Abstract
Laboratory evolution provides phenotype–genotype mappings and quantitative analysis of selective pressures, giving important insights about evolutionary dynamics. Moreover, parallel laboratory evolution clarifies which phenotypic and genotypic changes are inevitable for adaptive evolution. Such parallel experiments, however, remain labor-intensive. In this study, to facilitate massive parallel laboratory evolution, we developed an automated culture system that can maintain hundreds of independent culture series in exponential growth phase under various culture conditions. We demonstrate the performance of this automated culture system using the laboratory evolution of
Introduction
Laboratory evolution is a promising approach for understanding the nature of evolutionary dynamics. 1 Recent advances in several omics analyses and high-throughput sequencing have enabled us to elucidate phenotypic and genotypic changes fixed in laboratory evolution, providing valuable information on the detailed mechanisms of adaptive evolution by identifying the sets of beneficial mutations that contribute to fitness. 2 These studies have also provided important insights into the characteristics of evolutionary dynamics, including second-order effects during evolution and the interrelations among population size, robustness, and evolvability.3,4 Moreover, the impact of laboratory evolution has gone beyond the field of evolutionary biology and into engineering, as exemplified by the construction of microbial cells for biotechnological applications. For example, with the aid of in silico design of a metabolic network, microbial strains optimized for specific bioproductions were constructed by laboratory-evolution experiments. 5
In general, two types of long-term culture methods are used for laboratory-evolution experiments: chemostat culture and serial transfer culture. In chemostat (continuous)-culture experiments, fresh medium is continuously supplied to a bioreactor (jar fermenter), and an equal amount of medium, which includes cultured cells, is discarded to keep the culture volume constant. The advantage of the chemostat culture is its ability to keep a constant growth rate, population density, and culture condition based on the pH and dissolved oxygen. A major drawback, however, is the cost of operation, which limits the number of culture series for laboratory-evolution studies.
In serial-transfer cultures, at a regular interval (usually daily), an aliquot of cell-containing culture medium is transferred to a new tube or well with fresh medium for an additional round of growth. This simple experimental system enables massive parallel experiments, which provide valuable information on adaptive evolution. For example, evolution experiments under various environmental stress conditions revealed the molecular mechanisms for evolutionary trade-offs and cross-protection. 6 In another example, parallel-evolution experiments under identical conditions revealed which phenotypic and genotypic changes are inevitable for adaptive evolution and which occurred by mere chance. 7 Nevertheless, the serial-transfer culture for laboratory evolution suffers from its periodic (daily) changes in population density and environmental conditions, such as nutrient concentration. When the culture reaches the stationary phase, the competition among cells is not negligible, and evolutionary dynamics in such a varying environment often results in various unexpected phenomena, such as the coexistence of cells with different phenotypes. 8 One possible strategy to avoid environmental changes in laboratory evolution is to keep the cell concentration low. In other words, the culture should be maintained in the exponential-growth phase, when the cell–cell competition is negligible. Achieving this condition by daily serial transfer, however, requires relatively large amounts of culture volume, often compromising massive parallel laboratory-evolution experiments. For example, when the minimum transfer volume is 1 µL and the specific growth rate is 0.5 (1/h), the culture volume needs to be more than 20 mL.
In the present study, to overcome the limitation of manual laboratory-evolution experiments, we describe an automated culture system that achieves rapid (e.g., 2 to 6 h intervals) serial transfer using a 96-well microplate. A lab-automated system is placed in a clean booth and connected to a microplate reader and a shaker incubator. This system enables us to maintain hundreds of independent culture series in the exponential-growth phase. We demonstrate the performance of this automated culture system by observing the laboratory evolution of
Materials and Methods
Strain and Media
Automation
The automated culture system ( Fig. 1 ) consists of a Biomek® NX span8 laboratory automation workstation (Beckman Coulter, Tokyo, Japan) in a clean booth connected to a microplate reader (FilterMax F5; Molecular Devices, Tokyo, Japan), shaker incubator (STX44; Liconic, Mauren, Lichtenstein), and microplate hotel (LPX220; Liconic). These instruments are integrated as shown in Figure 2 . The shaker incubator and microplate hotel are combined with the deck of Biomek NX by a transfer unit (slide transfer station for double extended transfer reach; Liconic), and microplates are transferred between these instruments. All instruments are connected to communication ports of process-control computer by serial cables. The capacity of the incubator is 44 microplates, and the microplate hotel has 10 racks, each of which contains up to 22 plates or 7 tip boxes. The cell density is measured using optical density at 620 nm (OD620) by the microplate reader, in which OD620 in a blank well is subtracted as background. Fresh medium and stressors are kept on the bench using 24-well and 96-well microplates, respectively, at room temperature. All operations were controlled by our in-house program, which was developed in Biomek Software Version 3.3 (Beckman Coulter). The process flow of our method is summarized in Figure 3 .
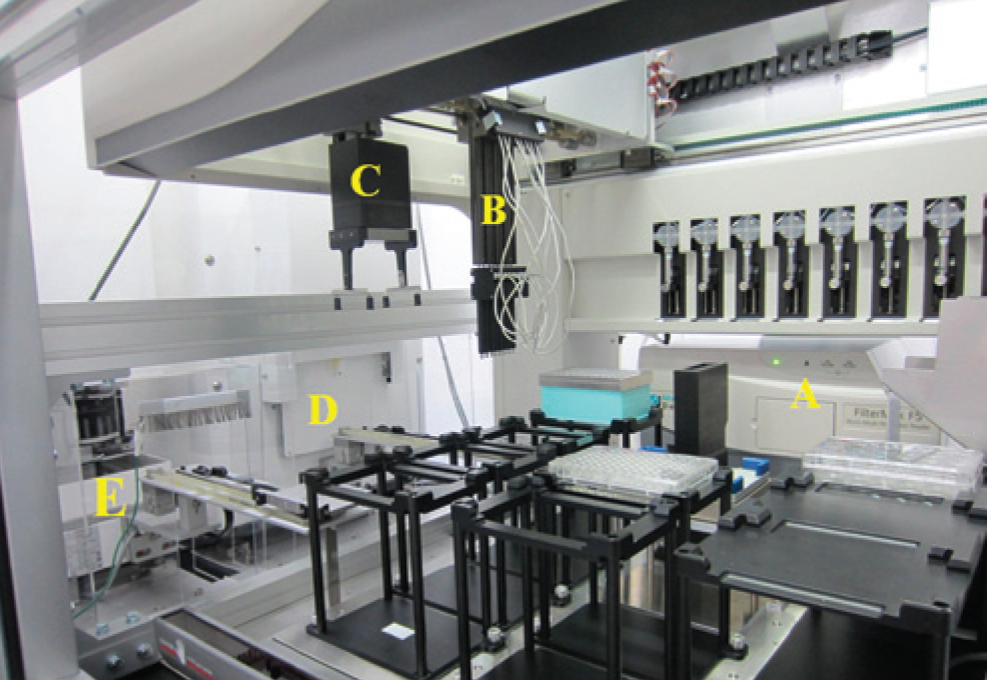

The automated culture system for laboratory evolution. (
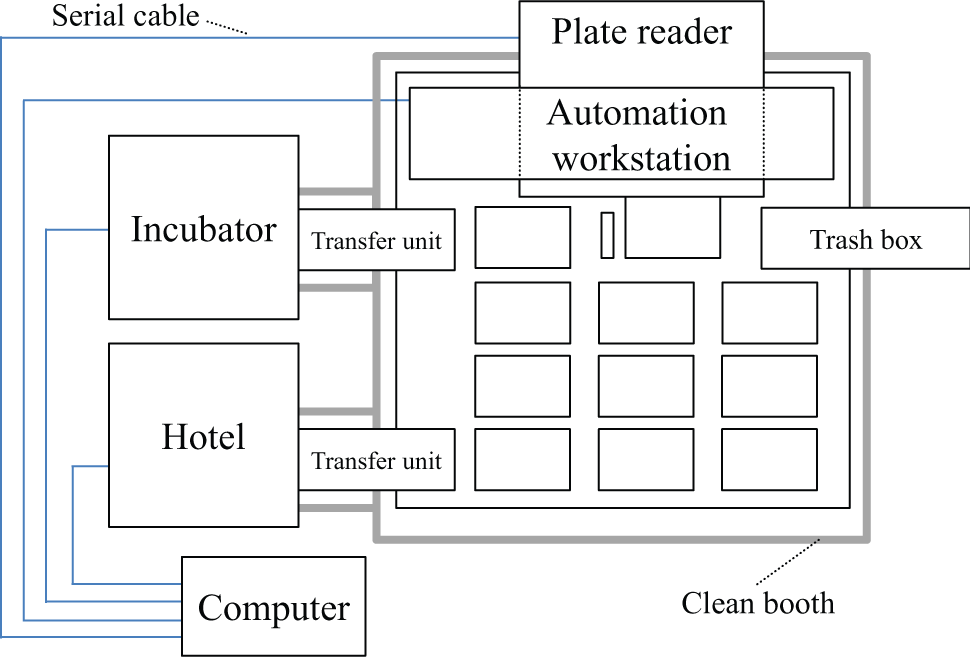
Schematic representation of the system integration. The shaker incubator and microplate hotel are connected with the deck of a Biomek NX, and microplates are transported between these instruments by a transfer unit of Biomek NX. All instruments are connected to the communication ports of a process-control computer by serial cables and are controlled by our in-house developed program.
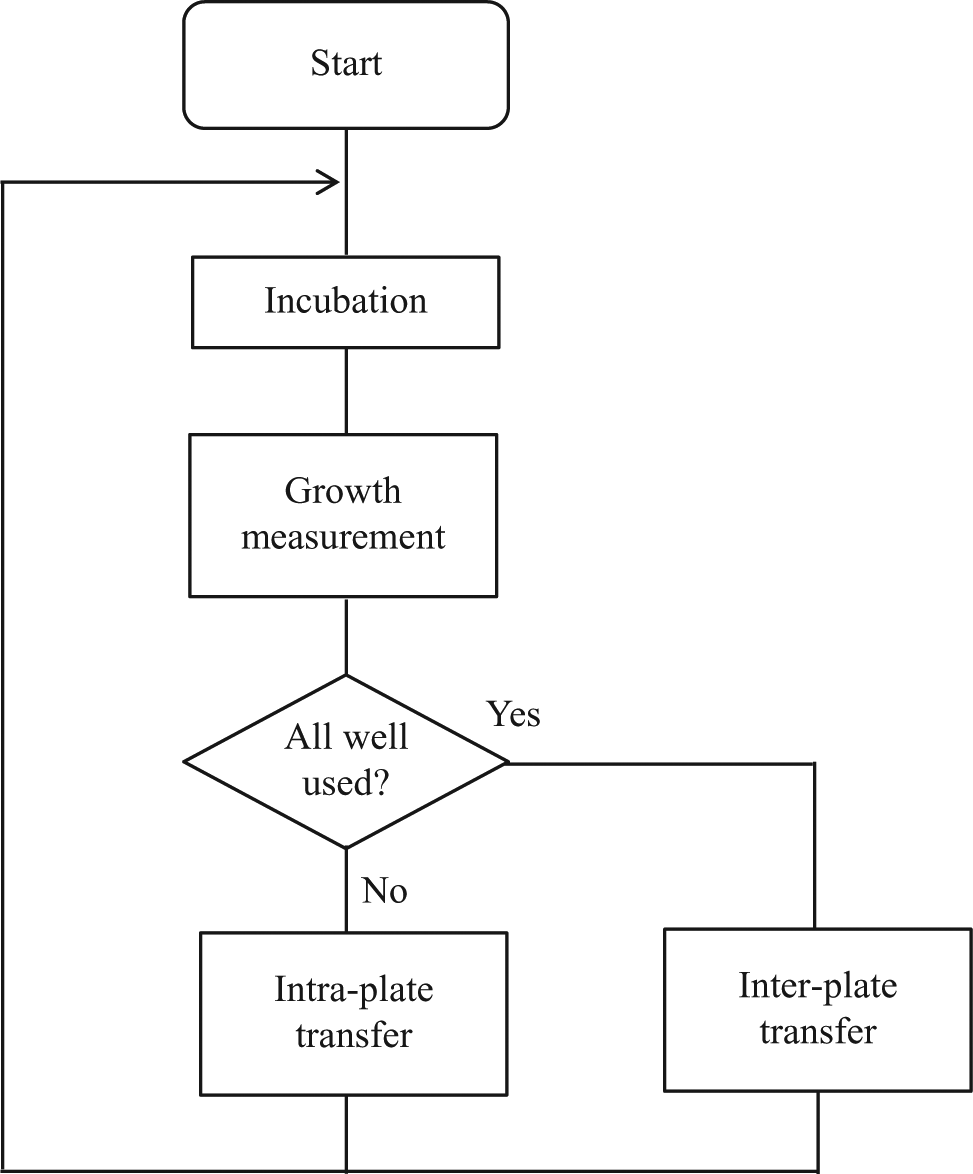
Process flow for the automated culture system. A microplate is transferred from the incubator at the constant time interval. Optical density is measured by the microplate reader and based on the measured cell density, cells are transferred into an adjacent well. If all wells in a 96-well microplate are used, new plate is carried from the microplate hotel, and the cells are transferred to a well in the new plate. These cycles of cell transfers are repeated.
Laboratory Evolution Experiments
Starting from the initial cell population, the medium with cells is transferred into fresh medium at a regular interval, Δt. We assume that in the
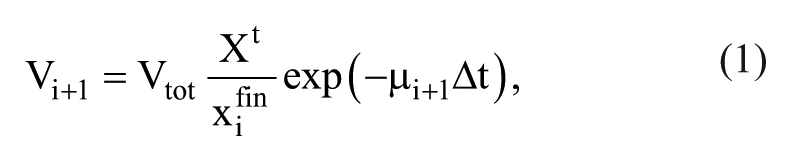
where Xt is the target cell density after Δt,
Results and Discussion
Batch Culture
To determine the range of cell densities in which
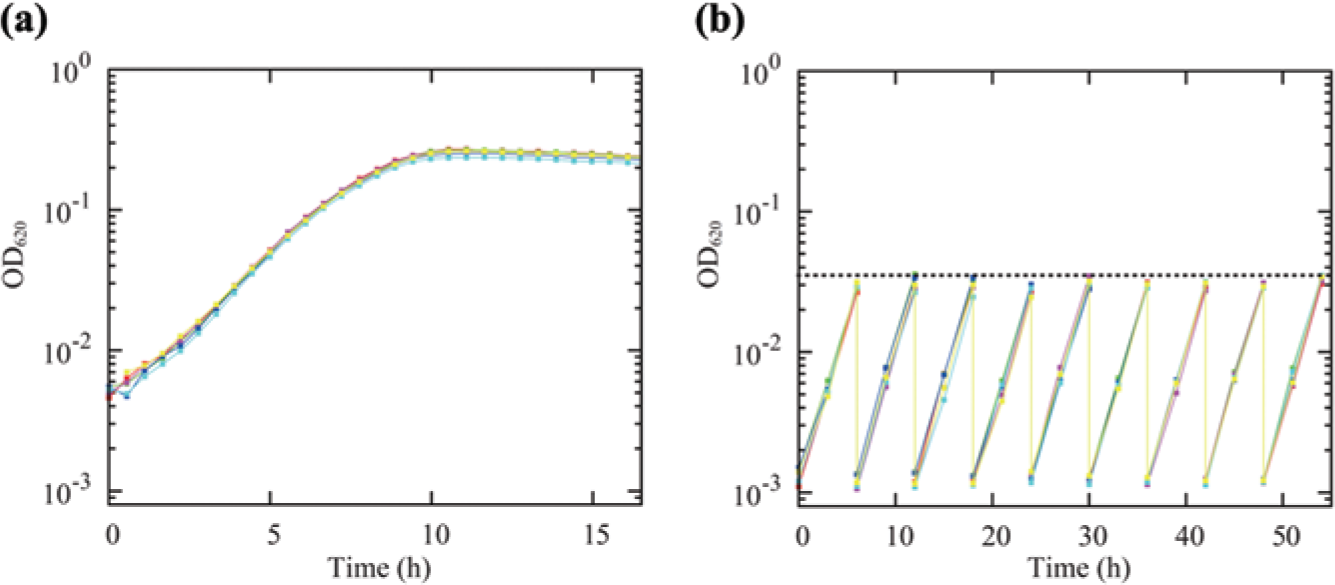
Growth measurement in the automated culture system. (
Laboratory-Evolution Experiments
To demonstrate the performance of our automated culture system, we performed laboratory evolution under one nonstress and five stress conditions, including high salt stress (400 mM sodium chloride), acid stress (44 mM lactate or 30 mM malate), alkali stress (32.5 mM sodium carbonate), and alcohol stress (1.25%
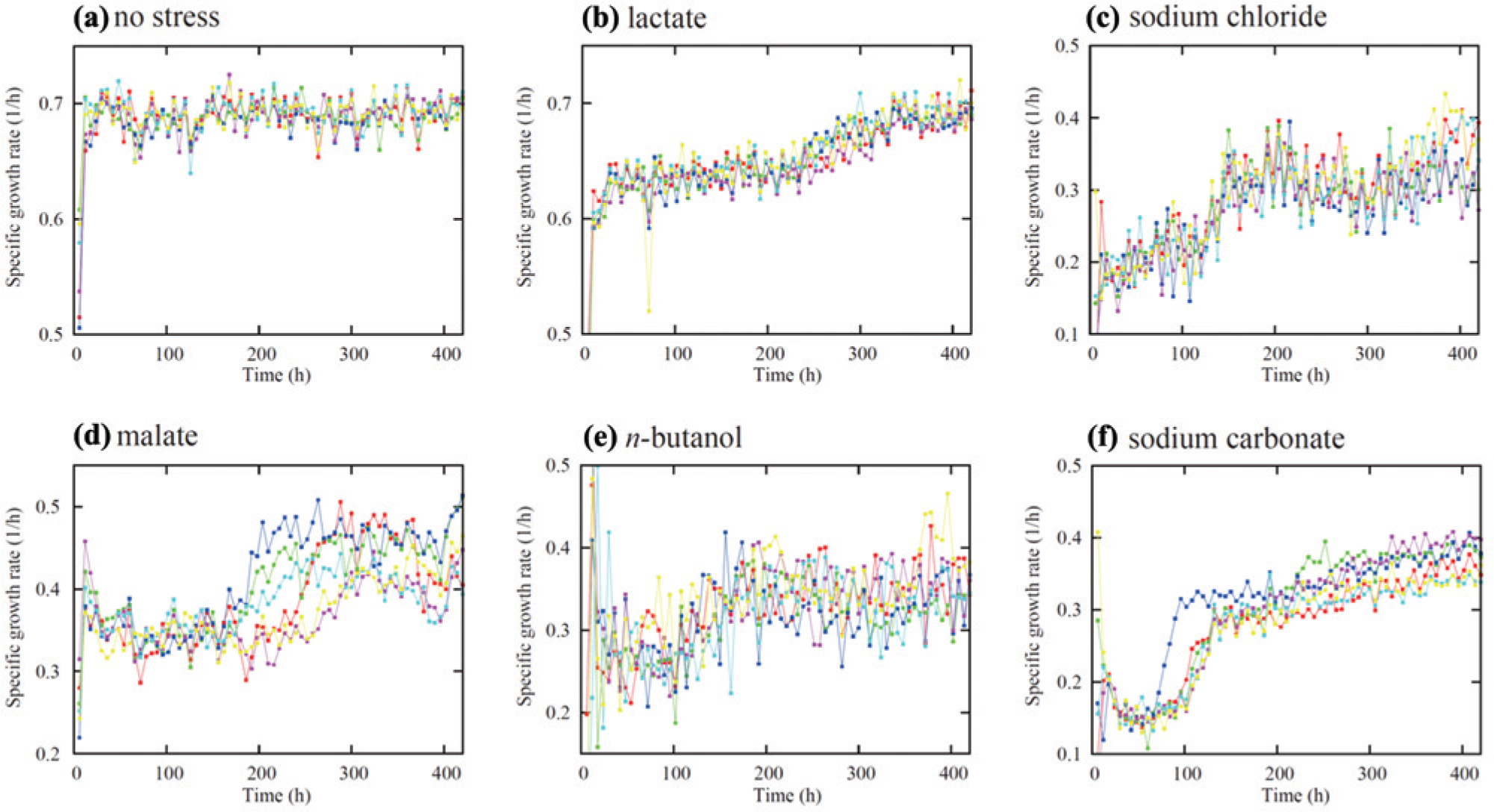
Growth increases in the laboratory evolution of
In our automated system, the process of cell transfer includes the following steps: (1) carrying a microplate from the incubator to the microplate reader, (2) optical density measurements, (3) the transfer of medium with cells to an empty well, (4) the addition of a stressor, and (5) carrying the microplate to the incubator. A movie of this transfer process is presented in
Even in cultures with identical conditions, the specific growth rate shown in Figure 5 fluctuated during time due to experimental error. In the case of the nonstress condition, the coefficient of variance of the specific growth rate was approximately 2%, which is similar to that obtained in the manual transfer of cells. In the case of stress conditions, the fluctuations in the growth rate were often larger than those of the nonstress condition. This result is presumably due to fluctuations in the amount of added stressors, because in some cases the volumes were as high as a few microliters.
Because the cells are maintained for a relatively long time, one possible problem in laboratory evolution experiments is contamination, including cross-contamination of
Summary
Our culture system is the first system for fully automated evolution experiments. The unique integration of a culture system (incubator) and measurement system (microplate reader) by the Biomek NX automated system enables us to maintain growing cells in a constant environment by a rapid serial transfer. This system can trace the evolutionary dynamics of hundreds of independent culture series under various environmental conditions. Phenotypic and genotypic analyses of the evolved organisms reveal detailed mechanisms of the adaptive evolution to various stress environments, providing valuable information for both evolutionary biology and bioengineering fields. Although we used
Footnotes
Acknowledgements
We thank Dr. Peter Karagiannis for proofreading of the manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported in part by Grant-in-Aid for Young Scientists (A) (23680030 to C.F.), Grant-in-Aid for Scientific Research (A) (24246134 to H.S.) from Japan Society for the Promotion of Science, and Grant-in-Aid for Scientific Research on Innovative Areas (25128715 and 23128509 to C.F.) from the Ministry of Education, Culture, Sports, Science and Technology, Japan.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
