Abstract
Two automated platforms using immunomagnetic separation technology were compared for detecting and recovering Escherichia coli O157 in ground beef and sprouts and Shigella flexneri in green onions. The foods were inoculated with <20 CFU/25 g and tested at 5 and 24 h postincubation. Immunomagnetic beads were mixed with food enrichments, processed through the Pathatrix Auto or KingFisher Flex, and tested by real-time PCR (qPCR) and recovery on selective agars. At 5 h, the Pathatrix Auto detected E. coli O157 in 90% and 60% of the ground beef and sprouts samples and S. flexneri in all of the green onion samples. It also recovered E. coli O157 in all the samples but could not recover S. flexneri in any of the green onion samples. In comparison, the KingFisher Flex detected E. coli O157 in 80% and 30% of the ground beef and sprouts samples and S. flexneri in all of the green onion samples. It also recovered E. coli O157 in 90% of the ground beef samples but none of the sprouts samples and S. flexneri in 20% of the green onion samples. At 24 h, both platforms detected and recovered the target bacteria in all of the samples.
Outbreaks of human illnesses due to Escherichia coli O157–contaminated sprouts or ground beef 1 or Shigella flexneri–contaminated green onions2,3 have been frequently reported. Although rapid methods such as PCR and enzyme-linked immunosorbent assay (e.g., VIDAS) have made screening for pathogenic bacteria in foods easier, it is still necessary for regulatory laboratories to obtain an isolate to prove that the bacteria are viable and pathogenic. In addition, a viable isolate can be subjected to further testing to confirm the identity of the isolate, strain type the isolate to match food as well as clinical and environmental samples, and further characterize the virulence factors to distinguish between pathogenic and nonpathogenic bacteria. Selective agars that can distinguish target bacteria are helpful in obtaining viable isolates, but food matrices with a high background of microbiota may make isolation challenging. Immunomagnetic separation (IMS) uses antibodies to a specific target bacteria bound to paramagnetic beads to aid in isolating and concentrating bacteria. The immunomagnetic beads can be separated by applying magnetic force to the sample and washed to reduce nonspecific binding before plating on selective agar. Here we describe and compare two IMS systems, the Pathatrix Auto and the KingFisher Flex for detecting and isolating E. coli O157 and S. flexneri from food samples.
Pathatrix Auto
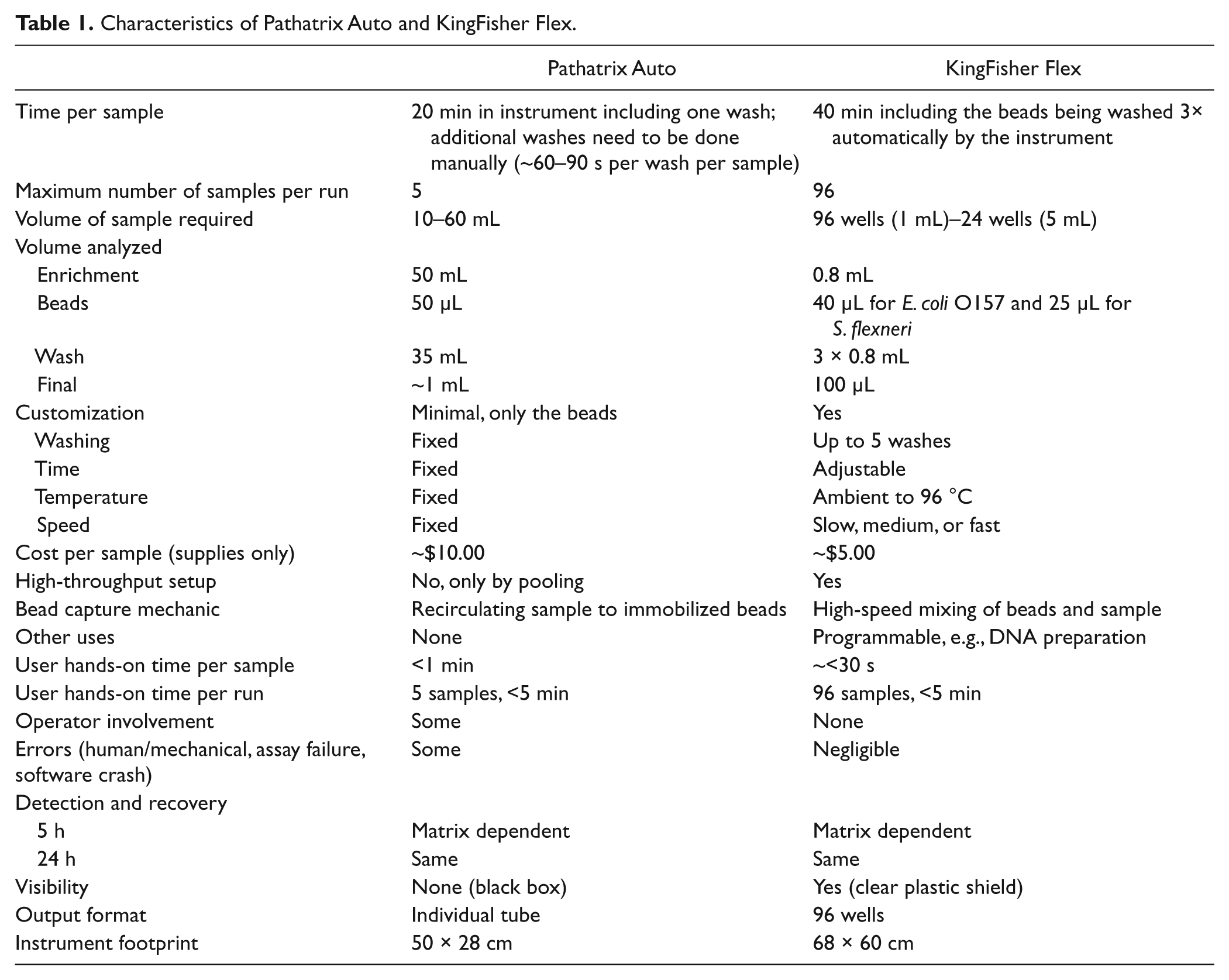
Matrix MicroScience (Golden, CO) created the Pathatrix Auto in 2009, an IMS method that uses paramagnetic beads coated with antibodies to selectively isolate the target of interest. As shown in Figure 1 , the paramagnetic beads are added to 10 to 60 mL of food enrichments before a Pathatrix Auto closed tube syringe apparatus is attached and inserted into a cartridge that is inserted in the Pathatrix Auto. Pathatrix Auto’s flow-through immunocapture technology pumps the sample enrichment past a dual capture magnet that immobilizes the paramagnetic beads and captures the target of interest. After the capture phase is completed, the user must initiate a phosphate-buffered saline (PBS) rinse of the beads to reduce any nonspecifically bound organisms and PCR inhibitors. The paramagnetic beads are released from the magnets and can be collected. The sample can then be further analyzed using selective plating and/or PCR with presumably reduced background microbiota interfering with detection. Characteristics of the Pathatrix Auto are presented in Table 1 .
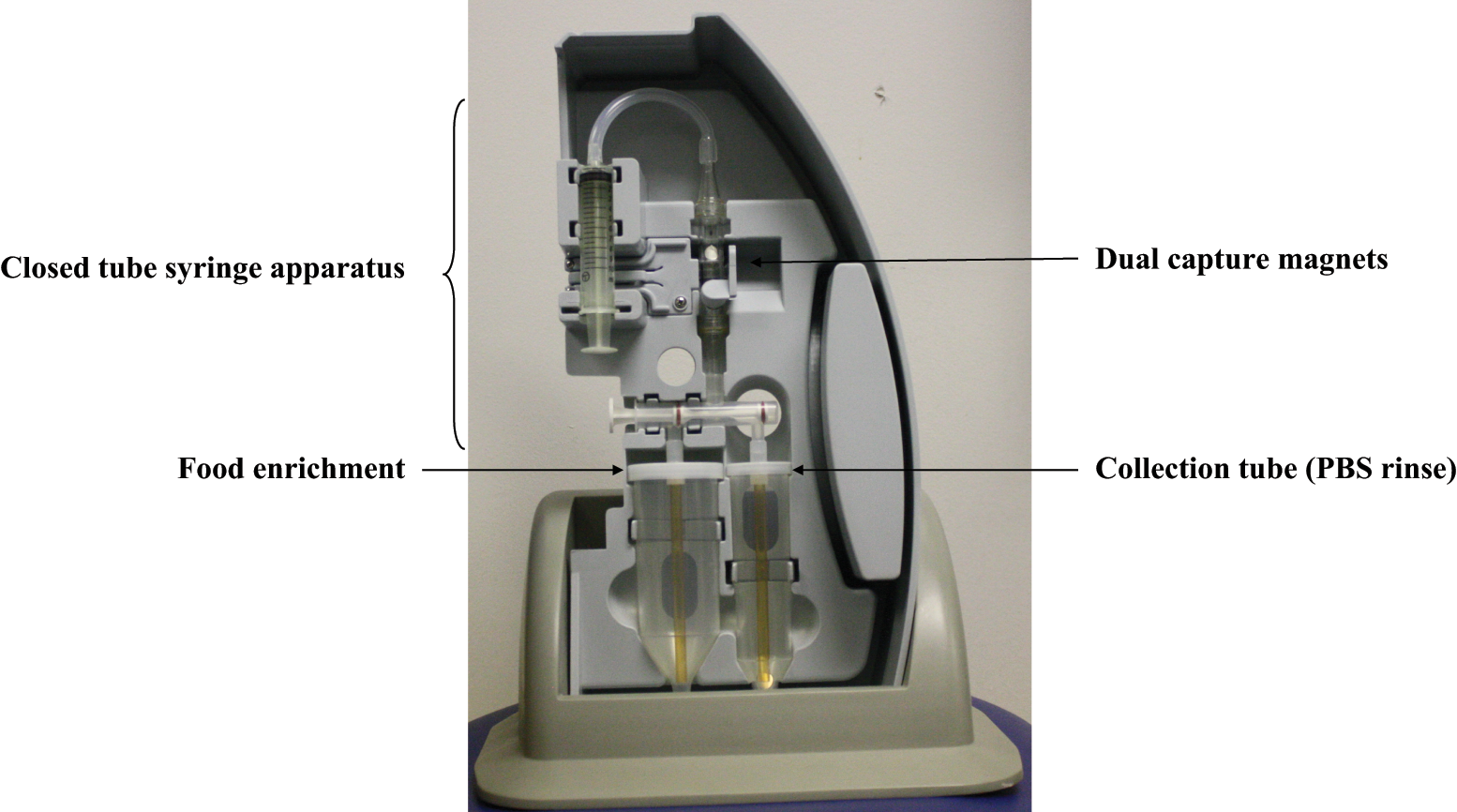

Pathatrix Auto cartridge (photograph taken by author Henry Lau).
Characteristics of Pathatrix Auto and KingFisher Flex
KingFisher Flex
KingFisher Flex is a magnetic particle processor robot from Thermo Fisher Scientific (Waltham, MA) capable of processing DNA/RNA, protein, or live cells. KingFisher Flex can be programmed by the user for different needs, such as concentration, extraction, or purification. As shown in Figure 2 , the unit sits on a circular plate with eight slots for 96-well plates. Each slot rotates through the interchangeable magnetic head for processing. There are magnetic heads available for 96-well plates, 96-deep-well plates, and 24-deep-well plates. The volume that the KingFisher Flex can process ranges from 20 µL to 5 mL. The machine can automate the binding, washing, and elution of the immunomagnetic beads and also minimizes user intervention and contamination. Therefore, it can generate 96 consistent results in one test run compared with traditional IMS manual methods. Characteristics of the KingFisher Flex are presented in Table 1 . Using the ThermoFisher’s BindIt software, the KingFisher Flex is customizable in temperature, time, and speed.
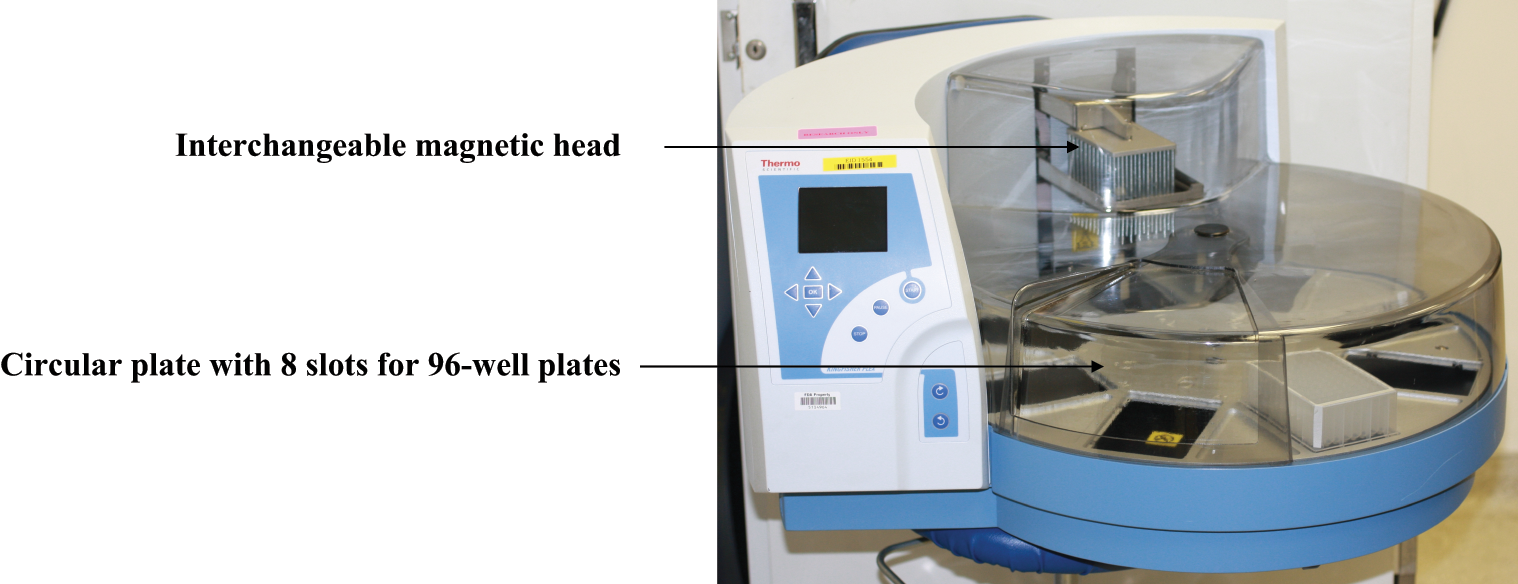
KingFisher Flex (photograph taken by author Henry Lau).
Materials and Methods
Sample Preparation
Sprouts, ground beef, and green onions were purchased from grocery stores. A total of 25 g of the samples (n = 10 ground beef, n = 10 sprouts, and n = 6 green onions [due to equipment constraint]) were aseptically weighed, placed into sterile Whirl-pak bags (Nasco, Modesto, CA), and inoculated with either frozen E. coli O157 (ATCC 43894) or S. flexneri 4 spikes. The concentrations of inocula were 3 to 11 CFU/25 g of E. coli O157 for ground beef, 11 CFU/25 g of E. coli O157 for sprouts, and 15 CFU/25 g of S. flexneri for green onions. Those concentrations were used because the infective doses of E. coli O157 and Shigella are estimated to be in the range of 10 to 100 and 10 to 200 cells, respectively. 2 No-spike samples were prepared as controls. For the ground beef and sprouts, 225 mL of modified buffered peptone water (10 g peptone, 5 g sodium chloride, 3.6 g sodium biphosphate, 1.5 g monopotassium phosphate, 5 g casamino acids, 6 g yeast extract, 10 g lactose, 1 g sodium pyruvate in 1 L) supplemented with cefsulodin-vancomycin (10 mg/L and 8 mg/L, respectively) were added, and the samples were incubated at 37 °C for 5 and 24 h. 5 For green onions, 225 mL of Shigella broth (20 g tryptone, 2 g dipotassium phosphate, 2 g monopotassium phosphate, 5 g sodium chloride, 1 g glucose, 1.5 mL Tween 80 in 1 L) with a final concentration of 0.5 µg/mL of novobiocin were added to each sample. 6 The samples were incubated at 42 °C anaerobically for 24 h. After 5 and 24 h of incubation, aliquots of enrichment were taken for analysis on both the Pathatrix Auto and KingFisher Flex.
Pathatrix Auto
Pathatrix Auto immunomagnetic extraction’s protocol cannot be customized and has to be carried out as described in the manufacturer’s 2009 operating manual. For the Pathatrix Auto analysis, 50 mL of the enrichments at 5 and 24 h were processed using 50 µL of anti–E. coli O157 (Matrix Microscience) or anti-Shigella–coated magnetic beads (see Table 1 for a summary of volume comparison) as instructed in the manufacturer’s instructions. Shigella antibodies (Matrix Microscience) were conjugated to the paramagnetic beads using the Custom Coating Kit (Matrix Microscience) to a final concentration of 1.25 mg antibodies/mL. E. coli O157 antibodies were purchased commercially through Matrix Microscience as conjugated beads. Thirty-five milliliters of PBS was added to the elution vessel. The addition of the Pathatrix Auto sterile, single-use kit connects the enrichment vessel and the elution vessel together into a closed system. The machine was run for 15 min, in which the sample enrichment with IMS-specific bead passed in an up-and-down direction through two magnets. After the target organisms were captured by the beads, the magnetic beads were washed with the 35 mL of PBS and collected in approximately 1 mL of PBS in the elution vessel. The beads were manually separated to the side of the vessel on the magnetic collection stand for 1 min, the supernatant was removed, and the beads were resuspended in 100 µL of elution buffer and mixed and transferred to a 1.5 mL microcentrifuge tube (Eppendorf AG, Hamburg, Germany) as described in the manufacturer’s instructions. The eluted beads were then spread on CT-SMAC (Sorbitol McConkey Agar supplemented with 0.05 mg/L cefixime and 2.5 mg/L potassium tellurite) for E. coli O157 and Shigella Rainbow Agar (Biolog, Hayward, CA) for S. flexneri and incubated at 35 °C for 24 h. Typical colonies were subsequently confirmed using the Remel latex agglutination kit (Remel, Thermo Fisher Scientific, Lenexa, KS) for E. coli O157 and with real-time PCR (qPCR) and/or Triple Sugar Iron (TSI; 10 g pancreatic digest of casein, 10 g peptic digest of animal tissue, 5 g sodium chloride, 10 g lactose, 10 g sucrose, 1 g glucose, 0.2 g ferric ammonium citrate, 0.2 g sodium thiosulfate, 0.025 g phenol red, 13 g agar in 1 L) slants for S. flexneri. Colonies that were positive by both tests (i.e., latex agglutination and qPCR for E. coli O157 and TSI and qPCR for S. flexneri) were considered positive for recovery of the organism of interested. For both systems, half of the eluted beads were boiled for 10 min for qPCR screening. 7
KingFisher Flex
For this study, the protocol for the KingFisher Flex IMS processing included binding, washing, and elution steps. A 96-deep-well magnetic head was used for the extraction in addition to corresponding 96-deep-well plates. The materials needed were a 96-deep-well rod cover, one heat-resistant 96-deep-well plate for the enrichment, three 96-deep-well plates for the washes, and one 96-well standard plate for elution. A total of 800 µL of the enrichments was placed in a 96-deep-well plate along with either 40 µL of anti–E. coli O157 DynaBeads (Dynal Biotech, Oslo, Norway) or 25 µL of anti-Shigella–coated Pathatrix paramagnetic beads. The amounts of beads used were previously optimized by Weagant et al. 5 for E. coli O157 and our laboratory (data not shown). The KingFisher Flex processing consisted of a binding step, three washing steps using 0.8 mL PBS-T (0.1% of PBS with 0.05% Tween 20) for E. coli O157 and 0.8 mL Butterfield’s phosphates-buffered water with 0.05% Tween 20 for S. flexneri, and an elution step using 100 µL PBS for E. coli O157 and 100 µL alkaline peptone water (10 g peptone and 10 g sodium chloride in 1 L) for S. flexneri ( Table 1 ). Fifteen microliters of the eluted beads were then spread on selective agar as described above, and 55 µL was boiled for 10 min and analyzed by qPCR.
Results and Discussion
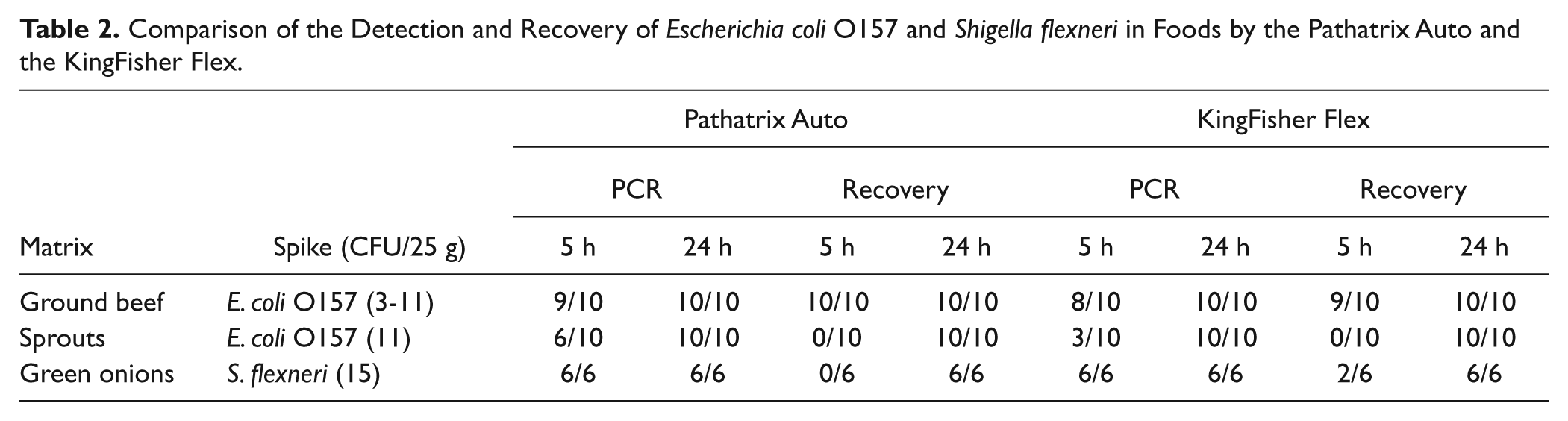
The data are presented in Table 2 . At 5 h of enrichment, the Pathatrix Auto was better at detecting E. coli O157 than the KingFisher in both ground beef (i.e., 9/10 vs. 8/10 for qPCR and 10/10 vs. 9/10 for plating) and sprouts (i.e., 6/10 vs. 3/10 for qPCR and no recovery for plating on either platform). The KingFisher Flex, however, was better at recovering S. flexneri than the Pathatrix Auto in green onions (i.e., total detection [i.e., 6/6] by qPCR on both platforms and 2/6 vs. 0/6 for plating). Not recovering the organism 100% of the time is considered an assay failure by regulatory agencies. Therefore, the 5 h data on both platforms are meaningless to compare to one another as both platforms failed at recovering the organisms in all of the tested samples. At 24 h of enrichment, total recovery by both plating and qPCR was observed for both platforms.
Comparison of the Detection and Recovery of Escherichia coli O157 and Shigella flexneri in Foods by the Pathatrix Auto and the KingFisher Flex
Conventional IMS involves incubating 1 mL of enrichment with immunomagnetic beads before collecting the beads and plating, 8 which works well for small sample demand. When demand exceeds 10 samples, its use becomes prone to human errors and is tedious. On the other hand, the Pathatrix Auto instrument involves pumping 50 mL of 5 h or 24 h enriched sample through the immunomagnetic beads, greatly increasing the sampling size and thus, theoretically, the sensitivity of the technique. However, it sacrifices automation and the amount of samples per run; as such, it takes 20 min for 5 samples. Also, the bead collection may vary from one to another sample cartridge, causing the recovery of isolates to be inconsistent. Specifically, visual inspection of the bead concentration recovered by the Pathatrix Auto seemed very inconsistent from sample to sample (data not shown). Although the Pathatrix Auto is an automated system, there are still many hands-on steps in its process: the connection of the tubing kit, the initiation of the washing steps, and the inconsistent elution volumes that require further adjustment by the user. The KingFisher Flex instrument excels at automation compared with the Pathatrix Auto (i.e., 96-well plate format versus one sample per cassette). The approximately 40 min process through the KingFisher Flex is very consistent for either 1 or 96 samples, as each of the 96 samples is processed simultaneously versus each Pathatrix Auto cassette being processed independently (data not shown). The output of the instrument is the immunomagnetic beads that are ready for postanalysis processing in a predetermined volume.
By using the Pathatrix Auto or the KingFisher Flex, a portion of the 5 h enrichment could be used for E. coli O157 or Shigella IMS concentration for detection, whereas the remaining enrichment can return for overnight enrichment if additional incubation is needed. Although this 5 h method can generate a screening result (i.e., detection by qPCR) in less time, it is not always desirable. First, the 5 h method can generate only the screening result (i.e., detection by qPCR) and cannot yield isolation of the bacteria of interest 100% of the time. Second, depending on when the sample is set up and started, a 5 h incubation will not be logical if the samples were received in the afternoon, and thus the screening result (i.e., detection by qPCR) will not be ready until late in the evening. The 5 h IMS concentration of both platforms gave inconsistent results in the recovery of the targeted bacteria ( Table 2 ). For more consistent recovery across the samples, the 24 h enrichments on both platforms gave the best results. From the results in this study and also depending on the difficulty of the matrices, the Pathatrix Auto may be very useful in detecting a low concentration of targeted bacteria at 5 h in low throughput (<10 samples), whereas the KingFisher Flex may be more useful with high-throughput (>10 samples) sampling at 24 h.
The characteristics of the Pathatrix Auto and KingFisher Flex are presented in Table 1 . Although the KingFisher Flex takes twice the time per sample (i.e., 40 min vs. 20 min on the Pathatrix Auto), it washes the beads automatically three times, thus minimizing the operator involvement. In fact, the Pathatrix Auto requires the operator to manually wash the beads as part of the platform’s instruction. Another advantage of the KingFisher Flex is that it can analyze up 19.2 times more samples at once than the Pathatrix Auto (i.e., 96 vs. 5 samples). Theoretically, the Pathatrix Auto can analyze a larger sample volume (i.e., 60 mL vs. up to 5 mL on the KingFisher Flex), which can aid in recovery of isolates at 5 h (i.e., the larger volume analyzed increases the chances of recovering at a low concentration of the targeted bacteria). At the inoculum that was spiked, there was some evidence, although not convincing, that the larger volume analyzed by the Pathatrix Auto might lead to better detection and recovery of the targeted bacteria at a lower concentration (i.e., sprouts: 6/10 detected vs. 3/10 on the KingFisher Flex; ground beef: 9/10 detected vs. 8/10 on the KingFisher Flex). The KingFisher Flex can customize the temperature, time, speed, and washes, whereas the Pathatrix Auto cannot. Running one sample on the Pathatrix Auto costs twice as much as on the KingFisher Flex (i.e., $10.00 vs. $5.00). Adding to that cost is the time the operator spends washing the beads manually on the Pathatrix Auto, whereas it is done automatically on the KingFisher Flex. The KingFisher Flex has a high-throughput setup, in which 96 samples can be analyzed simultaneously. The Pathatrix Auto though does not allow high throughput, except by pooling, in which a smaller aliquot (i.e., 5–10 mL) of several samples is combined to make up to a 60 mL sample volume. One disadvantage of this pooling method is that when a pooled sample is positive, the operator will not know which subsample is positive and will need to go back to test the positive samples individually. Although the Pathatrix Auto can strictly be used only for bacterial isolation, the KingFisher Flex is programmable and can be used for DNA preparation as well.
Depending on the laboratory’s needs, one may choose to use either one or the other platforms based on the characteristics presented in this article. However, because regulatory agencies such as the Food and Drug Administration and the U.S. Department of Agriculture–Food Safety and Inspection Service require 100% confidence in their results, testing after 5 h does not provide such confidence on either platform. Also, having a 5 h incubation is not convenient when analysts work an 8 h day. Therefore, if more than 10 samples need to be analyzed (as in the case of an outbreak), the KingFisher Flex should be the platform of choice.
Footnotes
Acknowledgements
The authors would like to thank David K. Lau and Thomas H. Sidebottom at FDA SAN LAB for their logistical support. We would also like to thank Dr. Chorng-Ming Cheng at FDA PRL-SW and Dr. Wen S. Lin at DFS for generously sharing their reagents and Dr. Keith Lampel at the Center for Food Safety and Applied Nutrition for the Shigella culture.
Declaration of Conflicting Interests
The views presented in this article do not necessarily reflect those of the Food and Drug Administration. The authors declare no potential conflicts of interest with respect to the research, authorship, and/or publication of this article. Use of trade names and commercial products in this article is solely for the purpose of providing scientific information and does not imply recommendation or endorsement by the Food and Drug Administration. The Food and Drug Administration is an equal opportunity employer.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
