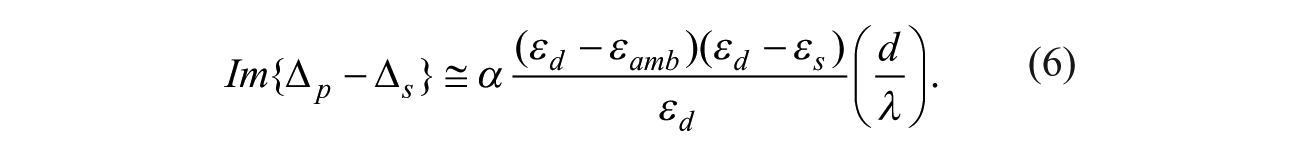
Therefore, by monitoring
As shown in Figure 8 , an OI-RD microscope works by mechanically and/or optically scanning the microarray-bearing slides in two dimensions. With a scanning pixel of around 10 µm, a microarray image is acquired to locate the spots. To obtain the kinetic curve of a particular element, the averaged signal of its two neighboring substrates is subtracted from the signal of the spot. Reaction rate constants are estimated by globally fitting a set of binding curves at different probe concentrations. After reaction, another image is taken to identify specific bindings.
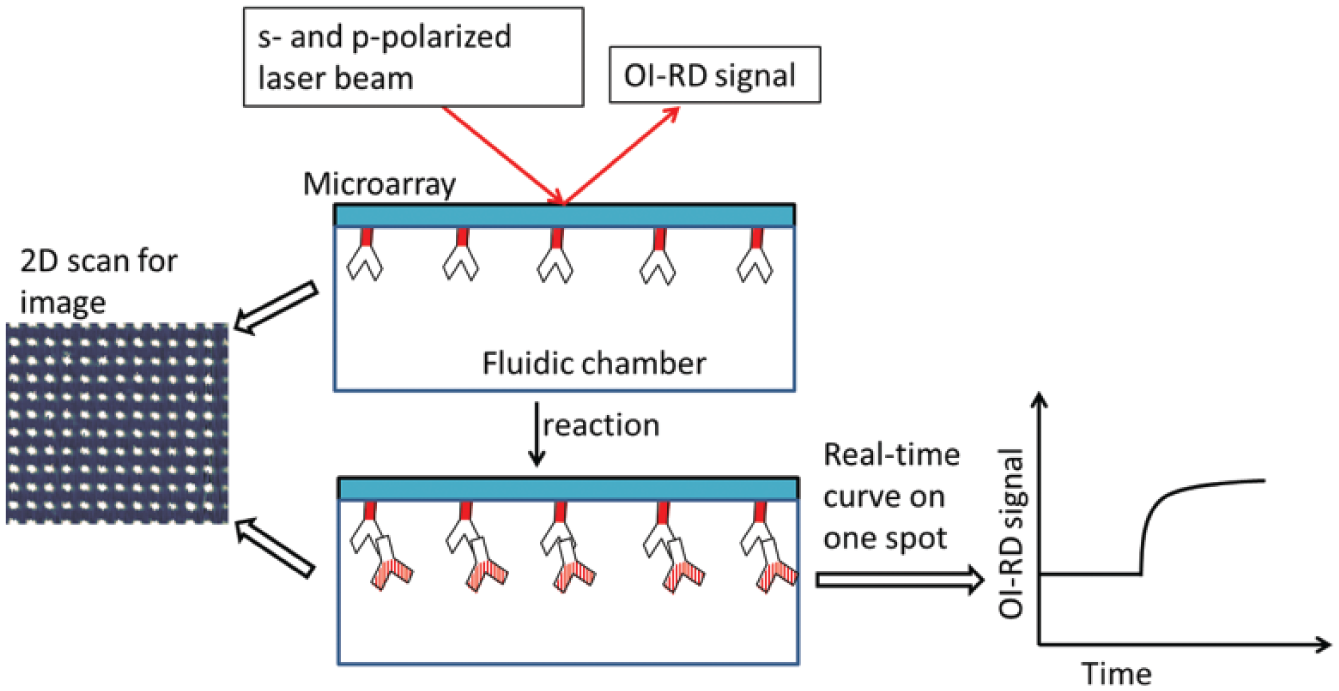
The oblique-incidence reflectivity difference (OI-RD) microscopy. Middle: The OI-RD signal is the difference between the reflection coefficients of the s- and p-polarized components of the laser beam. Left: An OI-RD image of a microarray is acquired via 2D scan. Right: The real-time binding curve of a selected microarray spot. Reprinted from Sensors and Materials, Vol. 25, No. 9, Ellipsometry-Based Biosensor for Label-Free Detection of Biomolecular Interactions in Microarray Format. 195
OI-RD Instrumentation
In OI-RD microscopy, an s-polarized laser beam passes through a photoelastic modulator (PEM) and a phase shifter (PS), and then it is focused onto the microarray surface by a lens. The PEM makes the output beam oscillate between p- and s-polarization at a modulation frequency Ω = 50 kHz. An adjustable phase
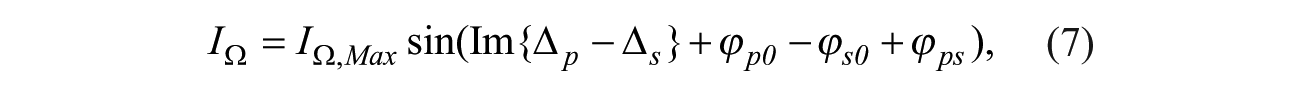
where IΩ,Max is the maximum of IΩ depending on the laser power; and
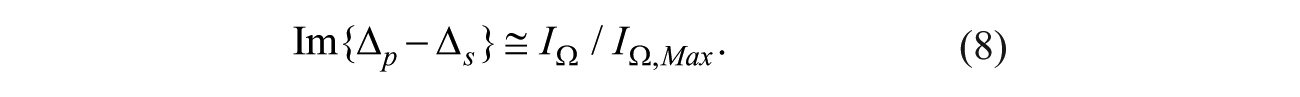
For the past 10 years, various modifications have been done to improve the sensitivity, throughput, and dynamic range of the OI-RD microscopy. To enhance to the signal-to-noise ratio, a prism was used to attain a TIR geometry, in which the laser beam was totally reflected from the biomolecule–glass interface, and almost 100% of the incident power was reflected back to the photodetector (without TIR, only 0.5% is reflected). To increase the throughput, translational stages and photodiode arrays were applied to acquire 2D images. For example, a 152-element line photodiode array detector (PDA) was built for multichannel detection. Besides, Fei et al. reported incorporating a rotating mirror into the OI-RD microscope to accelerate the scanning speed. 175 In this design, the sample-holding stage moved in the x-direction, and the y-directional scan is performed with the combination of a scanning mirror and an f-theta lens. Rotating the mirror to reflect incident light onto different locations of the f-theta lens took much less time than moving the sample stage itself. And the f-theta lens was capable of focusing these incident lights onto different locations of the microarray slide.
Examples
OI-RD microscopes were first developed for label-free detections of DNA microarrays. In 2004, Landry et al. reported fabricating oligonucleotide microarrays on poly-L-lysine-coated glass substrate.
173
After UV crosslinking, these microarrays (consisting of 8-spot 60-base oligonucleotides) were hybridized with their complementary binding partners. The results showed that the OI-RD microscope could detect specific DNA hybridizations, and after calculation, an
After a successful combination of OI-RD microscopy and DNA microarrays, this technique has been applied to detecting protein microarrays for applications in proteomics. Zhu et al. demonstrated using an OI-RD microscope to image an 800-spot bovine serum albumin (BSA) microarray within 14 min.
37
A 4 × 4 protein microarray containing BSA, rabbit immunoglobulin G (IgG) antigen, mouse IgG antigen, and human IgG antigen was reacted with goat antibody against rabbit IgG, and specific antigen–antibody bindings were observed. To illustrate that the OI-RD technique is capable of real-time measurements, kinetic curves of BSA binding to an epoxy-functionalized glass slide was monitored. Shortly afterwards, Fei et al. constructed a new OI-RD microscope by combining a scanning mirror with a sample translational stage for high-throughput detection of protein microarrays.
175
A 2760-spot BSA microarray in an area of 18.3×6.4 mm2 was imaged with a pixel dimension of 17.8×18.7 µm2. To quantitatively study protein reactions with surface-immobilized biomolecules, Landry et al. used the OI-RD technique to analyze the amount of captured proteins as a function of target printing concentrations.
176
A 180-spot protein microarray consisting of BSA, rabbit IgG, mouse IgG, and human IgG at different printing concentrations was fabricated on epoxy-coated glass slides. The microarray was reacted with goat antibody against human IgG, and the OI-RD signals (in
Using an OI-RD microscope, Fei et al. measured the binding curves of seven plant lectins, with and without fluorescent labeling, to carbohydrate microarrays containing 24 carbohydrates. 5 All these carbohydrates were biotinylated for immobilization on streptavidin-coated glass slides. From binding curve-fitted reaction rates, it was found that (1) fluorescent labeling agents changed binding profiles of lectins; and (2) the dissociation rates of these carbohydrate–lectin reactions were large and easily missed in other fluorescence-based assays. The authors also reported fabricating carbohydrate microarrays for characterization of Helicobacter pylori (H. pylori) binding to blood group antigens. 178 Lewis glycans, Lea-HAS, Leb-HAS, Lex-HAS, and Ley-HAS, were covalently conjugated to human serum albumin for immobilization on epoxy-coated substrates. The presence of additional Leb binding adhesins on H. pylori was confirmed, suggesting that the OI-RD microscopy is applicable to real-time detection of intact microbes binding to host receptors. Recently, the same group developed a variable-temperature OI-RD microscope for kinetic measurements of lectin–carbohydrate reactions from 10 °C to 60 °C within ±0.1 °C. 179 Using the above-mentioned 24-carbohydrate microarray, it was found that these reactions could be enthalpy driven, entropy driven, or both, and that water molecules played an important role in the thermodynamics of these reactions.
To efficiently immobilize small molecule compounds on functionalized solid substrates, Sun et al. explored two macromolecules, BSA and polyvinyl alcohol (PVA), as scaffolds for small molecule–protein conjugates. 49 For demonstration, biotin molecules were conjugated to BSA and amine-derivatized PVA for subsequent immobilization on epoxy-coated glass slides through reactions of amine residues on BSA and PVA with surface epoxy groups. As shown in Figure 9 , a 70-spot small molecule microarray consisting of biotin–BSA and biotin–PVA at different biotin loadings and different printing concentrations was reacted with streptavidin. Derived reaction rate constants indicated that both biotin–BSA and biotin–PVA conjugates presented two types of binding sites: The first site had a higher affinity, with KD values having upper bounds of 20~100 pM; and the second site showed a much lower affinity, with KD values ranging from a few nM to tens of nM. These results showed that both BSA and amine-derivatized PVA could serve as effective and efficient scaffolds for conjugation of small molecules and subsequent immobilization as microarrays. Fei et al. reported fabricating small molecule microarrays containing 20 small molecules known to be strong ligand candidates to intracellular proteins of Jurkat cells. 180 These compounds were synthesized using the OBOC combinatorial chemistry, and they were biotinylated for immobilization on streptavidin-coated substrates. Real-time binding curves of a Jurkat cell lysate solution to this 20-compound microarray were measured. A similar strategy based on OBOC synthesis and a novel trilayer bead-partition scheme was used to generate encoded compound libraries for fabricating small molecule microarrays; 181 9216 synthetic compounds released from beads were immobilized on epoxy-functionalized glass slides via multiple amine residues on the handles of these molecules. These microarrays were then reacted with 11 proteins from human plasma, sequentially, to screen for potential protein-binding ligands. Recently, Landry et al. printed 7961 small molecule compounds from NCI (National Cancer Institute)/DTP (Developmental Therapeutics Program) plate sets onto isocyanate-functionalized substrates. 94 These small molecule microarrays were screened against vascular endothelial growth factor (VEGF) and one of its receptors, kinase-insertion domain receptor (KDR). Twelve inhibitors to VEGF–KDR interactions were identified with IC50 (half maximal inhibitory concentration) ranging from 0.3 to 60 µM. The inhibitory abilities of these inhibitors were also double-checked in cell-based assays.
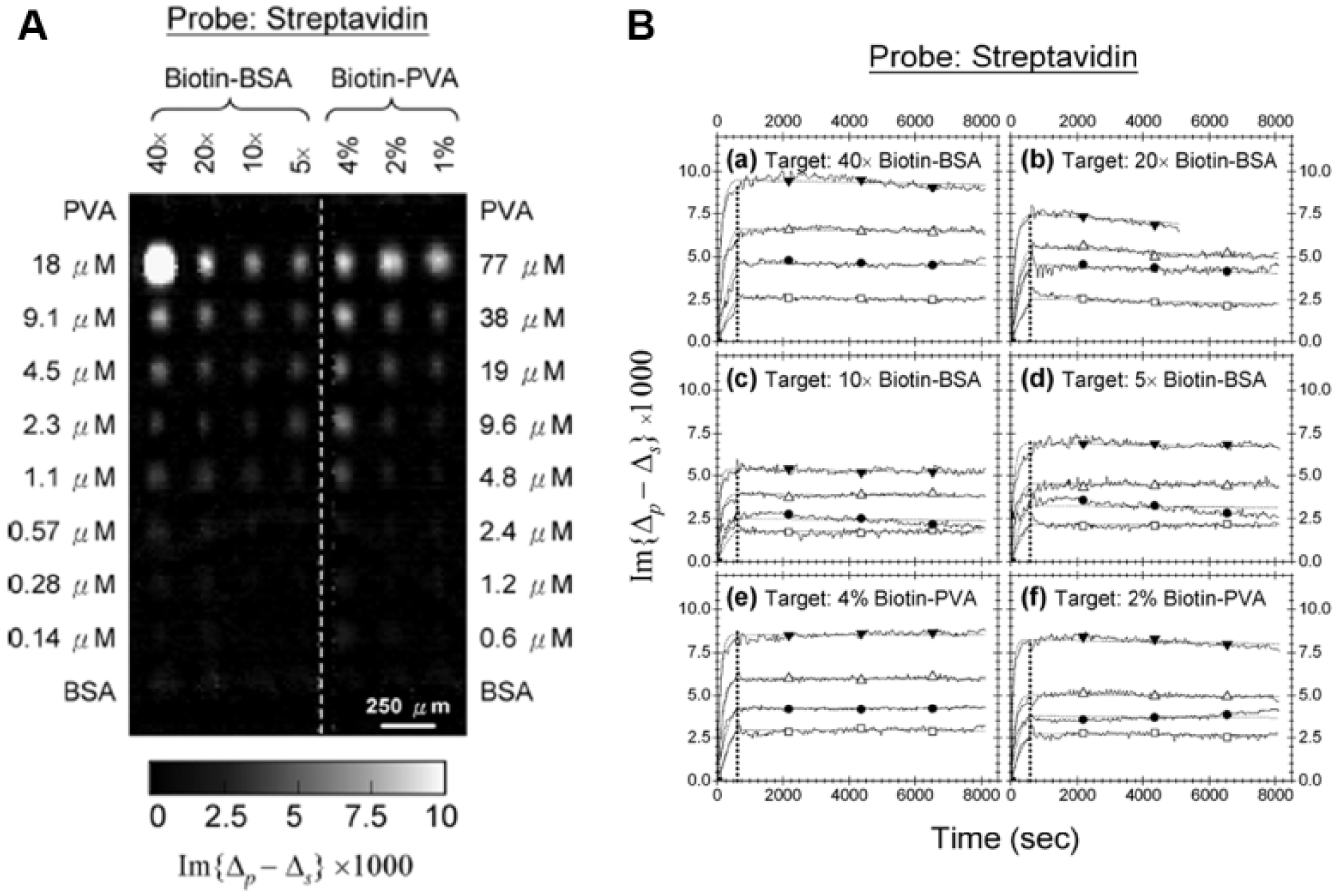
(
Lo et al. demonstrated using an OI-RD microscope for label-free detection of cell microarrays. 182 Mouse embryonic stem (mES) cells before and after differentiation, mouse-induced pluripotent stem (miPS) cells before and after differentiation, human embryonic kidney (HEK) 293 cells, and A19 fibroblasts were immobilized on aldehyde-coated glass slides in four replicates. The cell microarray was reacted with an antibody probe against the cell surface marker, stage-specific embryonic antigen 1 (SSEA1). The equilibrium dissociation constants were derived from real-time binding curves, and this information could be useful for determination of stem cell stages. The OI-RD microscopy is also capable of detecting the formation and disintegration of micelles. 183 Micelles, which are very efficient drug carriers made of hydrophilic heads and hydrophobic tails, were biotinylated for immobilization on streptavidin substrates. The microarray was reacted with micellar solutions at different concentrations to determine the conformation of micelles by monitoring the changes in OI-RD signals. The critical concentration above which micelles were formed was determined to be around 0.0006 mg/mL in this study.
Comparison between OI-RD and SPRi
Both OI-RD microscopy and SPRi microscopy apply microarrays in high-throughput detection of biomolecular interactions. They share many similarities: (1) Both techniques are capable of endpoint imaging and real-time measurements; (2) data from OI-RD measurements can be converted easily into SPR data and vice versa;34,184 and (3) when operated near the Brewster angle, OI-RD is as sensitive as SPR. 174 It is much easier to set up and operate a SPRi microscope than an OI-RD microscope because there is more know-how in designing the latter. As a result, there are several SPRi-based microscopes available on the market, whereas OI-RD-based microscopy has not been commercialized.
However, OI-RD provides some advantages over SPRi. First, the maximum throughput of OI-RD can be 10,000~20,000, about one order of magnitude higher than that of SPRi. Second, whereas SPRi is limited to Au- or Ag-coated surfaces, OI-RD is compatible with all flat substrates. This makes surface modification in OI-RD more flexible than that in SPRi. Third, in OI-RD, the number density of surface-immobilized targets can be well controlled to minimize the mass transport effect when reacting with solution-phase probes. 185
Concluding Remarks
A combination of microarrays and label-free biosensors has provided a platform for high-throughput, parallel detection of biomolecular interactions. Such integration has a number of applications in genomics, proteomics, glycomics, drug screenings, and even areas of food and environmental analysis. Much interesting research has been performed and reported, but there is still room for further improvement: (1) In clinical examinations, the amount of patient samples is usually very low. Therefore, the sensitivity should be increased. This can be done by improving the instruments themselves and/or modifying the microarray surface for more efficient analyte capture. In OI-RD microscopy, the sensitivity could be enhanced by a factor of 30 when the incidence angle is close to the Brewster angle θB, compared to that 10° away from θB. 174 Wang et al. applied the trilayered metallic structure (Au–Ag–Au) in a SPR imaging sensor, and more than 50% increases in sensitivity and signal-to-noise ratio than those of the single-gold-layered SPR chip were determined by SPR microscopy at 660 nm. 186 (2) To optimize the screening efficiency, the time required to image a microarray should be minimized. For example, in OI-RD microscopes, the mechanical and/or optical screening stages can be improved for faster imaging. As shown in Figure 10 , using a combination of a galvanometer y-scan mirror and an f-theta lens, a microarray with 10,804 spots covering an area of 2 cm × 4 cm could be imaged in 90 min. 180 Also, data acquisition, data processing, and data analysis can be accelerated via better computer hardware and software. (3) To immobilize a variety of biomolecules, especially small molecule compounds, surface chemistry remains a big challenge. Modifying either the probe molecules and/or the microarray substrates requires knowledge in chemistry as well as a lot of trial and error. For example, 7961 small molecular compounds from NCI/DTP (National Cancer Institute/Developmental Therapeutics Program) were immobilized on isocyanate-functionalized glass slides for screening against vascular endothelial growth factor (VEGF) with an OI-RD microscope. 94 Also, using the OBOC method, a library containing 9261 encoded compounds was generated for small molecular microarray fabrication in high-throughput OI-RD screening assays. 187 Recently, Singh et al. reported using the 3D photo-cross-linking strategy based on the use of α-cyclodextrin as an alternative surface chemistry for the preparation of small molecule microarrays. 188 Small molecules of diverse structures containing different functional groups were covalently immobilized on a single slide for SPRi detection. (4) Sometimes, the probes to be screened consist of many unknown molecules, so it will be of great use if one can combine the microarray–biosensor platform with mass spectrometry (MS) to identify what has been captured on the surface. Nedelkov reported for the first time using the combination of SPRi microscopy and MS to image a single high-content protein microarray. 189 Antibodies to five human plasma proteins were arrayed in a 10×10 microarray, and the subsequent protein–antibody interactions were monitored via SPRi microscopy. Following protein affinity retrieval, this chip was overlaid with MS and analyzed, producing protein-specific mass spectra from distinct spots on the array. Other groups have also shown coupling of MS to SPRi for identifying captured proteins.190–192 (5) To introduce different chemical or physical stimuli to the biomolecular reactions, incorporating microfluidics into the sensing system will definitely help. This can also minimize the amount of probes required in a reaction as well as increase the number of available channels of probes. Liu et al. explored the application of SiO2 thin films as an intermediate layer for irreversibly adhering polydimethylsiloxane (PDMS) to plastic substrate, and developed a hard-soft, compact, robust microfluidic-based biosensor flow cell for the SPRi-based multi-array immunoassay application. 193 Ouellet et al. reported an integrated microfluidic array for high-throughput SPRi-based detection and determination of binding affinities between antibodies and protein targets. 194 This device, consisting of 264 element-addressable chambers of a 700-pL volume, was combined with a serial dilution network for simultaneous interrogation of up to six different analyte concentrations.
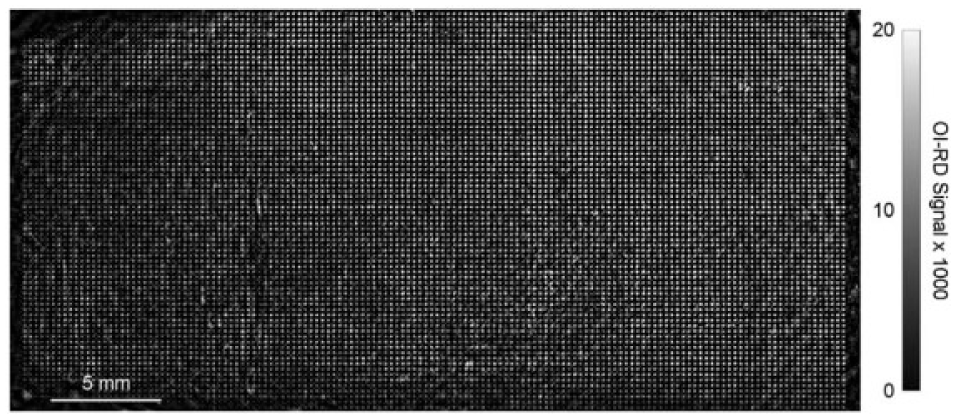

An oblique-incidence reflectivity difference (OI-RD) image of a 10,804-spot bovine serum albumin (BSA) microarray in buffer acquired with the f-theta lens-integrated scanning microscope. The image covers an area of 2 cm × 4 cm, and the pixel dimensions are 20 μm × 18.7 μm. Reprinted from J. Biomed. Opt. Vol. 15, No. 1, pp. 016018, Screening small-molecule compound microarrays for protein ligands without fluorescence labeling with a high-throughput scanning microscope. 180 Copyright © 2010 Society of Photo Optical Instrumentation Engineers.
Although some improvements are desired, it is believed that using microarrays in label-free biosensors has become a current trend in developing high-throughput screening platforms, and this will eventually lead us to a new era of the “-omics.”
Footnotes
Acknowledgements
The author is thankful for financial support from Taiwan MOST 101-2112-M-030-003-MY3.
Declaration of Conflicting Interests
The author declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The author is thankful for financial support from Taiwan MOST 101-2112-M-030-003-MY3.
