Abstract
Postmortem bacterial culture is controversial in human medicine, and veterinary-specific research in this area is lacking. To address this knowledge gap, we cultured liver, kidney, and spleen individually from on-farm calf mortalities to determine the number of bacterial species present, concordance between organ cultures, and agreement with gross and histologic findings. We hypothesized that the spleen, a filtering organ, would be the most useful organ with the least amount of postmortem contamination given that it does not have a direct conduit to a bacterial population. Fresh liver, kidney, and spleen were collected for culture from 30 calves 5–28-d-old with various causes of mortality. Bacterial growth of ≥2 species was observed in ~48% of cultures, with Escherichia coli and Streptococcus spp. being most frequent. One bacterial species was present in 20% of cultures, with E. coli predominating. No growth was observed in ~32% of cultures. In 43% of cases, there was agreement in the culture results for all 3 organs; however, the majority were mixed bacterial growth. The best agreement was observed when there were no gross and/or histologic septic lesions in target organs and no bacterial growth on culture. The spleen was not helpful in determining bacterial significance in comparison to kidney or liver.
Culture of abdominal organ samples collected at postmortem is performed routinely in veterinary medicine to identify a bacterial pathogen when clinical signs or gross lesions indicate an infectious disease. However, the diagnostic utility of postmortem bacterial culture is controversial in human medicine and largely unexamined in veterinary medicine. In human medicine, causes of, and ways to reduce, postmortem contamination have been investigated, and strategies to improve the accuracy of postmortem bacteriology have been explored.3,4,9 Approaches investigated include reduction in the time to sampling in the postmortem interval, minimizing cadaver manipulation, maintenance of body temperature at ~4°C, and performing autopsies using sterile surgical technique.3,4,12 Additionally, comparison of antemortem blood culture with postmortem culture has been used to differentiate contamination from diagnostically important growth.3,4,12
Despite strategies used to reduce contamination, the usefulness of postmortem cultures has been called into question by several studies in which culture consistently resulted in polymicrobial growth, largely indicative of contamination.1,7,9,12 A comparison of ante- and postmortem blood cultures from 111 patients showed that, in cases in which clinically important bacteria or fungi were identified antemortem, 35% of the postmortem blood cultures yielded the same result, but 50% identified only contaminants; 54% of postmortem blood cultures were positive for microbial growth despite a noninfectious cause of death. 12 Interestingly in that study, 15% of antemortem blood culture–positive cases identified other organisms considered significant to the patient’s clinical disease; however, this did not change the final diagnosis. 12 The ultimate conclusion of this comparison was that little was gained from postmortem culture given that it did not change or refine the diagnosis, or definitively aid in evaluation of appropriateness of therapy. 12 Taken together, the results from the literature that we reviewed indicated that postmortem culture has limitations for determining the presence or cause of infection.
Collectively, 42% of calf loss is attributed to infectious disease in the United States, usually in the form of respiratory or enteric disease, which can lead to septicemia and ultimately death.10,11 Given the severity of outcome, identification of a bacterial pathogen or pathogens is often attempted to aid treatment and guide farm management. However, in agricultural animal medicine, as opposed to human medicine, controlling postmortem variables is challenging. Field autopsy is common, and the postmortem interval is often prolonged by several hours or even days after death. Environmental conditions of the carcass are uncontrolled in field settings, and carcasses may have been moved by heavy machinery. Even when sample collection for bacterial culture is done in a manner that was as sterile as possible, the nature of the field autopsy means that environmental and postmortem contamination is highly likely. 5 Given these inherent challenges, the diagnostic value of postmortem cultures may be problematic and lead to overinterpretation of the significance of the bacteria identified. Given that treatment of in-contact herdmates may be based on postmortem culture findings, a better understanding of the usefulness of culture is essential to promote antimicrobial stewardship. If little actionable information results from abdominal organ culture, then resources, including the materials, time, and expertise needed for culture sampling, may be better directed in other avenues.
We assessed the diagnostic utility of abdominal organ culture by evaluation of the number of bacteria isolated per organ culture, agreement between organ cultures from the same animal, and agreement with gross and/or histologic indicators of infection. Additionally, the spleen has been proposed to be the most diagnostically useful organ for postmortem culture because it does not have a conduit to a contaminating environment, such as the biliary system in the liver or the ureters and urethra for the kidneys. 8 Based on postmortem research in human medicine to date, we hypothesized that the spleen would be the most diagnostically useful organ in calves to identify a bacterial pathogen. As well, we defined postmortem contamination as mixed bacterial growth (≥2 organisms detected).
We conducted a study of a commercial calf ranch in the western United States in May and June of 2018. Our research protocol was reviewed and approved by the Institutional Animal Care and Use Committee of Washington State University (ASAF 6414). The ranch housed ~25,000 Holstein, Jersey, and crossbred heifer calves from multiple dairy farms, with animals up to 200-d-old. We convenience-sampled 30 calves 5–28-d-old with a history of trauma, diarrhea, bloat, or behavioral changes. Calves that had died overnight (between 5 pm and 7 am) were autopsied by a veterinary pathology resident and trained 2nd-year veterinary student interns, using a standard protocol (developed by C. McConnel), the following morning (≤ 17 h after death) to avoid autolysis caused by summer daytime temperatures. Morbidity and mortality records included broad disease diagnoses, such as enteritis or pneumonia, and associated treatments, and were managed on-farm using electronic records (DairyComp 305; Valley Agricultural Software) with a secondary program providing record oversight and health data compilation (HEALTHSUM Syndicate). Antimicrobial therapy had been administered to 25 of the 30 calves and included one or more of the following: cephalosporins, fluoroquinolones, florfenicol, and/or sulfamethoxazole, depending on clinical signs and label indication. A modified death certificate was created to document the presumed cause of calf death6,11 and was completed by research staff and on-farm veterinary staff for each autopsied calf on the day of the autopsy; the certificate included details of treatments, gross findings, and significant postnatal issues or conditions contributing to the mortality phenotype. On-farm autopsy-based identification of gross lesions consistent with sepsis was determined with input from interns, on-farm veterinarians, and the principal investigator (C. McConnel) at autopsy if some or all the following criteria were met: perforating typhlitis; septic peritonitis; miliary lesions in kidneys, liver, or heart; interstitial or embolic pneumonia.
At autopsy, the abdomen of each study calf was opened with sterile instruments; samples of liver, kidney, and spleen were collected with sterile instuments, placed in sterile containers and transported to the Washington Animal Disease Diagnostic Laboratory (WADDL; Washington State University, Pullman, WA, USA). Samples were shipped on ice packs and arrived at the laboratory 24–48 h post-collection. The liver, kidney, and spleen were handled in a biosafety cabinet and sampled for bacteriologic culture on the day of arrival. Samples were seared and opened with sterile scissors to expose internal tissue. This tissue was plated on Columbia blood agar (CBA) and MacConkey agar (Hardy Diagnostics) per standard operating protocols. Plates were streaked and incubated for 18–24 h at 35°C, except that CBA plates were incubated in 5% CO2. Cultures were read by WADDL microbiologists per standard operating procedures at 24, 48, and 72 h, and all growth was recorded.
Bacteria were identified by mass spectrometry (MS; MALDI Biotyper, Bruker) following the manufacturer’s extended direct transfer protocol. 2 In general, confident species identification scores were 2.3–3.0, confident genus identification scores were 2.0–2.29, probable genus identification was 1.7–1.99, and no reliable identification was indicated by a score <1.7. Isolates unable to be identified by the MALDI-TOF MS system were Gram-stained and reported according to the Gram stain result and morphology.
Formalin-fixed samples (heart, lung, kidney, spleen, lymph node, thymus, liver, select segments of the gastrointestinal tract) were collected for histologic analysis. Histologic assessment was conducted by 1 of 3 board-certified veterinary anatomic pathologists on standard H&E-stained slides that were anonymized concerning cause of death. Overt histologic evidence of sepsis was defined as the presence of one or more of the following lesions: random suppurative or necrotizing hepatitis, suppurative interstitial nephritis or glomerulitis, interstitial pneumonia or pulmonary microthrombosis without broncho- or bronchointerstitial pneumonia, or acute infarction of various organs (such as the intestines).
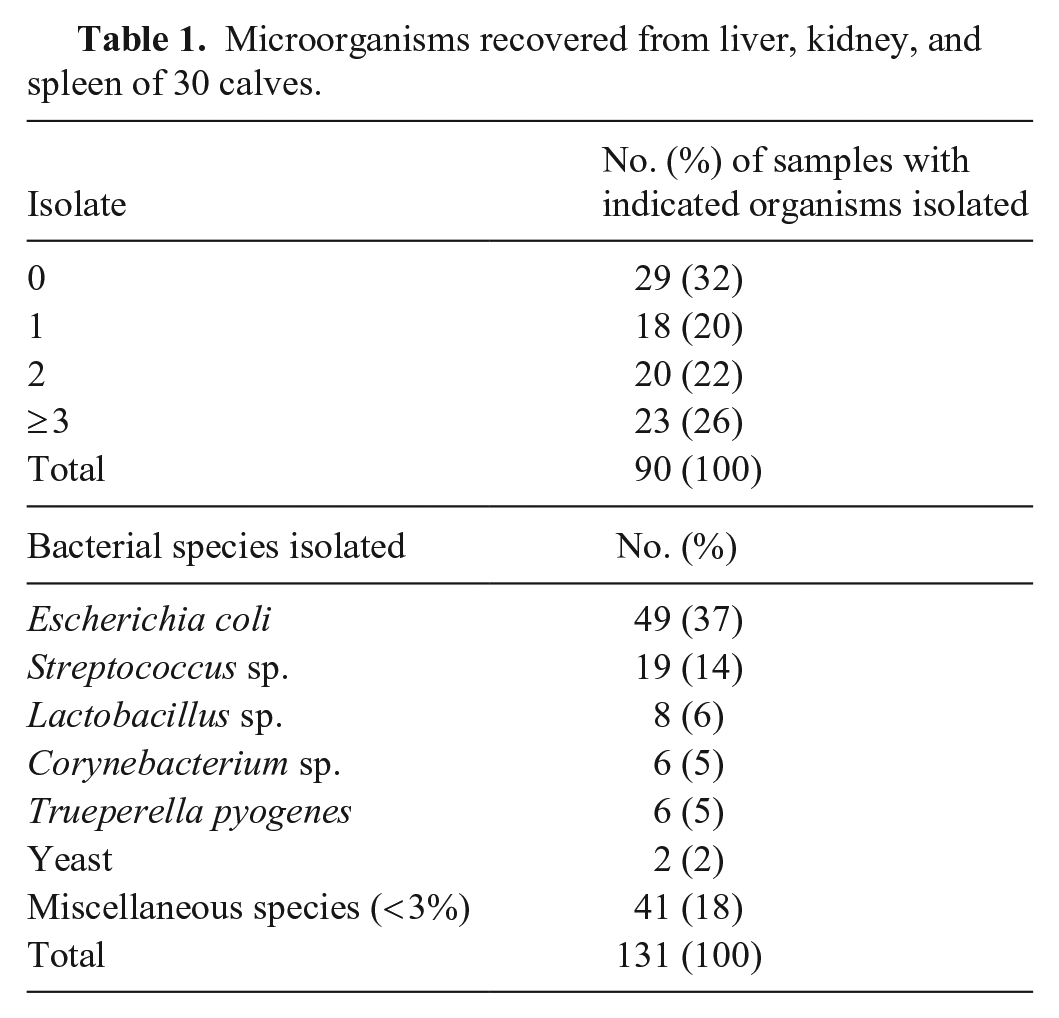
We cultured 90 samples (30 livers, 30 kidneys, 30 spleens) from the 30 cases; 18 cultures had pure bacterial growth, 20 had 2 organisms, 23 had ≥3 organisms, and 29 had no growth (Table 1). We isolated 131 organisms (39 unique) from the 90 samples, an average of 1.46 isolates per organ. Escherichia coli was the organism isolated most commonly (49 of 131; 37%) followed by Streptococcus sp. (19 of 131, 14%; Table 1). Either E. coli, Streptococcus sp., or both were present in 46 of 90 (51%) samples. Both E. coli and Streptococcus sp. were present in 15 of 46 (32%) samples. Consistent culture findings were observed across the 3 organs from the same animal in 12 of 30 (40%; Table 2) cases. In 7 of the 12 (58%) cases, all samples had mixed bacterial growth, which may or may not have been the same bacterial species but was ≥2 different species per organ. In 3 of the 12 (25%) cases, all samples had no bacterial growth. In 2 of the 12 (17%) cases, all samples had pure bacterial growth (Table 2). It is important to note that all organisms isolated would be considered opportunistic pathogens and/or commensal organisms. No primary pathogens were identified in either mixed or pure culture.
Microorganisms recovered from liver, kidney, and spleen of 30 calves.
Comparison of evidence of gross and microscopic sepsis to bacterial culture of abdominal organs of 30 calves.
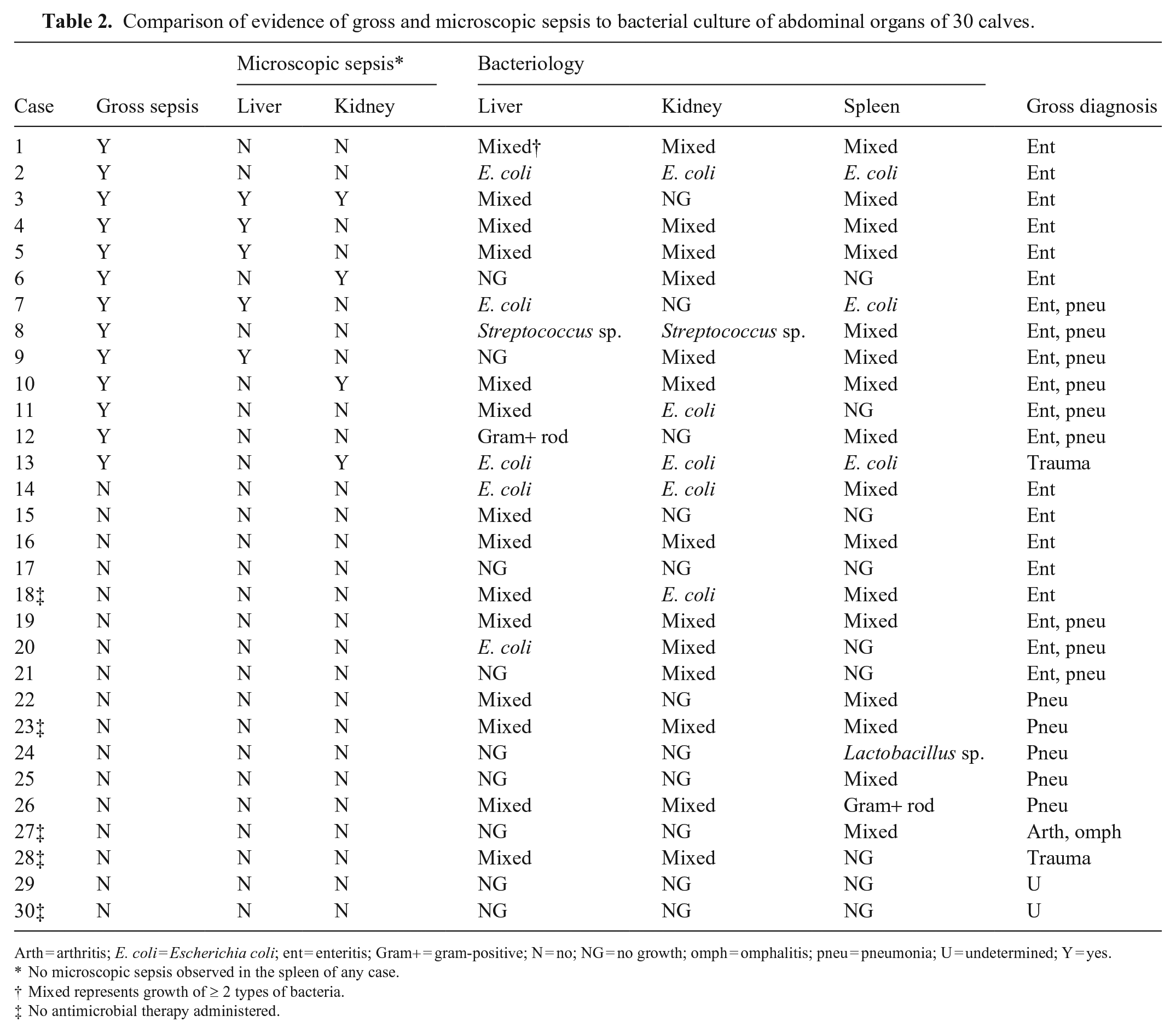
Arth = arthritis; E. coli = Escherichia coli; ent = enteritis; Gram+ = gram-positive; N = no; NG = no growth; omph = omphalitis; pneu = pneumonia; U = undetermined; Y = yes.
No microscopic sepsis observed in the spleen of any case.
Mixed represents growth of ≥2 types of bacteria.
No antimicrobial therapy administered.
Gross lesions indicative of sepsis were present in 13 of 30 (43%) cases, and were grouped by gross diagnosis, including enteritis, pneumonia, enteritis and pneumonia, trauma, not determined, and other (arthritis and/or omphalitis; Table 2). Histologic lesions indicative of sepsis were present in 8 of 30 (27%) cases. Neither gross nor histologic lesions consistent with sepsis were found in 17 of 30 (57%) cases (Table 2). In 2 of 30 cases, agreement was observed between gross and/or histologic lesions and pure bacterial growth (Table 2). In the 13 cases that had gross evidence of sepsis, only 8 (62%) cases also had histologic lesions in ≥1 sample, which suggests over-interpretation of tissue changes at autopsy. Conversely, histologic lesions were not observed in cases that did not have suggestive gross lesions.
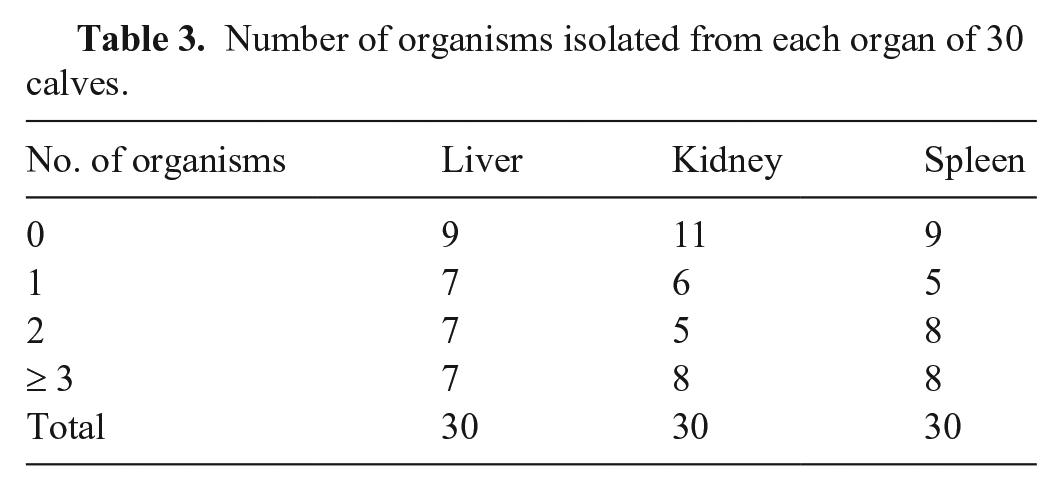
We had hypothesized that the spleen would be the most useful organ for evaluation of a bacterial cause of sepsis (defined as pure bacterial growth or no growth, in agreement with the presence or absence of gross and histologic lesions) with the least amount of postmortem contamination. The level of bacterial growth was similar across the organs, and no one organ had a dramatically different growth pattern than the other two. We did not find that the spleen was a superior organ for evaluation of a bacterial cause of sepsis compared with the liver or kidney. Ultimately, no organ had less postmortem contamination than the others (Tables 2, 3).
Number of organisms isolated from each organ of 30 calves.
In our study, pure bacterial growth was not correlated with gross or histologic lesions consistent with sepsis in postmortem samples of liver, kidney, or spleen from calves. In only 2 of 30 cases were there gross lesions consistent with sepsis, histologic lesions consistent with sepsis, and pure bacterial growth. However, pure bacterial growth was evident in 6 cultures (2 from each of liver, kidney, and spleen) that showed no gross or histologic evidence of a septic process. In addition, the discordance among organs of the same case introduces skepticism in determining a primary pathogen, even in the event of pure bacterial growth in one organ but not in others. The higher prevalence of pure bacterial culture when there was no gross or histologic evidence of a septicemic process further undermines confident identification of pathogens in the liver, kidney, and spleen. The prevalence of mixed bacterial growth poses additional difficulties in differentiation of contamination from a polymicrobial process that may very well be occurring if bacteria translocate across damaged mucous membranes.
In our cohort, antimicrobial treatment had been given to 25 calves as a result of various diagnoses; 13 with gross and microscopic lesions of sepsis, and 11 with no lesions of sepsis (Table 2). Both groups had similar culture results of mixed bacterial growth from one or more organs (10 vs. 9 calves). Interestingly, the 5 calves that did not receive antimicrobial therapy had no signs of gross or microscopic sepsis (Table 2). Antimicrobial therapy did not seem to impact the likelihood of mixed-growth cultures, but more data are needed. No-growth cultures were present in 22 of 51 (43%) samples when no evidence of gross and/or histologic lesions was present in the case, in comparison to those with gross or histologic lesions (7 of 36; 17%). Thus, negative bacterial culture may be more useful in ruling out sepsis as a potential cause of death.
Our results challenge the usefulness of the common practice in veterinary diagnostic medicine of routinely culturing abdominal organs of calves. Further studies are needed to identify appropriate case criteria for when to pursue culture, such as signalment, clinical signs, type of gross lesion(s), or pre-test probability. Additionally, standardized evaluation of sample quality, and stakeholder education in this area, are needed to ensure that high-quality samples are utilized. The use of antimicrobial drugs may influence pathogen detection, but is often not supplied in submission information. Our study modeled a real-life situation in which many calves have a history of antimicrobial use and, again, reinforces the potential for meaningless bacterial culture results. Applying appropriate clinical and sample integrity criteria when performing cultures of postmortem samples would increase the diagnostic utility of culturing and decrease unnecessary financial, time, and material costs to clients and the testing laboratory while maintaining good antimicrobial stewardship.
Footnotes
Declaration of conflicting interests
The authors declare no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Funding was provided by the American Jersey Cattle Association, the USDA National Institute of Food and Agriculture (Animal Health & Disease Research Capacity Grant 1014680; Agriculture and Food Research Initiative Competitive Grant 2019-68008-29897), and the Washington State University Caldwell Endowment.
