Abstract
In gold nanoparticle (GNP)–based colorimetric biosensors, the gradual color shift is often used to correlate with the concentration of target molecules, and therefore a UV-vis spectrometer is usually required for accurate quantification. Here, we present a critical coagulation concentration (CCC)–based salt titration as a generic and simple way to enable accurate quantification in GNP-based colorimetric biosensors without any analytical equipment. The titration is carried out by stepwise addition of a salt titrant to the premixture of sample and GNPs until the color changes rapidly from red to blue, determined solely by visual inspection. The number of titration steps or the final salt concentration required for a rapid color shift (i.e., CCC) is then used to quantitatively correlate with the concentration of target molecules in the sample. The salt titration–based quantification has been demonstrated with two previously reported GNP-based colorimetric biosensors. Compared with quantification based on the gradual color shift with a spectrometer, the visual quantification based on the rapid color shift in the salt titration eliminates the need for any analytical equipment without sacrificing the performance (i.e., sensitivity and accuracy) and therefore is highly suitable for applications in low-resource settings.
Introduction
Gold nanoparticles (GNPs) of various sizes can be prepared at low cost and high reproducibility. Due to their unique chemical and physical properties, GNPs have been widely used in various biosensors,1–4 especially colorimetric biosensors. The GNP-based colorimetric biosensors used the size- and distance-dependent optical absorption due to surface plasmon resonance.5,6 With the increase of particle size and/or the decrease of interparticle distance, the optical absorption peak of GNPs shifts to a longer wavelength, resulting in a visible color change of GNPs from red to purple or blue. Many colorimetric biosensors have been developed by using GNPs as a colorimetric indicator of target-induced aggregation (i.e., agglutination).
The GNP-based colorimetric biosensors can be classified into two major types based on the mode of target-induced aggregation: crosslinking and non-crosslink. For the crosslinking type of GNP-based colorimetric biosensors, target molecules mediate the crosslinking of GNPs, and labeling of GNPs with target-specific ligands is usually required. In the absence of target molecules, labeled GNPs will be well dispersed, thus maintaining a red color. However, in the presence of target molecules, target molecules will simultaneously bind to the ligands on different GNPs, thus crosslinking the GNPs, resulting in a target-specific color shift. By labeling GNPs with different types of ligands (e.g., antibodies, antisense oligonucleotides, and aptamers), detection of various nucleic acids,7,8 proteins, 9 and small molecules10,11 has been demonstrated. For the non-crosslinking type of GNP-based colorimetric biosensors, target molecules induce the aggregation of GNPs in a non-crosslinking manner, and no labeling on GNPs is usually required. These label-free GNP-based colorimetric biosensors rely on the effect of target molecules on the stability of GNPs. Depending on the design of the biosensors, the target molecules could stabilize or destabilize the GNPs in a direct or an indirect way. For example, based on the direct interaction of melamine and GNPs, our group has previously developed a colorimetric biosensor for melamine. 12 Based on the observation that oligonucleotides of different conformations could adsorb to GNPs in a different strength and thus stabilize the GNPs to a different extent, several label-free colorimetric biosensors have been developed for various molecular targets, including small molecules,13–15 nucleic acids,16,17 and proteins. 18 More specifically, oligonucleotide probes are carefully designed so that their conformations are sensitive to the presence of specific molecular targets. When the specific molecular targets are introduced, the oligonucleotide probes will undergo a target-specific conformational change. This will change the propensities of the oligonucleotide probes to adsorb on GNPs, resulting in a target-specific change in the stability of GNPs. Upon addition of appropriate amount of salts, a target-specific colorimetric signal can be observed.
In many of these GNP-based colorimetric biosensors, a gradual color shift is usually used to quantify the extent of the target-induced stability change of GNPs and thus the concentration of target molecules. The target concentration–dependent color shift can be observed with unassisted eyes. However, visual quantification based on a gradual color shift is not reliable. To achieve a reliable and accurate quantification, an absorbance reader or a more sophisticated scattering-based instrument19,20 is usually required for detection in the liquid phase, and image capturing and processing devices are often needed for detection on the solid substrate (e.g., thin-layer chromatography plate 21 ). Herein, we present a critical coagulation concentration (CCC)–based salt titration as a means to enable an accurate quantification in the liquid phase without any analytical equipment.
According to the DLVO (Derjaguin and Landau, Verwey and Overbeek) theory of colloid stability, stability of colloid is determined by the combined effects of the van der Waals attraction and the electrostatic repulsion between colloid particles.22,23 The total potential energy function (linear addition of the attractive and the repulsive potentials vs. particle separation distance) suggests that aggregation of colloid is always thermodynamically favorable. It is the primary maximum in the total potential energy that creates a kinetic energy barrier to aggregation. This kinetic energy barrier depends on many properties of both the particles and the media, such as particle size, surface potential, and salt concentration. Generally speaking, the small the barrier, the faster the aggregation. The CCC is the salt concentration where the kinetic energy barrier against aggregation just disappears and coagulation becomes rapid (i.e., diffusion limited). Compared with surface potential (or ζ potential, an experimentally accessible indicator of colloid stability), CCC could be a very sensitive and easily quantifiable indicator of colloid stability. For example, in colloidal systems with a low surface potential, the CCC is proportional to the fourth power of the surface potential. Therefore, we speculate that CCC-based salt titration could be a reliable way to visually quantify the target-induced stability change of GNPs and thus the concentration of target molecules in these GNP-based colorimetric biosensors. The titration is carried out by stepwise addition of a salt titrant to the mixture of sample, GNPs, and probe (if applicable) until the color changes rapidly from red to blue, determined solely by visual inspection. The number of titration steps (or the final salt concentration) required for a rapid color shift is then used to quantitatively correlate with the concentration of target molecules in the sample. In this article, the feasibility of this CCC-based quantification method is first demonstrated with two previously reported GNP-based colorimetric biosensors, and then effects of titrant type, titrant concentration, and titration interval are studied.
Materials and Methods
Materials
Melamine was purchased from Alfa Aesar (Ward Hill, MA). The 1000-ppm stock solution of melamine was prepared by dissolving melamine powder with deionized water and then stored at room temperature. Oligonucleotides were obtained from Integrated DNA Technologies (Coralville, IA). The 100-µM stock solutions of oligonucleotides were prepared by resuspending oligonucleotides in nuclease-free water and stored in aliquots at −20 °C until use. Tetrachloroauric acid (HAuCl4), trisodium citrate, sodium chloride (NaCl), potassium chloride (KCl), calcium chloride (CaCl2), and magnesium chloride (MgCl2) were purchased from Sigma-Aldrich (St. Louis, MO). If not specified, all the other reagents were also purchased from Sigma-Aldrich.
Synthesis of GNPs
GNPs were synthesized by reduction of gold hydrochlorate by citrate according to the literature.24,25 After 300 mL of 1 mM HAuCl4 was brought to boil while being stirred and refluxed, 30 mL of 38.8 mM trisodium citrate was quickly added into the boiling HAuCl4 solution. The solution was continuously heated and stirred until the color turned deep red. After the solution was cooled to room temperature, it was filtered with a 0.2-µm syringe filter. The filtered GNP solution was stored at 4 °C until use.
Characterization of GNPs
The UV-vis spectra of the GNP solution were obtained on a Synergy 2 microplate reader (BioTek, Winooski, VT) and then used to determine the size and concentration of GNPs according to the literature. 26 The size was estimated to be around 14.05 nm, and the concentration was estimated to be around 13.22 nM. Furthermore, the hydrodynamic size and the ζ potential of GNPs were determined using a Delsa Nano C particle analyzer (Beckman Coulter, Miami, FL). The dynamic light-scattering (DLS) analysis was performed in a disposable sizing cell at a scattering angle of 165° for 70 accumulation cycles. The ζ potential analysis was performed with a disposable ζ cell at a fixed voltage of 60 V for 10 measurement cycles. The DLS was repeated at least three times, and the ζ potential analysis was repeated at least six times. The hydrodynamic size of GNPs was determined to be 24.23 ± 0.46 nm, and the ζ potential of GNPs was determined to be −52.32 ± 3.26 mV.
CCC-Based Salt Titration
In the titration experiments for melamine detection, the serial dilutions of melamine from 0.3 ppm to 0.01 ppm were mixed with the 6.61-nM GNP (2-fold dilution of the GNP stock solution) in an equal volume ratio and incubated for 2 h at room temperature. Then, 160 µL of the melamine and GNP mixture was transferred to a breakable 12-well microtiter strip. The titration was performed by stepwise addition of the salt solution with an electronic pipettor. In each titration step, an equal volume of the salt solution was added, followed by a 5-s mixing. The number of titration steps required for a rapid red-to-blue color shift was recorded and then used to calculate the total titrant volume and the final salt concentration. For each sample, the salt titration was repeated at least five times.
In the titration experiment for DNA detection, the probe (5′- GAC CTT AGA CTT GAC ATG CTT CTC GAC GAC-3′) and the target (5′- GTC GTC GAG AAG CAT GTC AAG TCT AAG GTC-3′) were mixed in different molar ratios in 0.1× phosphate-buffered saline (PBS) supplemented with 15 mM NaCl and 5 mM KCl and hybridized overnight at room temperature. The probe and target mixture were then mixed with 6.61 nM GNP solution in a volume ratio of 1 to 2 and incubated for 1.5 h at room temperature. Then, 120 µL of the resulting mixture was used in the titration experiment. In each titration step, 10 µL of 350-mM NaCl solution was added, followed by a 2-s mixing.
Results and Discussion
Demonstration of the CCC-Based Salt Titration in Melamine Detection
To demonstrate the feasibility of the CCC-based salt titration as a reliable means of quantification in the GNP-based colorimetric biosensors, the CCC-based salt titration was first applied to a GNP-based colorimetric method for melamine detection, which was previously developed by our group. 12 As illustrated in Figure 1 , this method relies on the observation that melamine has a strong interaction with GNPs, and this strong interaction causes GNPs to aggregate in a melamine concentration-dependent manner. Even when melamine is present in a level as low as sub ppm, the melamine-induced color change of GNPs can be discerned with unassisted eyes. However, a spectrometer is still necessary for accurate quantification of melamine concentration because visual examination of the gradual color shift is not reliable.
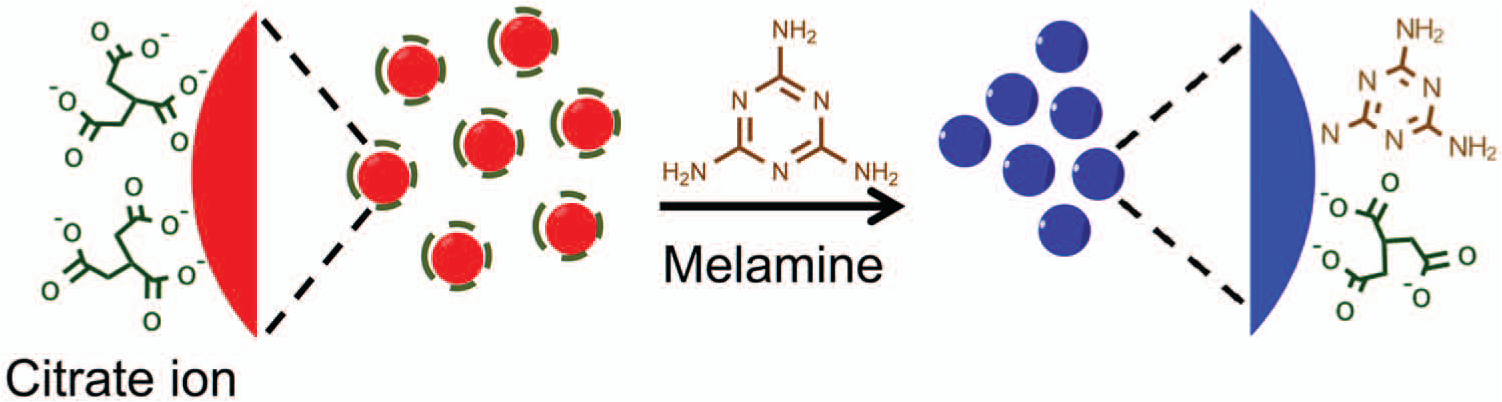
Schematic illustration of a previously reported gold nanoparticle–based colorimetric method for melamine detection.
A titration was performed with 100 mM NaCl on a series of premixtures of GNP and melamine. Then, 10 µL 100 mM NaCl was added to the premixture of GNP and melamine every 5 s until a rapid red-to-blue color shift was observed. The number of titration steps was then recorded and used to derive the total titrant volume and the final salt concentration required for a rapid color shift (i.e., CCC). As shown in Figure 2a – c , with the increase of melamine concentration, the number of titration steps, as well as the derived total titrant volume and the CCC, decreases, indicating the melamine concentration-dependent destabilization of GNPs. All three indicators (number of titration steps, total titrant volume, and CCC) can be used to quantitatively correlate with the melamine concentration. This has enabled us to visually quantify melamine as low as 0.02 ppm, which is one order of magnitude lower than that achieved using a spectrometer without the CCC-based salt titration ( Fig. 2d ). A comparison of CCC and ζ potential has verified that CCC is a much more sensitive indicator of GNP stability ( Fig. 2e ). For the melamine-destabilized GNPs, the CCC is roughly proportional to the fourth power of the ζ potential, as predicted by the DLVO colloid stability theory.
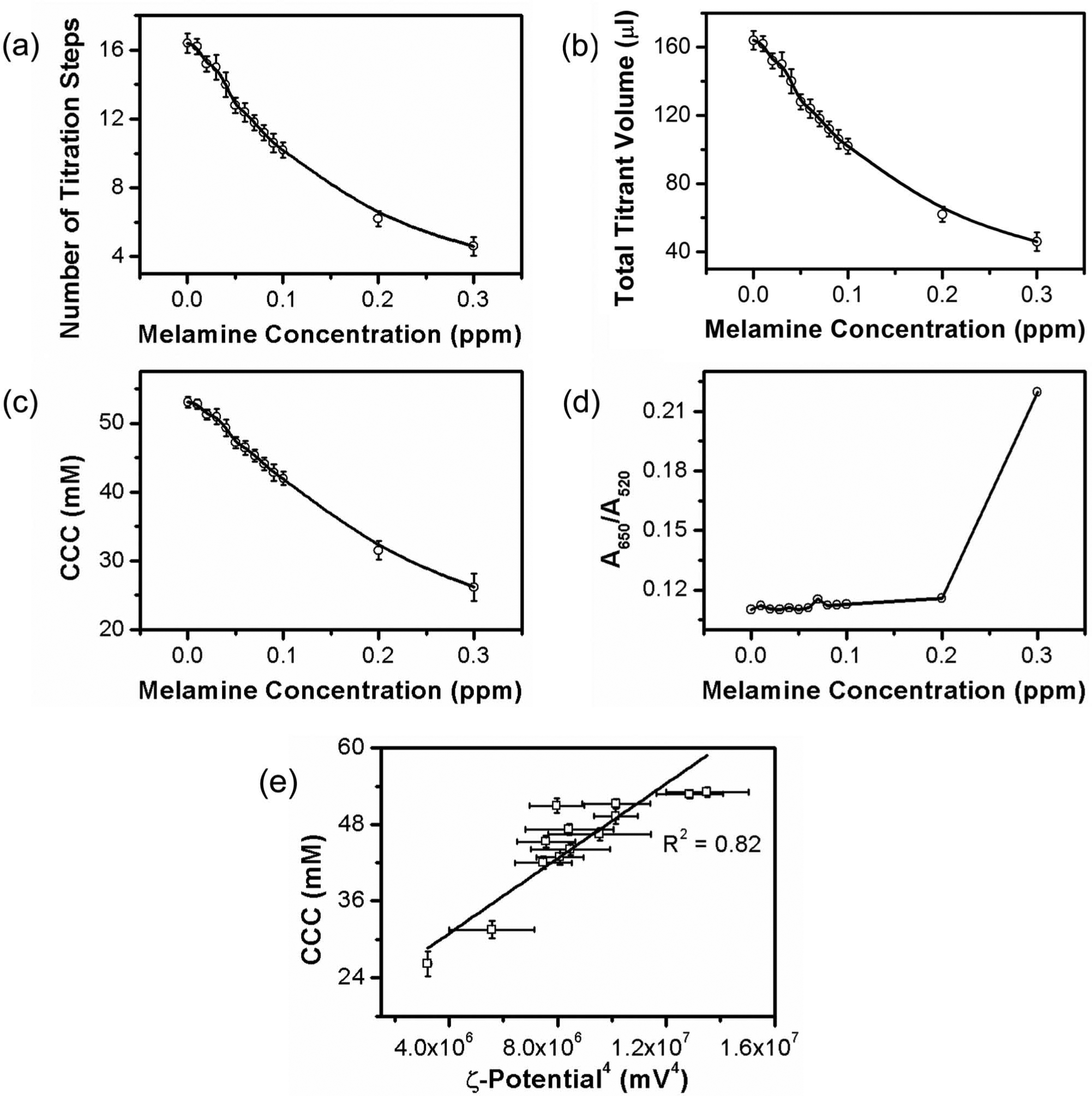
Visual quantification of melamine with the critical coagulation concentration (CCC)–based salt titration. (
Demonstration of the CCC-Based Salt Titration in DNA Detection
To examine the general applicability of the CCC-based salt titration to the GNP-based colorimetric biosensors, the CCC-based salt titration was then applied to a GNP-based colorimetric method for DNA detection, which was reported previously by Rothberg’s group.16,17 As illustrated in Figure 3 , this method uses the different effects of single-stranded DNA and double-stranded DNA on the stability of GNPs. An oligonucleotide probe complementary to the target DNA molecule is used along with unmodified citrate-capped GNPs. In the absence of targets, probes remain single-stranded. In the presence of targets, probes form double-stranded duplexes with targets. Compared with the double-stranded probe and target duplexes, the single-stranded probes bind more tightly to GNPs, thus stabilizing GNPs to a greater extent. When an appropriate amount of salts is added, the color of GNPs remains red if there are no targets and changes to blue if there are targets. Based on the extent of the color change, the concentration of targets can be accurately quantified with the help of a spectrometer.
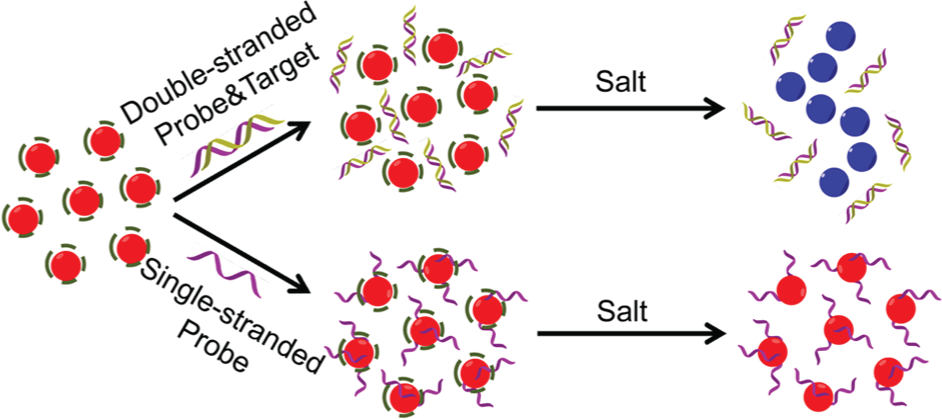
Schematic illustration of a previously reported gold nanoparticle–based colorimetric method for DNA detection.
Before the titration, the probe and the target were first prehybridized in different molar ratios (change the target concentration while keeping the probe concentration fixed at 562.5 nM) and then mixed with GNPs. The titration was performed with stepwise addition of 10 µL 350 mM NaCl every 2 s until a rapid red-to-blue color shift occurred. In Figure 4a – c , the number of titration steps recorded upon the rapid color change, the derived total titrant volume, and the derived CCC are plotted versus the molar ratio of the target to the probe ([Target]/[Probe]). All three indicators show a decreasing trend with the increase of [Target]/[Probe] (i.e., the target concentration), which indicates that the stability of GNPs against salt-induced aggregation decreases when the target concentration increases. The quantitative correlation between the three indicators and the target concentration has allowed us to visually quantify the target DNA even when the [Target]/[Probe] ratio is as small as 0.05. As a comparison, quantification based on the gradual color shift upon addition of salt was carried out. After a salt solution was added to the mixture of target, probe, and GNP in a 1:1 volume ratio, the consequential color change was characterized with the ratio of absorbance at 650 nm and 520 nm (A650/A520), which was obtained on a spectrometer. To differentiate the extent of a stabilizing effect on GNPs in the presence of different target concentrations, one needs to carefully choose the concentration of the salt solution. The results for four different salt concentrations are presented in Figure 4d . As expected, A650/A520 increases with the increase of the target concentration when the salt concentration is fixed, and A650/A520 increases with the increase of the salt concentration when the target concentration is fixed. Comparison between Figure 4a – c and Figure 4d reveals that visual quantification based on the rapid color shift in the CCC-based salt titration offers a sensitivity and an accuracy comparable to quantification based on the gradual color shift with a spectrometer. This has demonstrated that the CCC-based salt titration enables a visual quantification without sacrificing accuracy and sensitivity.
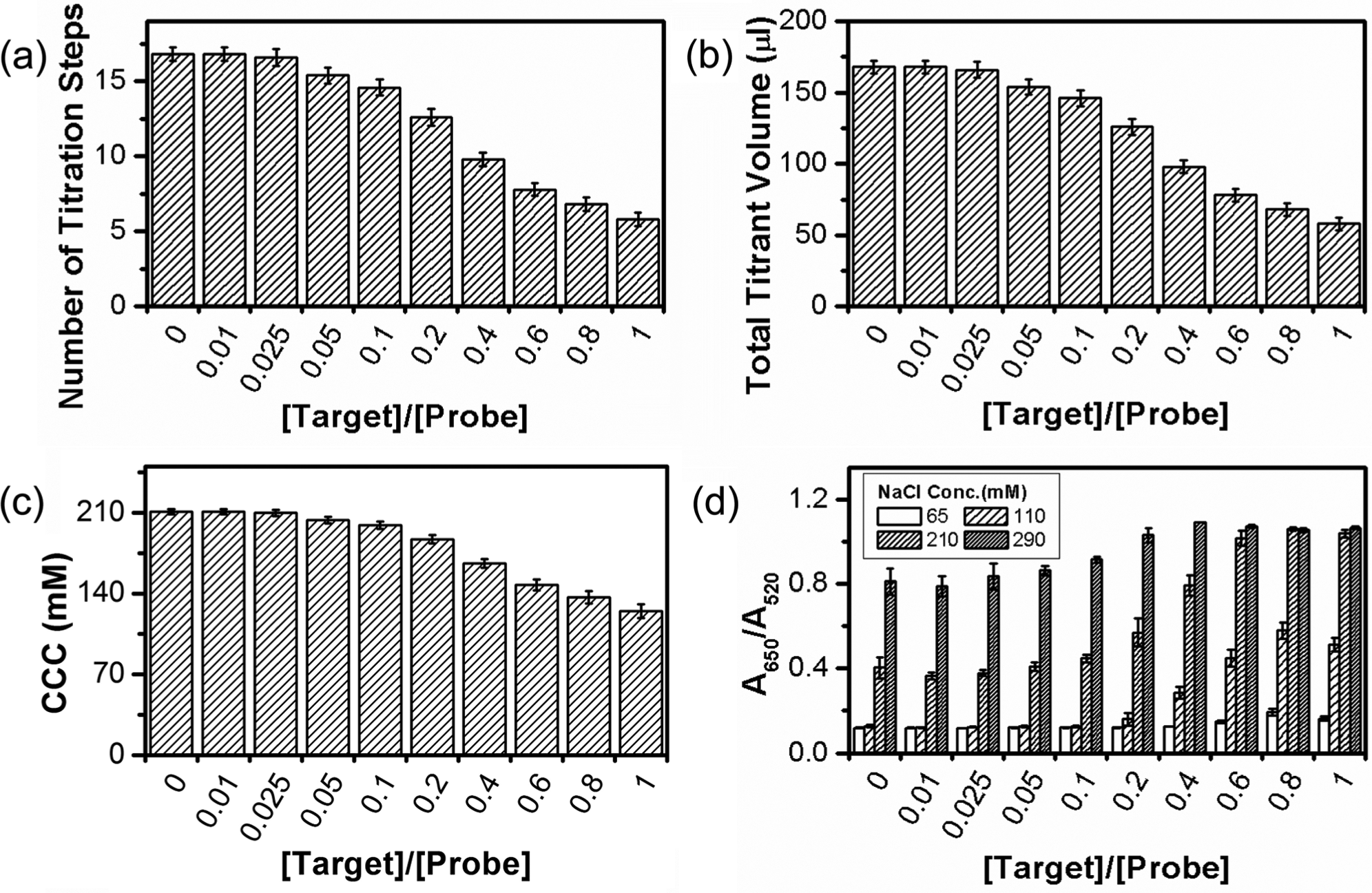
Visual quantification of DNA with the critical coagulation concentration (CCC)–based salt titration. (
Effects of Titrant Type, Titrant Concentration, and Titration Volume Interval
The performance of the salt titration–based quantification (e.g., sensitivity, dynamic range, and resolution) largely depends on the titration protocol. To obtain a better understanding of this, we studied the effects of titrant type, titrant concentration, and titration volume interval in GNP-based melamine detection.
Since CCC is more sensitive to the counterions (the ions with the opposite charge to GNPs), titrants containing different types of counterions were used to study the effect of titrant type. Four different types of chloride salts, including NaCl, KCl, CaCl2, and MgCl2, were compared. All titrations were performed with stepwise addition of 10-µL, 100-mM chlorides. With this titration protocol, the two divalent chloride salts could not provide a sufficient resolution to differentiate the melamine concentration-dependent destabilization of GNPs (data not shown). This is because CCC is very sensitive to the valence of counterions. As indicated by the Schulze-Hardy rule,27,28 for low surface potential colloid systems, the CCC is inversely proportional to the second power of the counterion valence; for high surface potential colloid systems, the CCC is inversely proportional to the sixth power of the counterion valence. Figure 5 shows the titration results with the two monovalent salts. For the same melamine concentration, number of titration steps required for the rapid color shift is more in the titration with NaCl than in the titration with KCl. This suggests that KCl has a stronger destabilizing power on GNPs than NaCl, which is not considered in the Schulze-Hardy rule, but agrees with the lyotropic series for flocculation of negative colloids. 29 It can be seen from Figure 5 that quantification via NaCl titration shows a higher accuracy and resolution than quantification via KCl titration. Use of titrant with a lower destabilizing power is expected to result in a more accurate and sensitive quantification; however, it would decrease the speed of quantification due to the increase in the number of titration steps. In addition to valence and destabilizing power, other properties of the titrant should also be taken into consideration when selecting the titrant. For example, the titrant should be inert to GNPs (no direct adsorption) and not affect the interactions between GNP, target, and probe (if applicable).
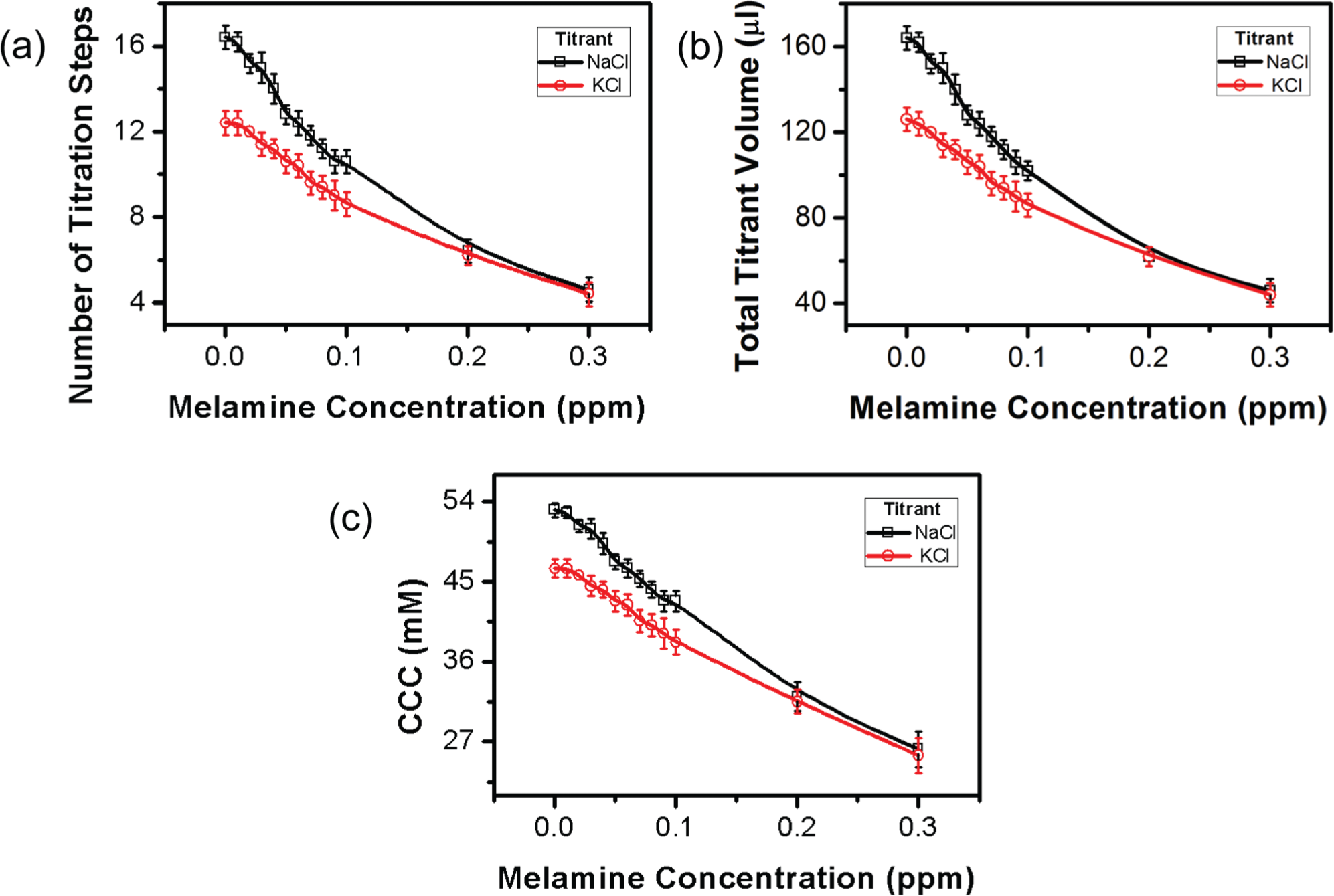
Effects of titrant type on the titration-based visual quantification. (
To study the effects of the titrant concentration, titrations were performed with three different NaCl concentrations (80, 100, and 120 mM). As shown in Figure 6 , number of titration steps and total titrant volume required for a rapid color shift increase with the decrease of the NaCl concentration, while the CCC remains almost the same. For a slightly different NaCl concentration, number of titration steps and total titrant volume are both significantly different, especially when the melamine concentration is low. This implies that the performance of the salt titration–based quantification is very sensitive to the concentration of titrant. For a given GNP-based colorimetric detection system, the titrant concentration should be fine-tuned for an optimal performance.
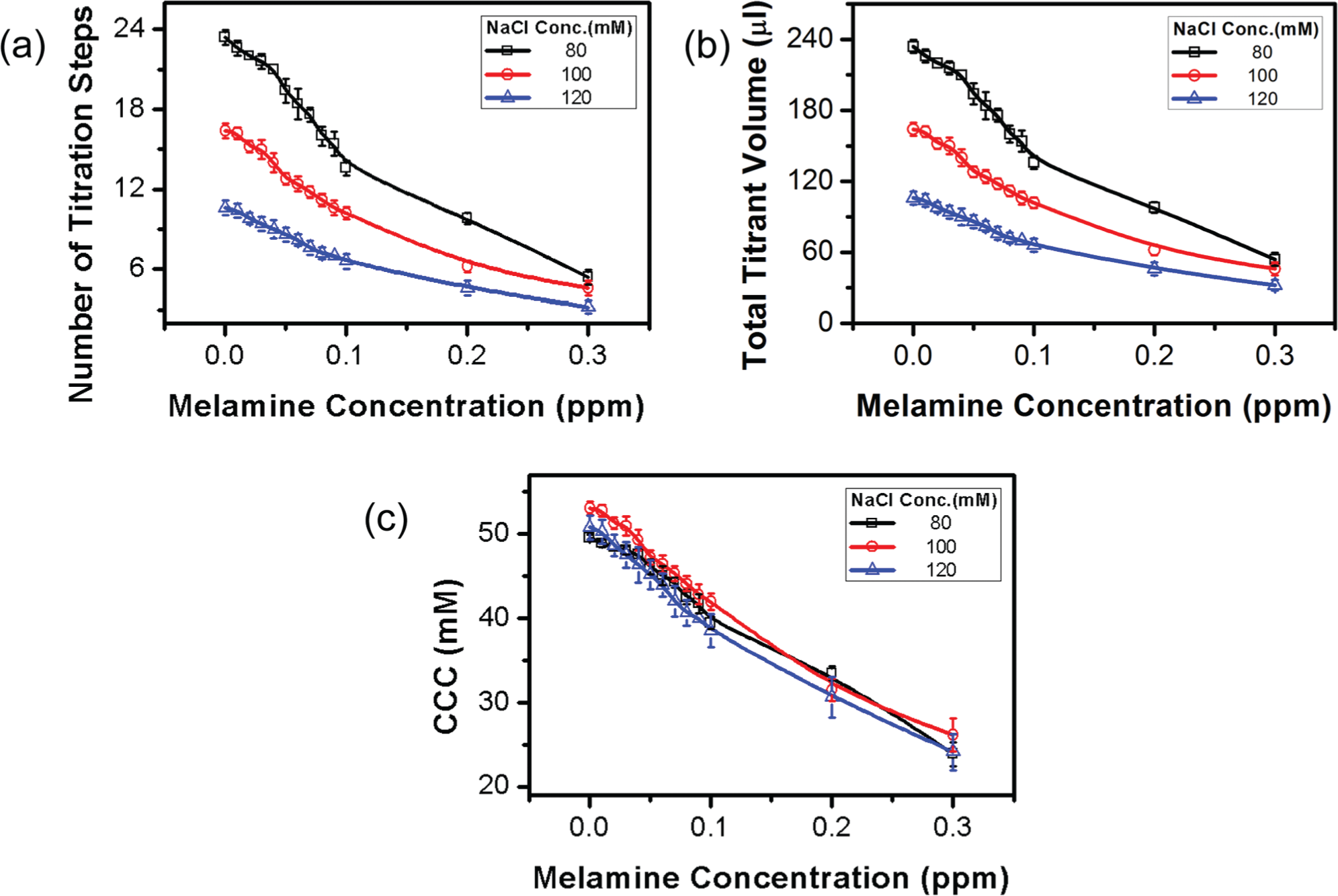
Effects of titrant concentration on the titration-based visual quantification. (
To study the effects of the titration volume interval (volume of titrant added in the each step), we compared three titration volume intervals (5, 10, and 20 µL) ( Fig. 7 ). With the increase of the titration volume interval, number of titration steps decreases while total titrant volume required for a rapid color shift and the CCC increases. The dependence of the CCC on the titration volume interval is more substantial when the melamine concentration is high. When the titration volume interval is small, the number of titration steps will increase and therefore it takes longer to titrate each sample. Because aggregation of GNPs is a kinetic process, the total amount of time required for conducting the titration will affect the final titrant concentration recorded upon the rapid color change (i.e., CCC). This effect is more significant when the GNPs are less stable in the presence of a high melamine concentration. As expected, a lower titration volume interval offers a better resolution and accuracy but a lower throughput in the salt titration–based quantification.
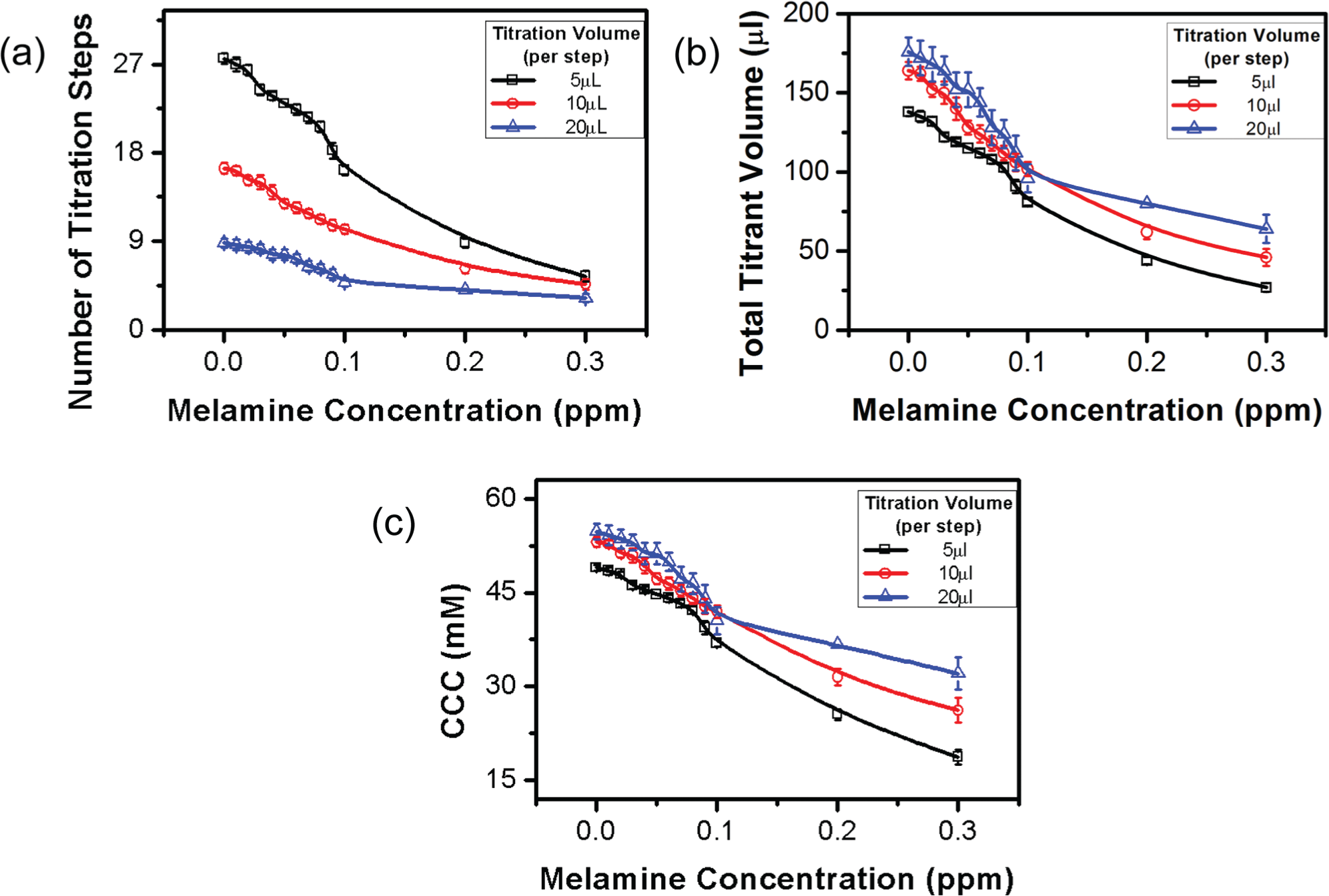
Effects of titration volume interval on the titration-based visual quantification. (
In conclusion, we developed a CCC-based salt titration for visual quantification in GNP-based colorimetric biosensors. By converting the qualitative, visual examination of gradual color change into a more quantifiable, rapid shift, it provides a simple yet reliable means to enable visual quantification in GNP-based colorimetric biosensors. The feasibility of this CCC salt titration–based quantification has been demonstrated with two previously reported GNP-based colorimetric biosensors. It has been shown that this CCC salt titration–based quantification has a similar (or even better) sensitivity and accuracy to the quantification based on the gradual color shift using a spectrometer. The effects of titrant type, titrant concentration, and titration volume interval on the performance of the CCC salt titration–based quantification were also investigated. Generally speaking, a titrant with lower destabilizing power, a lower titrant concentration, and/or a smaller titration volume interval would result in a more sensitive and accurate, but slower, quantification. An appropriate combination of titrant type, concentration, and titration volume interval should be chosen for each GNP-based colorimetric detection system.
Although the CCC salt titration–based quantification was demonstrated with two GNP-based colorimetric biosensors, it is expected to be generally applicable to other GNP-based colorimetric biosensors. With a performance comparable to that using analytical equipment, the equipment-free quantification method holds great promise to be used in low-resource settings.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the University of Miami through the Provost’s Research Awards.
