Abstract
Based on the principle of immobilized metal affinity chromatography (IMAC), it has been found that a Ni-Co alloy-coated protein chip is able to immobilize functional proteins with a His-tag attached. In this study, an intelligent computational approach was developed to promote the performance and repeatability of a Ni-Co alloy-coated protein chip. This approach was launched out of L18 experiments. Based on the experimental data, the fabrication process model of a Ni-Co protein chip was established by using an artificial neural network, and then an optimal fabrication condition was obtained using the Taguchi genetic algorithm. The result was validated experimentally and compared with a nitrocellulose chip. Consequentially, experimental outcomes revealed that the Ni-Co alloy-coated chip, fabricated using the proposed approach, had the best performance and repeatability compared with the Ni-Co chips of an L18 orthogonal array design and the nitrocellulose chip. Moreover, the low fluorescent background of the chip surface gives a more precise fluorescent detection. Based on a small quantity of experiments, this proposed intelligent computation approach can significantly reduce the experimental cost and improve the product’s quality.
Keywords
Introduction
Most manufacturing processes involve process parameters that specify sets of operating conditions typically derived from experiments. The set points of process parameters may be in conflict with each other, such that they can have a negative impact on the final product yields. Thus, proper design of process parameters is the key to success because it is significantly correlated with not only the cost but also the product quality.1–3 Intelligent computation is the approach to determine optimal fabrication conditions.
A protein chip is a powerful tool used to study protein abundance, functions, and interactions with other macromolecules in cells or organisms. The human body contains more than 100,000 distinct types of proteins that are essential for life. However, proteomes differ from cell to cell and are constantly altered through biochemical interactions. Understanding these complicated protein diversities in structures and functions will assist in developing effective therapies or potential new drugs for the treatment of disease. Besides the traditional methods and devices used in protein research, a protein chip, allowing high-throughput study of protein abundance and function, is the most common chip type presently developed.4–8 The analyte-specific reagents (ASRs; e.g., antibodies) are spotted on the chip surface. Each microspot works as the capturing molecule and can be incubated with a specific kind of analyte in the sample that usually contains multiple analytes. Various classes of capture scenarios (e.g., sandwich assay, antigen-antibody interaction, and protein-protein interaction) have been developed. 9 In the sandwich immunoassay, with the help of a second tagged molecule, an experimental readout based on fluorescence represents the relative amounts of the captured analytes. Recognizably, immobilization of proteins on solid surfaces is a prerequisite for the investigation of molecular interactions. Hence, an appropriate chip surface for the immobilization of protein samples is the key to protein chips. 10
Since immobilized metal affinity chromatography (IMAC) is one of the most popular methods for purification of recombinant proteins,11–14 it has been applied to the development of protein chips.15–21 In IMAC, a “tag” consisting of a series of histidine residues (His-tag) is usually attached to one terminus of the protein; moreover, an intermediate metal ion, such as nickel, is usually used to capture the His-tagged protein. As a result, the His-tagged protein binds the nickel and attaches to the chromatography column. In our previous work, a novel protein chip, with a Ni-Co alloy layer fabricated on the substrate of the printed circuit board by electrodeposition, was developed. 22 The Ni-Co alloy coating, by adding cobalt to form bimetallic deposit, has been shown to have better specific binding capability with the His-tagged protein than the Ni film. However, it is obvious that the proportion of Ni-Co composition on the chip will affect the affinity of the electroplated layer with the protein and, consequentially, influence the results of the immunoassay.
The aim of this article is to develop a systematic computation method for selecting optimal electroplating conditions based on a small quantity of experiments such that the Ni-Co protein chip has maximal immobilization ability to provide strong detectable fluorescence. Uricase was adopted as an example for experimental verification. Since uric acid is the purine metabolite of the human body, a person with a high concentration of uric acid might have more possibility of having gout. Uricase is an enzyme produced by the liver that contributes to catabolize uric acid. When enough uricase is present, uric acid can be oxidized to allantoin, which is harmless to humans; otherwise, uric acid may accumulate. Hence, the uricase protein is an important index for diagnosing gout. 23
Experiments
This proposed intelligent computation method for determining the optimal fabrication conditions involves the experimental design scheme of orthogonal arrays, artificial neural networks, and the Taguchi genetic algorithm (TGA). Based on a small quantity of experiments, the artificial neural network was used to establish the process model, and the TGA was used to search for optimal electroplating conditions.
Experiments Using an Orthogonal Array
In this study, a commercially available printed circuit board (PCB) was trimmed to 15 × 5 cm for the electroplating substrates. Besides the consideration of cost, the main reason to adopt PCB is that its conducting layer, typically made of thin copper foil, can directly serve as a plating seed layer. Thus, the fabrication process of the protein chip can be simplified. In the pretreatment step, the PCB substrates were immersed in 5% H2SO4 for 3 min to remove dioxide from surfaces and were washed by ultrasonic cleaning with a degreaser at 45 °C for 1 min.
Ni-Co electroplating was performed in a W 18 cm × L 25 cm × H 18 cm electroplating tank. Nickel cakes of 2 cm diameter and 5 mm thickness filled the slits of a metal Ti net (size: W 12 cm × H 14 cm × D 1.5 cm), which was placed on the anode. A 100-V/35-W pump equipped at the bottom of the tank provided a flow upward to the metal substrate to enhance the turbulence for promoting uniformity of the plating bath. To eliminate impurities and crystalline solids, a filter with 1-µm pores was installed above the pump. Furthermore, a digital power supply controlled by a PID controller supplied a 100-V/100-W output to a quartz heater for temperature control of the plating bath.
The necessary chemicals for electroplating included Ni(NH2SO3)2·4H2O, H3BO3, NiCl2·6H2O, (NH2SO3)2·4H2O, NH3·H2O, and Co(NH2SO3)2·4H2O. In addition, surface smooth additive (sodium dodecyl sulfonate, NaC12H25SO3) and low-stress additive (sodium saccharin, C6H4SO2NNaCO·2H2O) were used. All these materials were purchased from Blue Giant, Inc. (Taipei, Taiwan, ROC). The thickness of the coated Ni-Co film was 10 µm.
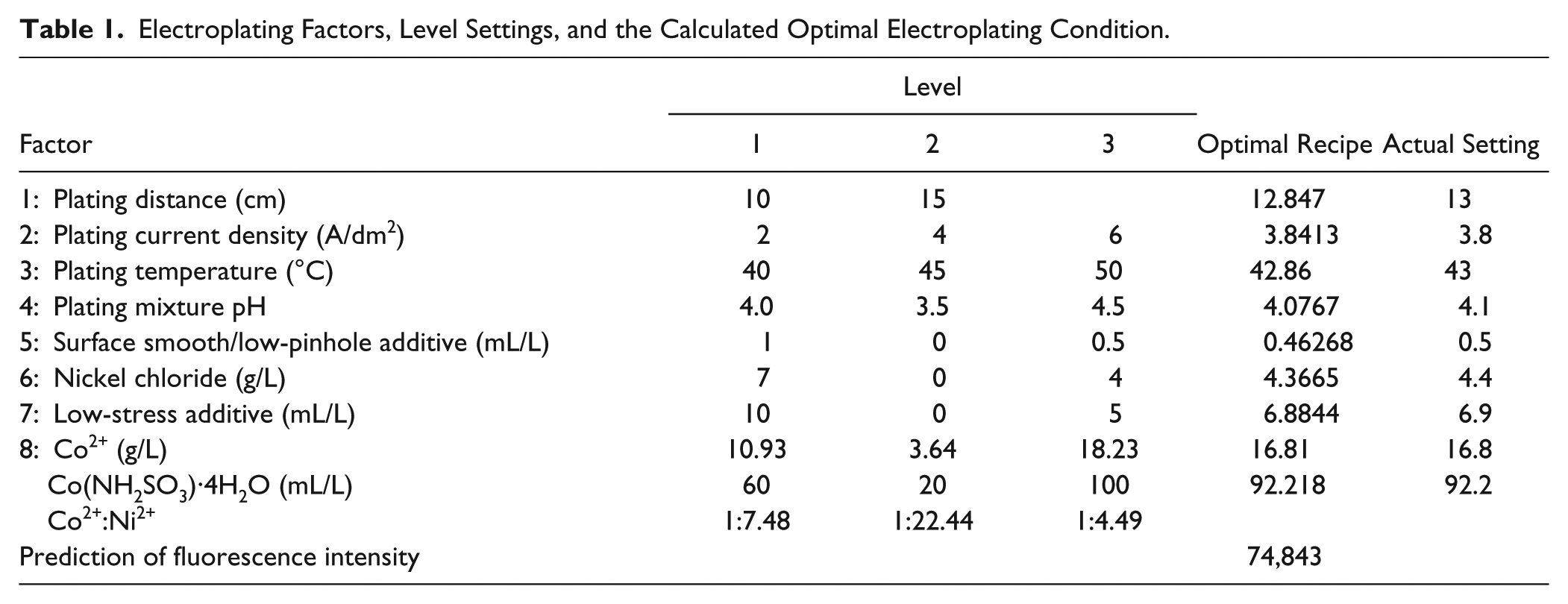
Eight factors were investigated in this study: (1) plating distance: determining the distance for metal ions to migrate, (2) plating current: affecting the crystallization and proportion of the deposited alloy’s composition, (3) plating temperature, (4) plating mixture’s pH: affecting the stress and brittleness in the coating layer, (5) surface smoothing additive: affecting surface smoothness in the coating layer, (6) nickel chloride additive: providing Cl ions to assist in the dissolution of nickel cakes in the anode, (7) low-stress additive: reducing the stress in the coating layer, and (8) Co2+/Co(NH2SO3)· 4H2O. Except for the plating distance, which had only two levels, the other seven factors had three levels, as shown in Table 1 .
Electroplating Factors, Level Settings, and the Calculated Optimal Electroplating Condition.
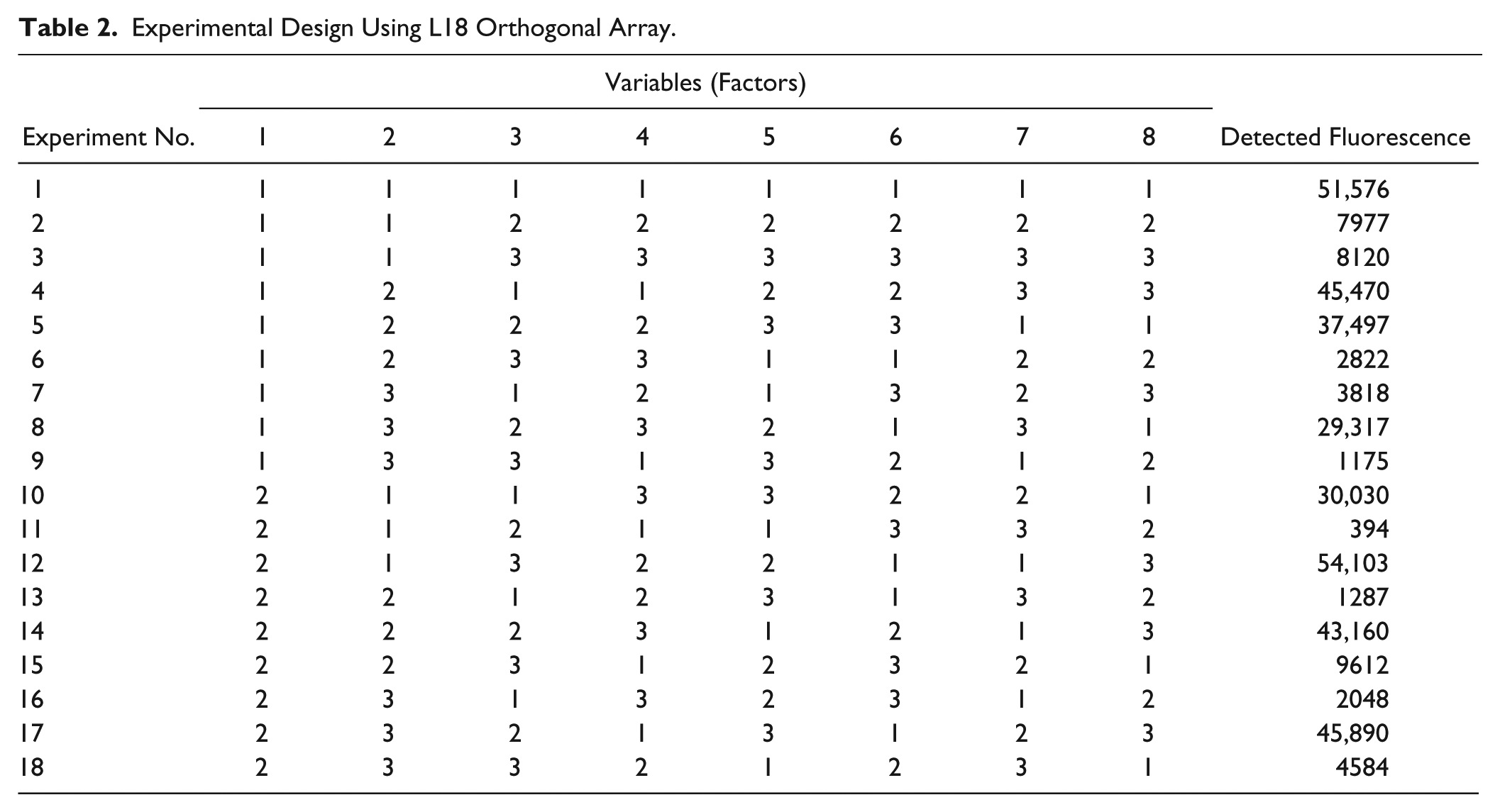
Experiments planned using an L18 orthogonal array, as shown in Table 2 , were performed first. Orthogonal array is a systematic and statistical way for conducting experiments. An orthogonality means that the design of experimentation is balanced and statistically independent. For any process factor, each level has an equal number of occurrences in the experimental matrix so that uniformly distributed coverage of the test domain can be provided. Each of the vectors conveys information different from that of any other vector in the sequence. That is, the effects of the factors can be separated from each other, and therefore, the experimental design can avoid redundancy. Moreover, due to this property of orthogonal arrays, a full factorial experiment to change one variable at a time is not necessary. The numerous experimental runs can be reduced without losing any vital information. Thus, the use of orthogonal arrays in this study fosters meaningful and cost-effective experimental designs.
Experimental Design Using L18 Orthogonal Array.
Bioexperiments
To verify the performance of Ni-Co–coated protein chips, bioexperiments were performed. The necessary biomaterials included the following:
Antigen: 6×His-tagged urate oxidase protein (uricase) purchased from PIERCE (Pittsburgh, PA)
Primary antibody: anti–uricase antibody obtained from Novus Biologicals, Inc. (Littleton, CO)
Immunofluorescence-stained secondary antibody (secondary antibody): Cy5-conjugated anti–goat antibody from Jackson ImmunoResearch Laboratories, Inc. (West Grove, PA)
Bovine serum albumin (BSA) purchased from Sigma-Aldrich (St. Louis, MO)
The principle of antigen/antibody binding specificity was applied to examine the performance of the Ni-Co–coated chips. The procedure is shown schematically in Figure 1 . First, a 6×His-Uricase was prepared by dilution in a 0.1 M PB (pH 7.4) coating buffer, with a 5 µg/mL concentration titration. The samples were then arrayed on the chips, followed by a 37 °C/2-h incubation for protein immobilization in a hot circulator oven. In addition, some spots without 6×His-tag were added as a negative control. After the immobilization step, the biochips were washed with a wash buffer (phosphate buffer with 0.1% Tween-20) three times for 1 min per wash remove the residual antigen. The first antibody was prepared by dilution in 1% BSA-PB (PB with 1% BSA) to a final concentration of 2 µg/mL and was then added onto the top of the antigen layer. One-hour incubation in a 37 °C hot circulator oven was followed by three washes in PB-T (1× phosphate buffer–Tween-20) for 1 min per wash. Next, the Cy5 fluorescently labeled anti–goat IgG secondary antibody was diluted in 1% BSA-PB to 5 µg/mL and added onto the top of the His-/anti-Uricase protein complex to find the result. The binding reaction was again incubated in a 37 °C hot circulator oven for 1 h, followed by the same wash procedure. Finally, fluorescent detection was performed by a GeneTAC LS IV laser scanner (Genomic Solutions, MI), and the intensity was analyzed by Gene Pix 4.1 software (Molecular Devices, CA), as listed in the right-hand column of Table 2 .
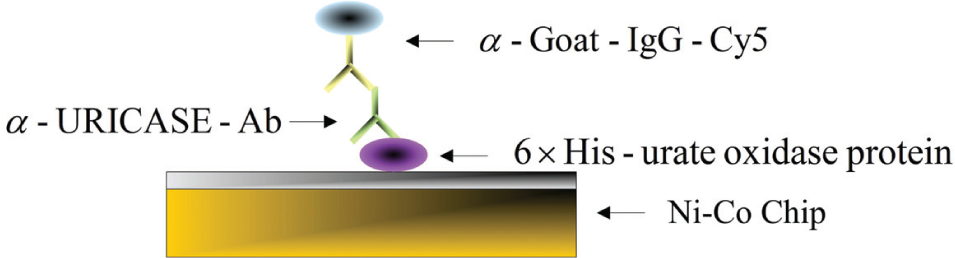
Immunoassay experiment using the principle of antigen/antibody binding specificity.
By means of electroplating, the surface of the protein chip was coated with Ni-Co duads in the form of alloy deposition. The fluorescent signal of immunoreaction tells the amount of affinity between the His-tagged proteins and metal duads. On the basis of the results of L18 experiments, we proceeded to develop a process optimization approach for chip fabrication.
Intelligent Computation Approach
Based on the orthogonal array, the mandatory experiment runs were reduced from 4374 (21 × 37) to only 18. Fewer experiments are necessary, making the experimental design more efficient. To promote the performance and repeatability of the protein chips, a systematic approach was developed using artificial neural networks to establish the process model and the TGA to search for the optimal electroplating condition. That is, a response surface is conceptually constructed from the discrete L18 experimental data by a neural network to serve as the process model, which links the input (the eight electroplating factors) and the output (the fluorescence intensity). By the encoding mechanism of the TGA, the eight electroplating factors are coded into a chromosome. Moreover, the evaluation function of the TGA is as a trained neural network to take a chromosome as input and to return a measure of the chromosome’s performance (i.e., the fluorescence intensity). The TGA finds good chromosomes by manipulating the material in the chromosomes to search for the peak of the response surface constructed by the neural network. Using the operators on chromosomes, the optimal electroplating condition can be obtained such that the Ni-Co protein chip has the best immobilization ability with His-tagged proteins. A maximal amount of His-tagged protein can be immobilized on the chip surface to incubate with the specific kind of analyte in the sample. Moreover, the His-tagged proteins would not be rinsed away during the washing step, eventually providing a precise experimental outcome.
A process model is a description of a process relationship between the inputs and the outputs. Modeling is a very important issue because the success of process optimization depends on whether the physical process is modeled properly. The most common approach to construct a model is to apply various physical laws to develop mathematical equations to describe the process. To establish a suitable model, a thorough understanding of the process is essential. Unfortunately, this might be difficult, time-consuming, or even impossible in many cases due to the complexity of the process. Alternatively, the black-box model developed by evolutionary algorithm techniques (e.g., the artificial neural networks [ANNs]) is a simpler approach.
In this study, the radial basis function network (RBFN) was adopted. The RBFN possesses good generalization capability with minimal network structure so that it offers a variety of applications in the curve-fitting (approximation) problems. This network, as shown in Figure 2 , consists of three layers, including an input, hidden, and output layer. The hidden nodes have Gaussian activation functions. To build the process model for the fabrication of protein chips, the input variables of the network used are the eight electroplating factors. In addition, the output, which implements a weighted sum of hidden-node outputs, is the associated fluorescent signal. The equation is given as follows. 24
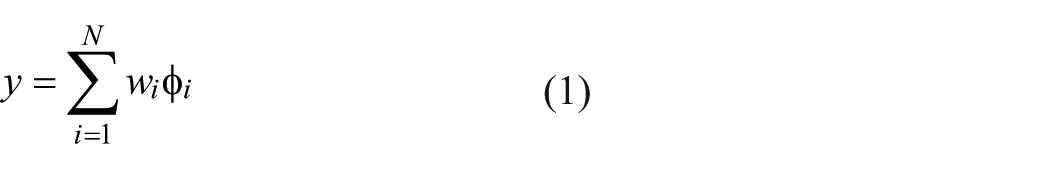
with
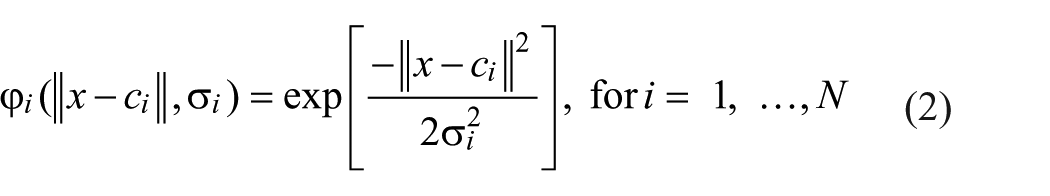
where x is the input vector; wi is the strength of the ith radial basis function ϕ
i
(usually called weight); ci and
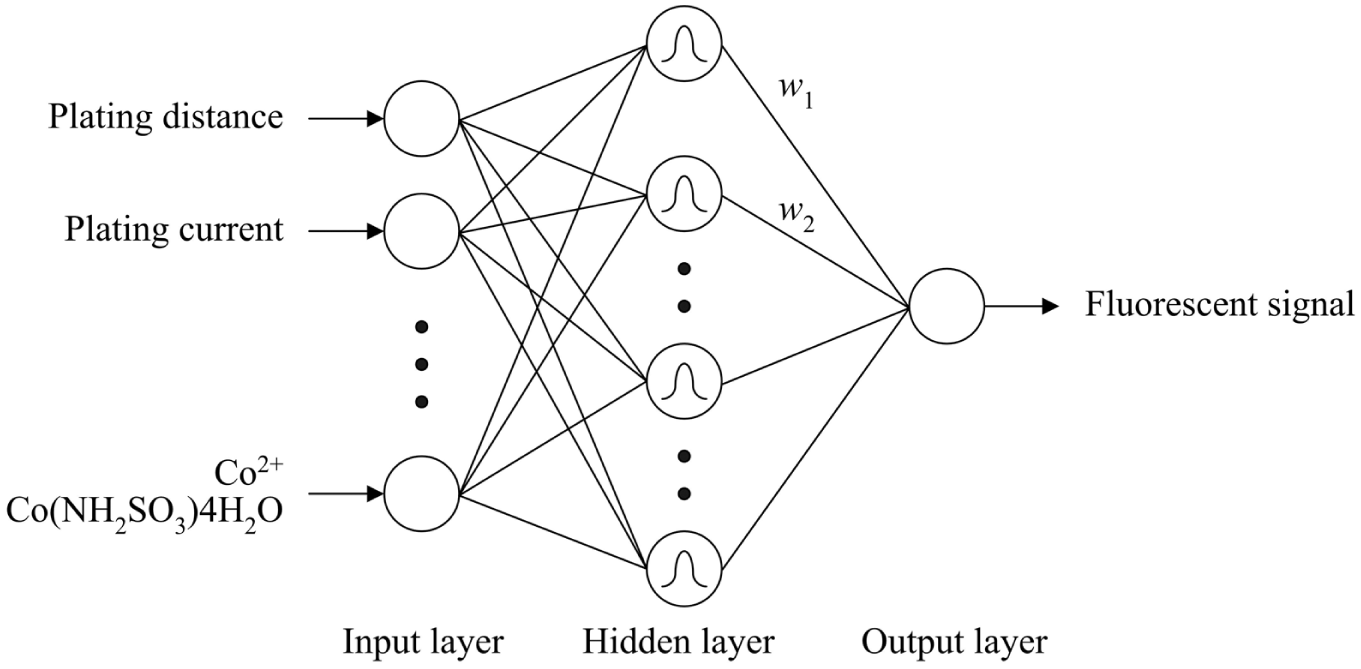
Radial basis function network.
Using L18 experimental data, the RBFN is trained by the unsupervised learning in the input layer and the supervised learning in the output layer. The former learning involves the determination of the centers ci and variances

where dk, k = 1, . . . , 18, denotes each fluorescence intensity of L18 experiments.
After training, the RBFN establishes the input-output dependence describing the relationship between the electroplating condition and the corresponding chip performance that emerges from the fluorescence intensity. Then, three chips, with electroplating conditions different from those of the L18 experiments, are fabricated to test the RBFN model.
Once the process model is established, the optimal settings of eight electroplating factors can be searched for iteratively by the TGA. The genetic algorithm (GA) is a heuristic search technique inspired by evolutionary biology, such as crossover, mutation, and eugenics. Currently, the GA has been widely used for optimization problems because it can generate the global optimum and prevent the search from being trapped into a local optimum. To simulate natural evolutions, the variables are coded into strings of binary bits called chromosomes. A fitness function replacing the objective function in the conventional optimization methods is used to evaluate the fitness of every candidate solution to decide whether it will contribute to the next generation of solutions. Hence, at the end of each generation, a new population with better fitness can be obtained and used in the next iteration of the algorithm. Finally, the optimal solution is sifted from the massive propagation of chromosomes through competition.
In comparison with a conventional genetic algorithm, the TGA includes the concepts of orthogonal experimental design and factor analysis that increase efficiency during chromosome crossovers. Therefore, TGA can execute global optimization more quickly. The procedure is elaborated as follows.
(a) Producing Chromosome Population
A chromosome consists of gene sequences. For recipe optimization of Ni-Co alloy-coated protein chips, each gene sequence is composed of 50 binary bits and denotes an electroplating factor. Thus, the gene sequence represents the parameter information transmitted during the procedures of crossover and mutation. The more binary bits a gene sequence has, the higher the resolution. For eight electroplating factors, a chromosome comprises 400 bits.
(b) Evaluating Fitness Value
For each chromosome, its fitness value must be evaluated. Decoding the chromosome to get the values of electroplating factors and substituting them into the trained RBFN supplies an output that represents its fitness. A larger fitness value signifies that the settings of the electroplating factors resulted in a stronger fluorescent signal. In the TGA optimization procedure, the chromosome with the greatest improvement in fluorescent intensity will be selected for the generation evolution.
(c) Orthogonal Crossover
In general, chromosomes interchange information by means of a crossover to avoid obtaining a local optimal solution during the optimization process. In TGA, the orthogonal array is used to learn the main effect of an individual factor. Therefore, an orthogonal crossover provides a systematic method to improve the efficiency of the genetic algorithm and may save computational time.
(d) Mutation
During the evolution process of chromosomes, the ones with the highest fitness value are kept. Moreover, under the consideration of calculation efficiency, chromosomes with lower fitness values are used for mutation action. By means of the recombination of individual genes, better chromosome evolutions are expected. However, the result of mutation can be influenced by random coding. The fitness values of chromosomes have the potential to evolve into either better or worse values. Thus, a mutation rate of 0.1 is employed to limit the percentage of chromosomes to be used for mutation.
(e) Eugenics
The evolution behavior in TGA keeps an optimal chromosome from the prior generation from competing with those in the next generation. This helps to maintain the optimization procedure and to reach the evolution purpose. In a eugenics process, it is common that at least one optimal chromosome will be retained without crossovers or mutations. The number of elite chromosomes will affect the speed of evolution.
Steps (a) to (e) of the TGA are repeated until the generated fitness value satisfies the requirement or the generations of evolution have been reached. At that moment, the global optimum of the electroplating condition is obtained.
Results and Discussion
In this study, the intelligent computation approach for enhancing the protein immobilization capability of a Ni-Co alloy was investigated. Furthermore, uricase was adopted as the test target. Since immobilization comes from the specific binding between a 6×His-Uricase protein and a Ni2+/Co2+ layered alloy surface, the result of an immunoreaction can explain the capability of a Ni-Co alloy-coated chip to capture a 6×His-Uricase protein.
For comparison, L18 experimental data were also analyzed using the regression kriging method. The results show that the plating condition of experiment 12 produces the maximal fluorescence. The detected fluorescence intensity of experiment 12 reached 54,103 a.u. (with a coefficient of variation of 2.2%). Then, the L18 experimental data were used in the proposed systematic approach for establishing the process model by RBFN. Moreover, the optimal electroplating recipe was searched for using the TGA. The computed optimal recipe with the expected fluorescence intensity, reaching 74,843 a.u., is shown in Table 1 . This result was validated experimentally.
Two sets of Ni-Co alloy-coated protein chips were fabricated, according to the optimal recipe, for the repeatability test (denoted as experiments 19 and 20). The actual setting of each electroplating factor is listed in the right-hand column of Table 1 . Four duplicate samples of a 6×His-Uricase protein were arrayed onto each chip. After the immunoreactions, it was observed that the four duplicate samples on the same chip had similar fluorescence intensities. The average values were 62,125 a.u. and 59,576 a.u. for experiments 19 and 20, respectively. The results are better than that of experiment 12. Moreover, the values almost approach the fluorescence detection limit of a GeneTAC LS IV scanner, which is 65,535 a.u. The results show that a better chip surface was produced using this process optimization method. It had stronger binding capability to immobilize His-tagged proteins, resulting in higher fluorescence intensities. Therefore, the feature allows this chip to be used under low concentrations of protein. Their coefficients of variation were 0.169% and 0.666%, respectively, which are also smaller than that of experiment 12. Under the optimal electroplating recipe, the coated chip surface has uniform characteristics so that the variance of the scanned fluorescence intensities of the four duplicate samples on the same chip is small. This fact explains that repeatability is better.
Bioexperiments always suffer from many variations, including materials and operations. In this study, the fluorescence intensities of experiments 19 and 20 did not agree with the prediction value. Two inferences can be drawn from the experiments. The protein samples were arrayed onto the chip surface using Pipetman (Gilson, WI) by manual operation. Due to affinity binding, the area of the Ni-Co layer that the His-Uricase protein occupied affected the amount of immobilized proteins. When the binding reached saturation, the redundant proteins could not be immobilized and, afterward, were removed during washing step. Furthermore, the areas of droplets were slightly different. These two reasons caused the difference in fluorescence intensities between experiments 19 and 20, as well as the variation with the prediction value. In addition, the maximal fluorescence was the target in our approach. The computed value of 74,843 a.u. was also beyond the detection limit of a GeneTAC LS IV scanner. However, the optimization target cannot be restricted to the detection limit of a scanner. Otherwise, the detected fluorescence will get weaker under the variation of bioexperiments. To use this proposed approach, overfitting in ANNs is another issue deserving notice. Cost function provides a way to improve the ability of an ANN to generalize from the training data to test data. 21
In addition, a set of experiments using a 3% nitrocellulose chip was performed for comparison (denoted as experiment 21). Immobilization of proteins on a nitrocellulose membrane is one of the most widely used techniques for protein analysis. A nitrocellulose membrane works as a universal blotting surface. With a porous fiber structure and electrostatic force, the membrane can provide nonspecific binding to capture samples.
Nitrocellulose powder was purchased from Sigma-Aldrich. Due to it being in a solid phase, the nitrocellulose powder could not be directly coated onto the surface of a chip. By adding the powder and 3-glycidoxypropyltrimethoxysilane (GPTS) to acetone and shaking the mixture at 100 rpm for 6 h, the powder fully dissolved to form an aqueous solution. The solution was deposited on a 1 × 3-in. glass by a spin coater with a rotational speed of 2000 rpm, followed by a 60 °C incubation for 1 h to form the nitrocellulose chip. Then, the bioexperiment was performed. The procedure was similar to the aforementioned procedure except that the sample-spotted nitrocellulose slide had to be immersed in a blocking buffer (1% BSA-PB) to reduce nonspecific binding of other proteins. A nitrocellulose membrane produces nonspecific binding with protein by the hydrophobic feature and oxygen bond. To obtain a better signal-to-noise (S/N) ratio, a blocking step is indispensable. After the experiment, the measured average fluorescence intensity was 22,484 a.u. Indeed, the immunoassay results revealed that the Ni-Co alloy-coated protein chip fabricated using the optimal recipe of the proposed approach had the best performance. The mean and standard deviation of fluorescence intensities of these three experiments are shown in Figure 3 . The fluorescent images give the intuitional evidence.
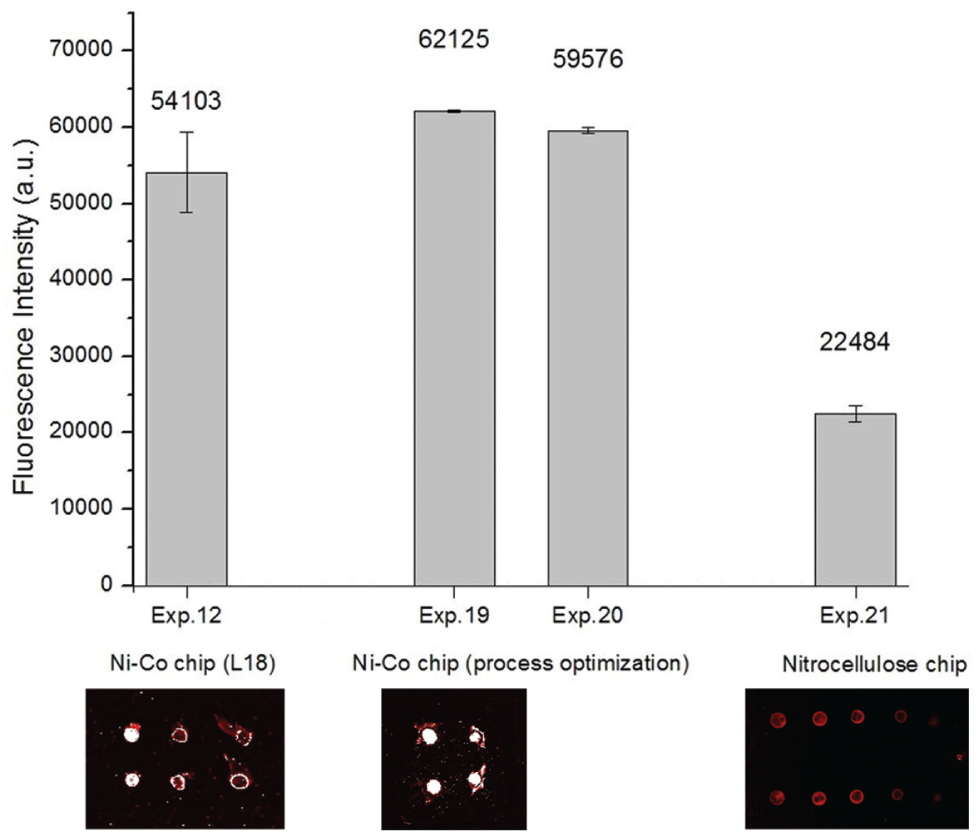
Statistics of fluorescence intensities and fluorescent images.
In conclusion, in this study, a systematic approach for developing the optimal Ni-Co alloy-coated surface to immobilize His-tagged proteins was investigated. This proposed approach used an artificial neural network and genetic algorithm to obtain the optimal fabrication condition. The experimental results showed that the proposed Ni-Co alloy-coated chip, fabricated using the intelligent computation method, had the best performance and repeatability compared with the Ni-Co chips of the L18 orthogonal array design and the nitrocellulose chip. Furthermore, this Ni-Co alloy-coated chip can provide a stable, inexpensive, and low-background chip platform for researchers to perform complex biochemical reactions. From the experimental results, we can conclude that this proposed intelligent computation approach, based on a small quantity of experiments, can significantly reduce the experimental costs and improve the product quality. It also provides the potential to be implemented in other manufacturing processes.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Ministry of Science and Technology (MOST) of Taiwan, R.O.C. (MOST 103-2221-E-033-039).
