Abstract
In the modern molecular diagnostic laboratory, cost considerations are of paramount importance. Automation of complex molecular assays not only allows a laboratory to accommodate higher test volumes and throughput but also has a considerable impact on the cost of testing from the perspective of reagent costs, as well as hands-on time for skilled laboratory personnel. The following study tracked the cost of labor (hands-on time) and reagents for fluorescence in situ hybridization (FISH) testing in a routine, high-volume pathology and cytogenetics laboratory in Treviso, Italy, over a 2-y period (2011–2013). The laboratory automated FISH testing with the VP 2000 Processor, a deparaffinization, pretreatment, and special staining instrument produced by Abbott Molecular, and compared hands-on time and reagent costs to manual FISH testing. The results indicated significant cost and time saving when automating FISH with VP 2000 when more than six FISH tests were run per week. At 12 FISH assays per week, an approximate total cost reduction of 55% was observed. When running 46 FISH specimens per week, the cost saving increased to 89% versus manual testing. The results demonstrate that the VP 2000 processor can significantly reduce the cost of FISH testing in diagnostic laboratories.
Introduction
To accommodate an ever-expanding menu of molecular tests, pathologists and cytogeneticists are constantly seeking new approaches for increasing assay throughput while decreasing assay and labor costs. Automation of molecular testing can decrease the hands-on time required for highly skilled laboratory personnel to perform various assays while increasing testing volumes in a standard laboratory shift. In addition, automation can also allow for increased standardization as well as a reduction of reagents used per run, which increase the overall efficiency of molecular laboratories. Fluorescence in situ hybridization (FISH) is often considered a labor-intensive and hence costly molecular assay for labs to implement.1–4 Nonetheless, FISH is considered the gold standard for numerous molecular analyses and has become an integral part of many molecular laboratories’ assay portfolios.2,4–6 This is underscored by the emergence of novel targeted therapies in recent years.7,8 The current study aimed to examine the cost-effectiveness and reproducibility of FISH testing by automation with the VP 2000 processor (Abbott Molecular), a deparaffinization, pretreatment, and special staining instrument that can accommodate up to 50 FISH specimens in a single run. Over a 2-y period, between 2011 and 2013, the Pathology and Cytogenetics Laboratory at the General Hospital of Treviso in Italy monitored the time required (and associated labor costs) as well as the reagent volumes required to process a variety of FISH specimens (from lower throughput, such as 6 assays per week, to higher volume testing, such as 40+ assays per week). Results indicated that even for experienced laboratory technicians, the cost associated with FISH testing scaled linearly as testing volumes increased. In contrast, FISH automation with the VP 2000 processor provided flexible automation for batch sizes up to 50 FISH specimens per run, at no incremental cost. Even a lower throughput of specimens (four to six FISH tests per week) achieved cost saving, where an approximate total reduction in FISH testing costs, including hands-on time and reagents, was on the order of 55% when processing 12 FISH assays per week, 77% when 24 FISH assays were run per week, and up to 89% when more than 46 FISH assays were processed per week.
Materials and Methods
FISH Specimens
Specimens for the study were routine formalin-fixed, paraffin-embedded (FFPE) tissue preparations sent to the pathology and cytogenetics laboratory at General Hospital in Treviso, Italy, between 2011 and 2013. The specimens covered a wide range of sample types including breast cancer (surgical resections, n = 320; ago-biopsies, n = 90; and lymph nodes, n = 7), gastric cancer (surgical resections, n = 6 of stomach; esophageal biopsies, n = 18), non–small-cell lung cancer (surgical resections, n = 40; bronchial biopsies, n = 98), gliomas (glioblastomas, n = 9; medulloblastomas, n = 6; astrocytomas, n = 24; oligodendrogliomas, n = 44; and mixed oligoastrocytomas, n = 42), sarcomas (myxoid liposarcoma, n = 59; Ewing sarcoma, n = 94; alveolar rhabdomyosarcoma, n = 12; synovial sarcoma, n = 61; endometrial stromal sarcoma, n = 23; dermatofibrosarcoma protuberans, n = 20), and non-Hodgkins lymphoma (lymph nodes from mantle cell lymphoma, n = 7; follicular lymphoma, n = 14; mucosa-associated lymphoid tissue (MALT) lymphoma, n = 4; and Burkitt’s lymphoma, n = 14). Specimens were processed with various FISH assays using laboratory-validated methods, following manufacturer-recommended protocols.
FISH Probes
Vysis LSI DDIT3 Break Apart FISH Probe Kit (used for 18 myxoid liposarcomas), Vysis LSI EWSR1 Break Apart FISH Probe Kit (used for 94 Ewing sarcomas), Vysis LSI FOXO1 Break Apart FISH Probe Kit (used for 12 alveolar rhabdomyiosarcomas), Vysis LSI FUS Break Apart FISH Probe Kit (used for 9 myxoid liposarcomas), Vysis LSI SS18 Break Apart FISH Probe Kit (used for 61 synovial sarcomas), Vysis 1p36/1q25 and 19q13/19p13 FISH Probe Kit (used for 44 oligodendrogliomas and 42 oligoastrocytomas), Vysis EGFR/CEP 7 FISH Probe Kit (used for 13 astrocytomas and 9 glioblastomas), Vysis CDKN2A/CEP9 FISH Probe Kit (used for 7 astrocytomas), Vysis PTEN (10q23)/CEP10 FISH Probe Kit (used for 4 astrocytomas), Vysis LSI MYC Dual Color Break Apart Rearrangement Probe (used for 14 Burkitt’s lymphomas), Vysis C-MYC Spectrum Orange FISH Probe Kit (used for 6 medulloblastomas), Vysis IGH/CCND1 DF FISH Probe Kit (used for 7 mantle cell lymphomas), Vysis IGH/BCL2 DF FISH Probe Kit (used for 14 follicular lymphomas), Vysis BIRC3/MALT1 DF FISH Probe Kit (used for 4 MALT lymphomas), Vysis IGH/MYC, CEP8 Tri Color Dual Fusion Traslocation Probe (used for 14 Burkitt’s lymphomas), and Vysis ALK Break Apart FISH Probe Kit (used for 138 NSCLCs) were obtained from Abbott Molecular (Abbott Park, IL). SPEC MDM2/CEN12 Dual Color Probe Zytolight (used on 32 myxoid liposarcomas) and SPEC JAZF1 Dual Color Break Apart Probe Zytolight (used on 23 endometrial stromal sarcomas) were obtained from Zytomed (Bremerhaven, Germany). YWHAE Break Apart Probe Set (used on 23 endometrial stromal sarcomas) and PDGFB Break Apart Probe Set (used on 20 dermatofibrosarcoma protuberans) were obtained from Empire Genomics (Buffalo, NY), and HER2 FISH Pharm Dx (used on 417 breast cancers and 24 gastric cancers) was obtained from Dako (Glostrop, Denmark).
Other Reagents and Instrumentation
Paraffin Pretreatment Reagent Kit I, VP 2000 Reagents Protease I, VP 2000 Reagents Protease Buffer, 20XSSC, NP40, DAPI I, and the VP 2000 Processor were obtained from Abbott Molecular. Xylene was obtained from Histo-Line Laboratories (Pantigliate, Italy). Ethanol was obtained from Sigma Aldrich (Milan, Italy). Formalin was obtained from Mondial (San Giorgio Monferrato, Italy). Hydrochloric acid was obtained from Carlo Erba Reagents (Cornaredo, Italy). Coplin jars were obtained from Vetrotecnica (Padova, Italy). Rubber cement (Royal Talens, Netherlands) was used to seal the cover glass to the microscope slides during the co-denaturation phase of the protocol.
Transferring Procedures from Manual FISH to Automated FISH
A number (six in total) of automated protocols were investigated prior to realizing the final protocol described below. To facilitate this, the manual protocol was compared with the automated protocols using the same specimens in various runs, which included two breast cancer specimens (one surgical resection and one ago-biopsy), one glioma, two sarcomas, and one lymphoma. There was enough material from these specimens to compare various VP 2000 and the manual protocols to find an optimal procedure to proceed in the study. Although it is beyond the scope of the current work to describe each of the pretest protocols in detail (the final standardized protocol is described below), various conditions on the VP 2000 instrument were examined, including length of time and/or temperatures for xylene deparaffination, baking of slides prior to pretreatment, protease digestion, and buffer and ethanol washes. Once a protocol was achieved that resulted in equivalent specimen processing and FISH signal intensities, that protocol was used for the duration of the study (described and discussed below).
Cost Analysis
Reagent costs were calculated as a result of the volumes and amounts of reagents required to process 6, 12, 18, 24, 30, 36, 40, and 46 FISH specimens, manually or on the VP 2000 processor. As the amount of FISH probe used did not differ between manual and automated protocols, the cost of FISH probes was not factored into the costs here. Costs for hands-on time were calculated assuming a standard Italian laboratory personnel cost of 19 EUR per hour (category D3 technician), multiplied by the actual time the technician was manually handling the specimens (6, 12, 18, 24, 40, and 46 FISH specimens, manually or on the VP 2000 processor). Times for specimen analysis and interpretation were not considered in this study, as it was focused on the pretreatment automation of FISH testing. While the VP 2000 processor can process up to 50 slides, 46 specimens was the maximum number examined to allow for direct comparison to the manual method (i.e., the number of specimens that can be fit within multiple Coplin jars comfortably).
Results
Labor Costs to Run FISH
The hands-on time for laboratory personnel to run a FISH assay varied to some degree, depending on the experience of the technician. The time for processing FISH manually included the setting up of buffers and reagents and the cleanup time upon procedure completion, along with a typical pretreatment workflow that was validated at the Anatomical Pathology and Cytogenetics Laboratory at the General Hospital of Treviso. Manual pretreatment involved baking the slides for 15 min at 74 °C, deparaffination of the FFPE specimens in xylene (3 × 15 min), ethanol washing to remove the xylene (2 × 5 min), drying the specimens (3 min at 47 °C), pretreating the specimens to reverse formalin cross-links (20 min in 0.2 N HCl, 3 min in distilled water, 3 min in wash buffer, 30 min in pretreatment solution at 80 °C), buffer (2 × 5 min) washing to remove the pretreatment solution, protease digestion (30 min at 37 °C) to make the chromosomal DNA accessible to the FISH probes and to remove chromatin compaction, washing buffer (2 × 5 min) to remove excess protease, drying the specimens (3 min at 47 °C), formalin dehydration of the specimens (10 min), washing of excess formalin with buffer (2 × 5 min), and finally drying the specimens (3 min at 47 °C). This concluded the manual workflow for what we termed the “first day” of the manual FISH protocol (209 min total). FISH probes were placed on the specimens, following the manufacturer’s recommended protocols, and covered with microscope cover slides, and co-denaturation and subsequent hybridization of probe and target DNA were performed in a ThermoBrite Programmable Temperature Controlled Slide Processing System (Abbott Molecular) overnight, a step that is not considered in these calculations as it was not labor intensive and was required for both the VP 2000 and manual protocols. The next day the, cover slips were removed, excess probe was washed from the specimens (2XSSC 0.3% NP-40 at 73 °C for 5 min followed by 2XSSC 0.3% NP-40 at room temperature for 2 min), slides were dried at room temperature for 3 min, DAPI counterstain and coverslips were added, and the specimens were henceforth ready for interpretation and analysis. This concluded the hands-on processing requirements for what we termed “day 2” of the manual FISH protocol (10 min total time). In total, the hands-on time for the manual processing of one batch of FISH specimens was 219 min, costing a total of €69.34 for labor. The cost of the first day alone (209 min) was on the order of €66.18.
Day 1 of VP 2000 automated FISH processing required a total of 35 min of hands-on time. The steps of the assay were the same as described above but were processed automatically on the VP 2000. The time requiring personnel here involved preparing the buffers and reagents, filling the various buffer basins of the VP 2000 processor, and cleanup following the procedure. Day 2 of the VP 2000 procedure required 10 min total hands-on time as described above for the manual procedure. As the second day of the procedure was identical between the manual and VP 2000 protocols, day 2 was not further factored into the calculations below. Hence, we considered a pretreatment time for manual FISH processing of one batch to be 209 min (€66.18 labor cost), versus 35 min (€11.00 labor cost) for the VP 2000.
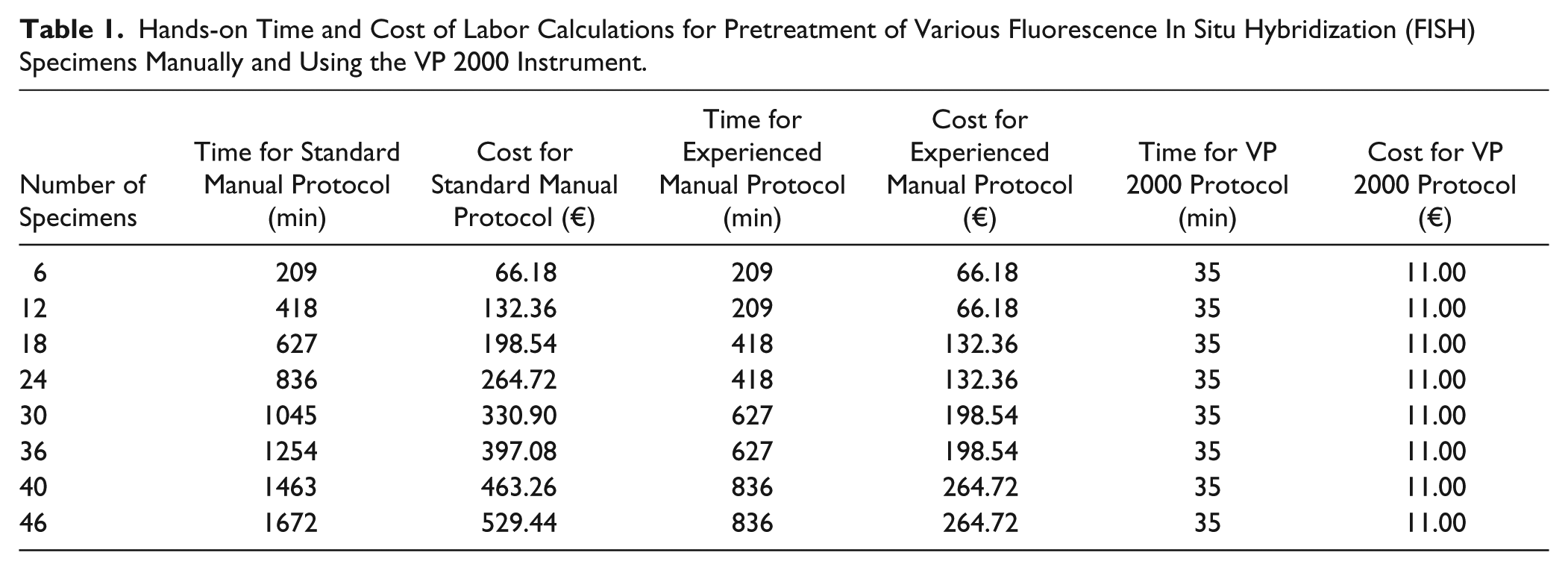
The manual procedure (209 min) described above was applied to the processing of six specimens, which can be comfortably processed by one technician. The time and cost doubled for every additional six specimens for a standard laboratory laborer (considered the “standard manual process time”; Table 1 ). In the case of highly experienced FISH laboratory workers, an equal amount of time was required to process 6 and 12 (209 min), 18 and 24 (418 min), 30 and 36 (627 min), and 40 and 46 specimens (836 min; considered the “experienced manual process time”; Table 1 ). The total time required for the VP 2000 did not change with specimen number as only one run was required regardless of 6 or 46 specimens being processed. Assuming an average hourly cost for laboratory personnel of €19, Table 1 summarizes the times and costs of processing various numbers of FISH specimens manually and with the VP 2000. It should be noted that both protocols then proceed to a step in which the FISH probe is added to the pretreated specimen, a cover glass is added to the slide and sealed with rubber cement, and the probes are hybridized (following the manufacturer’s recommended protocol) on a FISH probe hybridizer overnight. As this step of the protocol is vital and identical between the automated and manual procedures, it is not included in our calculations or discussions below. Addition of FISH probes and rubber cement and starting the hybridizer required only 1 to 2 min of hands-on time.
Hands-on Time and Cost of Labor Calculations for Pretreatment of Various Fluorescence In Situ Hybridization (FISH) Specimens Manually and Using the VP 2000 Instrument.
FISH Reagent Costs
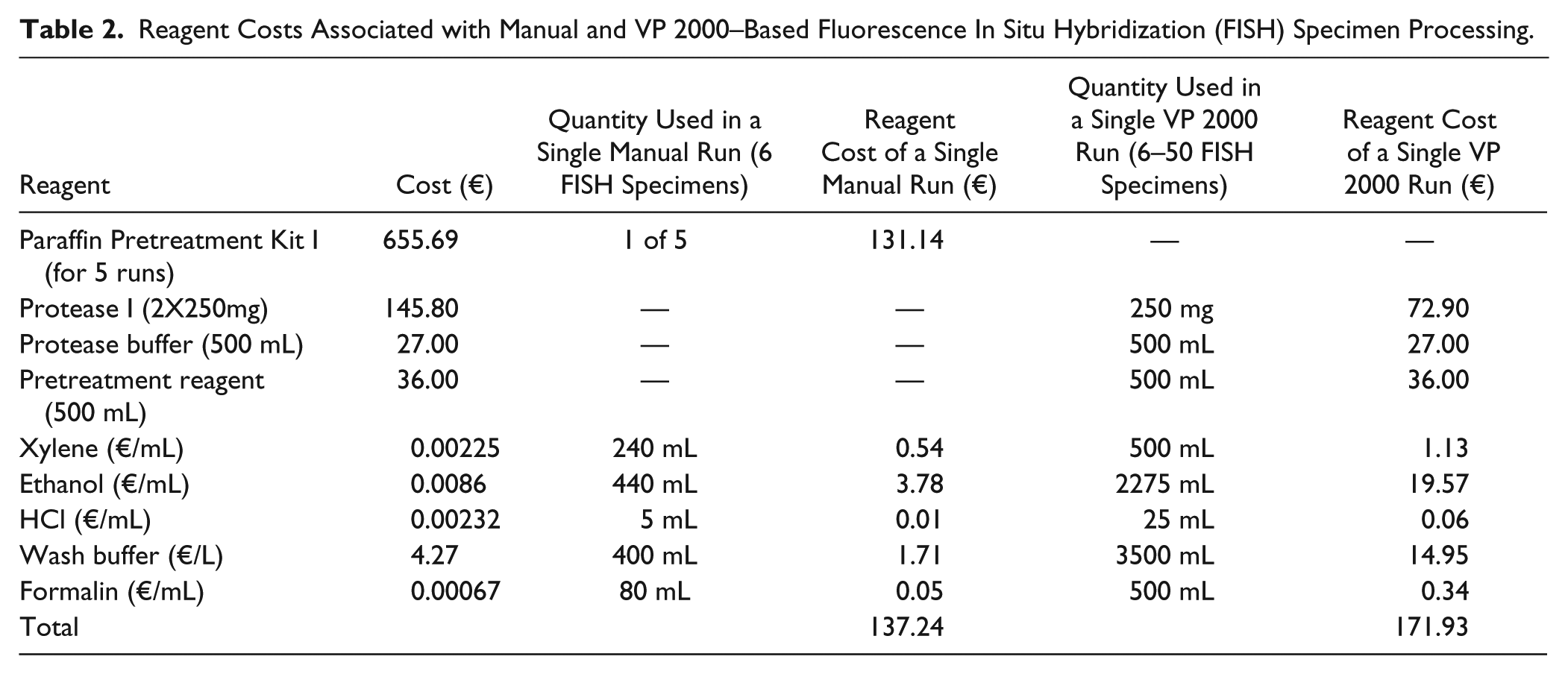
Reagent costs also come into consideration. The VP 2000 instrument uses large buffer basins to accommodate up to 50 FISH specimens. Compared with lower-throughput manual methods, in which a single Coplin jar may be used for up to six FISH specimens, we sought to compare the costs of reagents between manual and batched VP 2000 pretreatment of FISH, as the minimal volume of reagents required can differ significantly between the two methods. For manual processing, the Paraffin Pretreatment Kit I (Abbott Molecular) was used. A single kit can accommodate up to five runs (i.e., one run uses one batch of six FISH specimens). In contrast to the manual pretreatment kits, the VP 2000 reagents are available in larger volumes. Table 2 outlines the various reagent costs, in Euros, used by the laboratory for the manual versus the VP 2000–based protocols. The overall reagent cost of a single VP 2000 run cost on the order of €171.93 compared with a single manual FISH run (of six specimens) costing €137.24. It is important to note that for each additional batch of six specimens, the reagent costs of the manual protocol increases by €137.24, while the VP 2000 instrument can accommodate up to 50 specimens for a fixed cost of €171.93. Therefore, while it is slightly more expensive, from the perspective of reagents, to use the VP 2000 for a small batch of six samples, once more than six specimens are processed, the VP 2000 becomes the more cost-effective option. Although some laboratories choose to reuse reagents for a number of FISH runs, the laboratory in Treviso renews all reagents for each batch processed (both manually and with VP 2000). This practice has resulted in the most robust results for the laboratory.
Reagent Costs Associated with Manual and VP 2000–Based Fluorescence In Situ Hybridization (FISH) Specimen Processing.
Combined Cost of Labor and Reagents for Processing Various Batch Sizes of FISH Specimens
Summarizing the data from
Tables 1
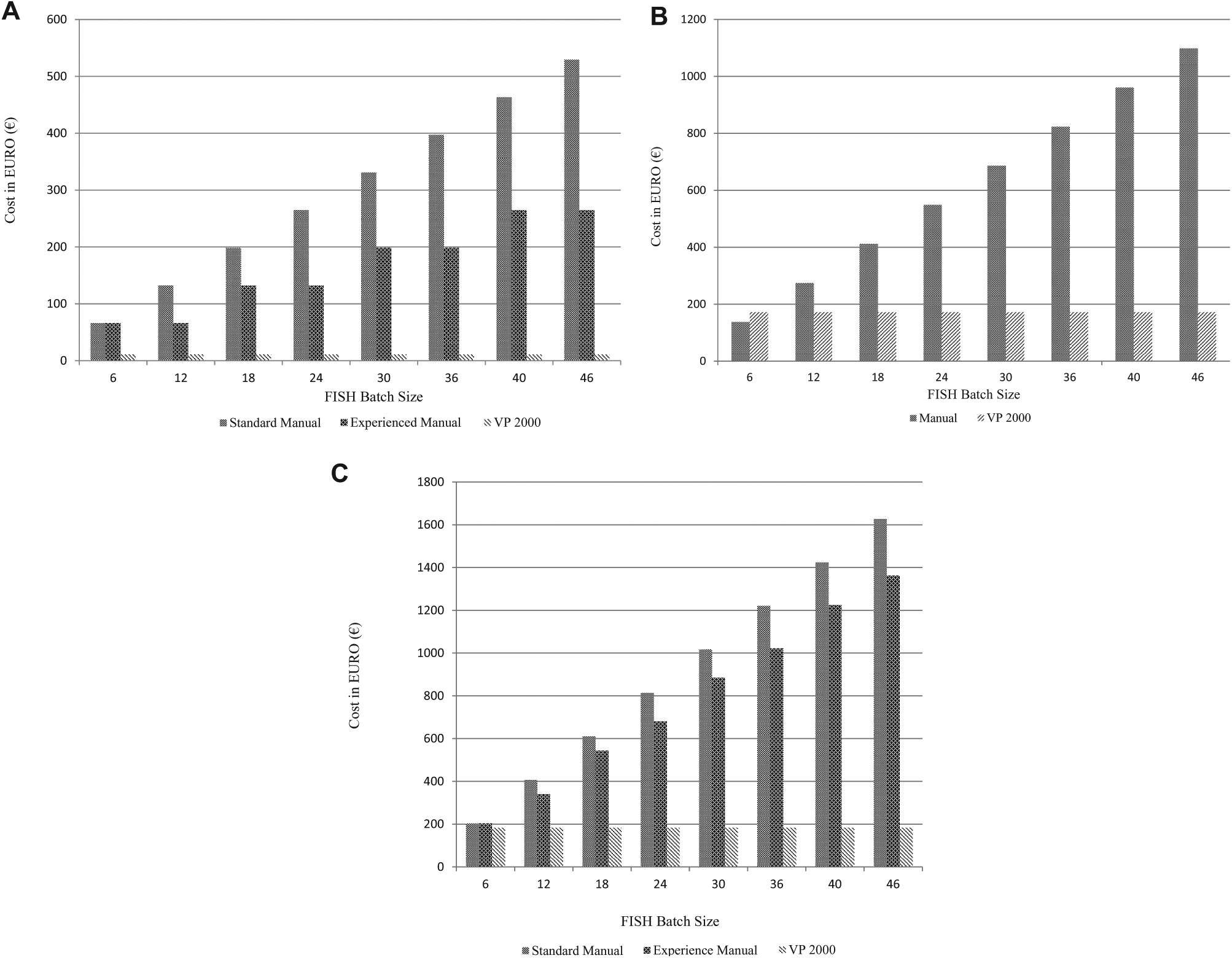
Graphical representations of labor and reagent costs for fluorescence in situ hybridization (FISH) comparing manual versus VP 2000–based automated protocols. (
Discussion
The current study compared the cost of labor and reagents for manual FISH methodology versus automated processing with the Abbott VP 2000 instrument, for various batch sizes ranging from 6 to 46 specimens. Interestingly, the automated instrumentation, which allows for simultaneous processing of up to 50 specimens at one time, facilitates significant cost saving from the perspective of both labor and reagents when more than 6 specimens are processed per week. Processing 12 specimens per week demonstrated an approximate cost saving of 46% to 55% compared with manual protocols, while high specimen throughput yielded even more significant cost saving (on the order of 86%–89%). Use of the VP 2000 instrument allowed for hands-free, walk-away time on the order of 174 min (209 min for manual pretreatment versus 35 min hands-on time for the VP 2000 setup), which gives laboratory staff time to perform other activities and protocols while the tissue pretreatment protocol is executed. These results demonstrated a significant increase in FISH throughput and efficiency in high-volume FISH laboratories but also suggest that lower-volume FISH labs may also benefit from such laboratory automation. The laboratory in Treviso used more than 20 different tissue FISH probes in the course of this study, suggesting that the protocols used are highly flexible for the needs of most pathology and solid tumor FISH laboratories. These results are significant as FISH is often considered an expensive and labor-intensive methodology, and therefore cost saving, reproducibility, and ease of use are high priorities for FISH users.2,3 Nonetheless, for many applications, especially gene rearrangements, deletions, and insertions, FISH remains the gold standard technique and has become a mainstay in modern molecular pathology laboratories.1,2,4–6
The manual protocol used here was on the order of 209 min, which is quite a lengthy protocol. The Pathology and Cytogenetics laboratory at the General Hospital of Treviso is a certified lab, and the protocols used were also certified. While the lab was aware that shorter protocols exist, the protocol used gave the most robust and reproducible results and hence was the protocol of choice. It is therefore important to note that the cost savings observed in Treviso cannot necessarily be directly assumed for all other centers. The amount of cost saving experienced with automation of FISH with the VP 2000 is directly related to the hourly wage of a laboratory worker and the amount of hands-on time required for manual FISH processing. The current study is a specific example of what was observed when automating FISH with the VP 2000 instrument in this particular Italian laboratory. Nonetheless, the authors believe a similar trend would be observed for most intermediate- to high-throughput FISH laboratories.
One aspect of the protocol not discussed in the current work is the posthybridization wash. This part of the protocol can also be completed on the VP 2000. In fact, the laboratory in Treviso demonstrated that the manual protocol “second day” posthybridization wash would cost on the order of €2.83 for a minimum batch of six specimens, whereas the VP 2000 protocol cost on the order of €17.74. Similar to the “first day” pretreatment protocol, the VP 2000 reagent costs for the posthybridization wash were more expensive compared with the manual protocol, due to larger volumes of buffer required to fill the basins of the instrument. Although it may not be practical to use the VP 2000 for posthybridization washing for limited specimen amounts, it does also become cost saving when higher specimen amounts (>36) are used (data not shown). The hands-on time for the second day posthybridization washes is on the order of 10 min for both the manual and VP 2000 protocols. Indeed, as more specimen batches were processed, the hands-on time for every additional six specimens added an incremental 10 min to the manual protocol, whereas the VP 2000 automated protocol remained unchanged (data not shown). Nonetheless, the major drivers of FISH cost come from the labor and the first day pretreatment, and hence the reagent costs for the posthybridization washes are negligible in the context of overall cost.
An important aspect that requires consideration when new molecular protocols are developed and/or automated is the specimen quality and assay failure rate. Indeed, as those labs that perform FISH are aware, there are cases in which specimens yield faint or nonanalyzable FISH fluorescence signals. This can be due to either failures in the assay protocol or issues with tissue quality due to the preparation of the specimen. 2 The laboratory in Treviso is a major FISH testing hub receiving numerous specimens from various medical centers and hospitals in the area. As such, there is little standardization of specimen preparation and storage between centers, making a direct comparison of failure rates between the manual and automated FISH protocols difficult. Over the course of the study, the percentage of nonanalyzable FISH specimens was 9% for the manual protocol and 7% for the VP 2000 automated protocol. Although these failure rates may suggest that the automated protocol is more robust, the results are difficult to compare in a controlled manner, as the specimens used over the course of the study originated from different centers and hence differences in specimen quality likely contributed to variations in failure rates. The laboratory did not notice any significant differences in the FISH signal fluorescence intensities when comparing manual versus VP 2000–prepared specimens (data not shown). Specimens that failed automated processing via VP 2000 were reprocessed manually; however, with the exception of a select few cases, manual reprocessing did not salvage any of these specimens. This suggests that the majority of nonanalyzable FISH specimens indeed resulted from specimen preparation issues, rather than FISH assay protocol failure.
Finally, it is also important to consider the costs of the instrumentation (i.e., the VP 2000 instrument itself) along with the maintenance. The cost of the instrument varies from country to country (approximately €35,000 list price in Europe). Although this represents a significant investment for many laboratories, considering the cost savings on reagents and labor, this one-time instrument cost can be remunerated with time. In addition, the service costs and intervals of maintenance also vary from country to country and instrument to instrument. In Treviso, there is a yearly preventative maintenance; however, this cost is coupled with the ongoing purchase of probes, and hence, we are unable to produce a specific maintenance cost for the system. Nonetheless, the system operates robustly and thus far has not malfunctioned or required any significant maintenance.
The current study demonstrates that laboratories can potentially save significant labor and reagent costs for FISH testing using the VP 2000 instrument. In this particular study, cost reductions of greater than 55% were observed when the laboratory processed 12 or more specimens per week. These results are significant as modern molecular pathology laboratories are continuously requested to accommodate higher testing volumes and increasing numbers of biomarkers and technologies. 8 Through robust automation protocols, laboratory staff can accommodate more assays, produce more reliable results, and increase hands-free time for other important tasks.
Footnotes
Acknowledgements
The authors would like to thank Dr. Karin Pfeifer for critical reading of the manuscript.
Declaration of Conflicting Interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: Dennis Merkle was an employee and stockholder of Abbott Laboratories at the time of submission of this article. He is no longer an employee or stockholder of Abbott at this time.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
