Abstract
Sepsis is a rapidly progressing, severe inflammatory response to infection, causing more than 200 000 deaths per year. Rapid, specific pathogen identification is important to guide sepsis treatment. In this review, we describe and compare currently available commercial products for sepsis diagnosis and pathogen identification, based on microbiological, molecular, and mass spectrometric technologies. Microbiological techniques, the current “gold standard” in sepsis pathogen identification, include blood culture followed by subculturing and pathogen identification via biochemical or microscopic means. These methods have been automated but nevertheless require several days to generate results. Alternative technologies, including highly multiplexed PCR-based methods and mass spectrometric approaches, can decrease the required turnaround time. Matrix-assisted laser-desorption ionization time-of-flight–based systems have recently become an attractive option to rapidly identify a broad spectrum of sepsis pathogens with good sensitivity and specificity. Effectively integrating rapid sepsis pathogen identification into the hospital workflow can improve patient outcomes and can reduce the length of hospitalization and cost per patient.
Introduction
This review aims to provide a brief overview of sepsis epidemiology, pathophysiology, and current practices for diagnosis and patient management, as well as causative agents and treatment options. The review then focuses on describing and comparing currently available commercial products for sepsis diagnosis and pathogen identification, based on microbiological, molecular, and mass spectrometric technologies.
Sepsis is a profound inflammatory response to infection caused by bacterial, viral, fungal, or parasitic pathogens. 1 About 750 000 Americans are diagnosed with sepsis annually, with this condition responsible for greater than 200 000 deaths per year.2,3 High-risk groups include individuals with a weakened immune system, such as infants, geriatrics, and people suffering from chronic diseases. 2 Sepsis persists as one of the most common hospital-acquired infections and one of the leading causes of in-hospital deaths in the United States.4,5 Hospitals are held financially liable for preventable infections acquired by patients while staying at the facility.3,6 Hospital-acquired sepsis was estimated to incur annual health care costs greater than $17 billion.7,8
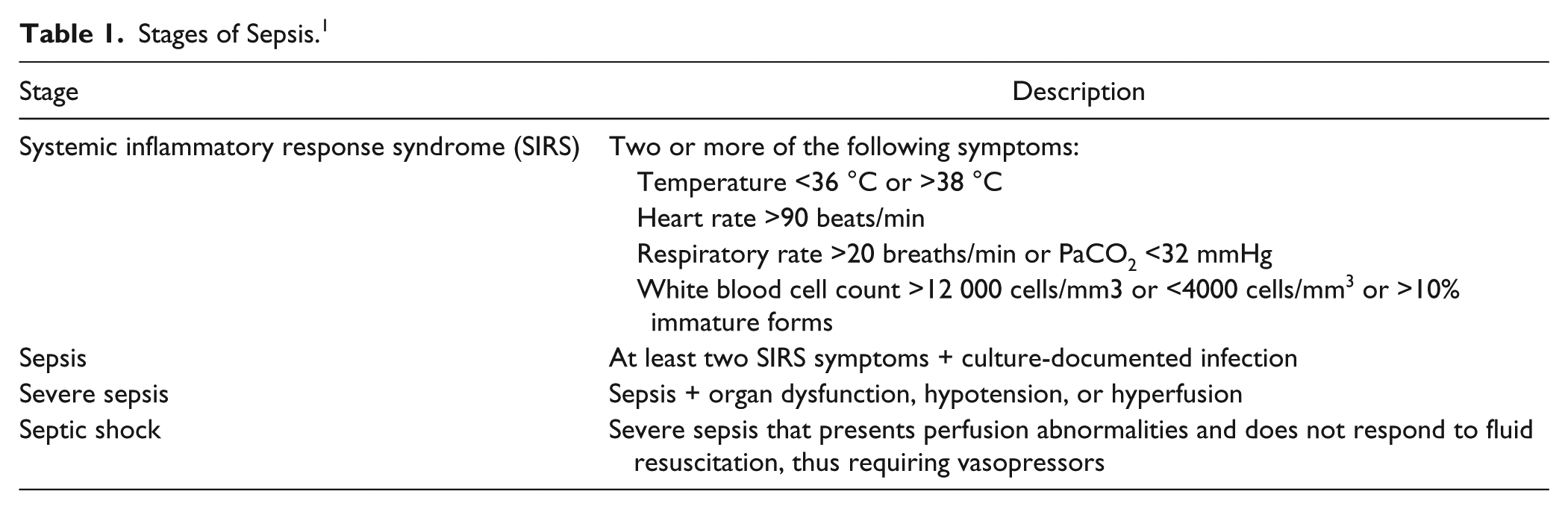
Symptom progression of sepsis is rapid, with an individual’s probability of survival declining with each hour of onset.9–12 If untreated, an individual suffering from sepsis can develop circulatory complications leading to organ failure.1,2 Health care professionals categorize sepsis into four progressive stages of severity ( Table 1 ): (1) systemic inflammatory response syndrome (SIRS, an overwhelming immune response that is often considered the precursor to sepsis), (2) sepsis, (3) severe sepsis, and (4) septic shock. 13
Stages of Sepsis. 1

Mortality rates increase significantly with sepsis severity ( Fig. 1 ). Furthermore, severe sepsis results in hospitalizations three times longer than those of lower-grade sepsis cases.8,14 Therefore, early differential diagnosis is essential to facilitate effective treatment and prevent disease progression. Rapid identification of the causative pathogen enables appropriate antibiotic treatment, which improves survival rates and overall patient outcomes and reduces hospitalization duration and overall hospital expenditures. 12
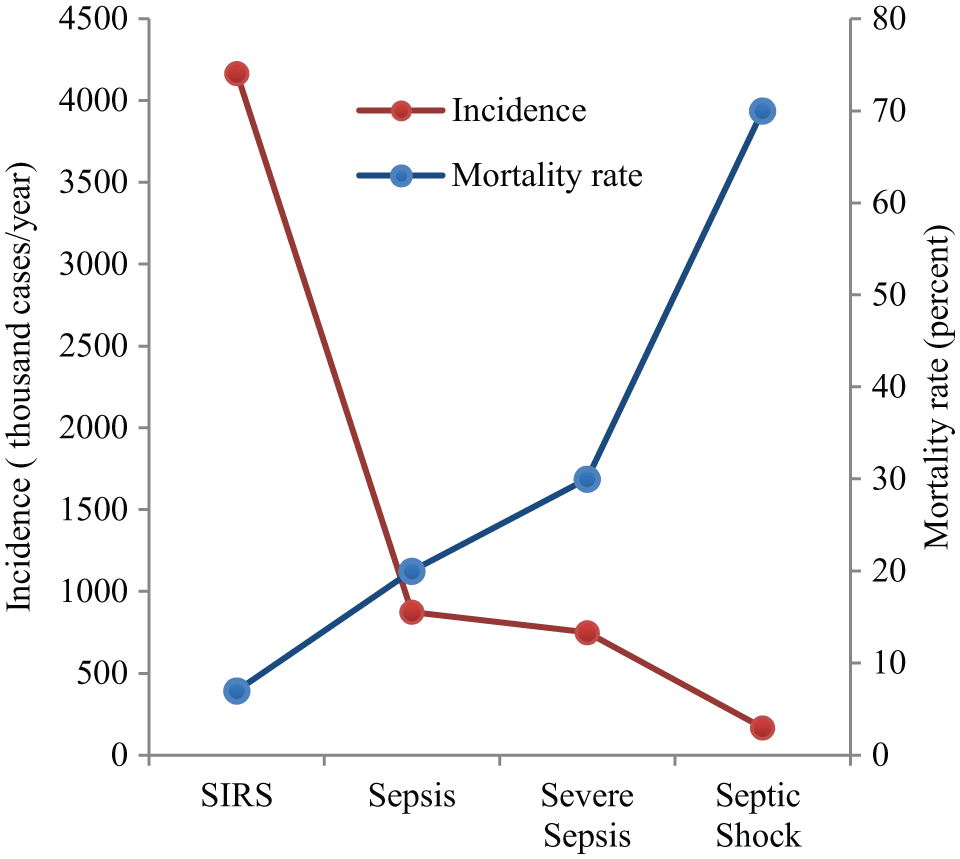
Sepsis mortality rate and incidence for different stages of sepsis severity. Systemic inflammatory response syndrome (SIRS) is often considered a preliminary stage for sepsis. As sepsis severity progresses to septic shock, incidence rates decrease, but mortality rates drastically increase. Mortality rates reflect the upper limits for each stage.2,15-18
Pathophysiology
The detrimental effects of sepsis are caused by pro-inflammatory and subsequent anti-inflammatory pathophysiological responses ( Fig. 2 ). 19 Infections can trigger a multitude of pro-inflammatory responses of the innate immune system. For example, gram-negative bacteria produce lipopolysaccharides as endotoxins, which activate toll-like receptors on macrophages and other cells, eliciting the release of pro-inflammatory cytokines and chemokines. Constant stimulation of toll-like receptors leads to damage of vascular endothelial cells and causes blood-clotting abnormalities, eventually resulting in tissue injury, multiple organ dysfunction, and death.19–21
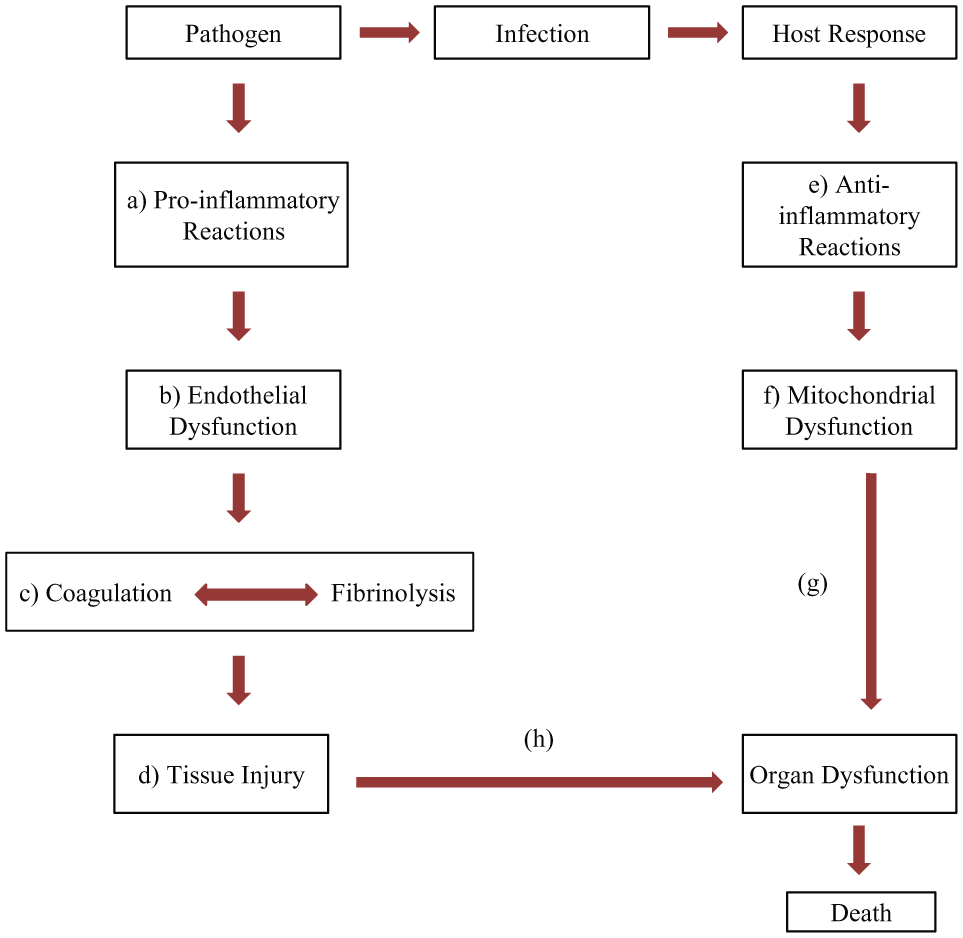
Sepsis pathophysiology. (
Anti-inflammatory pathways are activated in response to pro-inflammatory reactions to return the patients’ bodies to homeostasis ( Fig. 3 ). 25 Anti-inflammatory responses induce immunosuppression, thus increasing the patients’ susceptibility to secondary infections, which can worsen the patient’s condition.20,21,25
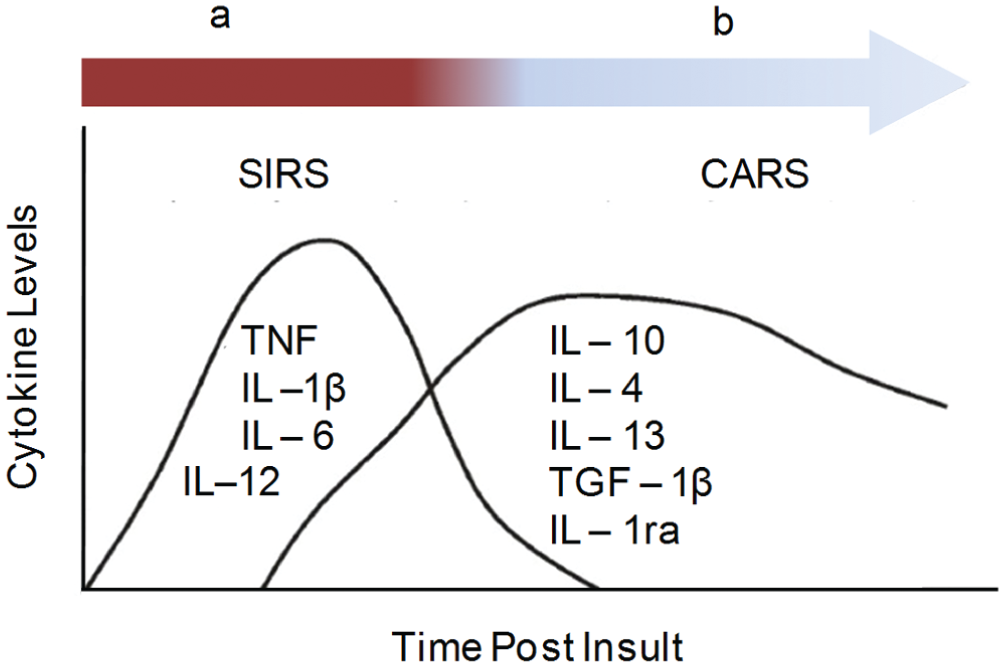
Cytokine expression levels in sepsis. (
General Diagnosis
Lack of appropriate early differential diagnosis is a major obstacle for effective sepsis treatment. 28 Currently, sepsis diagnosis relies mainly on clinical symptoms ( Fig. 4 and Table 1 ). Certain protein biomarkers 29 can aid in sepsis diagnosis, assessment of sepsis severity, and determination of the effectiveness of sepsis therapies. For example, rising blood levels of C-reactive protein indicate a general inflammatory response but cannot differentiate inflammation from infection. Procalcitonin (PCT) levels can be used to differentiate SIRS due to noninflammatory causes from bacterial sepsis. 29 Furthermore, PCT levels can be used to differentiate bacteremia caused by gram-negative versus gram-positive bacterial infections, with a positive predictive value of 83.0% and a negative predictive value of 74.0%. 30 However, these predictive values are relatively modest, and PCT cannot identify pathogens at the genus and species level.
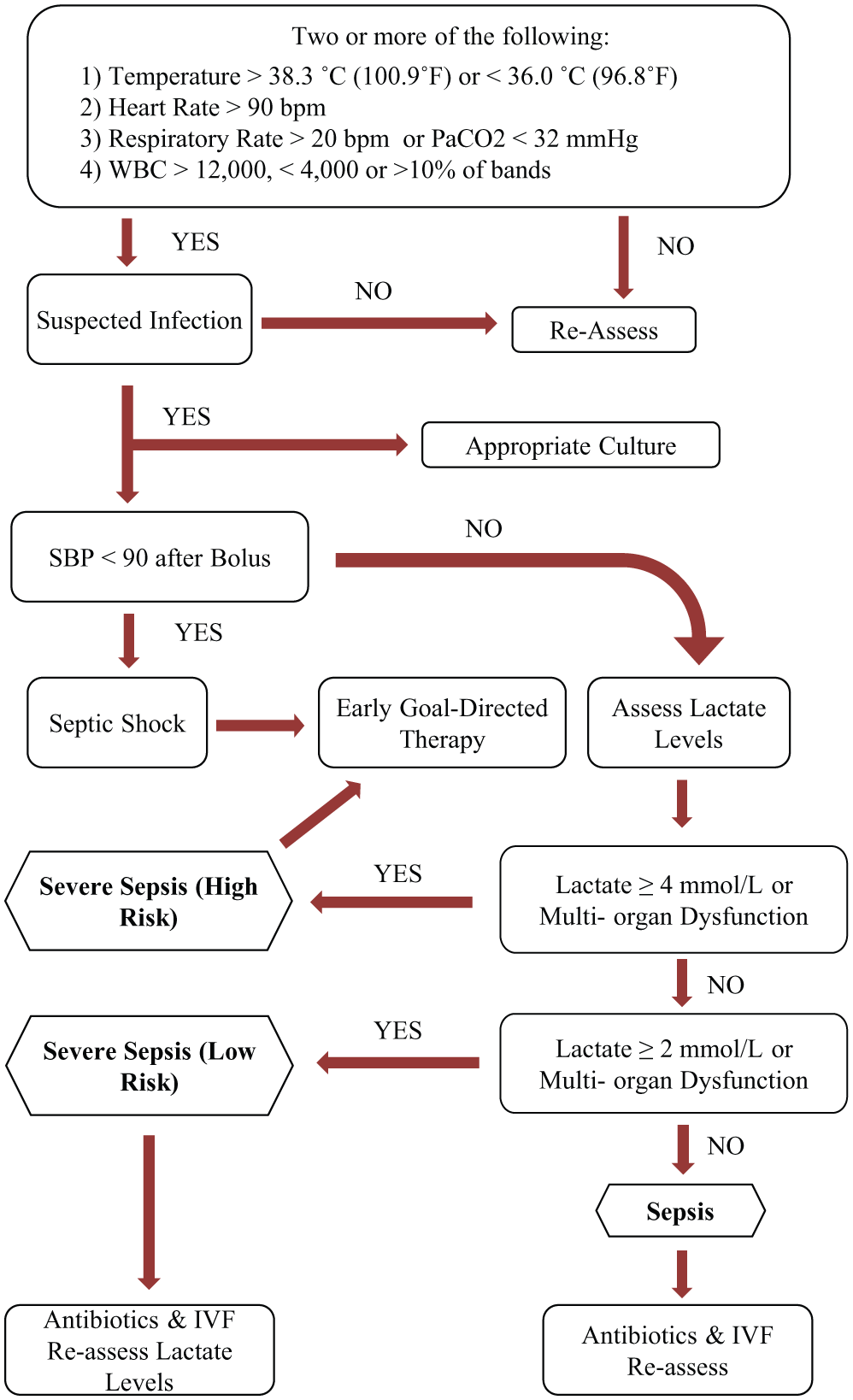
For all patients with suspected infection, to identify the causative pathogen, the clinician will submit blood samples or other body fluids such as urine or cerebrospinal fluid for laboratory testing. 29 However, multiple days are typically required for definitive pathogen identification (ID) and antimicrobial susceptibility testing (AST). 12 For the initial treatment period, sepsis treatment bundles have been developed,31,32 which focus primarily on stabilizing patients with septic shock or high-risk severe sepsis, by maintaining appropriate blood pressure, fluid levels, and oxygen saturation, for example, via administering intravenous fluids or vasopressors or through mechanical ventilation or blood transfusion. 31 These general sepsis treatment bundles include initial presumptive antibiotic therapy, but they do not address the adequacy of antibiotic treatment, which requires pathogen ID/AST.33,34
Causative Agents
Administering appropriate antibiotic treatment within the first 4 h of sepsis onset has been shown to decrease hospitalization by 2 d, decrease absolute mortality by 24%, and ultimately lead to better patient outcomes. 31 Every hour of treatment delay correlated to a 7.6% decrease in chance of survival. 31 However, due to the long turnaround time for pathogen ID/AST, the physician must prescribe presumptive antibiotic treatment before the causative agent is known. The appropriateness of treatment selection is assessed later on, and antibiotic therapy is adjusted if necessary.
The causative pathogens of sepsis include gram-positive and gram-negative bacteria, anaerobes, fungi, and parasites ( Fig. 5 ).35,36 Clinicians select the initial broad-spectrum antibiotics based on the patient’s background and the location of the initial infection. Broad-spectrum antibiotics are given in cocktails to target multiple mechanistic paths. For example, initial combination treatment with an aminoglycoside, a linezolid or vancomycin, and an antipseudomonal ®-lactam targets both gram-positive and gram-negative bacteria. 37 This initial empirical treatment, however, often is inadequate. In one international cohort study ( Fig. 6 ), 36 only 29% of 1156 sepsis patients were adequately treated with initial empirical monotherapy, and an additional 23% of the patients were adequately treated with combination therapy. For 47% of the patients, the initial empirical treatment was found to be inadequate.
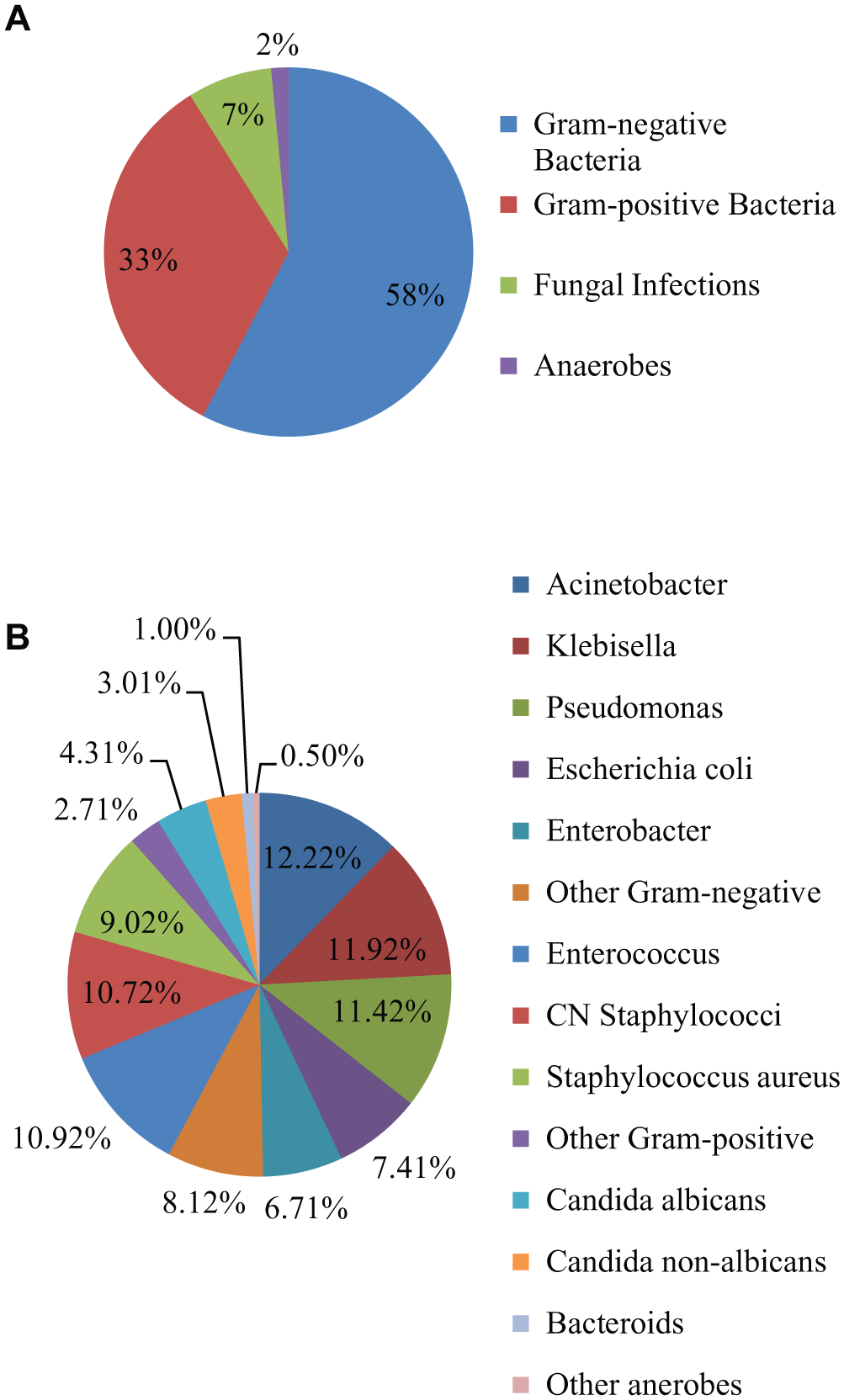
Causative pathogens of sepsis by (
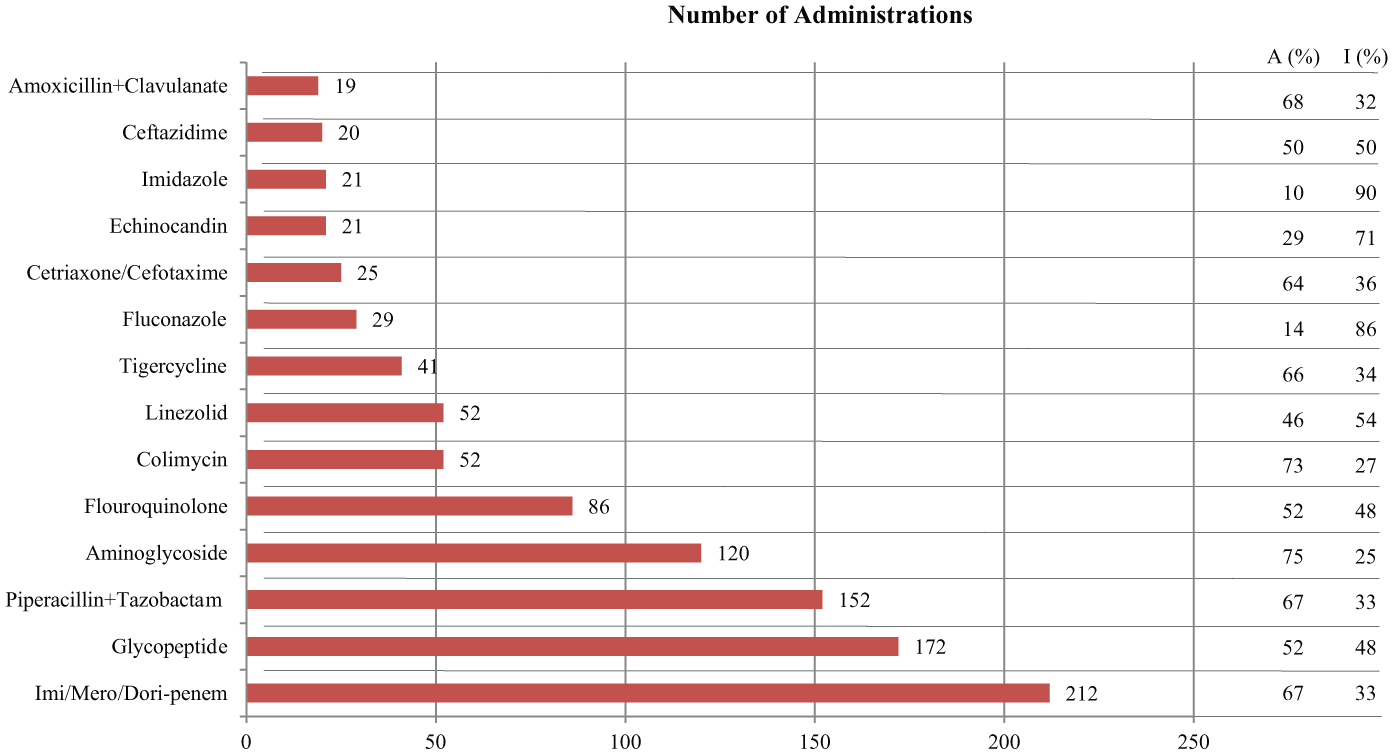
Common antibiotic treatments used in the first 12 h prior to and up to 24 hours after a positive blood culture has been obtained. 36 A% and I% indicate the percentage of antibiotic administrations that were found to be adequate or inadequate, respectively based on subsequent pathogen identification/antimicrobial susceptibility testing (ID/AST). Data includes antibiotic administrations as monotherapies and combination therapies. Figure adapted from Tabah et al. 36
The low adequacy of initial treatment with broad-spectrum antibiotics dramatically reduces the overall efficacy of sepsis treatment, 38 requiring multiple treatment adjustments. In addition, broad-spectrum antimicrobial agents target an extensive spectrum of beneficial microorganisms, causing additional adverse consequences for the patient. Inadequate use of antibiotics also leads to the emergence of drug-resistant bacterial pathogens. 17 Sepsis persistence is largely due to these shortcomings of current antibiotic treatment. 35 Improved methods for pathogen ID/AST are needed to guide appropriate treatment decisions. 12
Technical Product Analysis
Currently available technologies for sepsis pathogen ID39–42 discussed in the following sections include traditional microbiological/biochemical methods, automated microbiological methods, genotypic methods, and mass spectrometry (MS). Most of these methods also enable AST, which are described as appropriate, but the focus of this review is on pathogen ID.
Microbiology Methods of Detection and ID
Microbiology techniques such as cell culturing, Gram staining, and biochemical tests have been the “gold standard” in sepsis pathogen ID/AST. These techniques have been incorporated into automated systems, speeding up the process and preventing contamination. 43
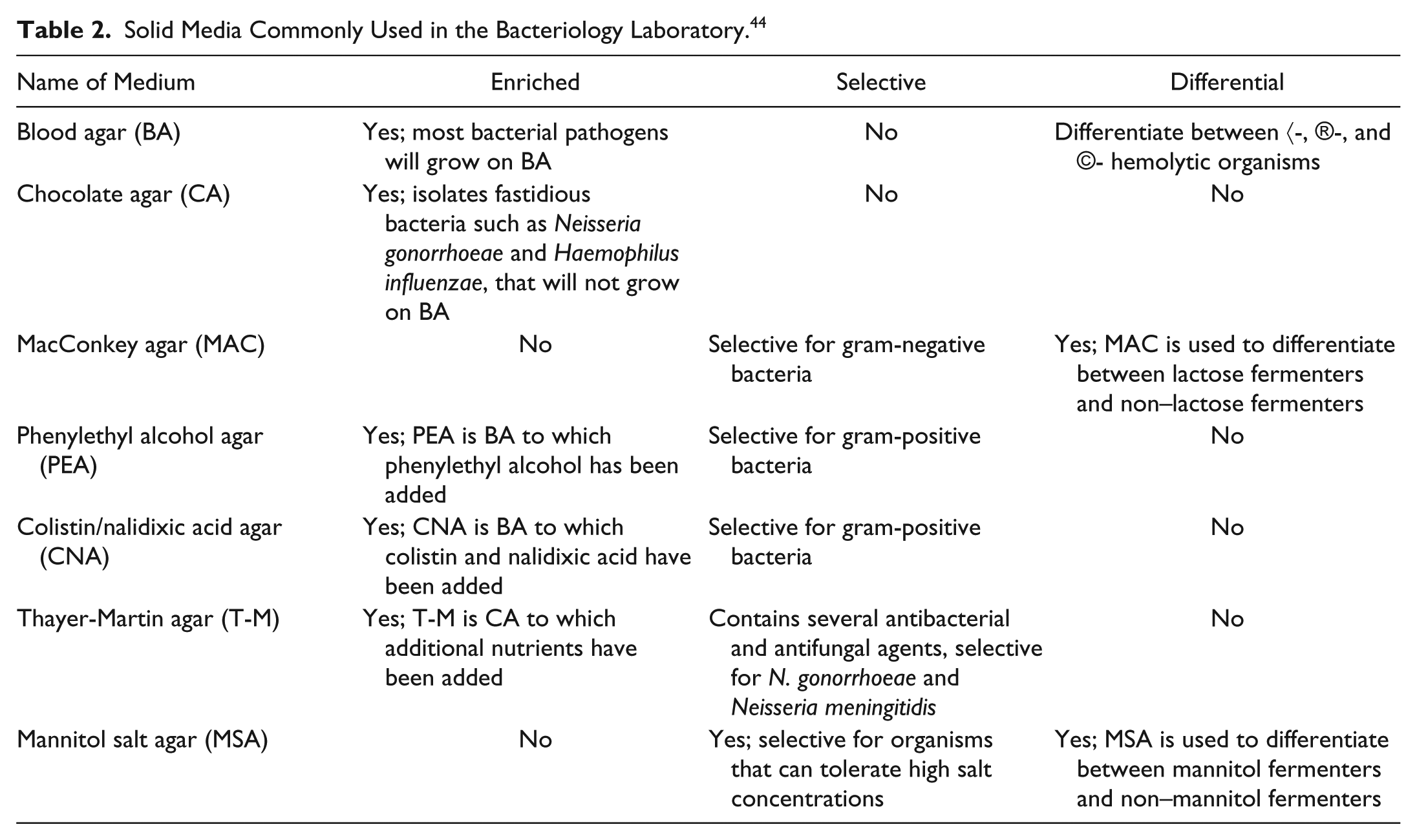
Blood cultures are the primary method of initially detecting the presence of an infectious agent in the bloodstream. Blood samples are cultured in bottles containing liquid media, which are then incubated. If the initial blood culture is positive, further tests are required to facilitate pathogen ID/AST. 44 Subculturing, which involves plating an aliquot from the initial blood culture onto solid selective media ( Table 2 ) and incubating for growth, enables isolation, amplification, and further identification of the pathogen. 44
Solid Media Commonly Used in the Bacteriology Laboratory. 44
The Septi-Chek (Becton Dickinson, Franklin Lakes, NJ) and Oxoid Signal (Thermo Fisher Scientific, Inc., Waltham, MA) blood culture systems are manual methods that incorporate a sensor to detect bacterial growth. In the Oxoid Signal system, the blood sample is introduced into the Oxoid culture bottle. If a pathogen is present, positive pressure created from pathogen growth displaces media into a separate chamber.44,45 The Septi-Chek blood culture system enables pathogen ID by observing changes in turbidity and color of the media.44,45 It is a two-part culture system consisting of a bottle with liquid medium and an additional compartment containing a “slide,” attached on top of the bottle, containing three separate regions with specific growth media. 46 After the liquid culture has been inoculated with the sample, the culture is subjected to initial incubation, enabling bacteria to grow in the liquid compartment. Next, the bottle is inverted, which allows liquid culture media to flow into the second compartment, enabling pathogens to come in contact with the additional selective growth media on the slide surface. This acts as a subculturing method to help narrow the identity of the pathogen.44,45
Many microbiology laboratories automate blood culture tests using systems such as the BacT/Alert (BioMérieux, Craponne, France) ( Fig. 7 ).47,48 In this device, blood samples are aspirated into multiple BacT/Alert culture bottles containing various media. The bottles are then inserted into shelf racks of the BacT/Alert instrument, where they are agitated and incubated. Culture bottles are automatically checked for bacterial growth, based on colorimetric detection. CO2 produced as byproduct of pathogen growth causes a color change, which is detected via reflectance spectrophotometry.49,50 The instrument automatically detects growth and alerts laboratory personnel, who can then subculture the pathogen. 50
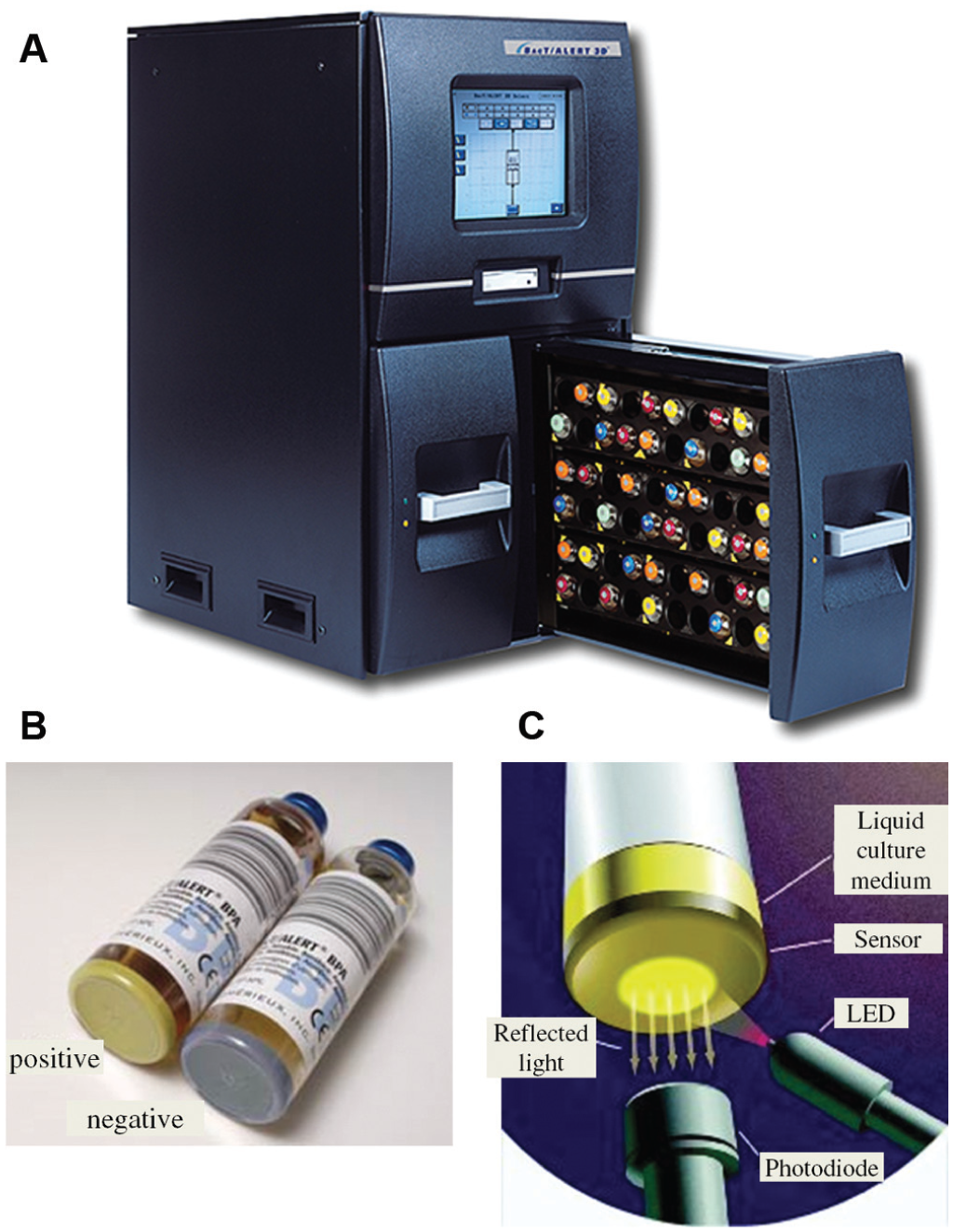
(
Differential staining is another method used to identify pathogens.44,51 For example, Gram staining differentiates between gram-positive and gram-negative bacteria based on differences in cell wall composition. Results are easily observed under a standard microscope. Biochemical methods (such as the catalase or coagulase assays) differentiate closely related organisms by determining if the microbial culture has specific enzymatic activity. 52 In addition, AST is used to detect possible drug resistance to help physicians select the most appropriate therapy. 53
Biochemical pathogen ID/AST methods can be automated on systems such as the BioMérieux Vitek 2 ( Fig. 8 ), to increase the efficiency of the sepsis pathogen ID/AST workflow. 53 For testing by the Vitek 2, a bacterial isolate obtained via subculturing is suspended in sterile saline solution. The automated system uses reagent cards ( Fig. 8A ) with 64 wells that contain specific substrates to conduct biochemical assays, or dry culture media and antimicrobial agents to conduct AST. 54 Each reagent card contains a transfer tube and a system of fluidic channels that can be used to fill the wells in a simple and automated manner. The operator suspends a bacterial isolate obtained via subculturing in phosphate buffer saline. A reagent card’s transfer tube is inserted into this suspension. A rack of cards and bacterial suspension tubes ( Fig. 8B ) are manually loaded into the vacuum chamber station of the Vitek 2 ( Fig. 8C ). The Vitek 2 then evacuates this chamber, thus removing the air from the wells and channels within the reagent cards. Back-flushing the chamber with air introduces positive pressure, pushing the bacterial suspension into the reagent card wells. The cards are then transferred into the incubation chamber of the instrument, where the transfer tubes are cut off, and the cards are sealed. During incubation, the cards are rotated and monitored every 15 min for changes in turbidity or absorbance at specific wavelengths.55,56 Changes in turbidity indicated bacterial growth, enabling determination of the minimal inhibitory concentration as part of AST. Changes in absorbance are due to conversion of chromogenic substrates by inherent enzymatic activity of the pathogen, enabling species-specific pathogen identification.
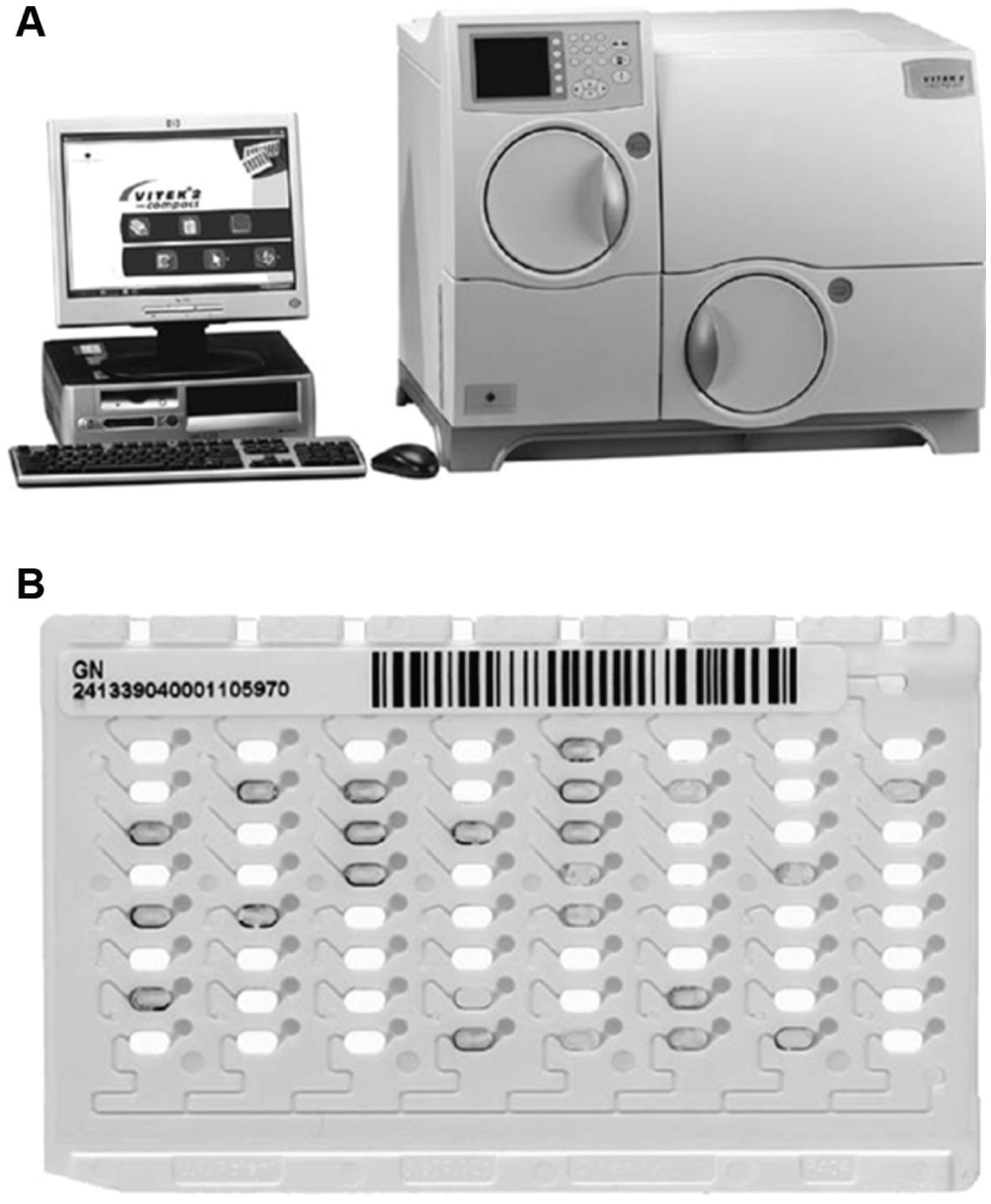

(
Although microbiological methods are currently the gold standard for sepsis pathogen ID/AST, these methods have a long turnaround time. According to a recent study involving 112 sepsis patients diagnosed via conventional methods ( Fig. 9 ), 12 on average, 15 h were required from the blood draw to obtain a positive blood culture result. Pathogen ID and AST required an additional 36 and 11 h, respectively. Therefore, following the blood draw, results were available on average after approximately 2.1 d for pathogen ID and 2.6 d for AST. Molecular diagnostic and MS methods described in the following sections have decreased the overall time to pathogen ID and/or AST.
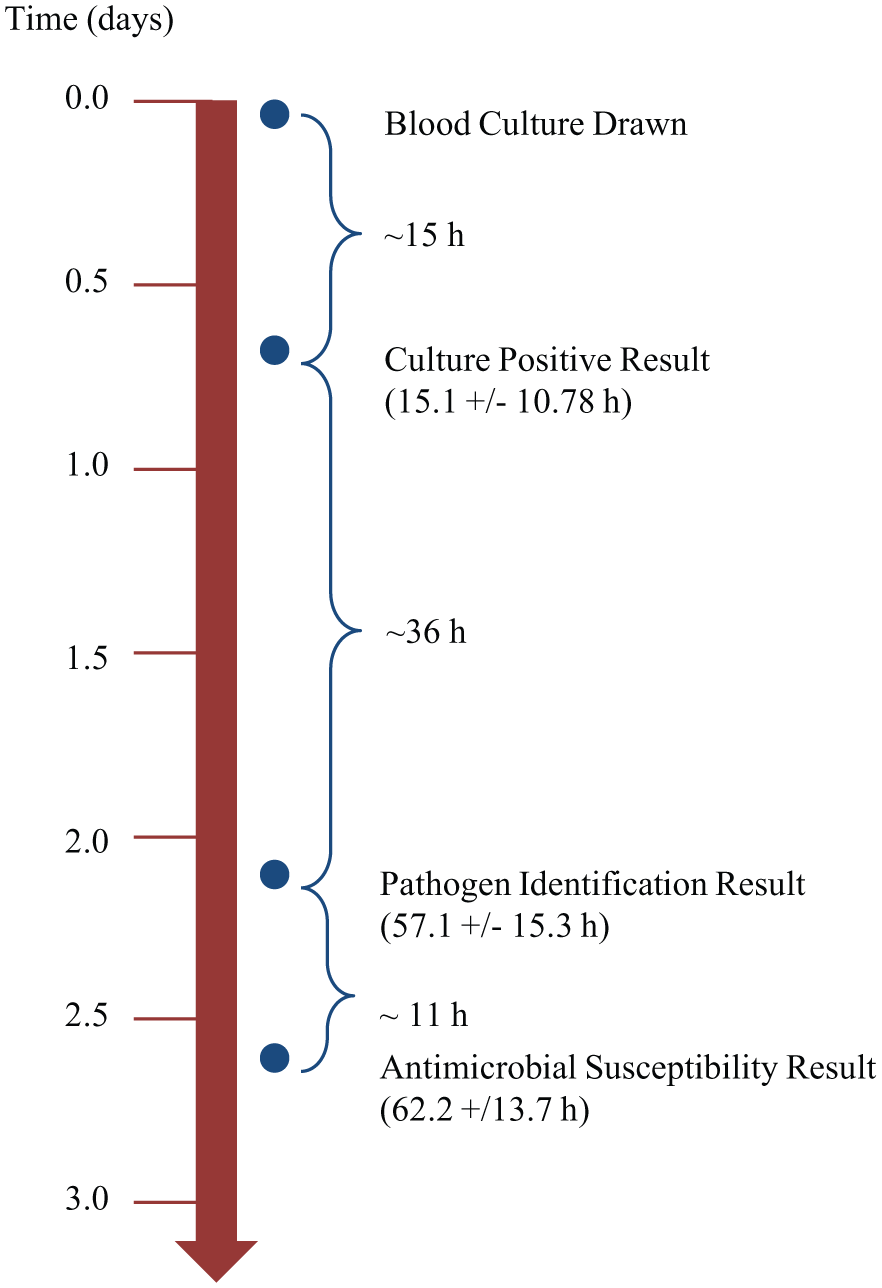
Timeline of conventional sepsis pathogen identification and antimicrobial susceptibility testing, according to Perez et al. 12 Blood culture results were obtained using a BioMérieux Bactec FX Platform. After subculturing on solid media, biochemical pathogen identification and culture-based antimicrobial susceptibility testing were performed on the Becton Dickinson Phoenix platform.
Molecular Diagnostic Methods:
Molecular diagnostic assays for sepsis pathogen identification include fluorescent in situ hybridization (FISH) and methods that use nucleic acid amplification, typically via the PCR, with real time or end point detection. 58 Molecular methods for sepsis diagnosis and pathogen identification have recently been reviewed. 59
FISH) Assays
FISH assays ( Fig. 10 ) can be used to identify bacteria or fungi in positive blood cultures, based on probes that detect specific ribosomal RNA (rRNA) sequences.60,61 From a positive blood culture, the fastest assay (QuickFISH, AdvanDx, Woburn, MA) requires about 26 min to generate a result,62,63 which is much faster than traditional microbiological methods. 12 Probes have been developed to target about 95% of pathogens causing bacteremia, 64 and AdvanDx sells FISH assays for reportedly more than 90% of sepsis-causing pathogens.65,66 However, each assay probes only for one to three pathogens at a time. Thus, multiple tests may be required to confidently identify a pathogen involved in sepsis.
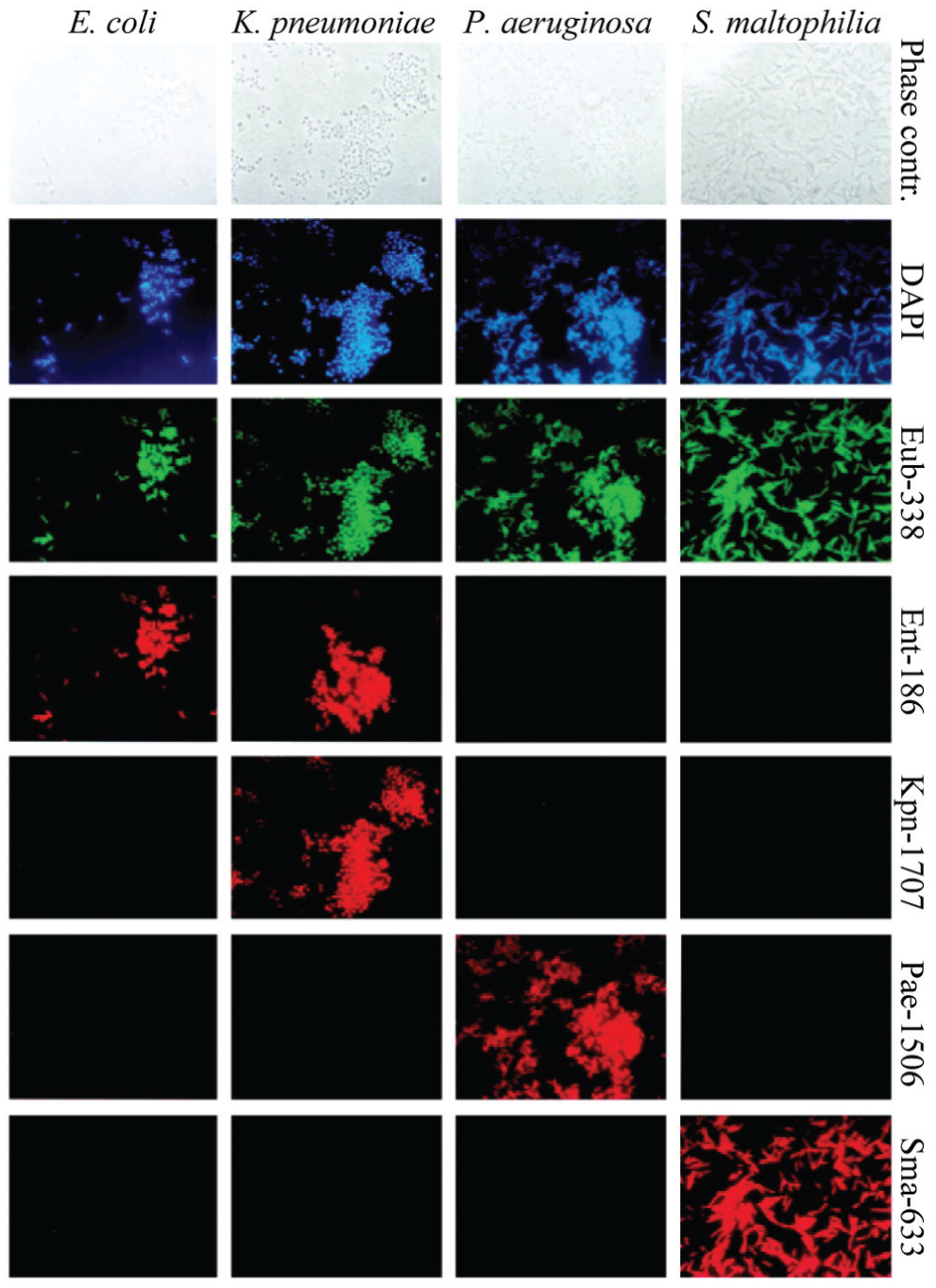
Fluorescent in situ hybridization (FISH) assay for identification of bacterial sepsis pathogens. A positive blood culture sample is smeared on a slide and treated with fluorescently labeled hybridization probes. After hybridization and stringency washing, slides are visualized under a fluorescent microscope. 67 All cells are visualized via DAPI (blue) nuclear staining. All bacteria are visualized via the FITC-labeled (green) EUB-338 probe that targets general bacterial ribosomal RNA (rRNA). Because this target is naturally abundant, FISH probes do not require the amplification step necessary in other genotypic assays. 61 Cy3-labeled (red) specific probes identify individual pathogens based on specific rRNA sequences. Figure from Kempf et al., 64 showing an earlier, precommercial FISH assay.
Peptide nucleic acid (PNA) FISH probes contain N-(2-aminoethyl)-glycine backbones that are electrically neutral. PNA probes are therefore able to transverse the bacterial cell wall and bind more stably to rRNA than do their DNA counterparts. Several proprietary PNA FISH assays are on the market today, most of which are from AdvanDx,67,68 although new FISH technologies are currently being pursued by Miacom Diagnostics. 69 PNA-FISH assays generally have high sensitivity and specificity ( Table 3 ).
Molecular Diagnostic Methods for Sepsis Pathogen Identification.

Data from Mancini et al 61 and additional sources, as indicated. AST, antimicrobial susceptibility testing; NA, not available.
Limit of detection (LOD) refers to CFU/mL in blood culture or whole blood, respectively, for assays performed on positive blood culture samples versus whole blood.
Data derived from FDA 510k summary statements.
Real Amplicon Detection (READ). To date, no literature exists describing this technology.
Nucleic Acid Amplification Tests
Molecular diagnostic tests based on nucleic acid amplification, typically via PCR, enable pathogen identification and detection of drug resistance markers with high sensitivity and specificity. 79 Many such systems have been reported, and the key characteristics of selected commercially available sepsis molecular diagnostic tests are summarized in Table 3 . Newer molecular diagnostic methods can shorten the time required to identifying the causative microorganism, compared to traditional microbiological methods, as well as FISH assays.43,91 This is especially true in cases where a test can be performed directly from a whole-blood sample, without the additional time required to obtain a positive blood culture ( Table 3 ). However, the low concentration of pathogens in fresh blood necessitates a relatively large sample input volume, which presents a technical challenge and typically results in the presence of a high background of human genomic DNA. To overcome these sample preparation challenges, the Vyoo sepsis assay (SIRS-Lab, Jena, Germany) includes a proprietary affinity chromatography medium to enable DNA extraction for pathogen identification from whole blood. This affinity chromatography medium removes 90% of human background DNA.61,76,76 After PCR amplification, products are analyzed via gel electrophoresis and matched to species-specific band patterns. 61 However, the sensitivity and specificity of this method are modest, and the end point detection format requires additional handling steps.
In contrast, in the LightCycler SeptiFast Test (Roche Molecular Systems, Mannheim, Germany), DNA is extracted directly from whole blood via silica-coated magnetic beads, followed by real-time PCR for pathogen identification in three parallel multiplex reactions for gram-positive, gram-negative, and fungal microbes. These PCR reactions amplify the internal transcribed space regions between the 16S and 23S rRNA genes for bacteria and between the 18S and 5.8S rRNA genes for fungi, with real-time detection followed by melt curve analysis, using hybridization probes. The system uses a four-color fluorescence readout, with one channel to detect the internal amplification control. Up to three pathogens are identified and differentiated in each of the other three channels based on differences in the melting temperature of the duplex formed between the hybridization probes and the target amplicon.61,70
The Magicplex Sepsis Real Time Test (Seegene, Seoul, Korea) likewise identifies sepsis pathogens via qPCR with melt curve analysis but with much higher levels of multiplexing, enabling the identification of more than 90 pathogens.72,73 Both the LightCycler SeptiFast Test and the Magicplex Sepsis Real Time Test are CE-IVD marked for commercial use in Europe but not U.S. Food and Drug Administration (FDA) cleared for use in the United States. Both assays have relatively high specificity (>90%) but moderate sensitivity (60-80%) and therefore cannot be used as the sole means for sepsis pathogen identification.
Higher sensitivity can be attained for nucleic acid testing performed from positive blood culture samples because, in this case, the pathogen is enriched via culturing. This increases the overall turnaround time but still enables faster pathogen identification compared to other automated microbiological methods such as the Vitek 2 because it alleviates the need for subculturing. In many of the currently available nucleic acid testing systems for sepsis identification from positive blood cultures ( Table 3 ), multiplexing is achieved via spatial encoding, that is, identification of different pathogens in distinct wells or on distinct spots of a microarray. For example, the Hyplex BloodScreen system (Amplex BioSystems, Giessen, Germany) uses multiplex PCR amplification with biotinylated primers targeting the pathogen’s species-specific housekeeping genes in two reactions, respectively, for four gram-negative and for six gram-positive bacteria. The latter includes testing for the Staphylococcal methicillin resistance gene. The amplicons are then identified via end point detection in distinct wells of a microtiter plate using an enzyme-linked immunosorbent assay–like sandwich assay format ( Fig. 11 ). 90 This system has relatively high sensitivity and specificity. However, the test entails a lengthy multistep protocol with significant amplicon carryover contamination risk.
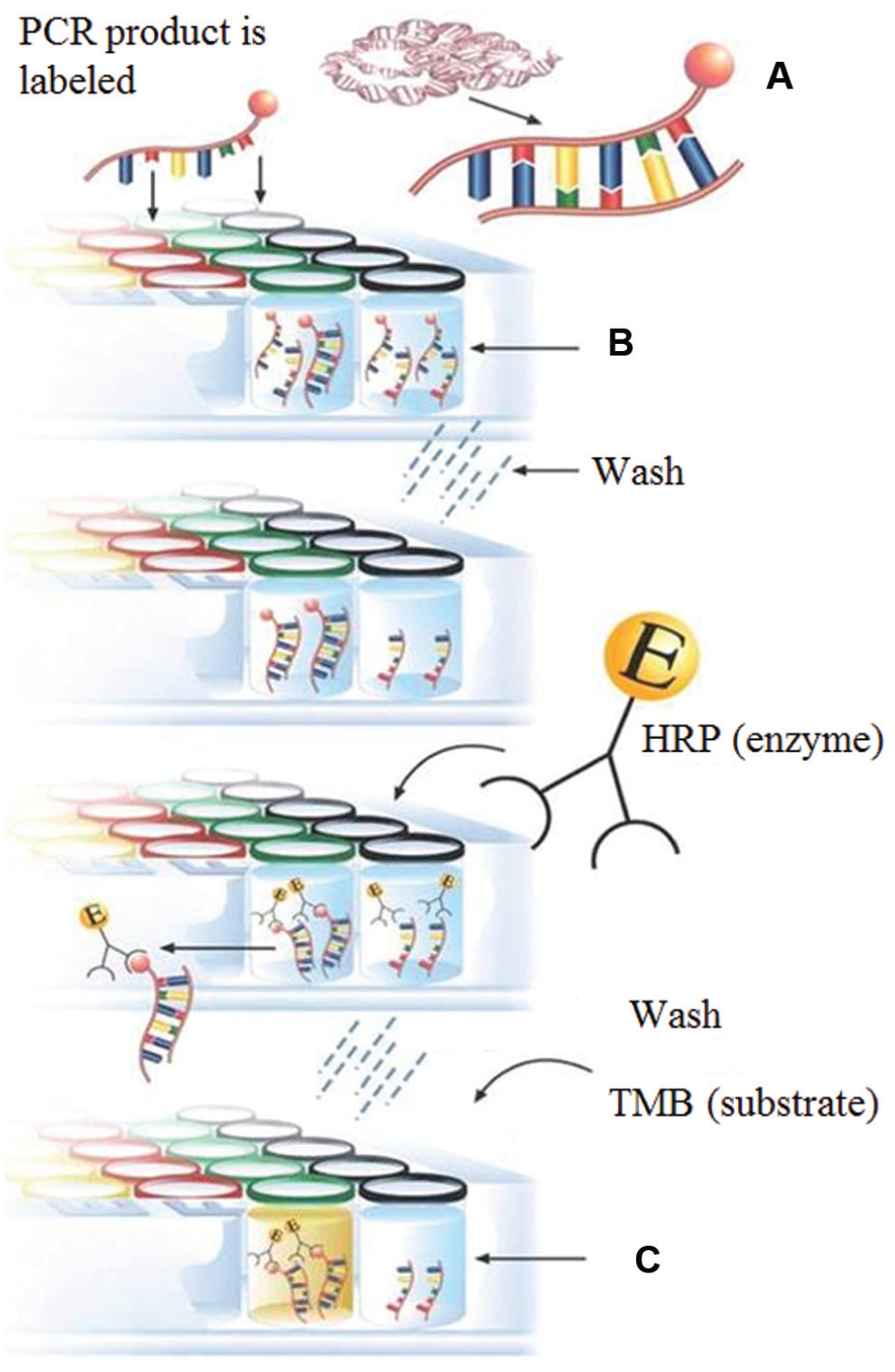
Hyplex sepsis blood screen assay. (
The Verigene bloodstream infection tests (Nanosphere Inc., Northbrook, IL) likewise employ multiplexed PCR amplification with end point detection through sandwich hybridization but on a microarray and in a more automated assay format, through an integrated cartridge and two instruments. The first instrument, the Verigene Processor SP, automates nucleic acid extraction from the blood culture sample, followed by PCR amplification. The amplified mastermix is then transferred into the Verigene Test Cartridge, where the amplicons are hybridized in a sandwich format to specific capture probes on a microarray and to gold nanoparticle-conjugated detection probes. Sensitive detection is achieved through silver staining, catalyzed by the gold nanoparticles on the microarray. The cartridge is then removed from the processor, and the microarray slide is detached and inserted into the second instrument, the Verigene Reader, which uses evanescent light scattering for detection.94–96 Nanosphere received U.S. FDA clearance for two tests to detect panels of gram-negative 95 and gram-positive bacteria, the latter including mecA, vanA, and vanB resistance genes.96,105,106 Nanosphere is further developing a Verigene yeast infection panel.
Higher levels of process integration and multiplexing can be accomplished using the CE-IVD–marked and U.S. FDA–cleared FilmArray blood culture identification panel (BioFireDx, Salt Lake City, UT), which identifies 24 gram-positive, gram-negative, and fungal pathogens, plus resistance markers to carbapenems, methicillins, and vancomycin.
The FilmArray system involves a pouch-based cartridge and a single instrument that executes sample preparation, nested PCR amplification, and real-time fluorescent detection in a fully integrated, automated, and easy-to-use format. The FilmArray cartridge ( Fig. 12 ) consists of a heat-sealed flexible pouch with multiple chambers and channels, connected to solid reservoirs for sample introduction and dry reagent storage along the top. The flexible pouch further incorporates a honey-comb shaped array of ~100 reaction chambers, with a gasket above the reaction chamber wells that acts as a flow restrictor. To execute the assay, sample and rehydration buffer is injected through septa into the cartridge. Because the cartridge interior is evacuated, the required volumes are aspirated automatically into the cartridge, without precision pipetting. Fluids are moved through the system via pistons and inflating bladders. Following sample introduction, pathogens are mechanically lysed via bead beating, and the liberated nucleic acids are extracted using silica-coated magnetic particles. The eluate-containing purified nucleic acids then reconstitutes mastermix reagents with broad specificity primers for the first round of the nested PCR reaction. The amplified mastermix is then diluted and combined with new mastermix reagents, upon which the solution is pumped into the wells of the honeycomb array. Each well contains a set of inner primers targeting a specific pathogen or genomic region. During the second round of PCR, amplicon formation is monitored in real time via an intercalating dye, with LED illumination and a CCD camera to image the wells. Multiplexed qPCR, therefore, in this case is based on monitoring a large number of single-plex reactions in parallel. The nested PCR format not only facilitates the specificity of the reaction, but it also increases the sensitivity, which would otherwise suffer in a split reaction scheme. The FilmArray can perform melt curve analysis to verify correct amplicon formation. The instrument performs automatic data analysis, and results can be obtained within approximately 1 h after sample introduction. All PCR products are contained in the sealed pouch, to mitigate amplicon carry-over contamination risk. 107
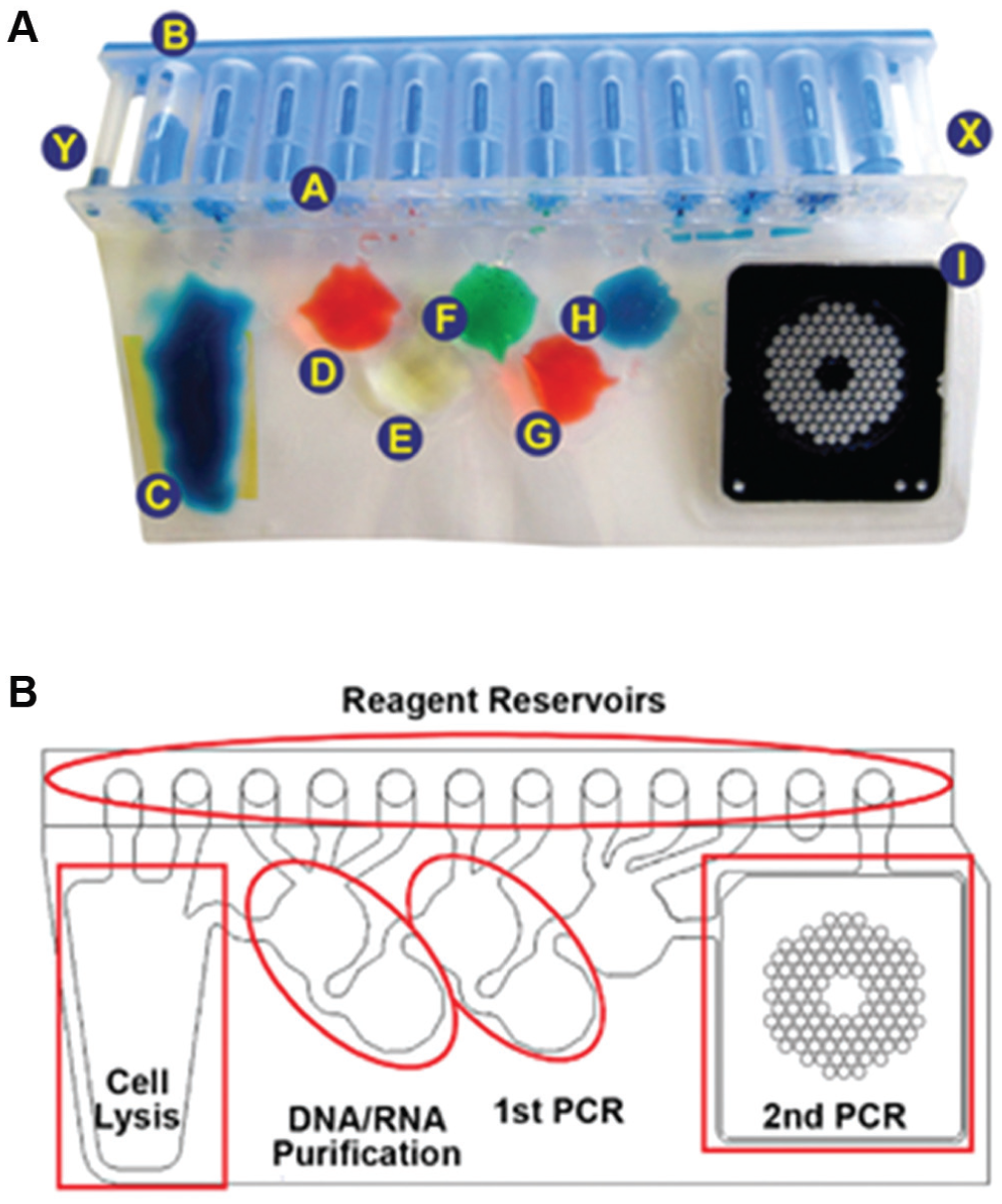
(
The clinical sensitivity of the FilmArray for pathogen identification appears to be slightly lower than are the sensitivities of the Hyplex and Verigene platforms ( Table 3 ), but all three systems have reasonable sensitivity and high specificity for pathogen ID. However, in these systems, the attainable level of multiplexing does not enable detection of all sepsis-causing pathogens, which means that only a subset of clinical samples can be properly classified. Furthermore, it is important to be able to readily expand the panel of pathogens that can be identified, due to the ever-changing set of pathogens responsible for sepsis. 17
Of all the systems listed in Table 3 , the highest level of multiplexing, and the ability to readily expand the panel of pathogens, can be achieved via the PLEX-ID BAC Spectrum blood culture assay. The PLEX-ID platform was developed by Ibis Biosciences (Carlsbad, CA) originally as the T5000, 99 and then acquired by Abbott in 2009. This system involves PCR amplification of specific regions within the pathogen’s genomic DNA, such as rRNA encoding genes and housekeeping genes, using broad range primers that target sequences conserved across multiple genera and species. The amplicons are then analyzed via electrospray ionization time-of-flight MS (ESI-TOF MS), to determine their molecular weight and base composition, which is then matched to a database of pathogenic base-composition signatures. 100 The PLEX-ID database reportedly contains signatures for all known bacteria, pathogenic families of fungi, and all families of disease-causing viruses. 99 This high inclusivity enables sepsis diagnosis with higher clinical sensitivity compared to other platforms with a more limited panel of targeted pathogens. Furthermore, the PLEX-ID can quantify the bacterial load of a sample. 101 However, the PLEX-ID platform is significantly more complex, larger, and more expensive than are other molecular diagnostic methods described herein. In addition to the ESI-TOF system, this system entails a separate bead beater for pathogen lysis, the Kingfisher Flex instrument (Thermo Scientific, Waltham, MA) for solid-phase nucleic acid extraction using magnetic beads, the EVO75 liquid-handling platform (Tecan, Mannedorf, Switzerland) for mastermix plate setup, plus a separate thermocycler for PCR amplification. MS is becoming more common in the clinical laboratory, and the performance of the PLEX-ID was found to be comparable to protein MS systems for sepsis pathogen identification, 102 discussed in the next section. However, the complex sample preparation and PCR amplification required upstream of MS analysis place the PLEX-ID at a significant disadvantage compare to protein MS.
Protein MS
In recent years, matrix-assisted laser-desorption ionization TOF MS (MALDI-TOF MS) has emerged as a novel technology to identify a wide range of pathogens that cause sepsis. MALDI-TOF MS for sepsis pathogen identification involves extracting acid-soluble proteins (2–20 kDa size range) from the pathogen, which are then analyzed by MS to generate a “mass fingerprint.” Pathogen identification (genus, species, or subspecies level) is based on matching MS patterns determined from the sample with mass spectra contained in a reference library.
A typical experimental flow for MALDI-TOF–based sepsis pathogen identification is shown in Figure 13 . Bacteria are isolated from positive blood culture media via differential centrifugation (to pellet all bacteria present in the sample) or via subculture. Next, formic acid protein extraction is applied to the isolated pathogen sample followed by spotting onto the MALDI plate. 109 Alternatively, colonies of single bacterial isolates obtained via subculturing can be applied directly onto the MALDI plate.110,111 In either case, the sample is overlaid with a MALDI matrix, such as an aqueous solution of 〈-cyano-4-hydroxycinnamic acid (HCCA). The sample is allowed to dry, and the plate is then inserted into the MALDI-TOF MS. A nitrogen laser pulse (337 nm wavelength) is used to excite the MALDI matrix molecules, which then pass energy to the protein molecules ( Fig. 14 ). The matrix aids in sample vaporization and improves ionization through proton transfer 110 and low-energy electrons generated within the matrix, 112 which contributes to ionization of protein molecules with limited fragmentation.
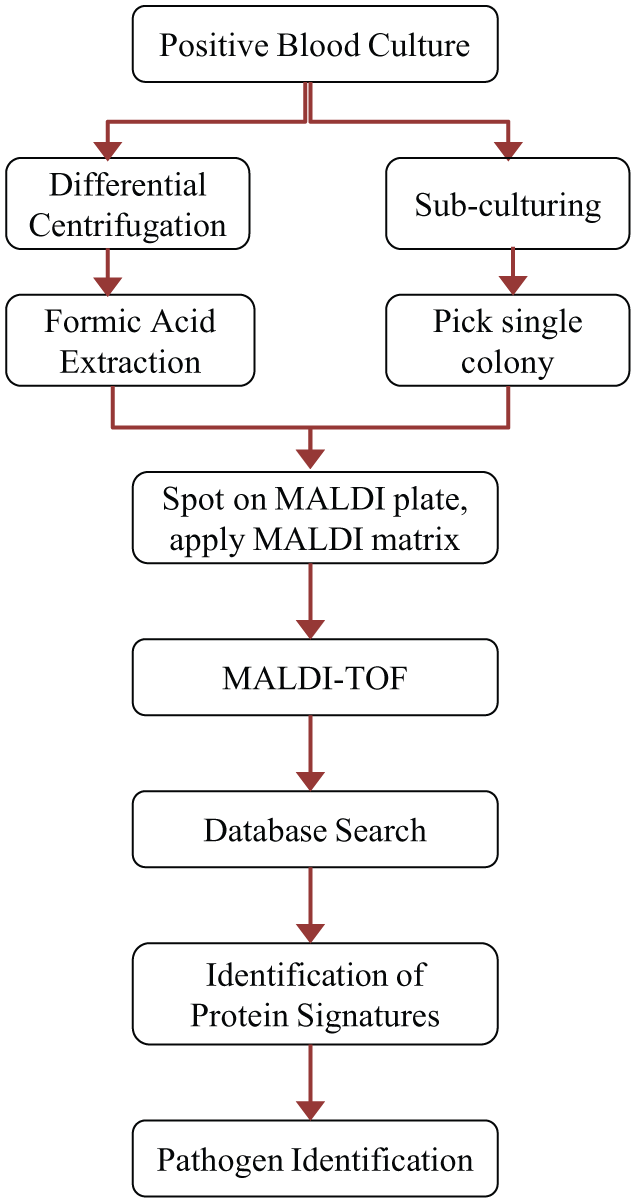
Sequence of steps used in sepsis pathogen ID using matrix-assisted laser-desorption ionization time-of-flight mass spectrometry (MALDI-TOF MS).
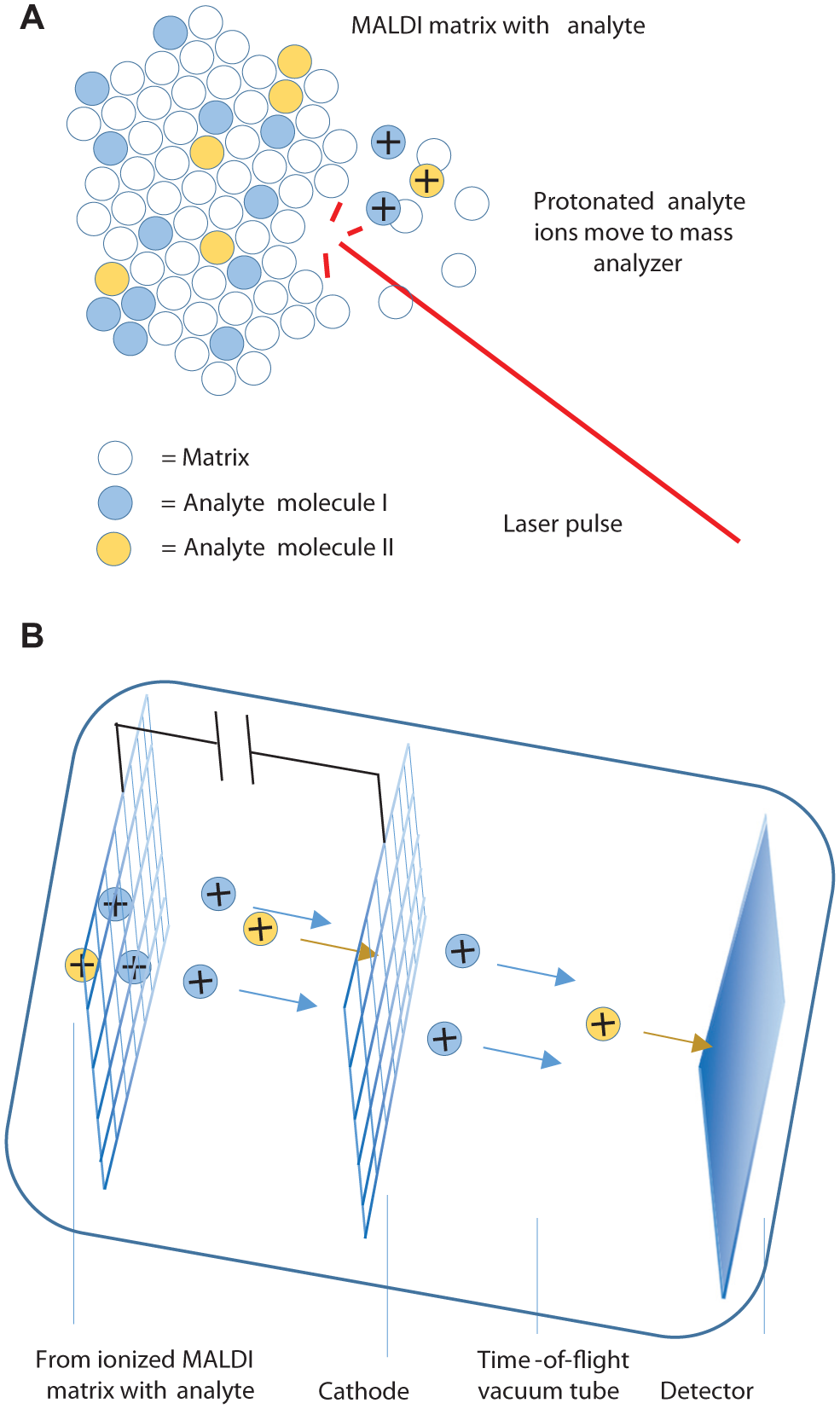
Principles of the matrix-assisted laser-desorption ionization time-of-flight (MALDI-TOF). (
Positive ions are accelerated through an electrostatic field and then travel through the mass analysis flight tube at a velocity inversely proportional to their respective masses. The detector records the time required for ions to reach the end of the tube. This value is used to calculate the ion’s corresponding mass-to-charge ratio and to generate the mass spectrum that effectively reveals the protein composition of the organism. 114 This mass spectrum is distinctive to the particular pathogen from which the proteins were extracted. The sample’s mass spectrum is searched against a reference database to enable pathogen identification.
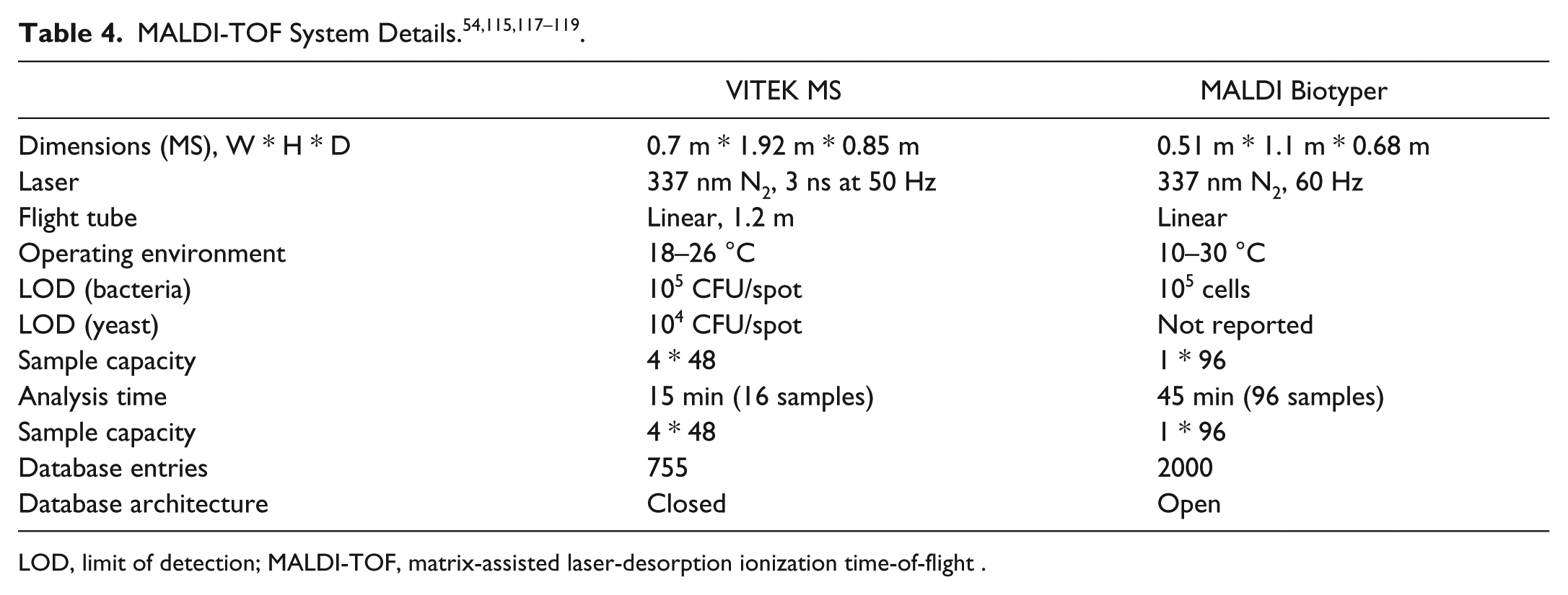
The two major MALDI-TOF MS platforms marketed worldwide for clinical microbiology are the VITEK MS (BioMérieux) and the MALDI Biotyper (Bruker, Billerica, MA), which are discussed in further detail herein. Both systems consist of a dedicated MALDI-TOF MS, various consumables, software used to input sample information and to process results, and a library of reference protein mass spectra.
The VITEK MS54,115 system is composed of the VITEK MS Prep Station, where bacterial colonies obtained via subculturing are spotted on barcoded MALDI slides; the VITEK MS (a rebranded Shimadzu AXIMA Assurance mass spectrometer and PC workstation) to perform the MALDI-TOF analysis; and the Myla data analysis server, which contains the reference database (Knowledge Base, generated from an average of 10 reference isolates per organism), to process the acquired data, and calculates identification results. BioMérieux uses a proprietary Advanced Spectra Classifier 116 that divides the m/z range from 3–17 kDa into 1300 “mass bins.” This makes MS library searches much less data intensive than in other systems that search against a complete mass spectral database, resulting in faster library searches. Other system details are listed in Table 4 . The VITEK MS is approved for clinical use in the United States (U.S. FDA cleared), Europe (CE-IVD marked), and several Asian and Latin American countries.
LOD, limit of detection; MALDI-TOF, matrix-assisted laser-desorption ionization time-of-flight .
The MALDI Biotyper system117,118 is composed of the MALDI Sepsityper kit, Bruker Microflex LT mass spectrometer, and the MALDI Biotyper Server/software bundle. As for the Vitek MS, colonies picked from a subculture plate can be applied directly onto the MALDI target plate. Alternatively, the Sepsityper sample preparation kit enables pathogen enrichment from a positive blood culture via centrifugation, followed by formic acid extraction and spotting onto the target plate. In either case, the dried sample spots are overlaid with the HCCA MALDI matrix. The plate is then inserted into the instrument for analysis. Sample lists (called “projects”) can be generated remotely using Biotyper client software communicating to the Biotyper Server. The Biotyper Server containing the reference database processes the acquired data and calculates results. Biotyper database entries are stored as “main spectra” that represent the average of multiple MS acquisitions from single-organism strains. The Biotyper database has an “open” architecture and is regularly updated with new spectra generated by researchers within the user community. Other system details are listed in Table 4 . The MALDI Biotyper is CE-IVD marked and approved for clinical use in Europe, Australia, New Zealand, several Asian countries, and Canada. Bruker recently received U.S. FDA clearance to market the Biotyper for identification of gram-negative bacteria from blood cultures in the United States.
The Vitek MS and MALDI Biotyper have each undergone extensive clinical validation ( Table 5 ), demonstrating accurate identification of a wide range of pathogens. In addition, recent third-party evaluations ( Table 6 ) directly compared the performance of these two systems. As the mass spectrometers are very similar, differences in identification accuracy most likely reflect differences between the two reference libraries and the algorithms used to generate a positive identification. Direct analysis of blood culture specimens was more difficult for both systems, which generally had an identification rate of 80–85%, compared to >90% for samples derived from solid culture media. Both systems also had significant difficulty with identification of anaerobes and enteric pathogens obtained from multiple clinical sources. 119 Polymicrobial blood cultures were especially problematic (identification rates of <25% 120 ). The VITEK MS was significantly more capable at distinguishing Streptococcus pneumonia from closely related nonpneumococcal species of the Streptococcus mitis group. 121
Individual Evaluations of the VITEK MS and the MALDI Biotyper.
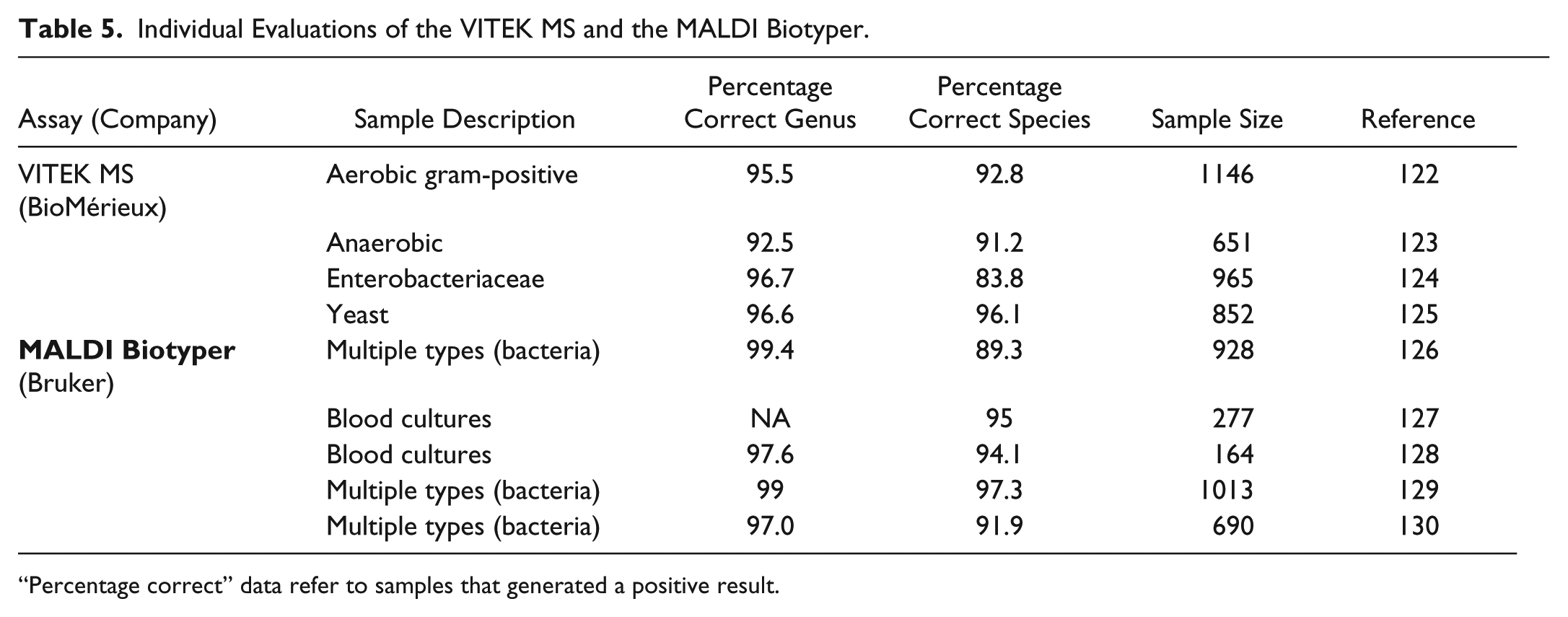
“Percentage correct” data refer to samples that generated a positive result.
Comparison between VITEK MS and MALDI Biotyper Performance.
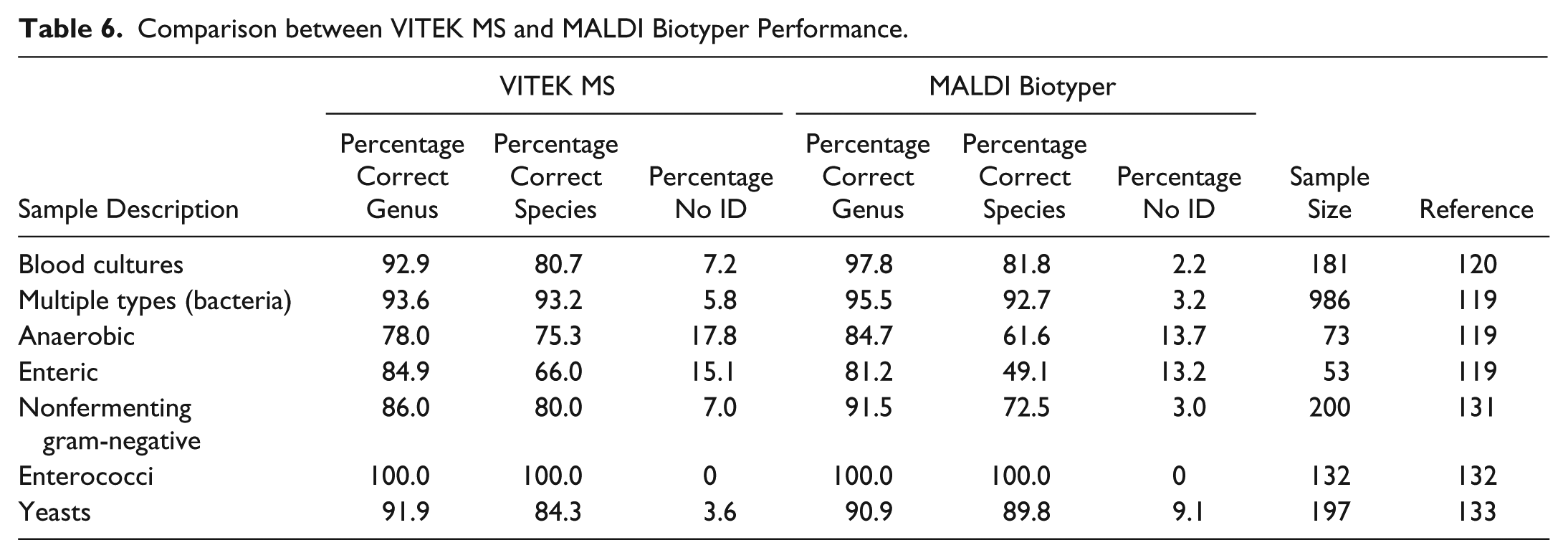
Discussion
Rapid identification of sepsis pathogens is critical to improving patient outcomes and is consequently a high priority for health care providers, payers, and patients. In addition to decreasing the time required for pathogen identification, systems requirements include the ability to identify a large panel of pathogens with high sensitivity and specificity, ideally using an automated and streamlined workflow. 134 Microbiological methods are very mature and universally accepted as the current gold standard. Tests and workflows readily accommodate mixed pathogens and lead to high-confidence identifications—but with substantial time required to obtain the final identification ( Fig. 9 ).
Molecular diagnostic methods can shorten the time to result, especially if the test can be performed directly from whole blood, without the need for blood culture. However, this is technically challenging due to the low pathogen concentration in blood and therefore tends to lead to low sensitivity and a low negative predictive value, meaning that nucleic acid amplification testing from whole blood cannot rule out sepsis. Better sensitivity for specific pathogen identification is obtained if testing is performed from a positive blood culture, which still reduces the time to result compared to traditional subculturing and pathogen identification ( Fig. 9 ). FISH technology can theoretically yield rapid identification. However, FISH assays are labor intensive and not amenable to high-level multiplexing; therefore, multiple tests have to be performed on each sample, extending the overall time to result.
The ability to achieve high levels of multiplexing is a key value driver for newer molecular diagnostic methods that have entered the market in recent years. Multiplexed PCR further enables characterization of mixed infections and identification of drug resistance markers. However, these PCR-based methods can be complex and labor intensive, unless fully integrated and automated as in the case of the FilmArray.
MS pathogen identification enables extremely high levels of multiplexing and can in theory identify all sepsis-causing pathogens in one experiment. Despite the substantial proteomic pathogen ID via complexity, MALDI-TOF MS is becoming established in routine clinical use. ESI-TOF MS has been coupled with nucleic acid testing in the PlexID platform, but proteomic methods essentially provide the same results with a much simpler and less expensive workflow.
BioMérieux and Bruker are the leading distributors of MALDI-TOF MS systems for sepsis pathogen identification; however, their core competencies and corporate strategies are quite different. Bruker is a leading provider of high-performance scientific instruments but is not a major IVD company. In contrast, BioMérieux is one of the top 10 global IVD companies and has been a market leader in the area of microbiological methods for sepsis pathogen identification and drug susceptibility testing, with platforms such as the BacT Alert and the Vitek 2. BioMérieux appears determined to maintain its position as market leader in this area, proactively pursuing new technologies to remain competitive. For example, BioMérieux acquired BioFire in Jan. 2014, adding the FilmArray molecular diagnostic platform to its portfolio. In 2010, BioMérieux entered into a partnership with Shimadzu to develop and launch a MALDI-TOF platform for sepsis pathogen ID, likely in response to the 2009 launch of the CE-IVD–marked Biotyper by Bruker for the same indication. This partnership resulted in the 2013 U.S. FDA clearance of the Vitek MS. According to Genome Web, 135 in mid-2013 there were approximately 400 VITEK MS systems installed worldwide compared to 800 MALDI Biotypers. Both systems are CE-IVD marked, but the VITEK MS has a current advantage in the U.S. market with its U.S. FDA approval for many important pathogen classes. Bruker recently gained approval for identification of gram-negative bacterial sepsis pathogens.
Despite the substantial instrument cost, MALDI-TOF–based pathogen identification is more cost effective on a per-test basis than are traditional microbiological or molecular diagnostic methods. MALDI-TOF MS can enable annual savings of $100 000 based on reduced reagent and labor costs, compared to the exclusive use of microbiological methods. 136 MALDI-TOF MS requires only generic and quite inexpensive consumables, such as organic solvents and MALDI matrix. These are provided as kits by the manufacturer of the platform but can also be readily obtained from alternative sources. For example, Chen et al. 120 compared data obtained using the Bruker Sepsityper Kit versus data obtained with in-house extraction protocol. Analytical performance was equivalent for the two sample-preparation methods, but the cost/sample was much cheaper with the in-house protocol, which used standard laboratory reagents ($15/sample for Sepsityper Kit, $1/sample for in-house protocol). Although advantageous for the consumer, this goes against the traditional business model in the IVD industry, where revenues are generated through markups on the consumables, whereas the profit margin on the instrument is modest.
Proteomic fingerprinting via MALDI-TOF MS significantly expedites pathogen identification, by 23–35 h compared to traditional culture-based methods.9,136,137 However, it is important to effectively integrate this technology with standard microbiology, especially with regard to AST. Although MALDI-TOF MS can enable AST, 138 these methods are still in development; thus, traditional culture-based AST has to be used in conjunction with MALDI-TOF pathogen identification, as both results are required to suitably adjust antimicrobial therapy. Furthermore, the overall time to result depends on how well MALDI-TOF pathogen identification is integrated into the overall hospital workflow. Perez et al. 12 assessed the impact of MALDI-TOF pathogen identification, effectively integrated within the hospital’s workflow and antimicrobial stewardship program, on the time to result, length of hospitalization, and health care expenditure for sepsis patients. 12 The time to sepsis pathogen identification following a culture-positive result was reduced from 36 h using traditional microbiology to 11 h using the MALDI-MS. In conjunction with more efficient result dissemination and timely treatment adjustment, this reduced the average length of hospitalization for the affected patients by 2.6 d and decreased the hospitalization cost per patient by almost $20 000. Therefore, MALDI-TOF MS is a cost-effective approach for sepsis pathogen identification, despite the high cost and complexity of the instrument.
Future Outlook and Conclusions
For sepsis diagnosis, overnight culture is typically required to selectively enrich the pathogen and increase the sensitivity, but overnight culturing prolongs the time to result. Novel microfluidic technologies have been used to sort and concentrate bacterial 139 or fungal 140 pathogens from whole blood. Such microfluidic pathogen enrichment can be combined with fluorescence detection. 141 Individual bacteria present in plasma can further be confined in nanodroplets, which localizes the signal and thus aids in detection. 142 Bacteria from blood can further be captured and imaged inside microfluidic channels. 143 These technologies are still in the early stages of development but show promise for rapid and sensitive sepsis diagnosis. To achieve specific pathogen identification requires high levels of multiplexing. It is currently not clear what level of multiplexing is attainable in such microfluidic devices. Furthermore, sepsis diagnosis typically requires processing of several milliliters of sample, which is a problem in most microfluidic devices.
Sepsis pathogen ID/AST methods based on nucleic acid amplification testing have significantly advanced in recent years ( Table 3 ). However, pathogen ID directly from whole blood with suitable clinical sensitivity and specificity remains a challenge. Another driver is increasing the attainable level of multiplexing to enable detection of all common causative pathogens, without significantly increasing the system’s complexity and cost. Next-generation sequencing technologies have evolved into systems that can be used in a clinical laboratory and may facilitate sepsis pathogen ID and genotypic drug resistance testing at very high levels of multiplexing, with reasonable cost and complexity. 144
Proteomic MS has emerged as a novel and powerful tool for sepsis pathogen ID. However, challenges remain related to identifying certain pathogens without a distinguishable unique spectral fingerprint 119 and properly diagnosing mixed infections. 120 Lipidomic MS may be used as a complementary tool to help resolve such borderline cases, 145 as unique lipid compositions are observed for different pathogens.146,147 In addition, it is likely that the capabilities of MS will expand beyond sepsis pathogen ID to also enable AST. 138
In conclusion, the field of sepsis pathogen ID/AST entails a broad range of established and emerging technologies. This field is expected to further evolve in the coming years, as novel technical approaches mature and transition to clinical use.
Footnotes
Acknowledgements
We thank Dr. Jim Osborne for helpful comments regarding the manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.

