Abstract
Ultraviolet (UV) mutagenesis is a widely used technique to increase bacterial mutation rates in laboratory experiments. UV mutagenesis requires fine regulation of UV dose, because the number of dead cells increases exponentially as the dose increases. Ignoring this hazard can cause extinction of UV-exposed populations. Therefore, an automated system that cooperatively conducts both growth measurement and UV irradiation is needed for efficient UV mutagenesis experiments. To address this task, we constructed an automated UV irradiation device for microbial cell culture. This device can measure cell density and irradiate the bacterial cells with UV light automatically according to the state of cell growth. We demonstrated that this growth feedback control avoided extinction and enabled accumulation of mutations in bacterial genomes at a rapid rate for a long period. Whole-genome sequencing revealed the high accumulation rate, neutrality, and spectrum of UV-induced mutations. These characteristics were all consistent with those obtained by manual UV irradiation. These results indicate that our automated device is useful in accelerating mutation accumulation over a long duration.
Introduction
Ultraviolet (UV) mutagenesis is a widely used technique to increase bacterial mutation rates in laboratory experiments. Because the occurrence of mutations is one of the rate-determining factors of evolution, UV irradiation can be applied for the purpose of rapid acquisition of adaptive traits in bacterial cells.1,2 Compared with various other methods of mutagenesis, such as administration of chemical mutagens or exposure to ionizing radiation, UV irradiation is considered to have advantages of safety and convenience. Therefore, it is important to keep investigating the types of mutations induced by UV irradiation, the rate of acquisition of these mutations, and the method of application of UV irradiation for accelerating laboratory experiments.
UV mutagenesis requires fine regulation of UV dose, because the number of cells killed by UV exposure increases exponentially as the dose increases. Overdose of UV light causes the cell population to die out, whereas too little UV radiation leads to cell saturation, which decreases the efficiency of the experiment ( Fig. 1 ). If an experimenter adjusts the UV dose while checking the status of the population, both extinction and saturation can be avoided. However, such a solution would require a tremendous amount of manual effort by the experimenter.
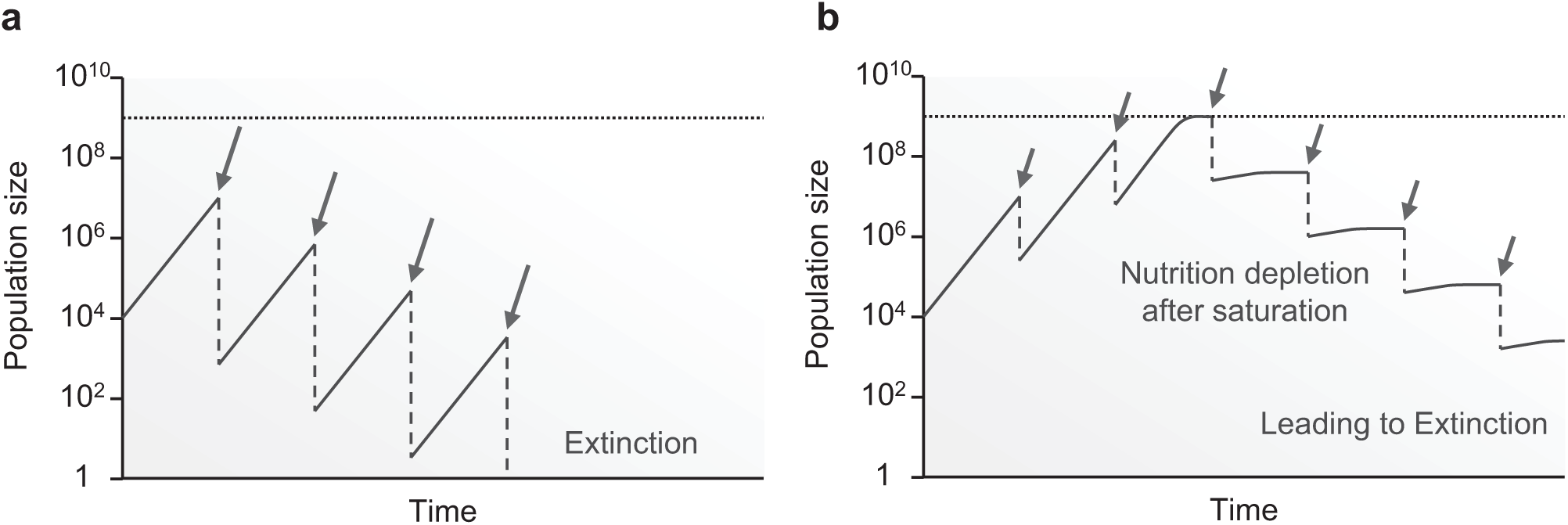
Schematics of the consequences of inappropriate dosages of periodic UV irradiation on bacterial growth. (
Therefore, an automated system that cooperatively conducts both growth measurement and UV irradiation is necessary for efficient UV mutagenesis experiments. Several studies have demonstrated that feedback between measurement and control of population growth is effective for avoiding extinction and saturation. For example, turbidostats 3 and automated serial transfer systems 4 can avoid saturation by controlling the rate and/or frequency of dilution of media based on the cell growth. Additionally, morbidostats 5 regulate the concentration of growth-inhibitory drugs on the basis of the growth status of the cell population. The feedback control embedded in these automation systems can be applied to a regulation of UV dose to achieve UV mutagenesis without extinction.
In this study, we designed a culture system for automated growth measurement and UV irradiation. The system consists of a turbidimeter, a UV emitter, and a specially designed control method. To demonstrate the performance of this culture system, we conducted mutation accumulation experiments in
Materials and Methods
Strain and Media
We used
Automation Device
The automated UV irradiating culture device consists of a fused-quartz test tube and a socket, which has a turbidimeter and a UV emitter ( Fig. 2a ). The turbidimeter consists of a visible LED and a phototransistor (typical wavelengths were 590 and 560 nm, respectively). These are positioned at a 135° angle to maximize the detection of scattered light. 5 As a UV emitter, we used a Deep-UV LED module (UV-EC910ZA, Panasonic Photo & Lighting, Takatsuki, Osaka, Japan; typical wavelength was 270 nm). The UV LED is placed in a manner such that the tube is irradiated from the bottom. A 3D-printed ABS resin housing holds the electronic elements at their respective appropriate positions. The electronic components are connected to a circuit ( Suppl. Fig. S1 ) and controlled by a microcontroller board (Arduino Uno Rev3, https://www.arduino.cc/), which is connected to a laptop computer.
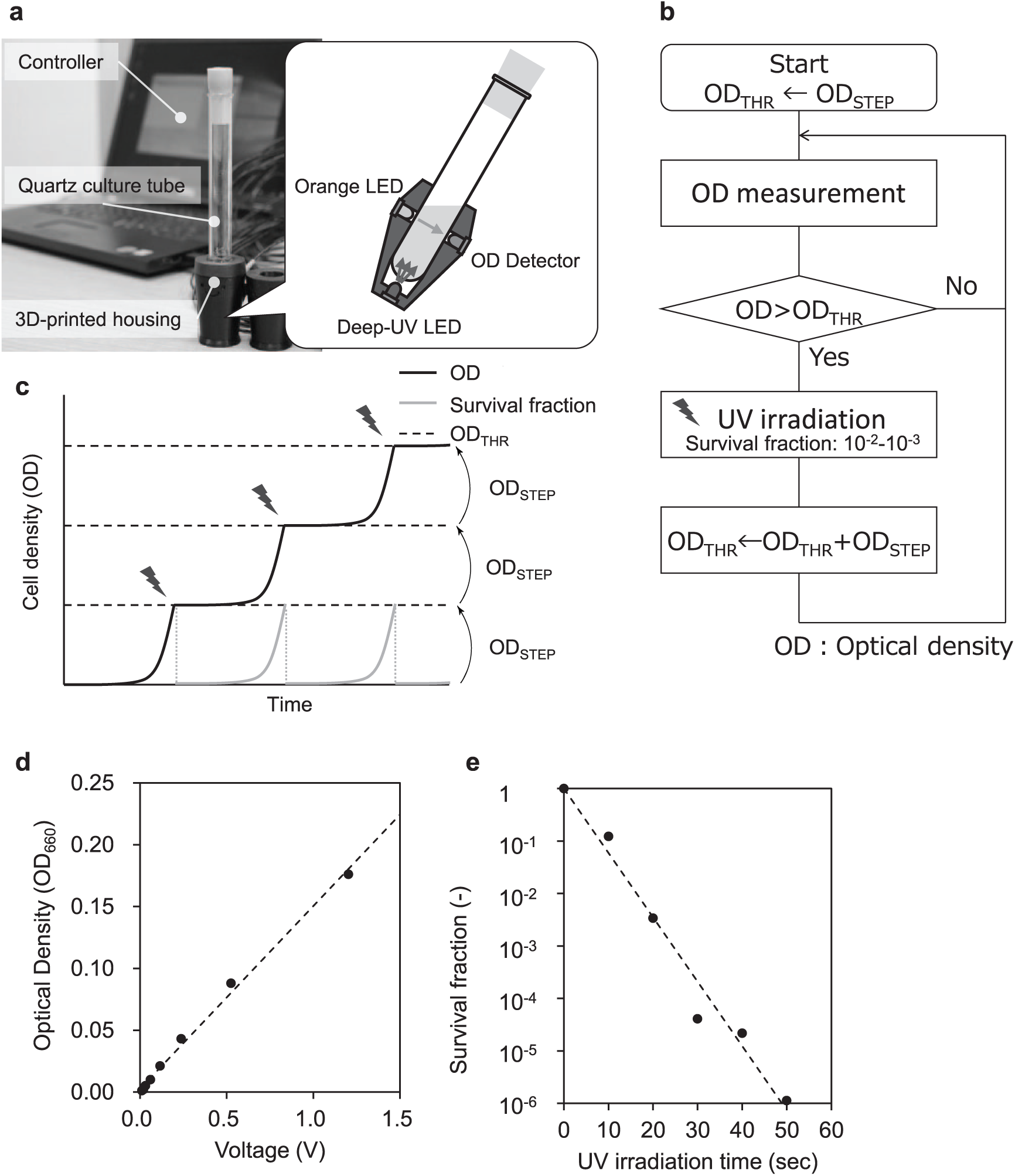
Automated UV irradiation device for bacterial culture. (
Operation Procedure
All operations are controlled by in-house programs, which were developed in Arduino IDE (v1.6.5, https://www.arduino.cc/) and Visual Studio 2013 (Microsoft Corporation, Redmond, WA). A process flowchart of our method is shown in Figure 2b . Optical density (OD) measurement and UV irradiation are implemented in repeated cycles. In each cycle, cells are incubated for a set time period (3 min), during which the OD is recorded. UV irradiation is conducted when the OD value is greater than a set threshold (ODTHR) in two consecutive cycles. Initially, ODTHR is set as ODTHR = ODSTEP. After irradiation, it is modified by ODTHR + ODSTEP, so that the next irradiation is conducted when the cell population recovers to the population size corresponding to ODSTEP ( Fig. 2c ). In this way, the cells keep growing exponentially while OD increases stepwise at intervals ( Fig. 2c ), enabling cells to avoid saturation for extended durations. In our experiments, ODSTEP was set to 0.0015.
Mutation Accumulation Experiment
The time duration for one UV irradiation is set so that the survival fraction of the wild-type cells is 10–2 to 10–3. The cells were serially transferred and glycerol stocks were stored for sequencing every 4 days. The experiment was replicated five times and continued for 56 days.
Whole-Genome Sequencing
We performed whole-genome sequencing for the transferred populations and their ancestral strain using Illumina MiSeq Desktop Sequencer (Illumina, Inc., San Diego, CA) and identified mutations according to the procedure reported previously. 8
Results
Specifications of the Device in Growth Measurement and Killing Ability
To determine a reliable range for OD measurement of our device, we compared the OD values obtained by our device with those obtained using a conventional commercial spectrophotometer (
Fig. 2d
). We prepared serial dilutions of
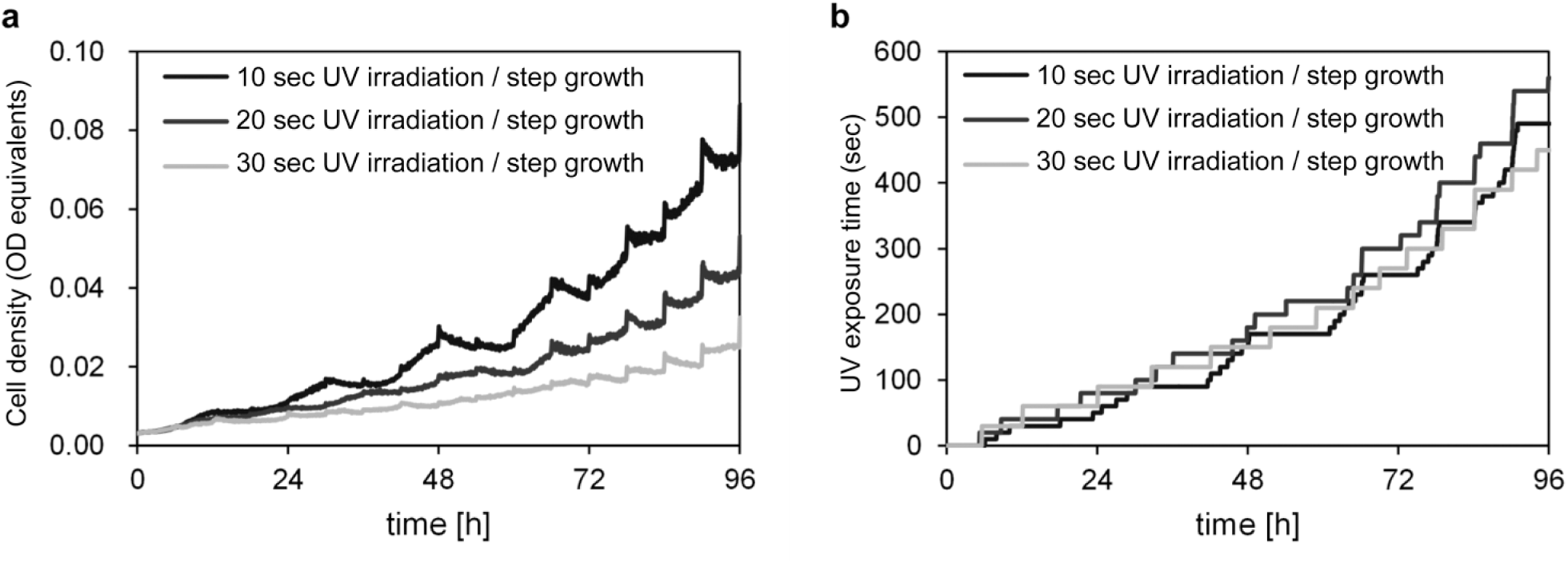
Growth measurement using the automated UV irradiation device. (
UV Irradiating Cell Culture Experiment
Next, we conducted a mutation accumulation experiment using our device. We established five independent replicate cultures from the ancestral strain. We cultivated the cells with UV irradiation in the device. The device was operated according to the flowchart shown in Figure 2b , where ODSTEP was 0.0015 and UV irradiation time was 20 s. After 4 days of cultivation, we serially transferred the cultures to fresh medium. We repeated these procedures for 14 rounds, yielding 56 days of cultivation in total ( Fig. 4a , b ). The growth curves and UV doses of the replicated lineages were similar to each other for at least the initial 4–16 days ( Fig. 4c , e ). However, some lineages markedly improved their growth in the device afterwards. In particular, the cells in lineage 1 after day 36 exhibited a strong tendency to aggregate on the wall of the culture tube. The cells grown on the wall were sporadically torn off the wall by shaking, resulting in a sudden increase in the cell density of the culture beyond the normal increase due to cell growth. Accordingly, the UV dose of such lineages also increased because of growth feedback-induced UV irradiation. As a result, both growth curves and UV doses in the latter period varied between the lineages ( Fig. 4d , f ). The maximum growth rates in the absence of UV irradiation changed only a little through the mutation accumulation experiment ( Suppl. Fig. S2 ). These results indicated that UV tolerance of the propagated cells in some lineages was improved during the cultivation. We note that there was significant variability in the growth rate in the absence of UV irradiation for lineage 1 at day 56 ( Suppl. Fig. S2 ), which might suggest potential variance of UV tolerance between individuals in this lineage.
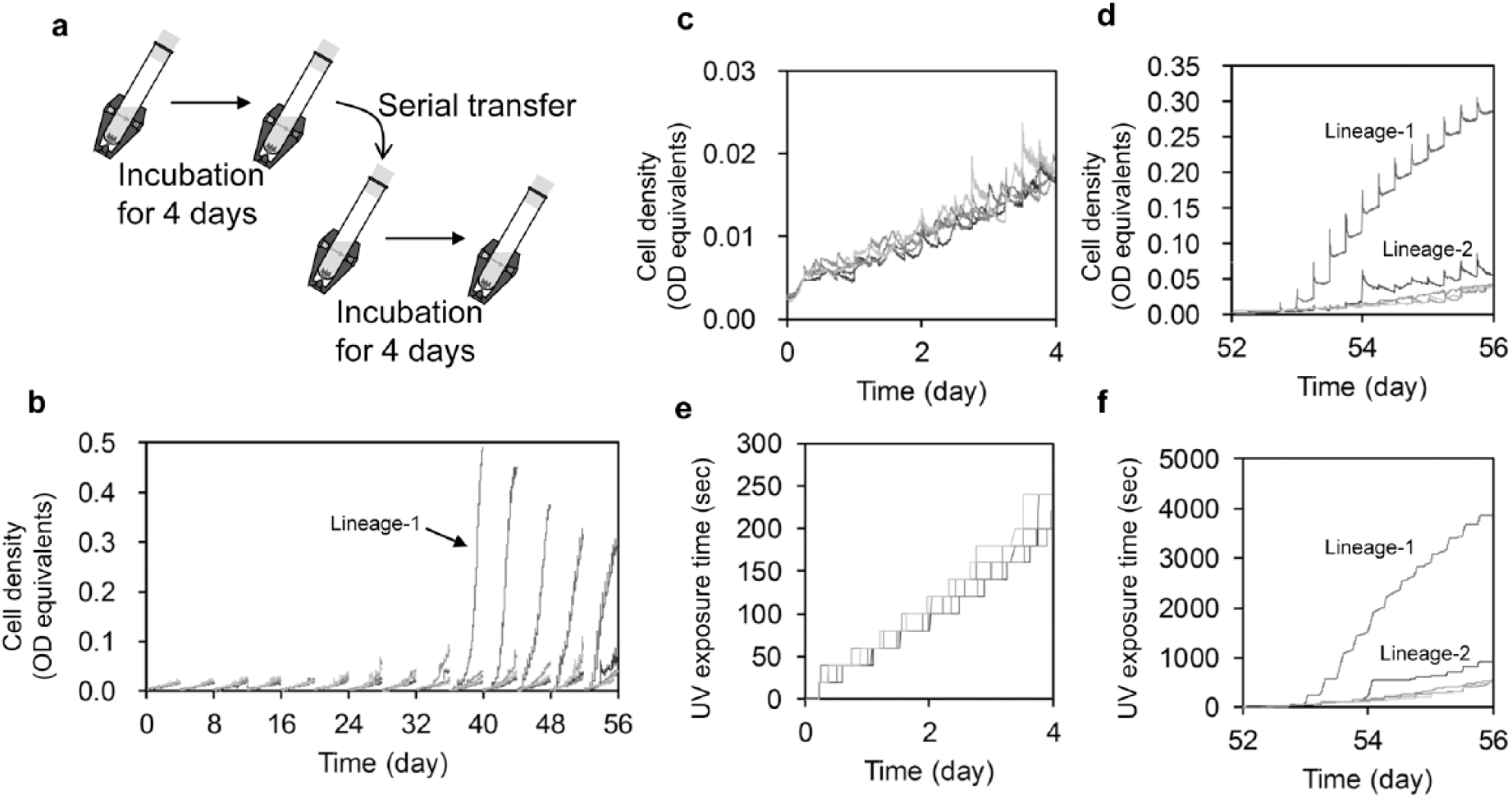
Growth curves in the repeated serial transfer with automated UV irradiation. (
Detection of Accumulated Mutations
Whole-genome sequencing revealed the number of genomic mutations accumulated in the populations. Analyzing the cell populations at the end of the evolution experiment (day 56), we detected 1389 base-pair substitutions (BPSs) and 33 small insertions and deletions (InDels) as fixed in the populations (278 BPSs per lineage and 7 small InDels per lineage on average). The accumulation rate of BPSs during the evolution experiments was calculated using the number of synonymous substitutions (
Fig. 5a
). As a result, the accumulation rates were 4–6 BPSs per genome per day, which was highly comparable to the previous results using manual UV irradiation (3 BPSs per genome per day; Shibai et al.
8
). These accumulation rates are approximately hundreds of times larger than the spontaneous mutation rates of a wild-type strain with proficient error correction or tens of times larger than the rates of a mutator strain with deficient DNA mismatch repair.
8
We also calculated the d
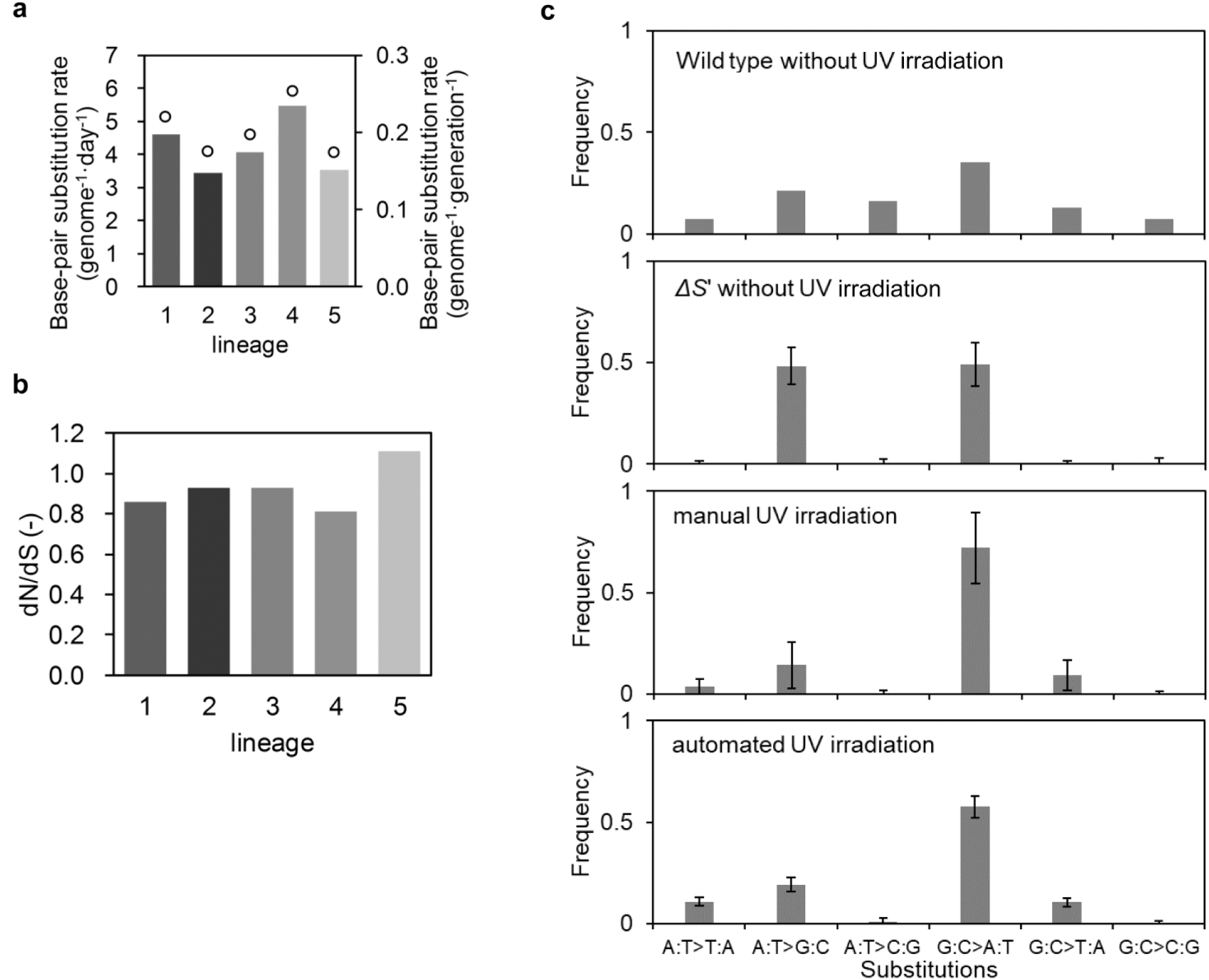
Accumulated mutations. (
Discussion
We constructed an automated UV irradiation device for microbial cell culture. This device can automatically monitor cell density and irradiate UV light to the bacterial cells according to the cell growth state. This growth feedback control enabled us to accumulate mutations at a high rate for a long period while avoiding extinction. The accumulation speed, neutrality, and spectrum of UV-induced mutations were consistent with those obtained by manual UV irradiation. These results indicated that our device is useful to accelerate mutation accumulation over long periods so as to explore genomic evolution in bacterial cells.
Variance in mutation accumulation rate was not particularly different between the lineages (4.8 ± 0.8 per genome per day), even though the UV dose varied greatly (
Fig. 5a
). There was no significant correlation between total UV exposure time and accumulated synonymous BPSs (
Suppl. Fig. S3
; ρ = 0.43,
Supplemental Material
Suppremental_Material_for_Automated_UV_Culture_by_Shibai,_et_al – Supplemental material for Development of an Automated UV Irradiation Device for Microbial Cell Culture
Supplemental material, Suppremental_Material_for_Automated_UV_Culture_by_Shibai,_et_al for Development of an Automated UV Irradiation Device for Microbial Cell Culture by Atsushi Shibai, Saburo Tsuru and Tetsuya Yomo in SLAS Technology
Footnotes
Acknowledgements
We thank Dr. Shota Nakamura and Dr. Daisuke Motooka for whole-genome resequencing.
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was partly supported by Japan Society for the Promotion of Science KAKENHI grants 15KT0078, 15H01682, and 26440200.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
