Abstract
The importance of biological assays spans from clinical diagnostics to environmental monitoring. Simultaneous detection of multiple analytes enhances the efficacy of bioassays by providing more data per assay under standardized conditions. Nevertheless, simultaneous handling and assaying of multiple samples, targets, and experimental conditions can be laborious, reagent consuming, and time intensive. Given these demands, microfluidic platforms have emerged over the past two decades as well-suited approaches for multiplexed assays. Microfluidic design supports integration of assay steps and reproducible sample manipulation across large sets of conditions—all relevant to multiplexed assays. Taken together, reduced reagent consumption, faster assay times, and potential for automation stemming from microfluidic assay design are attractive and needed multiplexed assay performance attributes. This review highlights recent advances in multiplexed bioanalyses benefitting from microfluidic integration.
Introduction
Since the late 1950s when radioimmunoassays were first demonstrated for the sensitive detection of insulin 1 and thyroxine 2 levels in human plasma, the application space for immunoassays has exploded. Modern workhorse formats are diverse, now spanning from enzyme-linked immunosorbent assays (ELISA) to Western blotting. From diagnostics for clinical questions to cell signaling studies in fundamental research, measurement needs are unified through the goal of delivering high-selectivity detection of biological or chemical entities in complex samples. A critical and oft-sought measurement capability is detection of multiple analytes while conducting just one single assay. Although not always the case, multiplexing can lead to faster results, 3 improved throughput, 4 and reduced sample volume requirements, thus reducing costs and biohazardous waste handling.5–8 In addition, multiplexed measurements enhance the confidence in assay outcomes. Microfluidic systems are of growing importance as platforms for multiplexed assays, owing to a capacity for integrated assay design, along with favorable material transport processes and limited material consumption. This review provides a brief context for multiplexed assays and then focuses on multiplexed assay developments realized with microfluidic design.
Multiplexed (or multiplex) assays address a wide range of applications: from measuring a disease biomarker in a biological specimen to understanding the proteomic state of cellular organisms to detecting chemicals in the environment. Specifically, multiplexed assays play a major role in clinical diagnostics. The assays answer questions regarding differential diagnosis or screening of disease biomarkers from correlated pathologies (such as hepatitis C virus [HCV] and human immunodeficiency virus [HIV] or other sexually transmitted diseases [STDs]). Measurement typically involves assessing the concentration of different biomolecules in a single biospecimen. For protein biomarkers, this may mean reporting multiple unique proteins,7,9 isoforms of one protein, 10 or different antibodies.11,12 In head trauma, for example, rapid comparative assays of biomarkers from blood and cerebrospinal fluid markers may allow severity diagnosis.13,14 For outbreaks involving bacterial infection, discrimination between multiple strains or determination of antibiotic resistance could reduce response times and inform drug therapy.15,16 In general medical practice, inflammatory marker assays, such as C-reactive protein (CRP) assays, are ordered in 4% of all consultations. 17 Although these inflammatory markers are nonspecific (e.g., cancer, cardiovascular disease, infectious viral or bacterial disease), 40% to 50% of assays are requested for a specific diagnostic purpose, 27% to 33% are requested for disease diagnostics, and 14% to 28% are requested for nonspecific diagnostic aims. Having multiplexed assays for the 70% of targeted assays would lead to faster and more accurate diagnostics. 18 For instance, verification of inflammation arising from a bacterial infection might be handled with antibiotic drug therapy with the length and the efficacy of therapy determined by monitoring CRP. 19 For diagnosis of a specific disease, multiplexed diagnostics are clearly important. 20 For example, multi-antibody Western blot and immunoblot assays provide confirmatory results after a positive screening result. Although labor intensive, confirmatory diagnostic assays offer high specificity and rule out false-negative screening results.21,22 Multiplexed assays also find use in drug screening,23,24 in particular testing for susceptibility, 25 cytotoxicity, or drug resistance.26,27 These formats can also be used in determining efficacy and toxicity of drugs.25,28 Multiplexing also underpins the fast genotyping afforded by nucleic acid tests, for genomic or epigenetic analysis.12,29 Multiplexing assays can support detection of diverse biomolecules, including combinations such as proteins and DNA, proteins and microRNA (miRNA), 30 or proteins and small molecules. 31
In fundamental life science inquiry, the ability to measure multiple targets can significantly enhance understanding of cellular function, including identification of sources of heterogeneity within cell populations. 32 Stochastic information in cellular response can be lost in ensemble averages inherent in analysis of bulk cell samples. 32 For example, monitoring of heterogeneity in cellular responses is critical in stem cell, tissue engineering, and regenerative medicine research. Analysis of cell signaling cascades involving multiple proteins and posttranslational modifications of these proteins are central to such investigations. Multiplexed assays can also provide spatial information associated with intracellular signaling,33,34 such as detail on analyte variation in 3D tissues 24 or subcellular localization. Multiplexed material handling can also assist complex analytical platforms (e.g., mass spectrometry) by increasing sample input throughput. 35
Historically, microtechnologies have played a role in the development of multiplexed analytical tools. In 1961, the first protein micro-spotting technology was demonstrated by Feinberg. 36 A microscope cover slide was coated with a thin layer of antiserum agar, and antigens were micro-spotted with fine capillary tubes. Following the incubation of the antigen-spotted cover glass in a moist environment, each micro-spot location was examined for precipitation. For strong antibody-antigen precipitation, signal was observed within 1 to 2 h or a day; for weak interactions, a 3-day-long incubation step was recommended. Then, in 1990, Erkins and colleagues37,38 reported micro-spot multianalyte detection systems. Soon after, technological advancements from the microelectronics industry began to percolate into assay design—in the form of microfluidic platforms—thus yielding increasingly sophisticated assay functions over the past 20 years, both in protein and nucleic acid measurement. Contributing substantially to advanced performance of microfluidic or “lab-on-a-chip” systems is the fine control of material transport and automation afforded, in turn leading to enhancement in assay reproducibility and quantitation. Fine control of fluid handing translates into lower required reagents volumes for assay operation (i.e., µL to nL regime), important in terms of reducing use of expensive or precious reagents. Fine fluid and material control also underpins integration of substantial sample preparation and material routing operations. 39 As anticipated from the microelectronics origins of microfluidic platform technologies, these miniaturized systems support integration of electronic components for sensing and data interpretation. 40 For interested readers, a summary of developments in the broad topic of microfluidic assays can be found in recent reviews.41–47
The next sections provide an overview of recent microfluidic design approaches for realizing multitarget detection. Both innovations from technology developers as well as necessities inherent in the application area dictate the solutions described. From surface patterning to bead-based systems, and from droplet-based systems to paper formats, challenges both unique and universal are encountered and surmounted. To provide a context for the reader, we have elected to detail approaches that are well established, as well as others that are budding with potential. Illustrative examples of recent multiplexed assay platforms are summarized in Table 1 .
Summary of Representative Microfluidic Assays with Multiplexing Capability.
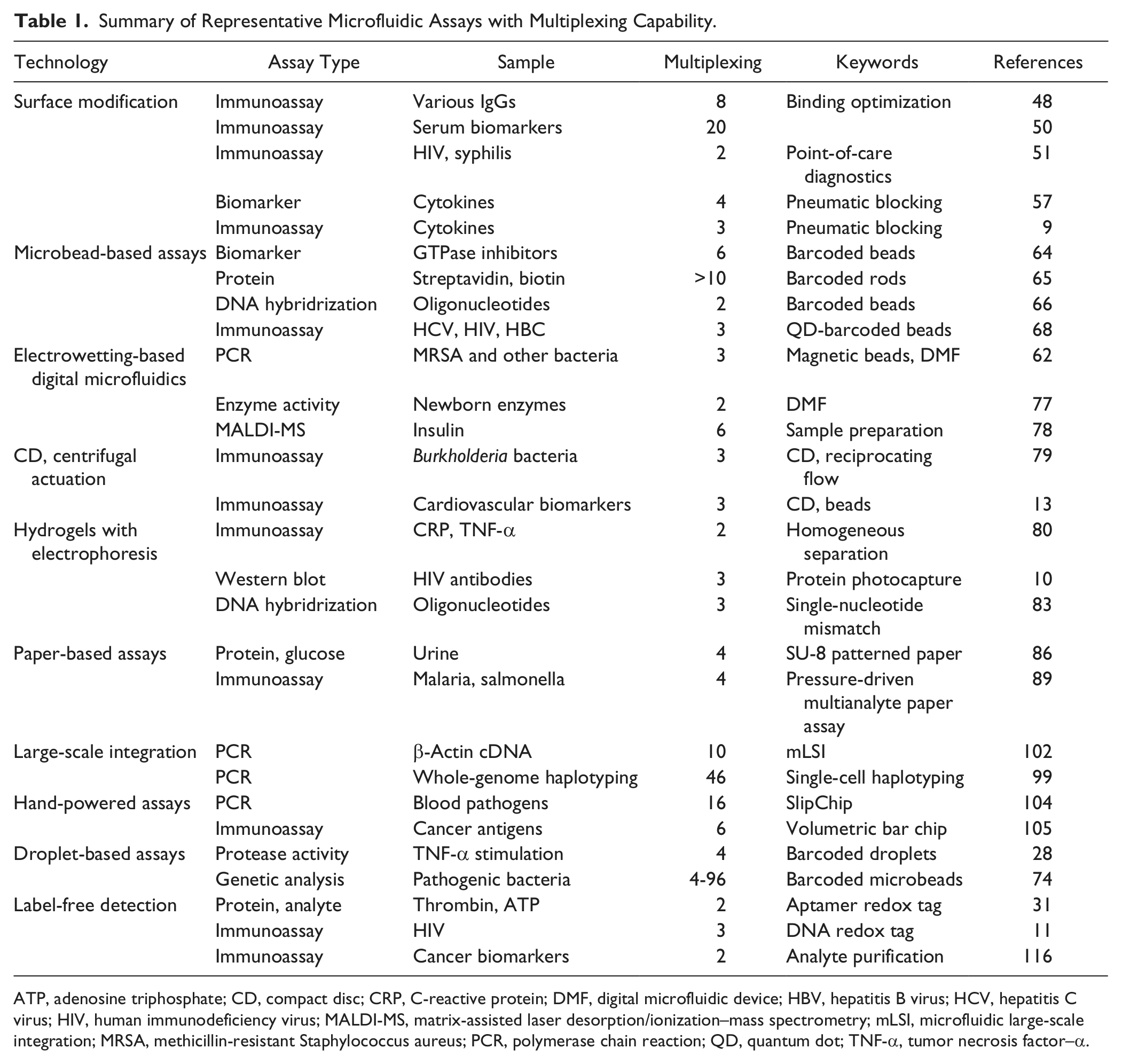
ATP, adenosine triphosphate; CD, compact disc; CRP, C-reactive protein; DMF, digital microfluidic device; HBV, hepatitis B virus; HCV, hepatitis C virus; HIV, human immunodeficiency virus; MALDI-MS, matrix-assisted laser desorption/ionization–mass spectrometry; mLSI, microfluidic large-scale integration; MRSA, methicillin-resistant Staphylococcus aureus; PCR, polymerase chain reaction; QD, quantum dot; TNF-α, tumor necrosis factor–α.
Surface-Patterned Assays for Facile Multiplexing
Adaption of heterogeneous multiplexed immunoassays to microchannel networks was driven in part by a need to reduce reagent and sample consumption, while conferring beneficial transport afforded by rational selection of operating conditions. The capacity to immobilize many reagents in a microchannel or microchamber enables detection of multiple analytes during one flow-through step. An early multianalyte heterogeneous immunoassay was demonstrated by Bernard et al. 48 In this work, a PDMS microfluidic channel network consisting of parallel channels was used as a fluid guide to pattern reagent stripes on a planar substrate. Following reagent patterning, the channel network was peeled off the substrate and a second similar network was placed orthogonal to the patterned reagent stripes. In this way, the new channel crossed multiple reagent stripes. Consequently, sample (flowed through the now-seated channel network) could be assayed against multiple immobilized reagents in one flow-through step. In total, the micromosaic assay allowed assessment of eight reagents against eight analytes (64× multiplexing) in a single assay. Based on a micromosaic patterning technique and a DNA-encoded antibody library technique, 49 Fan et al. 50 demonstrated a DNA-encoded antibody library (DEAL) blood barcode chip for the detection of up to 20 plasma proteins from whole blood within 10 min. Sample collection was performed via finger prick. The plasma proteins detected included cancer biomarkers and various cytokines and functional proteins involved in tumor growth, inflammatory response, or liver dysfunction. The DEAL barcode assay resulted in analytical sensitivities in the range of 1 to 30 pM depending on the sample. Most results were comparable with conventional ELISA, thus validating the assay. Although achieving appreciable multiplexing, the approach did require manual intervention between the patterning and assay stages.
With operation at the point of care as a design requirement, Chin et al. 51 demonstrated HIV and syphilis antibody detection in human serum using a microfluidic heterogeneous immunoassay. Both HIV and syphilis are STDs with a complex manifestation, including that the presence of one disease is likely to increase the risk of transmission of the other disease. 52 In a manner similar to the micromosaic strategies introduced earlier, immobilized antigens were enclosed in serpentine channels fabricated via injection molding in polystyrene substrates. Antibodies introduced through fluidic conduits were captured at the corresponding antigen zones. For detection and signal amplification, gold nanoparticles conjugated to anti–human antibodies and reduced silver growth over gold were employed to amplify readout signal. A key difference from the channel-based patterning of immobilized reagents was the use of manual or robotic spotting of the antigen, followed by sealing of the polystyrene channel network to a patterned polystyrene substrate. Use of plastics reduced the material cost to $0.10 per device. In a clinical field study, the duplex assays were tested in Rwanda with 67 patient specimens with known HIV and syphilis status. Each assay was completed within 15 min and yielded an overall clinical sensitivity of 100% and 94% and a clinical specificity of 95% and 76% for HIV and syphilis, respectively.
In a modular design approach, Henares et al. 53 used a PDMS microchannel network embedded with short segments of reagent-coated glass capillaries for the multiplexed detection of analytes. To create the modular device, reagents were first immobilized on the walls of 20- to 30-cm-long capillaries. Following the immobilization step, capillaries were cut into 2- to 3-mm-long segments and embedded in PDMS microchannels, thus creating discrete capture chambers with unique capture reagents in the PDMS channel network. By embedding multiple capillaries in parallel (each functionalized with one of anti–human, anti–chicken, and anti–goat antibodies), human, chicken, and goat antibodies were detected. For detection and signal amplification, anti–IgG–horseradish peroxidase (HRP) and substrate were used for a limit of detection (LOD) for human antibodies reported at 0.1 ng/mL. For human antibody samples tested at 100-pg/mL concentrations, multiplexing led to a 9% change in the single assay signal, attributed to cross-reactivity (3%) and use of multiple conduits (6%).
In a hybrid strategy, Didar et al. 54 combined microcontact printing with microfluidic patterning via laminar flow (laminar flow patterning). The fabrication approach was used to pattern silanized regions on a planar surface, followed by flow-based patterning of multiple antibodies. The microarray-in-a-microsystem assayed five different primary antibodies with secondary antibody-based detection specific to each immobilized primary antibody. Although the patterning step depended on the channel network design, the final assay was compatible with operation in a single channel, thus simplifying the assay workflow. However, since no clear boundary existed between the contiguous-patterned protein regions, extra care during the laminar flow–based patterning step was necessary to prevent false readouts when multispectral secondary labeling was not used.
Pneumatic actuation can be used to block the exposure of a liquid to a section of a microchannel by applying pressure to a membrane. 55 Zheng et al. 56 patterned distinct protein zones in microchannels and demonstrated a high-throughput assay used to detect a carcinoembryonic antigen in human serum with a 10-pg/mL LOD. Advancing the technique further, Garcia-Cordero et al. 57 patterned multiple analytes leading to multiplexed assays in a scalable, high-throughput platform for biomarker identification. Using this approach, they performed 384 multiplexed sandwich immunoassays. Each assay could analyze four different biomarkers, leading to a total of 1536 measurements. The platform was used to monitor the production of inflammatory cytokines from dendritic cells upon stimulation with 10 different adjuvants targeting Toll-like receptors. Each adjuvant was tested in various concentrations leading to 405 combinations in total. Two synergistic adjuvant interactions were identified and then validated with in vivo mouse models. In another study, Garcia-Cordero et al. 9 showed protein patterning of multiple species in a channel by tuning the pressure applied to a button membrane ( Fig. 1 ). The method enabled immobilization of three reagents in concentric annuli. As a demonstration of a multiplexed immunoassay, three cytokines (interleukin [IL–6], tumor necrosis factor [TNF–α], and IL-12p70) were captured at 100-pM concentrations. Proteins were patterned in a small footprint to simplify the final imaging step as well as enhance the scalability of the assay. Similar to the microarray system introduced by Didar et al., 54 contiguous-patterned protein regions (without any spacer or blank regions between protein regions) may lead to signal readout difficulties unless multispectral secondary detection methods are used. Although microfluidic systems provide enhanced reagent capture compared with bulk systems (due to higher surface-area-to-volume ratios) and offer favorable mass transport behavior, assays based on surface capture can be still limited by mass transport depending on the exact design and operation conditions used. 58
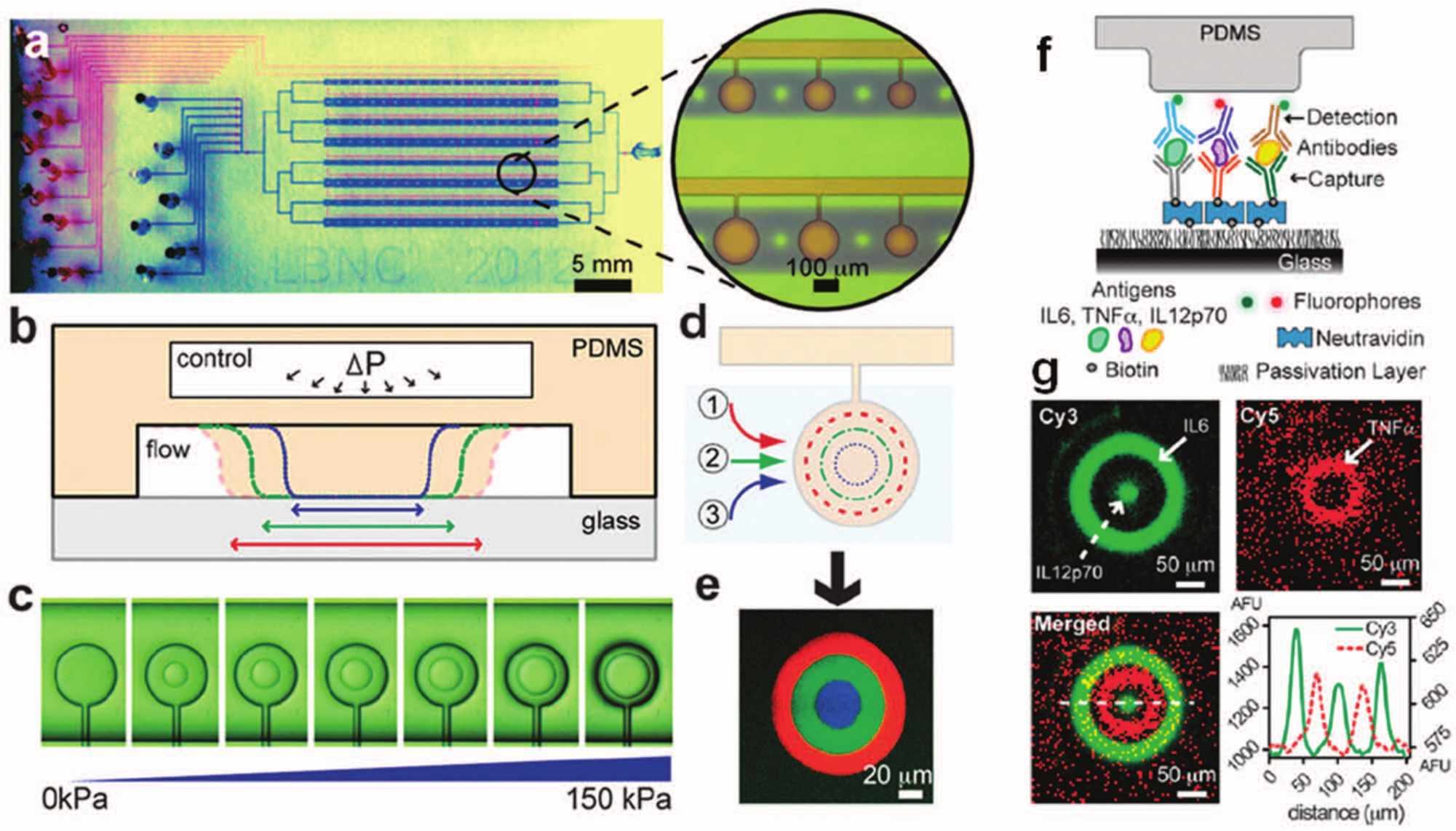
Protein patterning with a pneumatically controlled microfluidic button membrane. (
Microbead-Based Assays Take Advantage of Fluid Routing and Handling
Functionalized microbeads are a cornerstone of biotechnological assay design. Accordingly, myriad microfluidic approaches have been developed for the micromanipulation of microbeads using acoustical,59,60 optical, 61 magnetic, 62 or dielectrophoretic 63 interactions. With respect to multiplexed assays, functionalization of different beads with different analytes or spatially enclosing beads in different device compartments are the core strategies. Building on the infrastructure for flow cytometry, Surviladze et al. 64 reported a multiplexed assay testing the activity of six small GTPase proteins using microbeads. By screening six GTPases against 200,000 compounds, the authors identified two activity inhibitors. Custom microbead sets—each having different intensities of red fluorescence—were functionalized with a unique reagent. Correlation between the signal strength in the red fluorescence spectrum with the analyte signal in the green spectrum allowed identification of target analytes.
Building on the success of fluorescence labeling and size as encoding and identification approaches to multiplexed bead assays, barcoding has grown in importance. For example, Walton et al. 65 reported custom-fabricated microrods for barcoded and, thus, multiplexed detection of analytes. Fabricated on a porous alumina membrane, each microrod consisted of defined alternating metal layers that formed binary codes on the rods. Following fabrication, the barcoded rod was functionalized with a reagent. Alternating metal layers were visible under regular light microscopy and coded to the reagent patterned. Following the assay with the neutravidin-coated barcoded rods and fluorescently labeled biotin, comparison of a bright field and fluorescence image of a region of interest identified the target biotin. Availability of tens of unique barcode representations on microrod populations enabled the opportunity for multiplexed detection. In a similar vein, Lee et al. 66 demonstrated custom fabrication of color-coded magnetic polymer microbeads for multiplexed assays ( Fig. 2 ). Using the beads fabricated with an M-Ink method, 67 a multiplexed microfluidic DNA hybridization assay was demonstrated. Since the beads contained iron nanoparticles, the beads could be actuated by magnetic forces. Magnetic actuation was used to hold the beads in microfluidic chambers during the sample loading and washing steps and also facilitated rapid mixing via magnetic stirring. Incorporation of magnetic stirring enhanced the target analyte capture. Compared with the microrod-based system by Walton et al., 65 color-barcoded beads required a true color CCD camera for barcode identification.
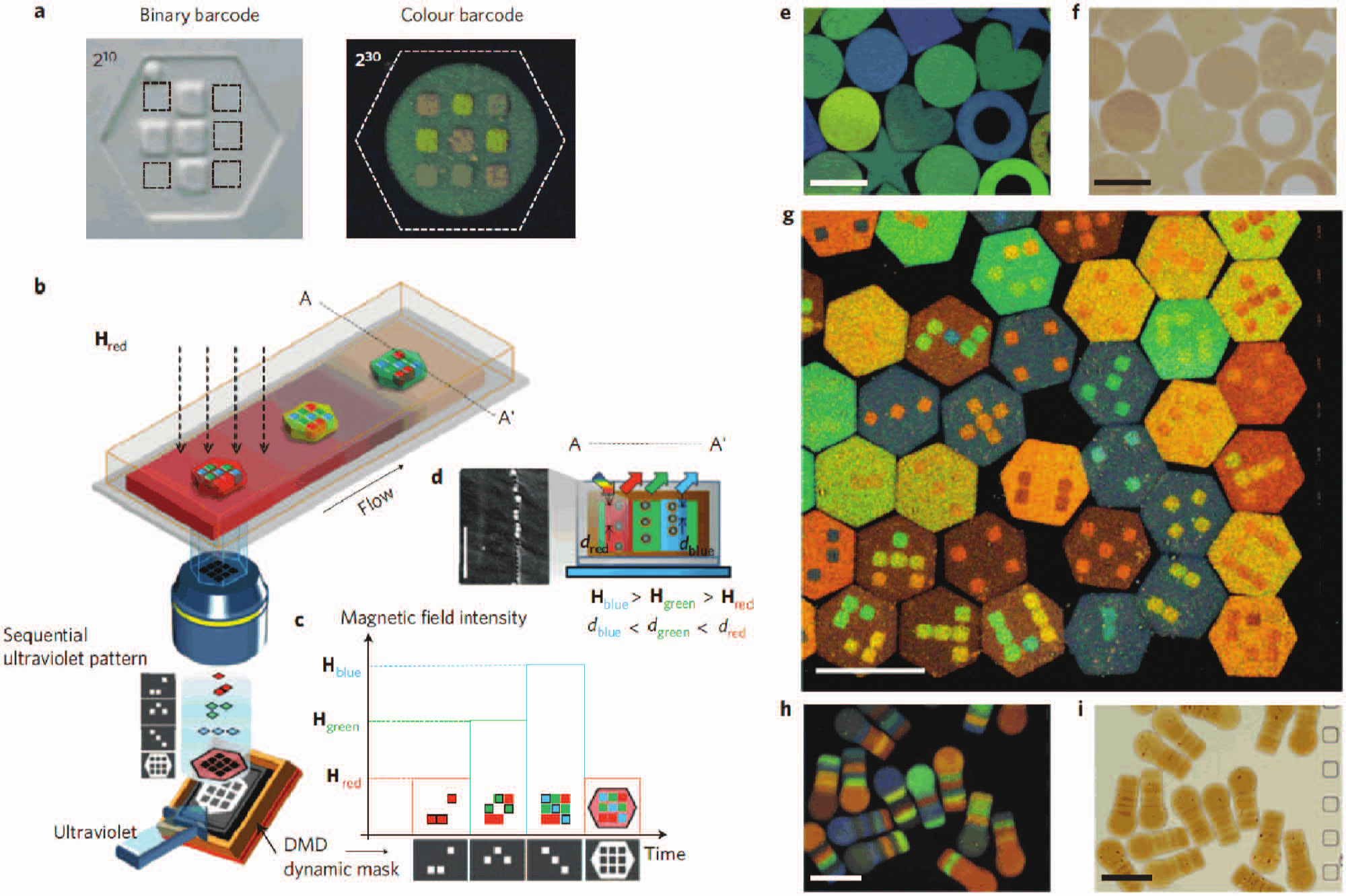
Color-barcoded magnetic particles. (
In another approach, spectrally multiplexed assays with quantum dot (QD) barcoded polystyrene beads functionalized with reagents have been reported in biomarker assays. For example, Klostranec et al.
68
demonstrated detection of HCV, HIV, and hepatitis B virus (HBV) antibodies spiked in human sera in a single assay offering picomolar sensitivity. These assays also detected genetic targets. Gao et al.
69
used four types of barcoded beads to detect HIV-1, HBV, and two biomarkers of
Microbead-based assays enable multiplexing and high-throughput performance, using well-established flow cytometry instrumentation and analysis methods.71,72 Since these applications require multispectral detection, these approaches are suited for established laboratory settings. Performance evaluations of few microbead-based commercial multiplexed assays and their comparison to established ELISAs can be found in the literature. 73 Microbeads have also played important roles as components of other microfluidic approaches such as compact disc (CD)–based, 13 droplet-based, 74 and digital microfluidic technologies. 75 Some of these approaches involving multiplexing capabilities will be discussed in the following sections.
Electrowetting-Based Digital Microfluidics for Sophisticated Sample Handling
Electrowetting phenomena were first reported arising from electric potential–dependent surface tension variation in early 1930s. 76 Electrowetting actuation is now a foundation of digital microfluidic devices (DMFs), wherein droplet actuation mechanisms depend on the displacement of a droplet of liquid on a hydrophobic surface that acts as a dielectric layer between the droplet and interdigitated electrodes. By applying an electrical potential to the electrodes, the surface tension of the region above the electrodes can be modified. The modified surface tension results in droplet transport from one electrode location to another. By manipulating multiple droplets over a checkerboard electrode array, multiplexed assays have been realized.
Using magnetic bead-based actuation and DMF, Hua et al.
62
reported a multiplexed assay based on polymerase chain reaction (PCR) amplification for methicillin-resistant
A similar assay strategy was used by Sista et al. 77 to demonstrate multiplexed detection of acid α-glucosidase and acid α-galactosidase (GAA) to screen for Pompe and Fabry disorders in newborns. Enzymatic assays conducted in disposable DMF cartridges detected insufficient enzymatic activities (an indicator of disease) within 6 h of sample incubation, whereas benchtop assays were reported to require 20 h. In addition to the decrease in assay time, a 3× enhancement in the assay analytical sensitivity was observed. In conjunction with a highly sensitive fluorometer, the authors reduced the assay incubation time to 1 h. The DMF assays required no pumps or valves and were developed to be scalable. Capacitive actuation led to low-power consumption (despite possible high voltages) and allowed isolation of sample from the electrodes, thus eliminating electrochemical reactions ( Fig. 3 ).
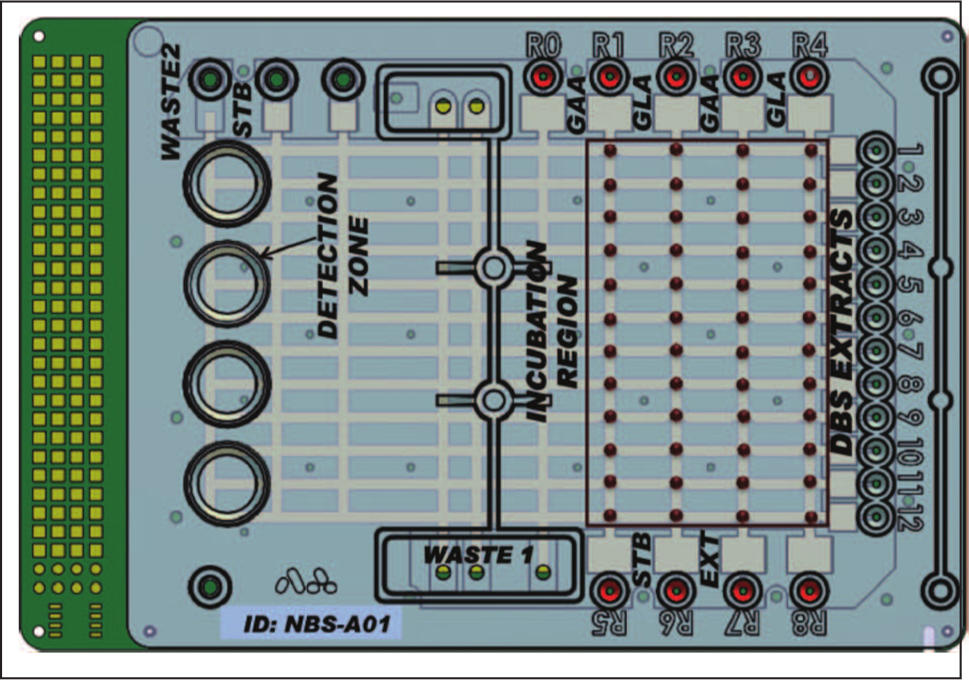
Disposable digital microfluidic cartridge for the multiplexed enzyme analysis assay based on electrowetting phenomena. Eleven droplets of GAA reagent from R1 were first dispensed to the channel. Then, these droplets were mixed with the 11 samples (dried blood spot extracts) dispensed from wells 1 to 11. The process was repeated for other reagents from R2 to R4. In the next step, all droplets were split into two droplets, and one was mixed with stop buffer (STB). Following a fluorescence measurement, all reaction droplets were incubated for 6 h at 37 °C. After incubation, each reaction droplet was mixed with STB and end-point fluorescence was measured and compared with the initial reading. The assay enabled testing of the 4-plex enzymatic activity of 11 newborn samples with automated fluid handling. Reprinted with permission from Sista et al. 77 Copyright 2011 American Association for Clinical Chemistry.
DMF devices have also been reported to assist sample preparation for mass spectrometry (MS) analysis. Reported by Moon et al., 78 droplets containing target molecules were cleared from impurities using DMF droplet manipulation with the resulting dried protein precipitate analyzed via matrix-assisted laser desorption/ionization (MALDI)–MS. By this method, six samples were subjected to purification and preparation steps in parallel for fast sequential analysis in MS.
Although powerful, DMF approaches do have limitations. Owing to the typical workflow for DMF assays—moving droplets over interdigitated electrodes—detection can be limited by droplet volume and the continuous nature of material transport across sensors (i.e., no accumulation of signal). As with many assays, nonspecific adsorption within DMF devices must be addressed with care, so as to reduce cross-contamination between droplets and assays.
CD Formats for Centrifugally Driven Fluid Actuation
CD-based microfluidic systems make use of centrifugal forces generated by rotating a “lab-on-a-disk” in a manner analogous to spinning CDs used for data storage. The rotation of the CD generates movement of liquid samples in networks of microchannels fabricated on the planar CD structure. Although a wide range of sample handling steps and assays have been demonstrated on CDs, Noroozi et al.
79
multiplexed a CD-based platform as shown in
Figure 4
. Antigens printed in a microarray on a 6 × 6-mm2 nitrocellulose membrane in the CD-based channel network served as the multiplexed capture substrate. Three antigens of
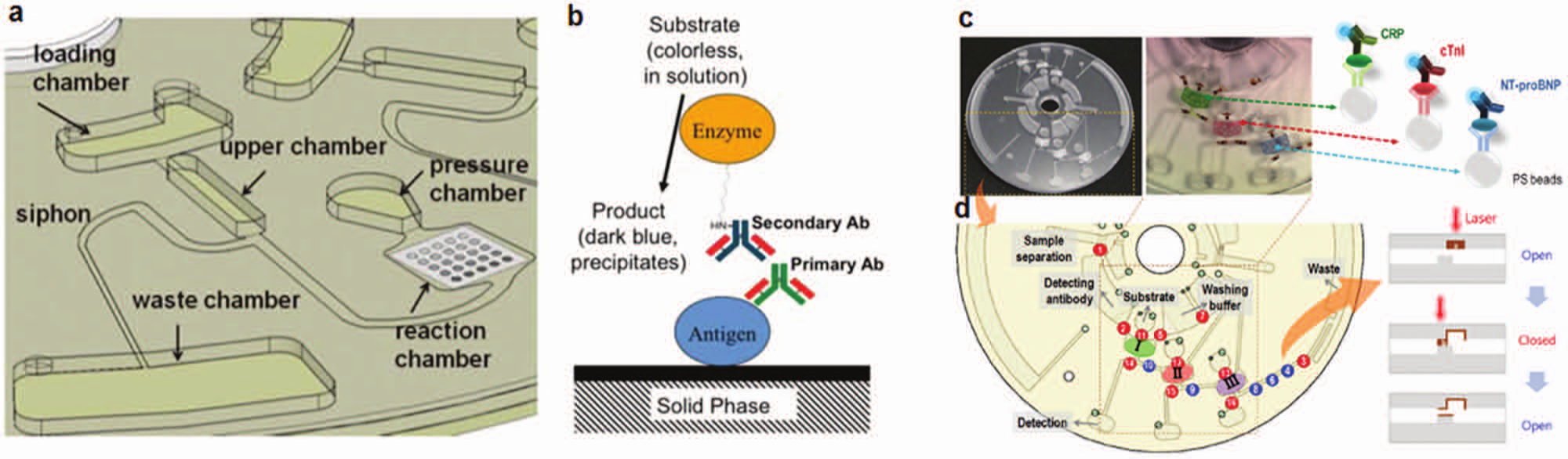
Centrifugal microfluidics on a disk. (
Lab-on-a-disk formats support sample preparation steps integrated with multianalyte assays. For example, Park et al. 13 demonstrated a multiplexed assay with integrated sample preparation for detection of CRP, cardiac troponin I (cTnI), and N-terminal pro–B-type natriuretic peptide (NT-proBNP) spiked into whole human saliva and blood for mock diagnosis of cardiovascular disease, as shown in Figure 4. To yield an ELISA, a blood sample was first directed through a centrifugal separation of plasma and cells. Subsequently, the plasma was assayed against three bead columns packed in the CD. The centrifugal force drove the plasma through the bead compartments, which were functionalized for detection of the target biomarkers. The method enabled two assays per disk. Given that each assay was used to detect three different analytes, six measurements were performed in a single run. In some applications, lab-on-a-disk platforms may have some shortcomings. For example, current CD assays use channel networks on relatively large footprint disks, needed to make the most of centrifugal forces. In addition, the technology depends on motors and aligned optical systems to detect assay results.
Hydrogels Used with Electrophoresis Support Homogeneous and Heterogeneous Assays
Lab-on-a-chip assays have been developed with the goal of harnessing efficient electrophoretic transport in microchannels housing hydrogels either as molecular sieving matrices or reactive supports. Homogeneous immunoassays for multiple targets have also seen advances in performance and capability with adaption to microfluidic formats. For example, for separation of bound and unbound species, polyacrylamide gel electrophoresis (PAGE) forms a high-performance format for mobility shift assays. Developed to allow low-power point-of-care electrophoresis assays, Hou et al. 80 reported a rapid multiplexed homogeneous immunoassay realized in ultra-short electrophoretic separation distances by using a discontinuous gel in PAGE. Contributing to appreciable electrophoretic separation between bound and unbound species (under often difficult native conditions), the sample stacked (concentrated) at the gel discontinuity as part of the PAGE separation. Through the use of multiple fluorophores, two antigens (CRP and TNF-α) were assayed against two antibodies. The in-chip gel shift assay completed in 10 s and required a 350-µm long separation distance. Similar slab gel PAGE separation times are on the order of an hour and require centimeter-long separation distances.
Using a photopatterning strategy in which polyacrylamide was used as both (1) a molecular sieving gel for electrophoretic separations and (2) a heterogeneous antibody-decorated capture surface, Tia et al. 81 demonstrated multianalyte Western blotting in a gel-patterned microchamber. In conventional Western blotting, probing one sample with multiple antibodies (reblotting) limits assay quantitation capacity and can degrade analytical sensitivity. Contiguous to the separation gel, multiple photopatterned blotting regions enabled simultaneous probing of all protein peaks with multiple antibodies. The fully integrated assay was completed in 5 min compared with the day-long conventional Western blotting workflow. In tackling the reblotting challenge with a different microsystem design, Hughes et al.10,82 demonstrated a photoactive hydrogel in which proteins separated by charge or size were immobilized in a benzophenone-decorated polyacrylamide gel matrix via UV irradiation (after separation) and probed by antibodies. The device, shown in Figure 5 , requires only one straight channel for the separation, photocapture (blotting), and subsequent antibody-based probing with multiplexed detection capability. In one example related to confirmatory diagnosis of infectious disease, HIV antigens separated via gel electrophoresis were immobilized, and corresponding HIV antibodies in positive human sera were detected.
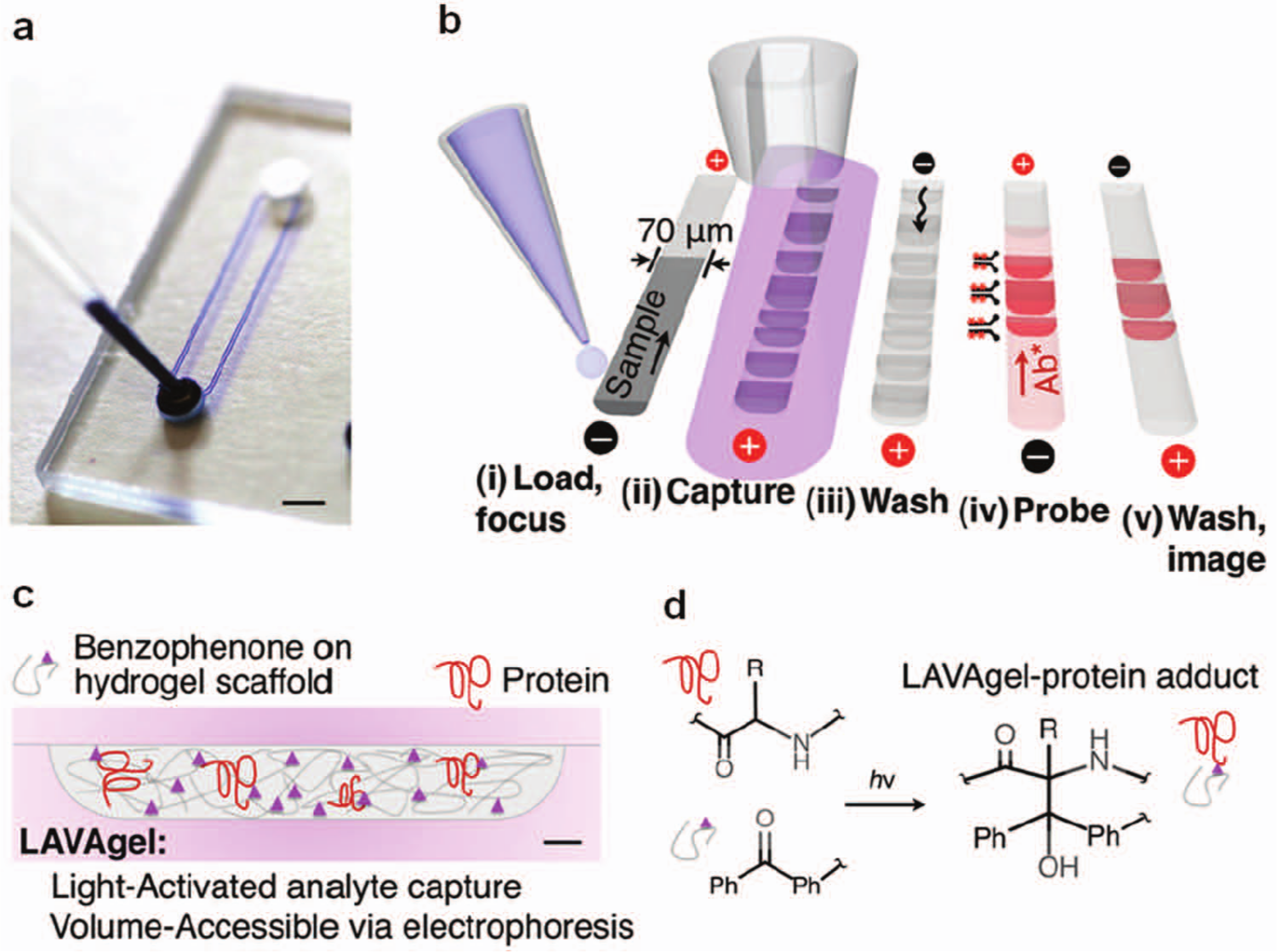
(
Assays that use reagent immobilization in gel matrices merge the benefits of heterogeneous and homogeneous assay platforms. Although these assays are heterogeneous, immobilization in the gel matrix enables capture analytes to sieve through the immobilized antigens in a 3D fashion. This enhanced interaction shortens the diffusion times for a capture event up to six orders of magnitude compared with reagents immobilized on the surface of a microchannel wall. 10 As an example of a multiplexed heterogeneous assay based on hydrogels, Bromberg et al. 83 analyzed DNA samples for single-nucleotide polymorphisms by electrophoresis of targets through a linear array of gels patterned in a microchannel. Each hydrogel element contained a high concentration of a unique oligonucleotide capture probe. Three different oligonucleotides were immobilized at three channel cross sections such that single base mutations were assayed for three oligonucleotides in a single run. Complementary oligonucleotide strands were then electrophoretically loaded to the channel and sieved through the regions with immobilized oligonucleotides. Capture was observed only when the complementary oligonucleotides were present without any mismatch.
Paper-Based Assays Move beyond Multiplexing Afforded by Lateral Flow Strips
Lateral flow tests and paper-based blotting assays are widely used diagnostics for screening and confirmatory assays.84,85 In recent years, paper assay formats have been designed with a focus on higher performance than conventional lateral flow assays. Multiplexed assay capability is one of the major design challenges tackled recently. Paper-based assays can be operated with low to no power, using wicking to drive fluid with high surface area to volume ratios supporting efficient heterogeneous reactions. Photoresist patterning or simply cutting the paper into patterns is an approach used to guide liquid flow into functionalized regions. Recently, Martinez et al.
86
demonstrated a 4-plex assay on multiple layers of SU-8 patterned paper. The assay measured glucose and protein levels in artificial urine samples. The detection range for glucose was in the mM range and, for bovine serum albumin (BSA), detection limits were in the µM range. Although detection was possible, the demonstrated sensitivity performance leaves room for improvement. The same group also reported a 96-well ELISA on paper patterned with photoresist.87,88 Lafleur et al.
89
demonstrated a multiplexed card assay for a malarial antigen (pfHRPII) and IgM antibodies to
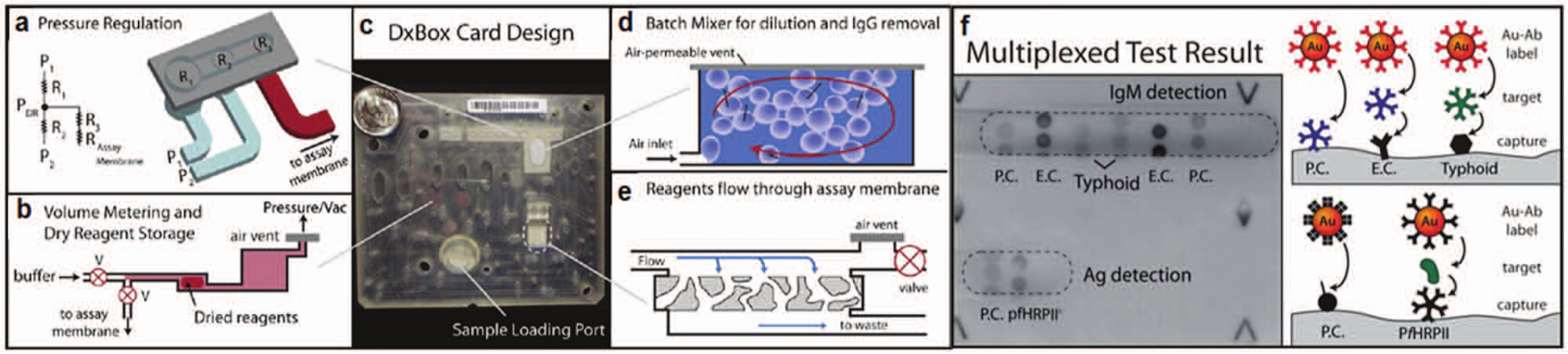
DxBox integrated microfluidic card. (
Large-Scale Integration of Microreactors for Heterogeneous Assay Multiplexing
Microfluidic multiplexing of samples and reagents has been performed in spatially distinct microreactors fabricated in microchannel networks. In one approach, fluid flow was controlled by pneumatic valves integrated into the devices using multilayer PDMS design. Similar to the role of the transistor in very large-scale integration (VLSI) technology, integrated microfluidic valves form a central function for microfluidic large-scale integration (mLSI) ( Fig. 7 ). A comprehensive review of mLSI devices and components can be found in recent reviews.93,94 An attribute of mLSI design relevant to multiplexed assays is the capacity for scale-up. Since the valve can use a binary combination multiplexor, just two log2N valve channels are required to control a network of N × N channels. 95 For example, 1024 flow channels can be controlled with 20 control valves. To support assays, valving has been used to trap functionalized beads in microreactors so beads serve as assay substrates. 96 The multiplexing capabilities of mLSI technology have been used in genetic analysis at the single cell level,32,97–99 epigenetic analysis and antibody characterization, 12 amino acid analysis, 100 and viral RNA detection for diagnostics. 101 In one example, Liu et al. 102 demonstrated 400 distinct PCR reactions in 41 pipetting steps—a conventional assay requires 1200 pipetting steps. The mLSI architecture in Figure 7 enables not only high-volume multiplexing and throughput but also simplifies assays by reducing handling steps. Similar to VLSI for electronic chips, mLSI systems can pose fabrication yield and cost challenges with the increasing fluidic circuit complexity, which can be partially handled similar to the VLSI experience by developing design rules, CAD tools, and foundries specialized with the fabrication of these complex devices. 93 In this regard, mLSI devices have gained in popularity for otherwise difficult single-cell studies, where the assay running costs are offset by valuable research findings. 99
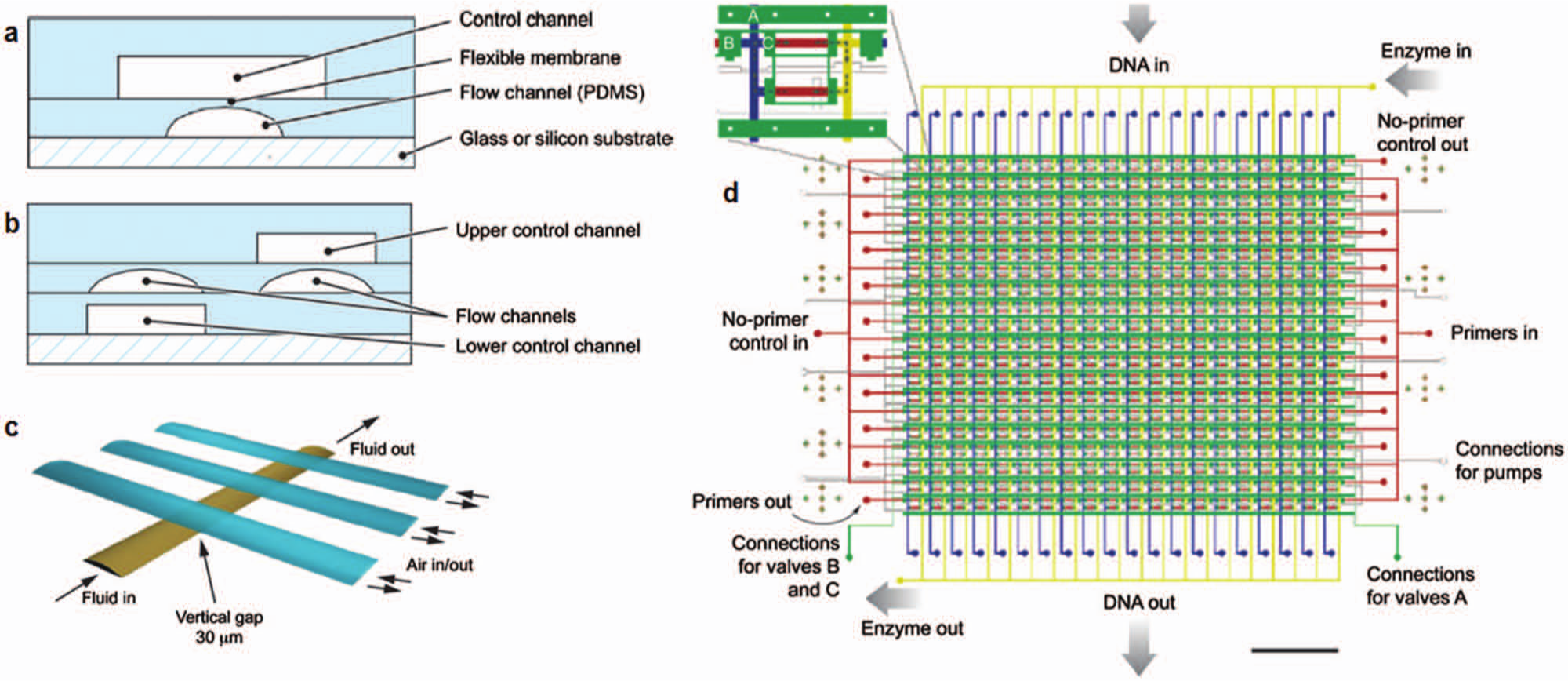
Microfluidic large-scale integration. (
Hand-Powered Multiplexed Assays for Point-of-Care Applications
Rather than using valves and pumps for mixing, movement of chip compartments has been developed to initiate reactions when the compartments merge. These “SlipChips,” shown in Figure 8 , involve fluidic channel networks and chambers sandwiched between two chips actuated (by hand) to slide over each other. 103 In one incarnation of a SlipChip-supported assay, the bottom layer of the SlipChip sandwich can contain preloaded dry PCR primers (or the reagents of interest). PCR sample mix is first loaded into the wells in the top layer, and the layers then move relative to each other, resulting in an overlap of sample wells with the wells with PCR primers. Using this approach, Shen et al. 104 reported a multiplexed PCR assay running in 384 wells on a chip with 20 different primer pairs to identify 16 blood-borne pathogens (bacterial and fungal species). Chips were assayed with five of these target pathogens. The presence of each pathogen led to a fluorescent signal in the corresponding zone. No cross-reactivity was observed. A rotational version of a SlipChip architecture—in which circular chips were rotated around a central axis—was used to compartmentalize and limit the viral load of a sample in a series of wells with varying volumes. 16 In each chip, up to five different samples were assayed. Each sample was distributed into various wells with volumes ranging from 1 to 625 nL. Experiments showed that when the viral load in the sample was high, signal was observed in most wells, including the 1-nL wells. In contrast, when the viral load was low, signal was obtained only in a few of the large wells. The reported LOD for the HIV viral load assay was 40 molecules/mL, with a four orders of magnitude dynamic range. For assays involving detection of single analytes, the dynamic range was extended up to five orders of magnitude.
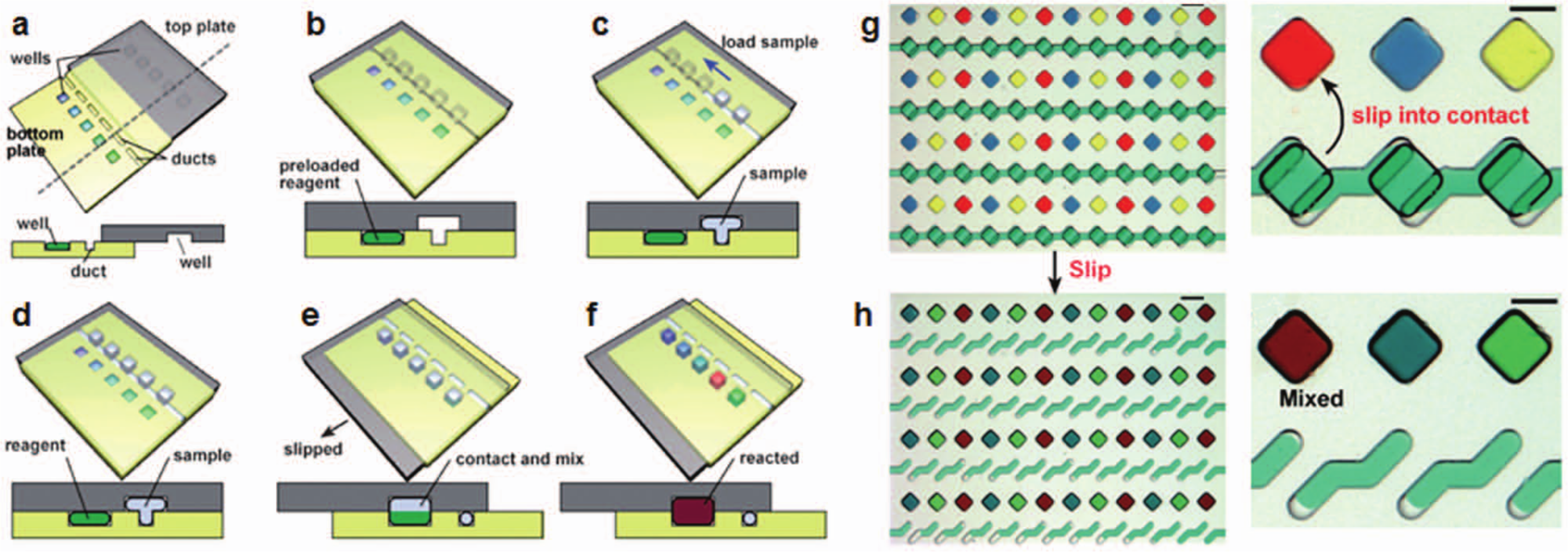
SlipChip assays: (
In subsequent developments, Song et al. 105 employed the SlipChip and developed a multiplexed volumetric bar chart chip. In this case, antibodies against cancer antigens in human sera were immobilized in the wells. The serum sample was delivered to the wells and target carcinoembryonic antigens (CEAs) were captured in the wells, and then the sample was washed and secondary antibody conjugated with HRP-coated nanoparticles was delivered. The resulting ELISA catalyzed the H2O2 substrate delivered in the final stage into H2O and O2. Red ink available in the interconnected channels was pushed down the channels, generating a bar chart readout of analyte level for up to 50 samples. The amount of ink movement in the bar channels was correlated with the target analyte concentration determined by standard clinical tests. LOD was reported to be 5 ng/mL for CEA, 10-fold higher than a typical LOD for CEA detecting ELISA (0.5 ng/mL). Currently, SlipChip technology uses 2D designs in which the number of assay steps is limited by the topology of having the wells and channel networks in two layers. Incorporation of a third dimension may open new application areas for even more sophisticated assays involving multiple steps. The motion of the layers has been typically controlled by hand, potentially contributing to variation in assay performance.
Droplet-Based Assays Streamline Manipulation in Multiplexed Assays
Since the introduction of microfluidic systems for the generation and control of droplets in the early 2000s, numerous approaches for the control, manipulation, merging, and separation of droplet emulsions have been reported.106,107 The reader is directed to recent reviews for details.108–110 In the context of multiplexed assays, generation of a barcoded droplet has been accomplished by preparing droplets separately from solutions barcoded with known nucleic acid sequences 111 or fluorophore combinations/concentrations in a manner similar to the fluorescent bead barcoding approaches detailed earlier. 112 Encoded droplets can be collected and subsequently used for multiplexed assays.
In this manner, Brouzes et al. 28 reported a single-cell toxicity assay ( Fig. 9 ). A fluorescently barcoded droplet library representing eight different concentrations of the drug mitomycin C allowed monitoring of cell viability over time. Recently, Chen et al. 113 showed a droplet-based microfluidic platform for multiplexed monitoring of protease enzyme activity in clinical samples. Four different protease substrates were encapsulated in droplet emulsions barcoded with specific indicator dye concentrations. After each droplet was serially merged with a biological sample in the pico-injector, protease activity was monitored at an observation chamber. In parallel, fluorescence increases due to substrate proteolysis were monitored. The assay monitored activities of MMP-2, MMP-0, ADAM-9, and ADAM-10 proteases secreted by 12 cells after stimulation with TNF-α. This method resulted in a 100-fold reduction in biological sample and reagent use. A similar assay was recently reported by Sjostrom et al. 114 showing multiplexed enzyme kinetic analysis in combinations of droplets generated containing various substrate and enzyme concentrations.
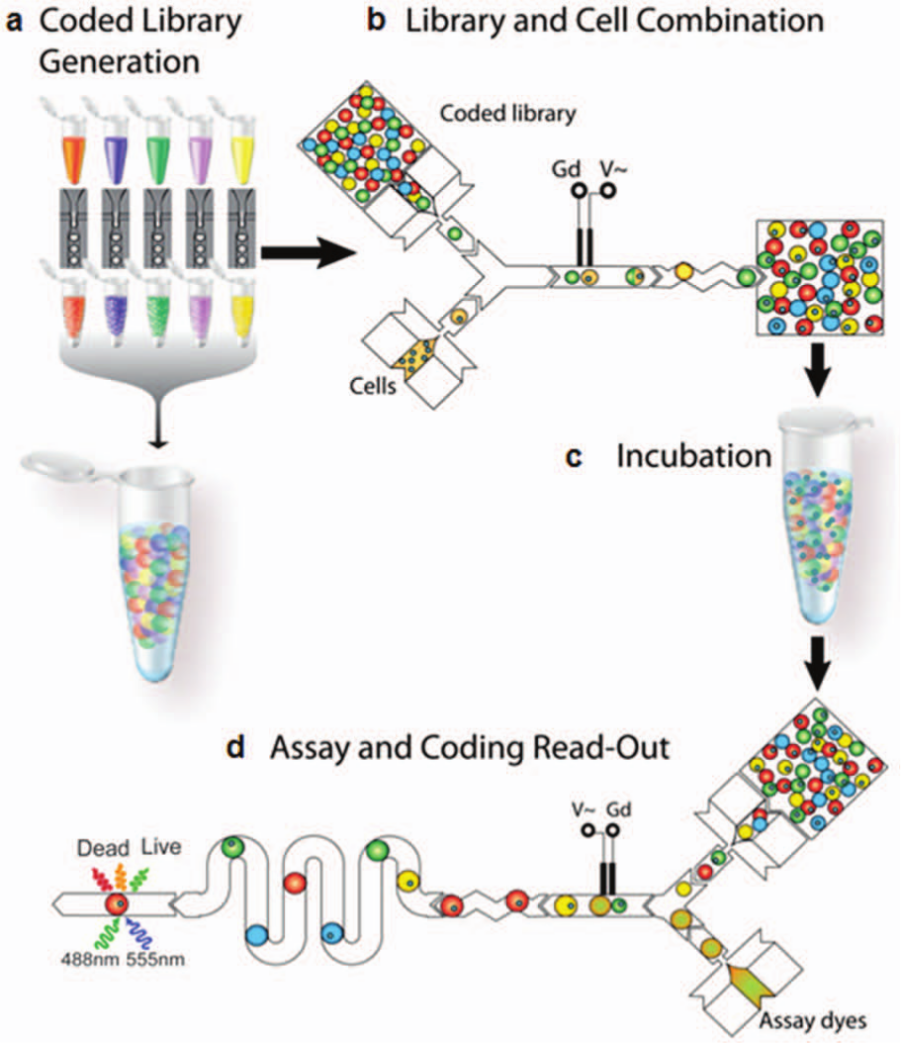
Droplet-based multiplexed assay for cytotoxicity assessment. (
Droplet emulsions have played a role in multiplexed genomic analysis of single bacteria. Zeng et al. 74 reported single-cell multiplexed PCR detection to identify mutant and pathogenic cells. The assay used microbeads functionalized with forward primers serving as a solid substrate for PCR amplification. Droplet emulsions consisting of individual cells, beads, and reagents were generated in a microfabricated emulsion generator array (MEGA). Upon PCR amplification, beads were separated and analyzed by multispectral flow cytometry. The LOD for detection of pathogenic bacteria yielded an analytical sensitivity of one pathogenic bacterium in 105 nonpathogenic mutants.
Label-Free Detection to Simplify Assay Operation
Motivated to simplify operation, researchers have adapted label-free detection strategies to multiplexed microfluidic assays. Xiang et al. 31 reported multiplexed detection of multiple biomarkers of different size (proteins and adenosine triphosphate [ATP]) in a label-free format using aptamer-conjugated redox tags and electrochemical sensors. As shown in Figure 10 , one aptamer strand was anchored to the electrode and conjugated to a different conjugation aptamer tagged with a specific redox reporter. When a target analyte was present, the complementary aptamer was bound to the target and displaced from the surface along with its redox tag. Removal of the tag led to a change in the readout signal. By using specific tags for each reagent, a 2-plex assay detecting thrombin proteins and ATP was realized. The LODs for thrombin and ATP were reported to be 15 nM and 0.5 µM, respectively. In comparison, enzyme precipitation assays (no multiplexing) have been reported with a LOD of 13.5 pM for thrombin. 115
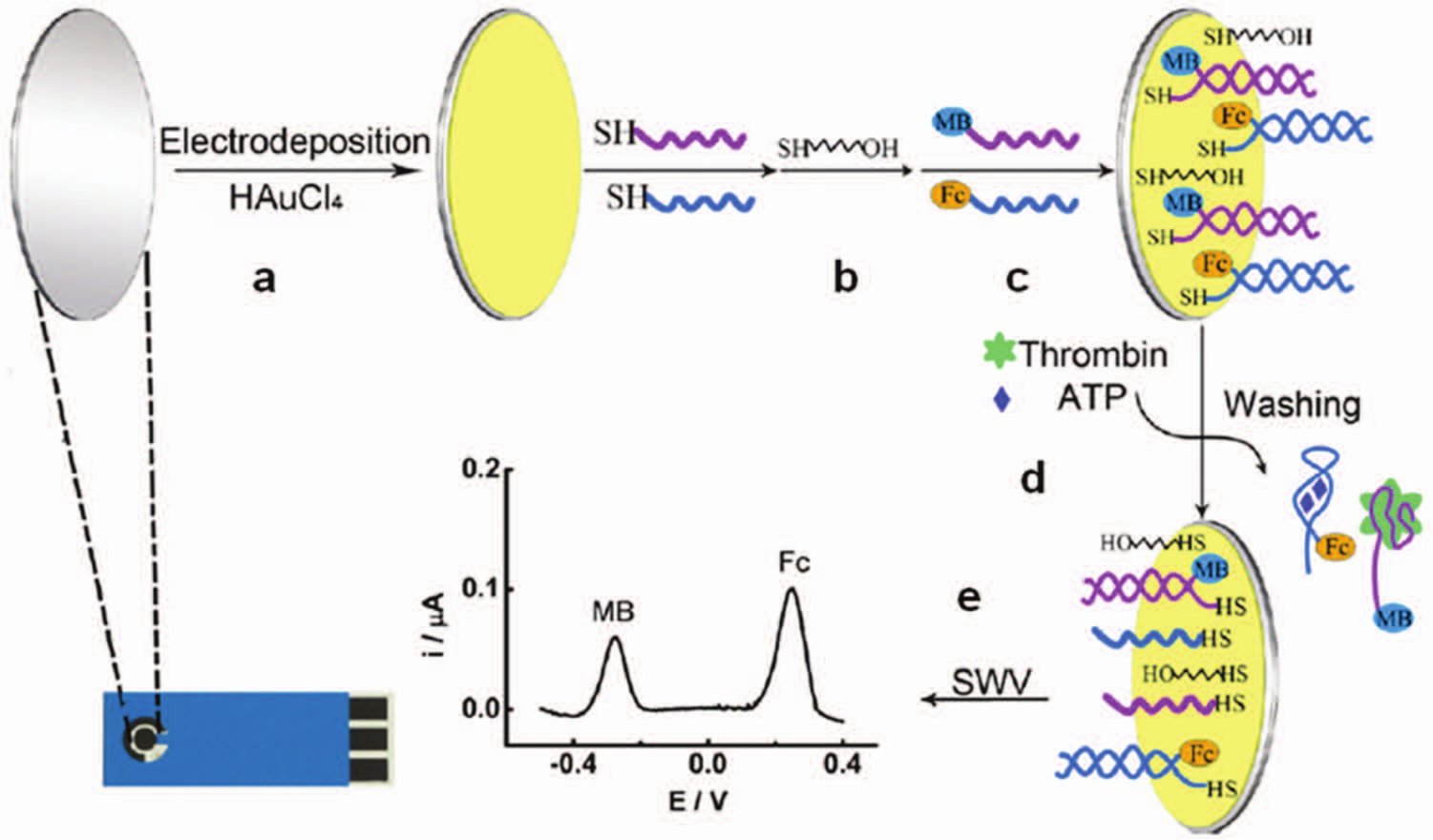
Schematic of the label-free multiplexed detection of adenosine triphosphate (ATP) and thrombin. (
By using a similar technique—now for point-of-care clinical diagnostics applications—White et al. 11 demonstrated a wash-free and label-free electrochemical detection assay by using a modified nucleic acid duplex as a signaling and recognition probe. One DNA strand attached to the substrate presented a redox tag at the tip of the DNA molecule. A second DNA strand was conjugated with a capture reagent (HIV antigen). With no target analyte present, the DNA strand moved freely and the redox tag interacted with the electrode surface transferring electrons. Whereas when a target analyte (HIV antibody) in the serum was captured by the antigen at the tip, the redox reporter could not interact with the surface, thus leading to a change in the electrochemical signal. The authors demonstrated multiplexed detection of three different antibodies against three different antigens as a general approach to rapid and specific quantitation of antibodies in biospecimens. The LOD for the assay was reported to be in the range of 1 to 10 nM. Although more sensitive microfluidic assays have been reported (Hughes et al., 10 50 pM), the 1- to 10-nM LOD aligns with clinically relevant levels for this application.
For applications where eliminating dependence on external detection equipment (e.g., microscopes or other compact readers) is important, label-free electrochemical detection has been incorporated in multiplexed microfluidic formats. Complications do arise with complex biospecimens, as the myriad proteins and metabolites found at high concentrations in these materials can result in nonspecific signals. In addition, the pH and electrolyte concentrations can vary from sample to sample. To overcome these challenges, Stern et al. 116 described a method in which preconcentration of multiple biomarkers in human blood was accomplished in a microfluidic chamber housing reagents specific to the target biomarkers. The reagents were immobilized in the chamber via a photo-cleavable, biotin-conjugated DNA strand. 117 Biomarkers for prostate and breast cancer (e.g., PSA and CA15.3, respectively) were immobilized in a capture chamber and then released by UV irradiation for downstream detection via secondary antibody immobilized on nanowire sensors. The multiplexed assay platform was tested with PSA and CA15.3 spiked into human blood at 2-ng/mL and 30-U/mL concentrations, respectively. Antigens were successfully detected within 20 min.
Conclusions
From high-throughput applications to highly sensitive and specific immunodiagnostic assays, multiplexed analysis is a desired and often necessary feature in assay platforms. With the maturation of microfluidic capabilities over the past two decades and recent emphasis on translational research, 118 microfluidic assays are beginning to offer practical and high-functionality solutions to addressing bioanalytical problems. In tandem, the number of companies offering assay technologies based on microfluidic platforms is increasing. 119 It may be fair to say that microfluidics platforms are becoming an important format for multianalyte assays, perhaps emerging as a standard for automated multiplexed analyses.
Our intent in this review has been to overview a set of core microfluidic technologies that form multiplexed detection platforms or methods. Given the constraints of space, we were not able to recognize all of the important work in this growing area but instead sought to highlight examples from a diverse range of approaches. Some of the developments we highlighted aimed at adapting existing multiplexed benchtop technologies such as ELISA or immunoblotting to microfluidic formats while offering advancements. These platforms bring the potential for faster genetic or protein-based assays in high-throughput or point-of-care platforms. High-throughput methods are also likely to have an impact in addressing unmet needs in biomarker discovery and validation.120–122 In addition to assays advancing on currently available assay platforms, we also have been witnessing the introduction of microfluidic technologies that allow measurements not previously possible. For instance, multiplexed analysis of the contents of single cells is a challenge that is uniquely poised to be addressed by microfluidics. These measurements provide a level of detail regarding cell population heterogeneity that has not yet been attainable and can be crucial to understanding cell-level decisions and response.
Challenges remain since most multiplexed microfluidic assays are in the early research stage, still at the proof-of-principle stage. Clearly, continued innovation coupled with focused development and application to a wider set of applications is on the horizon. In this regard, pursuit of promising approaches to translational gaps is likely to increase the performance, capabilities, and availability of these assay platforms. We look forward to the next decade, in which we will be seeing multiplexed microfluidic assays addressing—in the field—challenging medical and life sciences questions.
Footnotes
Acknowledgements
The authors acknowledge members past and present of the Herr Lab for critical discussion.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors acknowledge the following financial support for the research, authorship, and/or publication of this article: M.K.A. has received financial support from Novartis Diagnostics. A.M.T. is a 2012 UC Cancer Research Coordinating Committee predoctoral fellow. This work was supported by a New Innovator Award (DP2OD007294 to A.E.H.) from the Office of the Director, National Institutes of Health.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
