Abstract
Knowledge graphs and ontologies represent symbolic and factual information that can offer structured and interpretable knowledge. Extracting and manipulating this type of information is a crucial step in complex processes. While large language models (LLMs) are known to be useful for extracting and enriching knowledge graphs and ontologies, previous work has largely focused on comparing architecture-specific models (e.g. encoder-decoder only) across benchmarks from similar domains. In this work, we provide a large-scale comparison of the performance of certain LLM features (e.g. model architecture and size) and task learning methods (fine-tuning vs. in-context learning (iCL)) on text-to-graph benchmarks in two domains, namely the general and biomedical ones. Experiments suggest that, in the general domain, small fine-tuned encoder-decoder models and mid-sized decoder-only models used with iCL reach overall comparable performance with high entity and relation recognition and moderate yet encouraging graph completion. Our results also suggest that, regardless of other factors, biomedical knowledge graphs are notably harder to learn and are better modelled by small fine-tuned encoder-decoder architectures. Pertaining to iCL, we analyse hallucinating behaviour related to sub-optimal prompt design, suggesting an efficient alternative to prompt engineering and prompt tuning for tasks with structured model output.
Keywords
Introduction
Acquiring structured knowledge from text is a fundamental step in a complex process like reasoning and answering questions, whether such a process is carried out by a human or an artificial intelligence (AI) system (Tiwari et al., 2023). In natural language processing (NLP), structured knowledge is often handled via ontologies or knowledge graphs (Hogan et al., 2021; Paulheim, 2017; Peng et al., 2023). Knowledge graphs are typically organised as collections of
Knowledge graphs have seen a surge in their application in recent years (Chen et al., 2020; Ji et al., 2022). However, building them can be laborious and costly (Kejriwal et al., 2019; Peng et al., 2023). This has led to the development of numerous methods aimed at auto-generation of these graphs from text sources in various fields (Ji et al., 2022; Kejriwal, 2022; Liu et al., 2016; Peng et al., 2023). Until recently, extracting and manipulating knowledge graphs and other forms of graphs has been largely dealt with by small knowledge graph embedding models (KGEs) (Wang et al., 2017), which are lightweight but limited in capabilities, or different types of graph neural networks (GNNs) (Wu et al., 2023; Ye et al., 2022), such as convolutional GNNs (CGNNs) (Zhang et al., 2018), or gated attention GNNs (GAT-GNNs) (Veliˇković et al., 2018). Recently, many of these architectures have been replaced by transformer-based large language models (LLMs) (Vaswani et al., 2017a), which have shown great potential in modelling graph-based data.
Despite these advancements, current techniques still suffer from significant limitations concerning accuracy, completeness, privacy, bias, and scalability (Peng et al., 2023; Radulovic et al., 2018; Rashid et al., 2019). Therefore, generating a large-scale knowledge graph automatically from text corpora remains an open challenge (Hogan et al., 2021; Kejriwal, 2022; Peng et al., 2023). As shown by a consistent body of evidence (Jin et al. 2023; Liu et al., 2023; Pan et al., 2024), LLMs can be adapted to both extract knowledge graphs from a reference text (text-to-graph task), as well as to convert knowledge graphs into natural language while maintaining the semantic meaning (graph-to-text task). We are interested in the former and adopt two text-to-graph benchmark datasets, to be referred to as Web NLG (Gardent et al., 2017) and Bio Event (Frisoni et al., 2022). Web NLG is a popular benchmark containing text-graph pairs of multiple types or relations, maintaining a rather general domain, while Bio Event pertains to biomedical data by aggregating 10 popular biomedical datasets.
To adapt an LLM to a particular task, two popular task learning methods are fine-tuning and in-context learning (iCL) (Brown et al., 2020). Given a training dataset pertaining to the new task at hand, fine-tuning an LLM amounts to an additional training phase to update a subset of learnable model parameters to adapt to the new task. In-context learning, on the other hand, amounts to including a few task examples in the model prompt at inference time - a special case of few-shot learning. Typically, iCL provides weaker performance than fine-tuning and is computationally more expensive at inference time (Brown et al., 2020; Liu et al., 2022), yet it is highly flexible as it does not require any parameter updates. Both options involve a vast amount of design choices, from the quality and quantity of available training data to the amount of in-context examples to include in iCL.
While most work on knowledge graph extraction has focused on pushing the state-of-the-art in terms of performance or summarising the field in terms of different applications and formulations of scenarios and tasks, it remains unclear to the general AI practitioner what would be, given a specific dataset and computational resources, the best solution to approach a text-to-graph task, formulated as an end-to-end LLM-based solution.
Research Objective and Contribution
We direct this work to the general AI practitioner in the general or biomedical domain who aims to develop an end-to-end LLM-based knowledge graph extraction system from textual sources. We investigate how to best approach such task by examining various combinations of model design choices, assuming a fixed and accessible computational resource of a single NVIDIA Quadro RTX 8000 GPU. 1 The main variables under investigation are model architecture (encoder-decoder and decoder-only), model family (T5, BART, Mistral-v0.1, and Llama-2), model size (small (60M) to mid (13B learnable parameters)), task learning method (fine-tuning and iCL) and additional pre-training data (relation extraction data, conversation data, instruction data, and (bio)medical data). In brief, this article’s main insights are as follows:
In the general domain, we show that small fine-tuned encoder-decoder models and mid-sized decoder-only models adopting iCL achieve comparable results with high entity and relation recognition and moderate yet encouraging graph completion. We provide tentative evidence that biomedical knowledge graphs are substantially harder to model from textual sources than the general domain. Mid-sized decoder-only models adopting iCL show weak performance, while performance of small fine-tuned encoder-decoder models is robust compared to the general domain. However, we discover issues with the biomedical benchmark adopted from Frisoni et al. (2022) and a thorough revision is required to adopt it as gold standard for text-to-graph tasks. Only additional pre-training data on relation extraction tasks boosts model performance, while neither observing conversation data, instruction data nor (bio)medical data during pre-training makes a notable difference. We propose and experimentally prove the effectiveness of a simple truncation-based heuristic on model output to control for a specific type of hallucination of in-context learning, avoiding expensive prompt tuning and prompt design. Off-the-shelve LLMs in the zero-shot setting (i.e. no in-context examples, only a task instruction and reference text) show weak performance. Careful design choices and a task learning method are required to be suitable for such task, especially in safety-critical and domain-specific contexts such as the biomedical domain. We highlight several areas of importance.
Article Structure
This article is structured as follows. Section 2 introduces the related work, with a focus on architecture, tasks, and proposed benchmarks. Subsequently, Section 3 presents the methodology, as well as the used datasets (3.2), metrics (3.3), model architectures (3.4), task learning methods (3.5), and experiments’ set-up (3.6). Then Section 4 shows the experimental results, followed by Sections 5 and 6 to, respectively, discuss and conclude the article.
Related Work
Recent surveys suggest LLMs are of primary interest and hold potential for multiple types of graph-based tasks (Jiang & Usbeck, 2022; Jin et al., 2023; Liu et al., 2023; Pan et al., 2024). The task of automatically generating a knowledge graph from a reference text (text-to-graph) is closely related to the more general NLP task of relation extraction, traditionally composed of the two separated steps of named entity recognition and relation classification (Huguet Cabot & Navigli, 2021). Named entity recognition involves identifying and classifying entities in the text, which can be seen as nodes in the resulting knowledge graph (Yadav & Bethard, 2018). Sub-tasks include co-reference resolution and entity disambiguation. Relation classification, in turn, aims at identifying the relation between two given entities, as (often implicitly) expressed in the reference text containing the identified entities. In an LLM-based knowledge graph extraction system, both steps are potentially entangled in a single end-to-end solution.
Previous works have explored different approaches to text-to-graph tasks and the utilisation of LLMs (Babaei Giglou et al., 2023; Hofer et al., 2024; Neuhaus, 2023). However, the main focus has been on pushing the state of the art on specific sub-tasks and benchmarks, such as research data (Dessí et al., 2022; Kabongo et al., 2024; Kuhn et al., 2018), question-answering (Kacupaj et al., 2021; Kapanipathi et al., 2021), common-sense (Ilievski et al., 2021; Zavarella et al., 2024), biomedical (Himmelstein et al., 2023; Zietz et al., 2024), and other (Angioni et al., 2024). Moreover, unlike this work, the literature does not offer a systematic experimental comparison of the efficiency of contributing factors in an end-to-end text-to-graph task, as suggested by the summary proposed in Table 1. For instance, Bosselut et al. (2019) proposed COMET, a model that generates commonsense knowledge graphs from textual inputs. At the same time, the authors introduced the ATOMIC dataset, which is designed for commonsense inference, and utilised a transformer-based model to extract and generate the graph-based knowledge. The presented transformer model was only compared against existing LSTM-based solutions, which were already known to be subsumed by transformers.
Related Work Summary. Schematic Representation of the Most Relevant Related Work on Text-to-Graph, Focusing on Five Dimensions: Model Architecture (Encoder-Decoder vs. Decoder-Only), Main Benchmark (Name), Learning Method (Tuning vs. In-Context Learning (iCL)), and New (Yes vs. No, for Both Model Architecture and Benchmark).

Related Work Summary. Schematic Representation of the Most Relevant Related Work on Text-to-Graph, Focusing on Five Dimensions: Model Architecture (Encoder-Decoder vs. Decoder-Only), Main Benchmark (Name), Learning Method (Tuning vs. In-Context Learning (iCL)), and New (Yes vs. No, for Both Model Architecture and Benchmark).

Further work on the state-of-the-art includes Guo et al. (2020) introducing CycleGT, a two-loss model to learn from text-to-graph and graph-to-text tasks, based on an encoder-decoder pre-trained model (T5), by bootstrapping from fully non-parallel graph and text data, and iteratively back translating between the two forms. The authors propose a comparison of this solution and architecture on multiple general-domain datasets, using alignment and an unsupervised setting (Jin et al., 2020). In addition, Dash et al. (2021) propose a new text-to-graph model, called CUVA (canonicalizing using variational autoencoders), that addresses the redundancy and ambiguity of noun and relation phrases in open knowledge graphs. Unlike current methods that use a two-step process, CUVA simultaneously learns both embeddings and cluster assignments, resulting in better performance. Additional advancements in the field of knowledge graph extraction include Zhang and Zhang (2020), who demonstrate the effectiveness of pre-trained models for generating knowledge graphs from text when fine-tuned with graph-aware objectives. They propose a graph-augmented text representation model that significantly improves the performance of this task.
Other relevant work has focused on introducing new datasets and benchmarks. Wang et al. (2021) introduce Wiki Graphs, a dataset to benchmark text-to-graph and graph-to-text tasks. The study focuses on a single solution, a combination of GNN graph-transformers, compared against transformer models. The study then focuses more on the structure of the graph than its content and, as such, the dataset is generally stripped of the name entity in the task outputs. According to Colas et al. (2021), the authors introduce event narrative, an event-based text-graph and graph-to-text dataset. Similar to previous endeavours, the authors compare a graph-based transformer with two encoder-decoder pre-trained LLMs, namely T5 and BART, and find mixed performance. Frisoni et al. (2022) test a self-implemented version of these models on a new benchmark of biomedical graph-to-text and text-to-graph datasets.
In another study, Mihindukulasooriya et al. (2023) introduced a text-to-graph dataset,
The previous paragraphs suggest a disparity in the proposed approaches to the task under investigation, in terms of a lack of decoder-only solutions. Of note, Khorashadizadeh et al. (2023) propose a qualitative analysis of the abilities of multiple high-level and mostly decoder-only models, such as ChatGPT and Bard (now Gemini), in terms of knowledge graph completion and question answering. Focusing on biomedical queries, the authors conclude that ChatGPT might present a valuable asset in automatically extracting knowledge graphs, albeit at a significant computational cost. Hu et al. (2020), examined the impact of incorporating graph structural information into the encoding process of a decoder-only model. Their proposed model, GPT-GNN, combines the generative pre-trained transformer with GNNs to enhance the learning of graph representations.
To the best of our knowledge, this work proposes a first
Our work differs from previous studies in knowledge graph extraction in that most proposed either a new model, dataset, or comparison between relatively similar transformer models, while this study experimentally compares the effect of model architectures, family, size and pre-training data across two different domains, namely general and biomedical. As such the goal is not to push state-of-the-art performance, but to study the impact of each factor and describe best practices and important considerations, given a specific amount of computational resources.
This session presents the materials and methods, focusing on knowledge graph structure (Section 3.1), benchmark datasets (Section 3.2), evaluation metrics (Section 3.3), LLMs of various architecture, family, size and pre-training characteristics (Section 3.4), task learning methods (Section 3.5), and experimental setup (Section 3.6).
Knowledge Graph Structure
To ensure a stable and fair comparison across domains, we pre-process our benchmark datasets to match the following linearised text-graph structure. Formally, a dataset consists of two sets of strings 
Benchmark Datasets
We adopt two parallel text-to-graph datasets, to be referred to as Web NLG (Gardent et al., 2017) and Bio Event (Frisoni et al., 2022). Table 2 contains the basic descriptive statistics for both datasets.
Descriptive Statistics of the Web NLG and Bio Event Dataset.

Descriptive Statistics of the Web NLG and Bio Event Dataset.

The Web NLG dataset (we choose version 3.0) is a widely used text-to-graph dataset that contains text-graph pairs of multiple types or relations, maintaining a rather general domain. For each text-graph pair, the corresponding DBpedia category is available, pertaining to the topic of the Wikipedia article it is extracted from. Using this information, we divide the test set into categories that are either seen or unseen in the training and validation data, providing opportunity to test in- and out-of-distribution generalisation capabilities of LLMs (see Appendix C). Out of a total of 18 categories, the categories film, scientist and musical work are unseen in training and validation.
Contrary to the general domain, Bio Event pertains to biomedical data and aggregates 10 popular biomedical datasets, thus (potentially) representing an important domain-specific benchmark for text-to-graph tasks in healthcare. However, the overall quality of the Bio Event dataset requires further comments. Originally presented by Frisoni et al. (2022) as separate datasets for text-to-graph and graph-to-text tasks, we adopt the graph-to-text dataset for our text-to-graph task as it surprisingly includes a larger amount of unique text-graph pairs. Another surprising finding is how in this Bio Event dataset for the graph-to-text task, there are up to 78 unique sets of triplets corresponding to a single reference text. This contradicts the underlying assumption that knowledge graphs and reference texts only differ syntactically and not semantically. While the same knowledge graphs could correspond to multiple natural language reference texts with the same semantic meaning, a reference text naturally only has one corresponding knowledge graph that contains the full set of entities and relations described. Finally, there exists a large degree of dataset contamination, where the same reference text appears in both train and test set, yet connected to a different knowledge graph.
For the purpose of this work, we use a quick heuristic to clean up the Bio Event dataset, allowing us to use it as a biomedical alternative to the general domain of Web NLG. First, we refine the Bio Event graph-to-text dataset by removing any duplicate reference texts and breaking ties in favour of the text-graph pair pertaining to the longest linearised knowledge graph, assuming that the longest knowledge graph is the most complete description of the entities and relations described. This filters out 66% of the datapoints in the original Bio Event dataset, explained by the earlier observation that a single reference text in that dataset often corresponds to multiple unique sets of triplets, whereas we limit ourselves to strictly unique text-graph pairs. We further process the knowledge graphs to match the Web NLG setup by removing metadata associated with nodes and edges, and we finally obtain a train/validation/test set with an 80/10/10% split.
As shown in Table 2, the issue remains that there is a large number of unique entities and triplets compared to the number of available examples. Furthermore, as shown in Appendix A, manual inspection of cherry-picked examples reveals that reference texts and knowledge graphs do not always contain the same set of entities and relations, and thus they differ semantically. For example, consider the following tuple taken from the Bio Event test set,
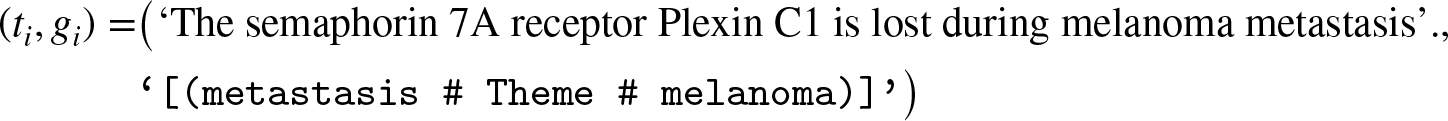
Producing the ‘correct’ knowledge graph in this instance requires (i) ignoring most of the text, and (ii) producing a relation that is absent in any grammatical form. The opposite is true for general domain graphs in Web NLG, which contain relations such as
Finally, we follow Keysers et al. (2020) in assessing the compositional generalisation aspect of the train, validation and test set. In brief, text, or knowledge graphs, in this case, can be categorised into
On Web NLG, we observe that the training and validation sets are similar in terms of both atoms and compounds, whereas the training and test sets differ substantially in terms of both atom and compound divergence. Not surprisingly, this is highest for unseen categories. How the distribution of Web NLG examples over train and test set originates is unclear, but a redistribution of examples over train, validation and test set in future work could be beneficial to more robustly test compositional generalisation. On Bio Event, we see low atom divergence and high compound divergence between the train, validation and test set, in line with the principles of compositionality. We recommend that the general practitioner pay attention to the quality of benchmark datasets, such as descriptive statistics and atom/compound divergence, since model performance is highly sensitive to the quality of the underlying data.
We evaluate a model’s performance with recall-oriented understudy for Gisting Evaluation (Rouge) scores (Lin, 2004). Originally introduced for summarisation evaluation, the set of metrics is transferable to our text-to-graph setup by identifying a graph as a single-sentence summary. We specifically focus on Rouge-
All scores are reported as the harmonic mean between recall and precision, making use of the implementation by Hugging Face.
2
The exact formulas to calculate Rouge-
Rouge-L metrics is based on the LCS between the candidate graphs (


The classical knowledge graph extraction pipeline consists of multiple stages, typically including co-reference resolution, named entity recognition, entity disambiguation and relationship classification. Using LLMs for knowledge graph extraction entangles all stages into one, therefore, complicating the process of ascribing model errors to one or more stages. Comparing the Rouge metrics employed here, however, allows some insights. For example, Rouge-1 is a direct measure of entity and relation recognition, although it does not distinguish between either. Furthermore, Rouge-2 and Rouge-L additionally take sequence order into account, such that the difference with Rouge-1 is a measure of entities and relation being in the right order. In both cases, however, entity ambiguity and co-reference problems might be confounding factors. Finally, we note that the longest common sub-sequence differs from the longest common sub-
How to best disentangle failure modes in knowledge graph extraction using LLMs remains an open question that other popular natural language evaluation metrics such as BLEU and cross-entropy do not provide an obvious answer to. Perhaps a more natural solution to evaluate knowledge graphs would be to extract entities, relations and triplets from structured natural language output and use knowledge-graph-specific metrics to measure model accuracy, coverage, coherence and succinctness. In this preliminary work, however, natural language in model output is not always rigidly structured and thus we pertain to natural language metrics. Furthermore, it is not obvious whether to assess knowledge graphs on entity or triplet level and how to deal with entity disambiguation and co-reference issues, such that natural language metrics on token level provide an easily accessible baseline.
This work focuses on the systemic comparison of two transformer architectures that allow text-generation capabilities, namely encoder-decoder, and decoder-only models. The next paragraphs introduce these architectures from a high-level perspective and discuss the specific pre-trained models adopted in the experiments. Importantly, all models mentioned below are fully open-source and accessible through Hugging Face by adopting the transformer library (Wolf et al., 2020). A summary of adopted models and their specifications are proposed in Table 3.
Models Specification. Summary of the Models Used for Our Experiments, and Their Basic Specifications.

iCL: in-context learning; T5: text-to-text transfer transformer; BART: bidirectional auto-regressive transformers; REBEL: relation extraction dataset during pre-training.
Models Specification. Summary of the Models Used for Our Experiments, and Their Basic Specifications.

iCL: in-context learning; T5: text-to-text transfer transformer; BART: bidirectional auto-regressive transformers; REBEL: relation extraction dataset during pre-training.
Encoder-decoder architectures are a generic class of transformer models (Vaswani et al., 2017b). At their core, encoder-decoder models leverage the power and computational scalability of self-attention mechanisms and feed-forward neural networks in transformer architectures. Both the encoder and decoder consist of a stack of multi-head attention layers, be it that the attention layers in the decoder are masked to prevent attending to future output tokens. Whereas the encoder learns a rich vector representation of model inputs, the decoder auto-regressively generates model output by attending to the encoded inputs and its own generated output. For a detailed description of encoder-decoder architecture and self-attention mechanisms, please refer to Vaswani et al. (2017b). We make use of two families of pre-trained encoder-decoder models,
T5
The T5 family of encoder-decoders was introduced by Raffel et al. (2020). The models undergo two pre-training processes of supervised and self-supervised learning. In the first self-supervised stage, often referred to as denoising language modelling, multi-word pieces of sentences are hidden from the input, and the model is trained end-to-end to spell out which tokens are missing. In the second supervised step, the model is trained to solve task-specific scenarios, added by instruction pre-fixes. We adopt three sizes in the T5 family, namely 60.5M parameters (
BART
The BART model (
For a more in-depth understanding of these differences and the underlying architectural designs of the BART and T5 models, the reader is referred to Lewis et al. (2020) and Raffel et al. (2020). In our experiments we consider a recent version of BART (
Decoder-Only Models
Decoder-only models omit the encoder module in encoder-decoder models, trading off a richer understanding of model inputs for the computational benefits of a streamlined architecture with less learnable parameters. We adopt two families of decoder-only models, namely
Llama-2
Upon release, the Llama-2 family of decoder-only models (Touvron et al., 2023) showed among the strongest performances in the field of LLMs. We include two sizes, one with 7B parameters (
Mistral-v0.1
A more recent introduction, the Mistral-v0.1 family of decoder-only models showcases a strong performance, outperforming the Llama-2 13B model with almost half its parameters (Jiang et al., 2023). Mistral-v0.1 models make use of a significant number of computational advancements, for example, sliding-windows attention (Beltagy et al., 2020), which allows the model to dramatically extend the number of tokens it can simultaneously process. We make use of three models in this family, namely the original 7B model (
Learning Methods
We adopt two distinct task learning methods, fine-tuning for the smaller encoder-decoder models and iCL (Brown et al., 2020) for the larger decoder-only models.
Fine-Tuning
All fine-tuning experiments are based on the
In-Context Learning
Under the iCL setting, each pre-trained model is queried with a simple prompt, containing a set of

Example prompt –
We omit time-consuming prompt engineering and computationally expensive prompt tuning to resemble common practice of end-users. However, we highlight the importance of such practice to prevent model hallucinations of the kind highlighted in Section 4.2.1 and more generally to prevent spurious features in prompt design along the lines of Sclar et al. (2023). To provide a fair estimate of iCL performance, we introduce a simple post-hoc hallucination-control heuristic to determine the end of the desired structured output (i.e. the end of a knowledge graph). Simply put, we truncate model output at the appearance of the tokens
The motivation behind this heuristic arises from preliminary experiments that showed our prompt design is sub-optimal. In various instances the model seems to misinterpret the specified text-to-graph task by continuously generating text-graph alterations along the lines of the in-context example sequence, instead of outputting one single graph corresponding to the final reference text (see Appendix A for some cherry-picked examples). There are ample known methods to engineer a prompt that optimises a given task, such as using delimiters to separate in-context examples, chain-of-thought prompting or more careful phrasing of the task description to maintain intent, but we avoid to engage with this time-consuming process and instead adopt the previously mentioned heuristic. An extensive error analysis of the hallucination phenomenon and our hallucination-control heuristic is given in Section 4.2.1.
This article consists of two sets of experiments, designed to unveil the approximate overall power of selected models and task learning methods, as well as to understand what impacts and shapes their performance. All our experiments are run on a single NVIDIA Quadro RTX 8000 GPU to resemble the experience and facilities of a general AI practitioner. We do recognise that assuming larger computational power could significantly improve results, especially by including large decoder-only models or by fine-tuning the mid-sized Mistral-v0.1 and Llama-2 families, which is out of the computational reach of the current setup.
General Comparison
The main goal of this experiment is to understand how LLM characteristics and task learning methods perform in our text-to-graph task, under fixed computational resources, adopting two task domains – general and biomedical. Throughout, we aim to guide the general AI practitioner to understand which combination is most suited for such task and to show-case how to navigate (part of) the vast and complex spectrum of model design choices. Given the fixed computational resources, we fine-tune the previously introduced set of smaller encoder-decoder models and compare performance to the set of larger decoder-only models in combination with iCL. This choice is framed in the context of a given computational resource such that fine-tuning is computationally infeasible for larger models. At the same time, the short context window of the T5 and BART families (1k tokens or below) prove iCL unsuitable. We adopt
Effect of the Number of In-Context Examples
The optimal number of in-context examples varies with the task at hand and perhaps with other unknown factors. In this experiment, we investigate the impact of
Results
General Comparison
The overall results of various combinations of model architecture, family, size, relevant pre-training data and task learning method are shown in Table 4. We describe general trends and subsequently compare Rouge metrics in Section 4.1.1. Note that for Web NLG, we compute Rouge metrics based on the full test set, and we refer to Appendix C for a comparison of seen and unseen categories.
General Experiment Results. We Report Rouge Scores Obtained by Various Combinations of Model Architecture, Family, Size (Learnable Parameters), Relevant Additional Data Seen During Pre-Training and Our Task Learning Method of Fine-Tuning or iCL on Two Benchmarks – Web NLG and Bio Event.
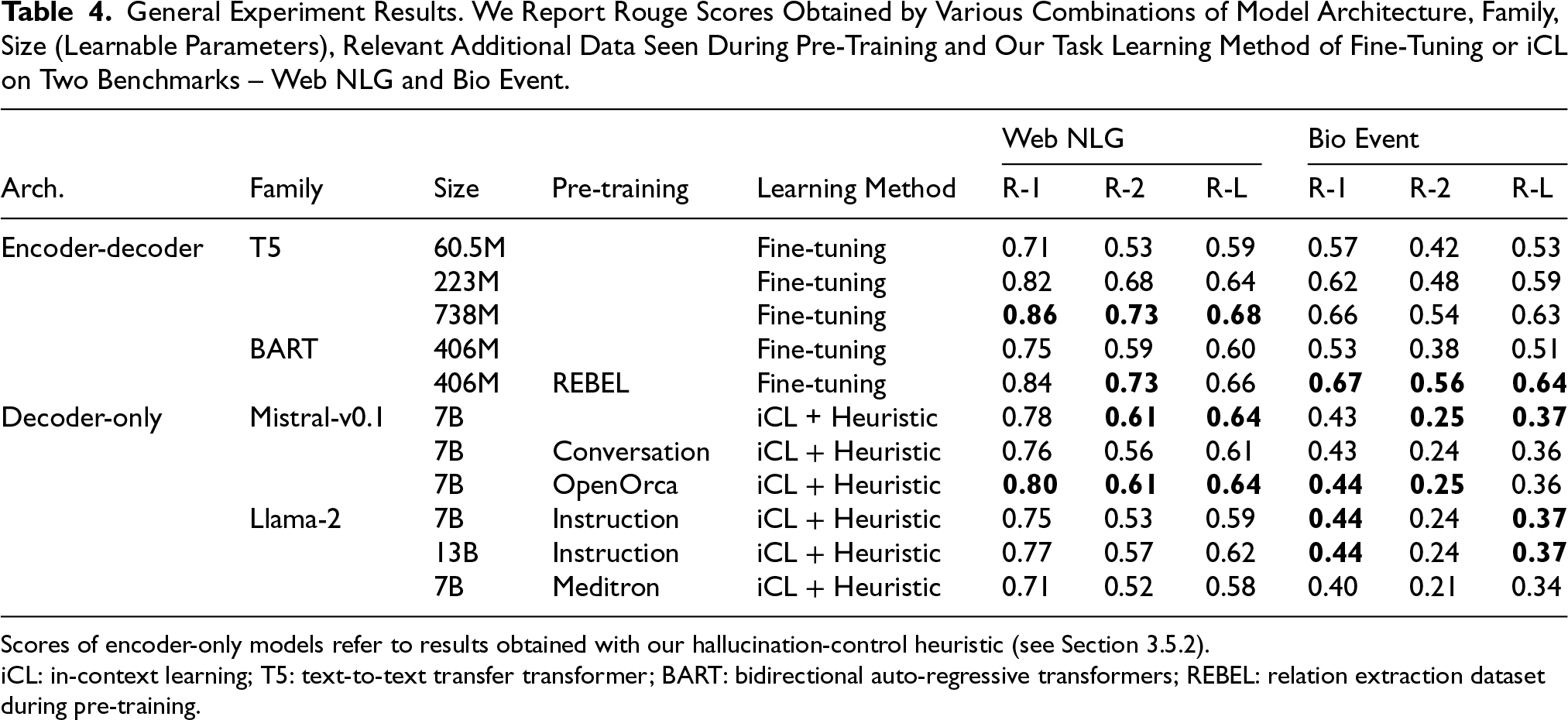
Scores of encoder-only models refer to results obtained with our hallucination-control heuristic (see Section 3.5.2).
iCL: in-context learning; T5: text-to-text transfer transformer; BART: bidirectional auto-regressive transformers; REBEL: relation extraction dataset during pre-training.
General Experiment Results. We Report Rouge Scores Obtained by Various Combinations of Model Architecture, Family, Size (Learnable Parameters), Relevant Additional Data Seen During Pre-Training and Our Task Learning Method of Fine-Tuning or iCL on Two Benchmarks – Web NLG and Bio Event.

Scores of encoder-only models refer to results obtained with our hallucination-control heuristic (see Section 3.5.2).
iCL: in-context learning; T5: text-to-text transfer transformer; BART: bidirectional auto-regressive transformers; REBEL: relation extraction dataset during pre-training.
First, we compare task learning methods. On Web NLG, we observe comparable results among smaller fine-tuned encoder-decoder models across metrics, as opposed to larger decoder-only models adopting iCL. On Bio Event, however, there is a clear benefit in fine-tuning smaller encoder-decoder models. We hypothesise this relates to issues regarding benchmark quality outlined in Section 3.2. For example, the high amount of unique entities and triplets in Bio Event shows a complex distribution of patterns in reference texts that is difficult to infer correctly from just eight in-context examples. Overall best performance on both benchmarks across metrics is reached with fine-tuned encoder-decoder models, that is, the largest model in the T5 family on Web NLG and BART + REBEL on Bio Event.
A visualisation of Rouge-L model performance is shown in Figure 2. Within both the T5 and Llama-2 family, we find a clear positive correlation between model size and LLM performance, which is in line with other research findings in the literature (Hestness et al., 2017). Focusing on the BART family, we see that adopting an additional relation extraction dataset during pre-training (REBEL) yields universally superior results. This is in sharp contrast to other pre-training additions, since neither conversation data nor instruction, OpenOrca or Meditron datasets seem to affect performance on either benchmark. We hypothesise none are particularly relevant to our text-to-graph task, although this is notably most surprising for the biomedical knowledge in the Meditron pre-training data.

The scatter plot visualises Rouge-L scores presented in Table 4, in addition to the results of in-context learning (iCL)
Assessing in Figure 2, the performance of our hallucination-control heuristic for iCL, we observe it yields a large performance boost, independent of benchmark, model architecture, family, size or pre-training data. To briefly reiterate, we avoided computationally and experimentally demanding prompt engineering or tuning by simply truncating model output at the tokens that signal the end of a knowledge graph (i.e.
Figure 3 presents the performance obtained by different combinations of model architecture, family, size, task learning method, and additional pre-training data, as evaluated by the set of Rouge metrics discussed in Section 3.3. To simplify the visualisation, results for models with the same amount of learnable parameters have been collapsed, reporting mean scores and an interval of one standard error both ways.
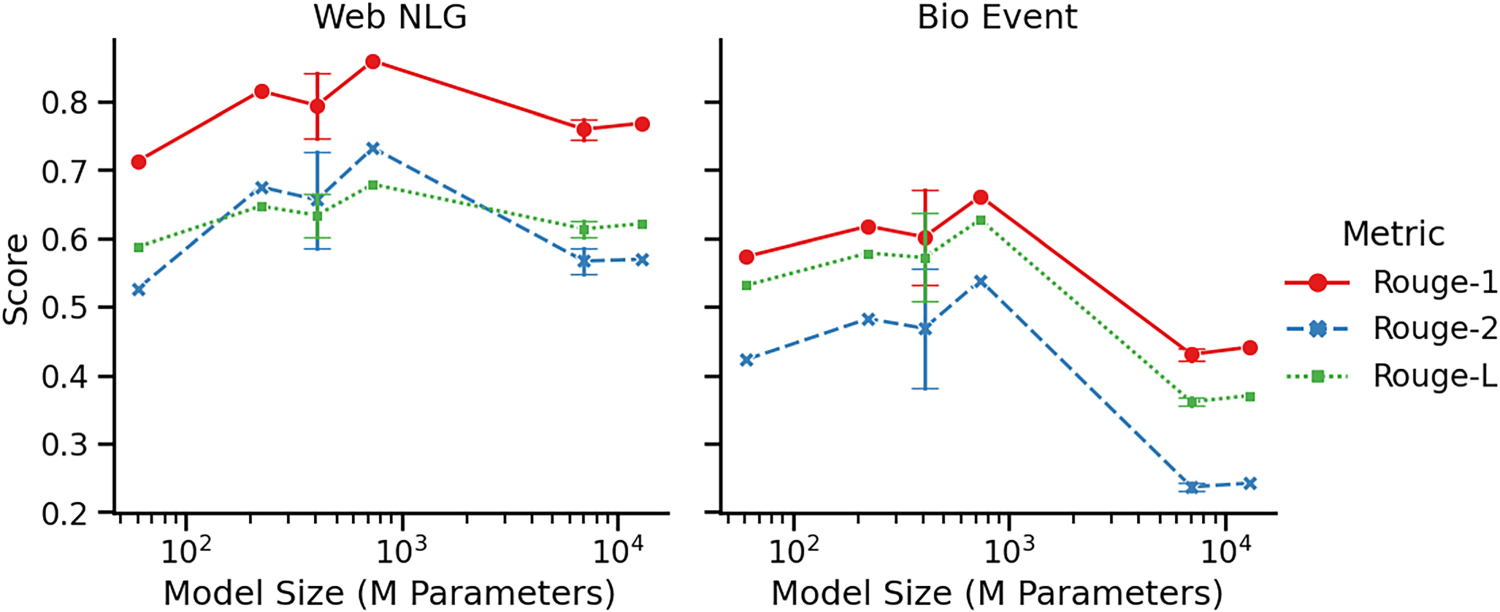
Metrics analysis. The lineplot visualises the Rouge scores in Table 4 as a function of model size. Hue and style refer to the different Rouge metrics (see Section 3.3). At 406M and 7000M learnable parameters we report mean scores, obtained by collapsing the results for models with the same amount of learnable parameters, while error bars report one standard error both ways.
We observe Rouge-1 score to be consistently high, especially on Web NLG, indicating strong entity and relation recognition. On Bio Event, recognising entities and relations poses a more difficult task, as there is a significantly larger amount of entities to be learned. On Web NLG, Rouge-2 and Rouge-L are in similar neighbourhoods with a consistent gap towards Rouge-1 across model sizes, indicating that where entities and relations are often well identified, models struggle to put them in the right order and provide the right triplets. This indicates the more difficult task of relation classification in the knowledge graph completion pipeline. On Bio Event, however, Rouge-L is consistently above Rouge-2 and close to Rouge-1, indicating the identified entities and relations are often in the right order, but certain entities or relations are missing such that correct 2-grams are lacking.
For the remainder of this section, we report only Rouge-L scores to simplify visualisations, while noting that performances relative to other metrics are robust to what is described here.
In the previous section, we have learned that complex knowledge graph consisting of a wide spectrum of entities and relations are hard to model using iCL, especially with exposure to only eight in-context examples. In this experiment, we aim to understand how the amount of in-context examples
In the zero-shot setting (

Effect of the number of in-context examples for the Mistral-v0.1
When evaluating models on their standard generated output, we observe a peak of performance at four to eight samples, preceded by a sharp increase, and then a slightly more moderate decrease. We go into further detail on the dynamics behind this phenomenon in the next section.
We now explore a quantitative analysis of model hallucinations and our proposed heuristic for controlling them. Although hallucinations are observed across all models included, we focus specifically on the Mistral-v0.1

Hallucination analysis: Boxplots for the number of tokens and percentage of graph output for the Mistral-v0.1 model fine-tuned by OpenOrca, as a function of various amounts of in-context examples
Figure 5 allows us to draw more general conclusions on the nature of this hallucinated text. The figure displays a boxplot of the percentage of graph-based tokens in the output text, measured as the number of tokens inbetween
Finally, Appendix E shows identical figures, except now our hallucination-control heuristic is applied. We observe all boxplots are close to the target distribution across
Overall, this work has been directed at the AI practitioner in the general or biomedical domain, aiming to develop an end-to-end LLM-based automatic graph extraction system from textual sources. Assuming a realistic computational baseline, we aim to contribute to the development of more effective and efficient pipeline for knowledge extraction and representation tasks by highlighting the impact of a plethora of design choices. Our large-scale comparison provided several empirical insights.
Foremost, we showed off-the-shelve LLMs together with a task learning method show strong entity and relation recognition, and on the overall task of knowledge graph completion results can be classified as moderate yet promising. Optimal performance of LLMs is likely higher than displayed here, e.g. due to prompt engineering/tuning, hyper-parameter tuning, more computational power and more model parameters. Without a task learning method, we showed off-the-shelve LLMs are not directly suitable for text-to-graph tasks and especially in the biomedical domain, zero-shot performance is weak. Comparing task learning methods, fine-tuning has proven more robust than iCL, since mid-sized decoder-only models adopting iCL show weak performance in the biomedical domain, while small fine-tuned encoder-decoder models achieve robust moderate results in both the general and biomedical domain. We hypothesise that expert knowledge contained in reference texts in the biomedical domain poses a more difficult knowledge extraction problem, such that iCL with a small amount of in-context examples is not sufficient to correctly learn said task. That is, knowledge graphs in the biomedical domain might require knowledge obtained
Due to computational constraints at inference time, we experimented with fine-tuning models up to 738M learnable parameters and due to context window constraints, we experimented with up to 32 in-context examples. As context window constraints are based on computational constraints during pre-training and the general practitioner designing a knowledge extraction pipeline typically starts with a pre-trained LLM, it only benefits from scaling up its computational resources when opting for fine-tuning as the preferred task learning method. When it comes to fine-tuning, we find that including additional datasets only boosts performance when the dataset directly pertains to the text-to-graph task. Among the options explored in this paper, only relation extraction data in the form of the REBEL dataset showed a significant boost in performance, while neither conversation data, instruction-tuning nor additional biomedical data made an impact. It is especially surprising that biomedical data does not make a difference on our biomedical benchmark.
We leave the general AI practitioner with three recommendations with regards to being mindful of small details in designing an end-to-end LLM-based knowledge graph extraction pipeline. First, an LLM-based system is highly sensitive to the underlying training and benchmark data. In the case of knowledge graphs, data should contain text-graph pairs that contain identical semantic meaning, but differ syntactically in their natural language and graph structure. Furthermore, benchmarks offering a sparse distribution of entities, relations and triplets poses a difficult knowledge extraction problem, since LLMs are given few examples to learn from. In addition, assessing atom- and compound-divergence between training and test set should be common practice to validate an LLM’s compositional generalisation capabilities and distinguish between in- and out-of-distribution generalisation. We outline several issues with the biomedical benchmark adopted in this article in Section 3.2, such that our findings in this setting should be regarded as tentative. A thorough revision of Bio Event is required to establish a gold standard benchmark in the biomedical domain. Also with regards to the general domain, we find the Web NGL benchmark does not adhere well to the principles of compositional generalisation, and a redistribution of examples over training and test set is recommended.
Second, the choice of evaluation metric for knowledge graph extraction using LLMs require careful consideration. We opted for a set of Rouge metrics for natural language evaluation in order to allow for unstructured model output, yet a post-hoc method to identify entities and triplets in combination with graph-specific evaluation metrics might allow to better distinguish between failure modes in the knowledge graph extraction pipeline. We discuss this in Section 3.3. Such a setup is applicable when LLMs can be trusted to adhere to rigid output structure, but we find LLMs to often violate our desired linearised graph structure. Using Rouge metrics, we find strong results for entity and relation recognition and moderate yet promising results for putting these together in the correct triplets.
Third, careful prompt engineering and prompt tuning are essential to avoid task misinterpretation and model hallucinations of the types described in Section 4.2.1. LLMs are known to be highly sensitive to small changes in input prompts and we provide empirical evidence for this in the case of text-to-graph tasks. To solve the type of hallucinations we encountered, we proposed a simple heuristic based on our linearised graph structure to truncate model output at the tokens signalling the end of the first knowledge graph. This proved highly effective in boosting model performance without time-expensive prompt engineering and computationally expensive prompt tuning, which is not necessary generalisable across subsets of the same dataset (Bertolini et al., 2022). These results suggest that when the output of a model follows a constrained structure, simple rule-based heuristics can be an efficient method to limit undesired output. Finally, we mention what seems to be a standard surgeon’s recommendation in the field of prompt engineering, to use delimiters to separate in-context examples from a new observation and to use careful phrasing to maintain intent of the desired task, such that post-hoc heuristics are redundant.
Conclusion
In this work, we examined the performance of LLMs to automatically generate knowledge graphs from reference texts in the general and biomedical domain. In an end-to-end fashion, we used LLMs in combination with a task learning method in the form of fine-tuning small encoder-decoder models or mid-sized decoder-only models adopting in-context learning. We obtained comparable performance in the general domain with high named entity and relation recognition and moderate yet promising knowledge graph completion. We show tentative evidence that knowledge graphs in the biomedical domain are harder to learn from textual sources than the general domain, independent of other factors considered. Moreover, in the biomedical domain, mid-sized decoder-only models adopting iCL show weak results, while small fine-tuned encoder-decoder models perform robustly. However, we find a gold standard benchmark of text-to-graph data in the biomedical domain is lacking. In the zero-shot setting, we obtained weak performance for all LLMs considered. Additionally, we found no connection between including additional datasets during pre-training that are not directly linked to the text-to-graph task, such as conversation-tuning, instruction-tuning and biomedical expert knowledge. Only including REBEL, a relation extracting dataset, showed a notable boost in performance. Finally, we proposed a simple heuristic to control for model hallucinations as a result of sub-optimal prompt design and provide evidence of its positive impact on performance. We hope these results guide best practices for implementing LLMs in automatic graph extraction and suggest that smaller fine-tuned models with domain-specific optimisations are preferable over large models adopting iCL. Future work could focus on refining these findings, especially by developing novel benchmark datasets and tools to evaluate structured knowledge graph output and disentangle failure modes in the knowledge graph extraction pipeline. The ultimate goal is to enhance knowledge extraction pipelines by utilising the power of LLMs for complex reasoning and text-to-graph AI systems.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
