Abstract
The low efficiency of in vitro differentiation of human embryonic stem cells (hESCs) or human-induced pluripotent stem cells (iPSCs) into insulin-producing cells is a crucial hurdle for the clinical implementation of human pluripotent stem cells (PSCs). Our previous investigation into the key factors for the differentiation of PSCs into insulin-producing cells suggested that the expression of GATA binding protein 6 (GATA6) and Gremlin 1 (GREM1) and inhibition of early growth response protein 1 (Egr1) may be important factors. In this study, we investigated the role of Egr1 in pancreas development. The transfection of small interfering RNA (siRNA) of Egr1 in the early phase induced the differentiation of iPSCs derived from fibroblasts (FiPSCs) into pancreatic endoderm and insulin-producing cells. In contrast, the downregulation of Egr1 in the late phase suppressed the differentiation of FiPSCs into pancreatic endoderm and insulin-producing cells. In addition, the overexpression of Egr1 suppressed the differentiation of iPSCs derived from pancreatic cells into pancreatic endoderm and insulin-producing cells. These data suggest that the downregulation of Egr1 in the early phase can efficiently induce the differentiation of iPSCs into insulin-producing cells.
Keywords
Introduction
Diabetes is caused by an absolute or relative insufficiency of insulin, which is secreted from pancreatic β cells, resulting in impaired glucose metabolism in the entire body 1,2 . The supplementation of the β-cell function is an effective therapeutic strategy, but the insufficient cell supply is a major obstacle to this intervention 3,4 . Therefore, vigorous efforts have been made to develop a stable source of pancreatic cells for clinical use. Human embryonic stem cells (hESCs) and human-induced pluripotent stem cells (iPSCs) are capable of differentiation into cells from the 3 embryonic germ layers that constitute the body. Indeed, a number of groups have generated immature pancreatic β-cell-like cells, which are referred to as insulin-producing or insulin-secreting cells, from ESCs/iPSCs in vitro 5 –9 . However, it is difficult to obtain the stable production of pancreatic lineage cells in a sufficient quantity for use in the clinical setting 10 .
We previously investigated the key factors for the differentiation of pluripotent stem cells (PSCs) into insulin-producing cells and found that the expression of GATA6 and GREM1 and the inhibition of early growth response protein 1 (Egr1) may be important factors for this differentiation 11 . The protein encoded by the Egr1 gene belongs to the EGR family of C2H2-type zinc-finger proteins. It is a nuclear protein and functions as a transcriptional regulator. Although 1 group reported Egr1 to regulate the transcription of the insulin gene 12 and the Pdx1 gene 13 , the relationship between Egr1 and pancreas development is unknown.
In this study, we investigated the role of Egr1 in pancreas development. We first investigated the efficacy of differentiation into pancreatic endoderm by the inhibition of Egr1 in iPSCs derived from fibroblasts (FiPSCs) and next investigated the efficacy of differentiation into pancreatic endoderm by the overexpression of Egr1 in iPSCs derived from pancreatic cells (PiPSCs). We used PiPSCs and FiPSCs in this study because the expression of Egr1 in FiPSCs was higher than in PiPSCs 11 , and PiPSCs differentiated into insulin-producing cells more efficiently than FiPSCs because of their epigenetic memory 14 .
Materials and Methods
Cell Culture
PiPSCs 11 and FiPSCs (201B7; RIKEN BioResource Center, Ibaraki, Japan) were maintained in ESC culture medium. These cells were cultured on mitomycin C-treated SNL76/7 (European Collection of Cell Culture, cat. No. 07032801, lot no. 08F009). The ESC culture medium comprised Dulbecco’s modified Eagle’s medium (DMEM)-F12 (Sigma-Aldrich, St. Louis, MO, USA) supplemented with 20% Knockout Serum Replacement (KSR) (Thermo Fisher Scientific, Yokohama, Japan), 2 mM L-glutamine (Nacalai Tesque, Kyoto, Japan), 1% nonessential amino acid (Thermo Fisher Scientific), 0.1 mM 2-Mercaptoethanol (Thermo Fisher Scientific), and 5 ng/mL Fibroblast Growth Factor 2 (FGF2) (Repro CELL, Kanagawa, Japan). For routine passaging, iPSCs colonies were detached with a CTK solution (2.5 μg/mL Trypsin, 1 mg/mL Collagenase IV, 20% KSR, 1 mM CaCl2/phosphate buffered saline [PBS], and 70% PBS) and split at a ratio between 1:3 and 1:6.
siRNA Transfection
Cells were transfected with siRNAs of Egr1 (Thermo Fisher Scientific) at a final concentration of 10 nM using Lipofectamine® RNAiMAX Transfection Reagent (13778030; Thermo Fisher Scientific) in accordance with the manufacturer’s protocol. For a 6-well plate (9.6 cm2), we used 4 µL of transfection reagent in 150 µL of OptiMEM (Thermo Fisher Scientific), 0.8 µL of 50 µM siRNA solution in 150 µL of OptiMEM, and 2 × 105 PSCs in 2 mL of Essential 8 medium on Matrigel-coated dishes (hESC-qualified, 354277; Corning, NY, USA). Silencer® Select Negative Control No. 1 (Thermo Fisher Scientific) was used as a control. The medium was changed after 48-h incubation.
Messenger RNA (mRNA) Transfection
For transfection of the messenger Egr1 in FiPSCs, synthetic mRNA for Egr1 (mEgr1) was made with the mMESSAGE mMACHINE T7 ULTRA Transcription Kit after amplifying the Egr1 ORF clones (RG209956; ORIGENE, Rockville, MD, USA) with polymerase chain reaction (PCR) primers (Table 1). The quantity of resultant mRNA was measured with a Nano Drop ND-1000 (Thermo Fisher Scientific). Lipofectamine Messenger MAX (Thermo Fisher Scientific) was used for mEgr1 transfection according to the instructions of the manufacturer. For a 6-well plate (9.6 cm2), we used 7.5 µL of transfection reagent, 2.5 µg messenger Egr1, and 3 × 105 PiPSCs in 2 mL of SNL-conditioned medium on Matrigel-coated dishes (hESC-qualified, 354277; Corning). Enhanced green fluorescent protein (EGFP) mRNA (TriLink BioTechnologies, San Diego, CA, USA) was used for control. The medium was changed after 24-h incubation.
List of Primers Used for Quantitative Reverse Transcription Polymerase Chain Reaction of Induced Pluripotent Stem Cells and the Synthesis of Egr1 Messenger RNA.
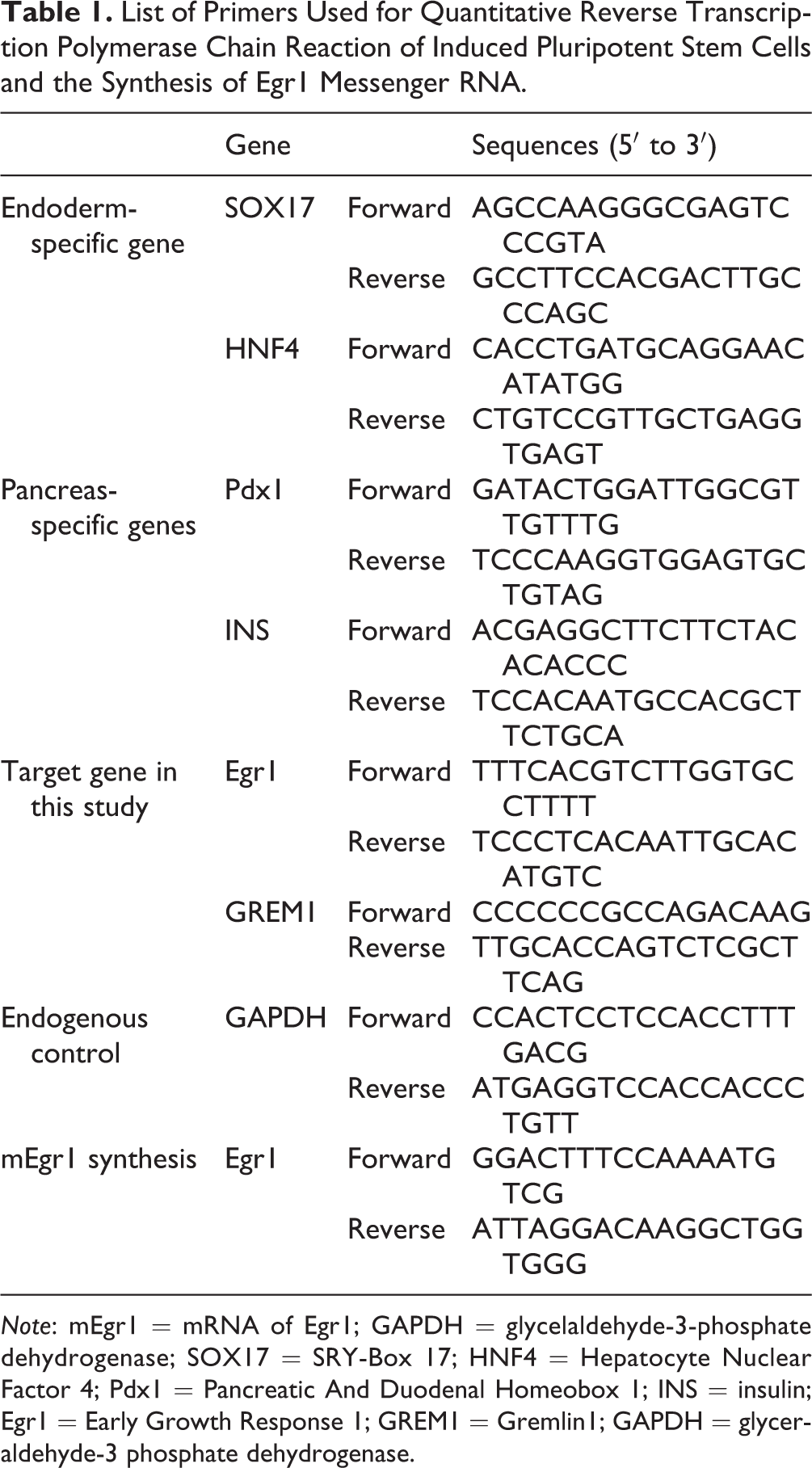
Note: mEgr1 = mRNA of Egr1; GAPDH = glycelaldehyde-3-phosphate dehydrogenase; SOX17 = SRY-Box 17; HNF4 = Hepatocyte Nuclear Factor 4; Pdx1 = Pancreatic And Duodenal Homeobox 1; INS = insulin; Egr1 = Early Growth Response 1; GREM1 = Gremlin1; GAPDH = glyceraldehyde-3 phosphate dehydrogenase.
Differentiation
Directed differentiation into insulin-producing cells was conducted as described previously 5 –8 , with minor modifications. For endoderm differentiation (stage 1), cells were treated with 25 ng/mL of Wnt3a and 100 ng/mL of activin A (Peprotech, Rocky Hill, NJ, USA) in Roswell Park Memorial Institute (RPMI) (Thermo Fisher Scientific) for 1 d, followed by treatment with 100 ng/mL of activin A in RPMI + 0.2% fetal bovine serum (FBS) for 1 d. For differentiation into primitive gut tube (stage 2), the cells were treated with 50 ng/mL of Keratinocyte Growth Factor (KGF) (Peprotech) in RPMI + 2% FBS for 1 d. Medium in stage 3 was not used in this study. For spontaneous differentiation (stage 4), the cells were cultured in DMEM + 1% (vol/vol) B27 supplement for 3 d.
Quantitative Reverse Transcription (RT) PCR
Total RNA was prepared using RNeasy Micro Kit (Qiagen, Tokyo, Japan) and treated with RNase free DNase (Qiagen). A total of 500 ng RNA was used for a RT reaction using the QuantiTect Reverse Transcription Kit (Qiagen). Quantitative RT-PCR (qRT-PCR) was carried out in triplicate using SYBR® Select Master Mix in StepOnePlus™ (Thermo Fisher Scientific) with the following PCR program: 95 °C for 10 min, 40 cycles at 95 °C for 15 s, 60 °C for 1 min, and 72 °C for 15 s. The specific primers are listed in Table 1. Relative quantification was calculated using the 2− ΔΔCt method after normalization to the glycelaldehyde-3-phosphate dehydrogenase.
Results
Increase in Pancreas-Related Genes by Transfection of siRNA of Egr1
To investigate whether or not the suppression of Egr1 induces the differentiation of iPSCs into insulin-producing cells, siRNA of Egr1 (siEgr1) was transfected into FiPSCs, which were derived from fibroblasts and expressed Egr1 more strongly than PiPSCs. FiPSCs were incubated with siEgr1 for 48 h (Fig. 1A). An expression analysis of Egr1, endoderm-specific genes, and pancreatic specific gene (Table 1) in FiPSCs was conducted using qRT-PCR. We used Silencer® Select Negative Control No. 1 as control of siEgr1 (siNeg). After 48-h incubation for transfection, the expression of Egr1 was reduced by about 60%, and endoderm- and pancreatic-specific genes were expressed to a greater degree in siEgr1-transfected FiPSCs than in siNeg-transfected FiPSCs (Fig. 1B).
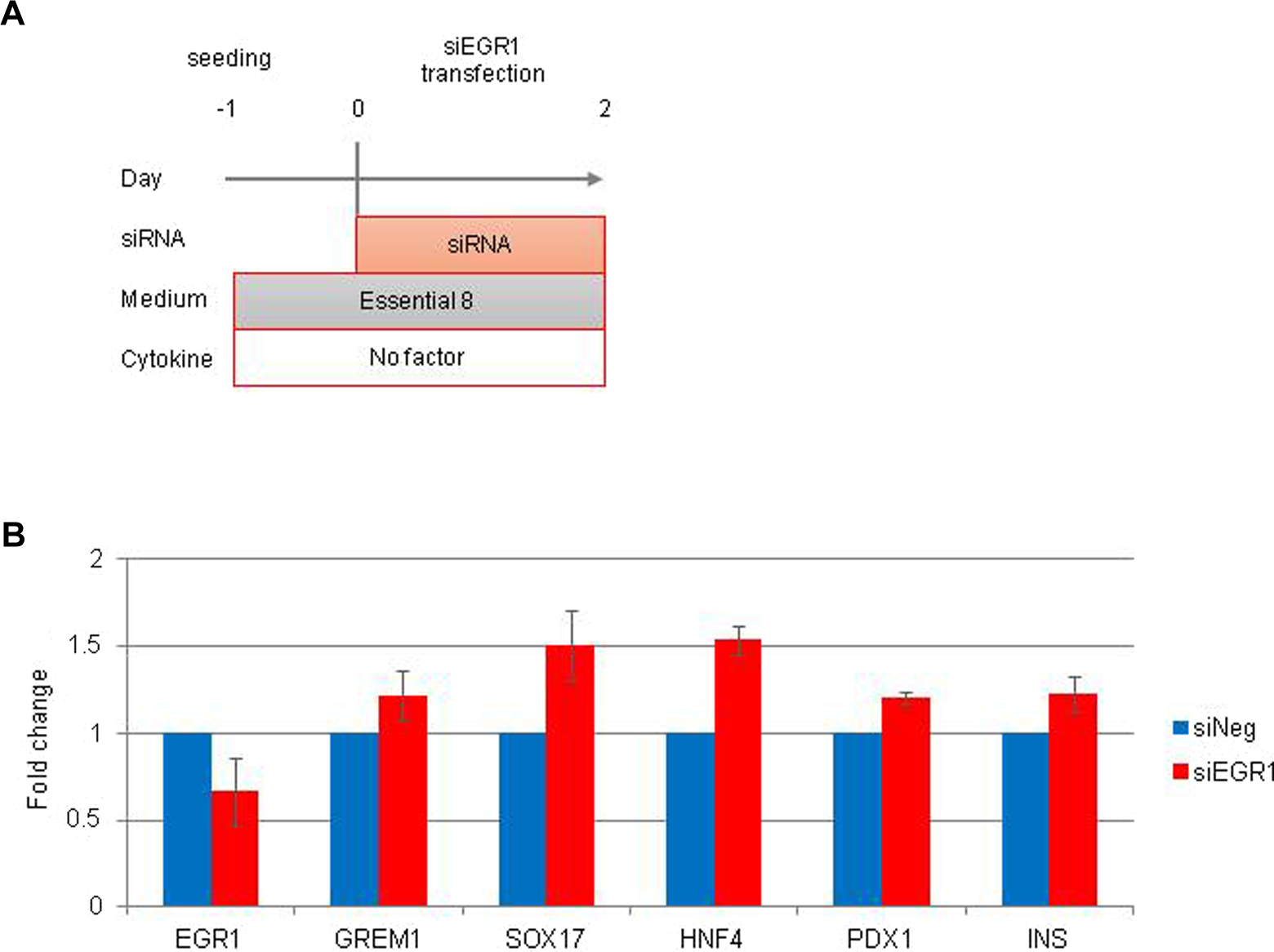
Transfection of early growth response protein 1 (Egr1) small interfering RNA (siRNA) (siEgr1) in human-induced pluripotent stem cells (iPSCs) derived from fibroblast (FiPSCs) in the early phase. (A) Time schedules for the induction of FiPSCs by siEgr1. (B) A quantitative reverse transcription polymerase chain reaction analysis of Egr1 and endoderm- and pancreas-specific genes in FiPSCs treated with siEgr1. The data are expressed as the genes to glycelaldehyde-3-phosphate dehydrogenase ratio, with that of FiPSCs treated with siNeg arbitrarily set at 1.
Increase in Endoderm Genes by siEgr1 Transfection and Treatment with Growth Factors
To investigate the effect of siEgr1 with growth factors on endoderm differentiation, FiPSCs were incubated with siEgr1 for 48 h and then were treated with Wnt3A and/or activin A (Fig. 2A). After 96-h incubation for siEgr1 transfection, the expression of Egr1 was reduced, and endoderm- and pancreatic-specific genes were expressed to a greater degree in siEgr1-transfected FiPSCs than in siNeg-transfected FiPSCs. In particular, the expression of SRY-Box 17 (SOX17) in FiPSCs with siEgr1 was significantly (5- to 6-fold) higher than that in FiPSCs with siNeg (Fig. 2B).
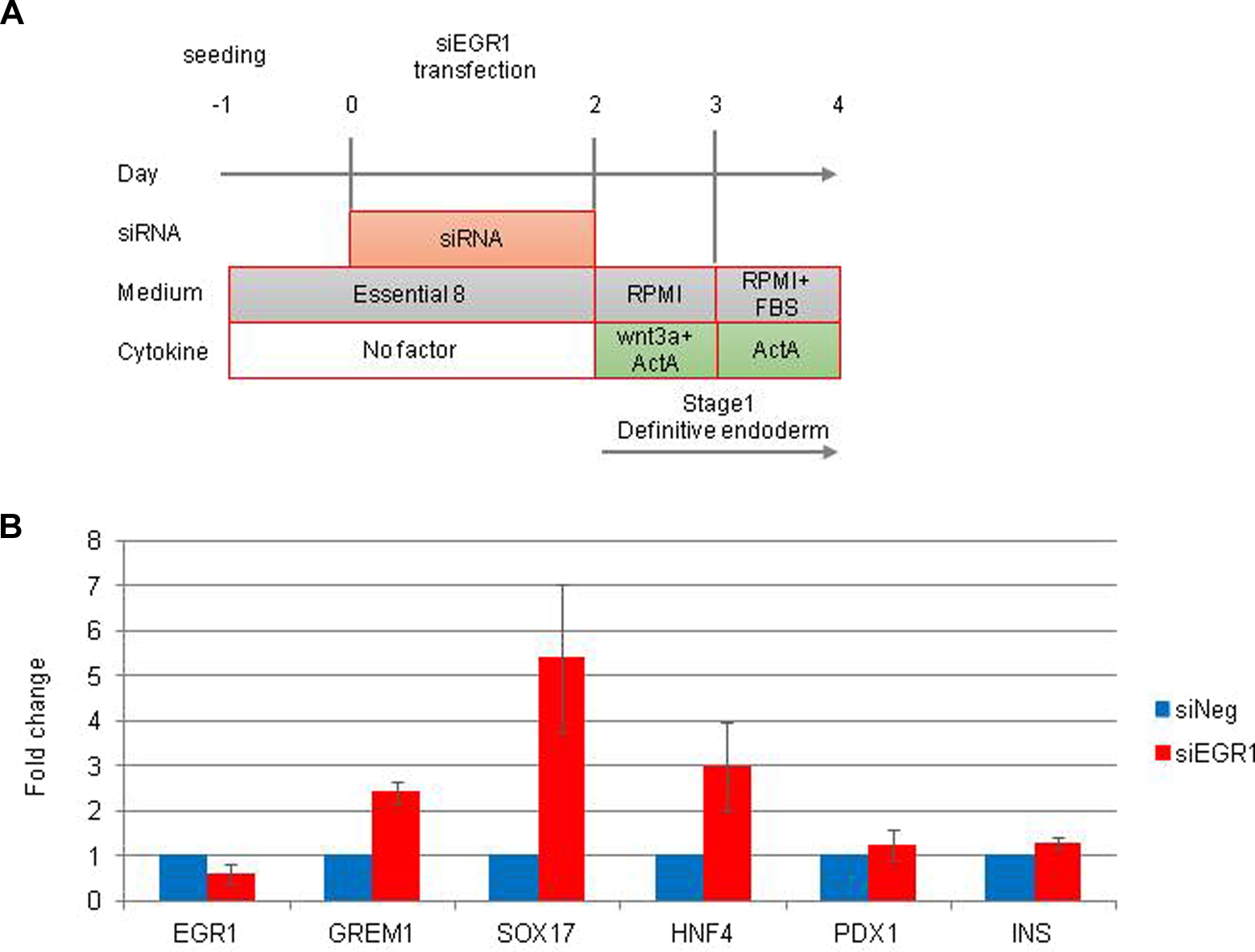
Transfection of small interfering RNA (siRNA) of early growth response protein 1 (Egr1) (siEgr1) in human-induced pluripotent stem cells (iPSCs) derived from fibroblast (FiPSCs) in the early phase and treatment with growth factors. (A) Time schedules for the induction of FiPSCs by siEgr1 and growth factors. (B) A quantitative reverse transcription polymerase chain reaction analysis of Egr1 and endoderm- and pancreas-specific genes in FiPSCs treated with siEgr1. The data are expressed as the genes to glycelaldehyde-3-phosphate dehydrogenase ratio, with that of FiPSCs treated with siNeg arbitrarily set at 1.
Increase in Insulin Genes by siEgr1 Transfection, Treatment with Growth Factors, and Spontaneous Differentiation
We next investigated whether or not the endoderm differentiation with the suppression of Egr1 led to pancreatic differentiation. FiPSCs were transfected with siEgr1 and incubated for 48 h. The cells were then treated with growth factors for another 48 h and cultured in DMEM + B27 (spontaneous differentiation) for 72 h (Fig. 3A). Seven days after siEgr1 transfection, the endoderm- and pancreatic-specific genes were expressed to a greater degree in siEgr1-transfected FiPSCs than in siNeg-transfected FiPSCs (Fig. 3B). Furthermore, the expression of pancreas-specific genes such as insulin in FiPSCs with siEgr1 was likely to be higher than without spontaneous differentiation (Figs. 2B and 3B). These data suggest that the downregulation of Egr1 in the early phase induced the differentiation of FiPSCs into pancreatic endoderm and insulin-producing cells.
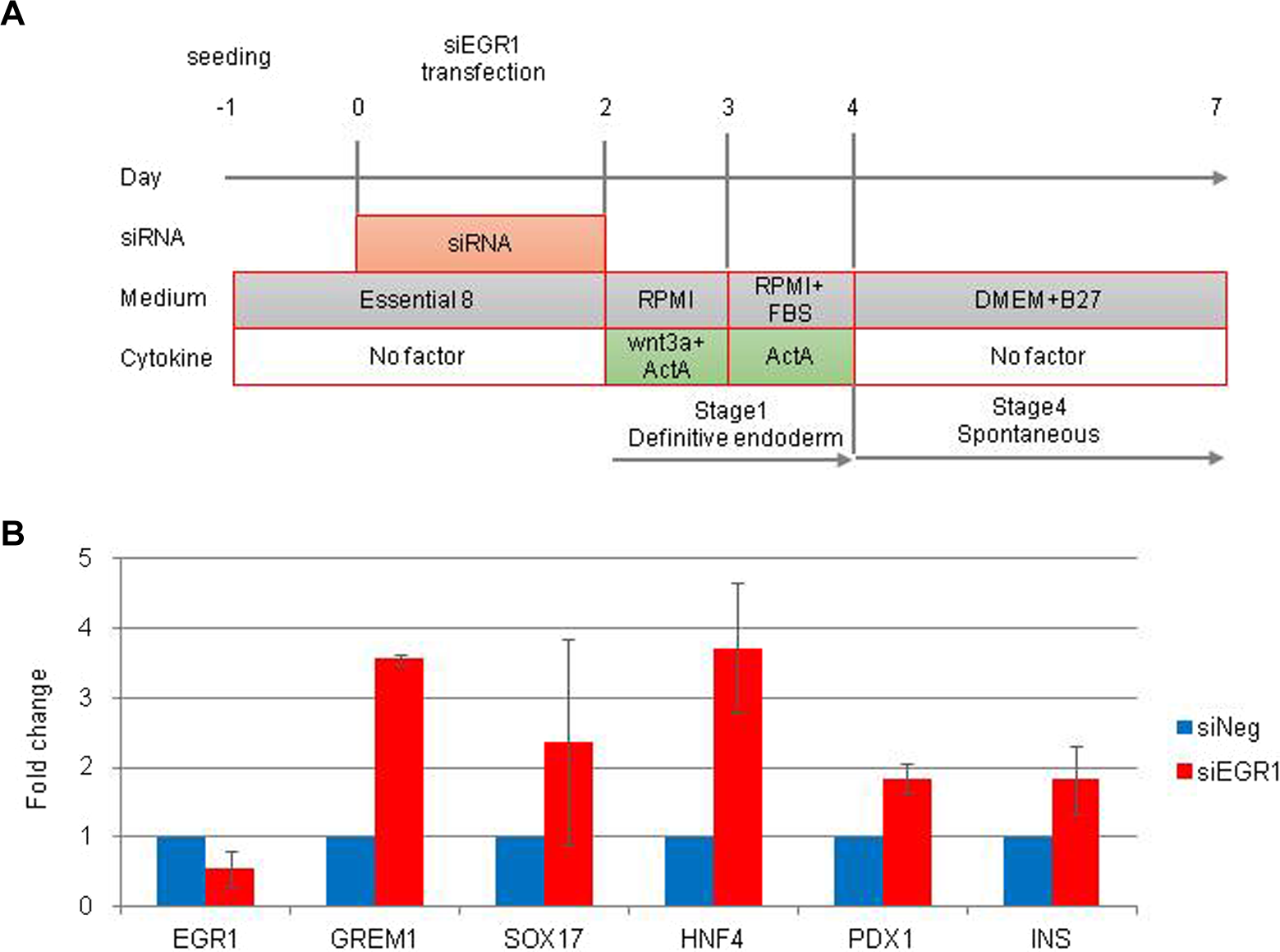
Transfection of small interfering RNA (siRNA) of early growth response protein 1 (Egr1) (siEgr1) in human-induced pluripotent stem cells (iPSCs) derived from fibroblast (FiPSCs) in the early phase, treatment with growth factors, and spontaneous differentiation. (A) Time schedules for the induction of FiPSCs by siEgr1, growth factors, and spontaneous differentiation. (B) The quantitative reverse transcription polymerase chain reaction analysis of Egr1 and endoderm- and pancreas-specific genes in FiPSCs treated with siEgr1. The data are expressed as the genes to glycelaldehyde-3-phosphate dehydrogenase ratio, with that of FiPSCs treated with siNeg arbitrarily set at 1.
Inhibition of Endoderm Differentiation by the Downregulation of Egr1 in the Late Phase
Since the downregulation of Egr1 in the early phase induced the differentiation of FiPSCs into pancreatic endoderm, we also investigated whether or not the downregulation of Egr1 in the late phase induced the differentiation of FiPSCs into pancreatic endoderm. FiPSCs were incubated with siEgr1 for 48 h and then treated with Wnt3A and/or activin A for another 48 h. Subsequently, the cells were transfected with siEgr1 again (Fig. 4A). After the second transfection of siEgr1, the endoderm- and pancreatic-specific genes were expressed significantly more weakly in siEgr1-transfected FiPSCs than in siNeg-transfected FiPSCs (Fig. 4B). These data suggest that the downregulation of Egr1 in the late phase suppressed the differentiation of FiPSCs into pancreatic endoderm and insulin-producing cells.
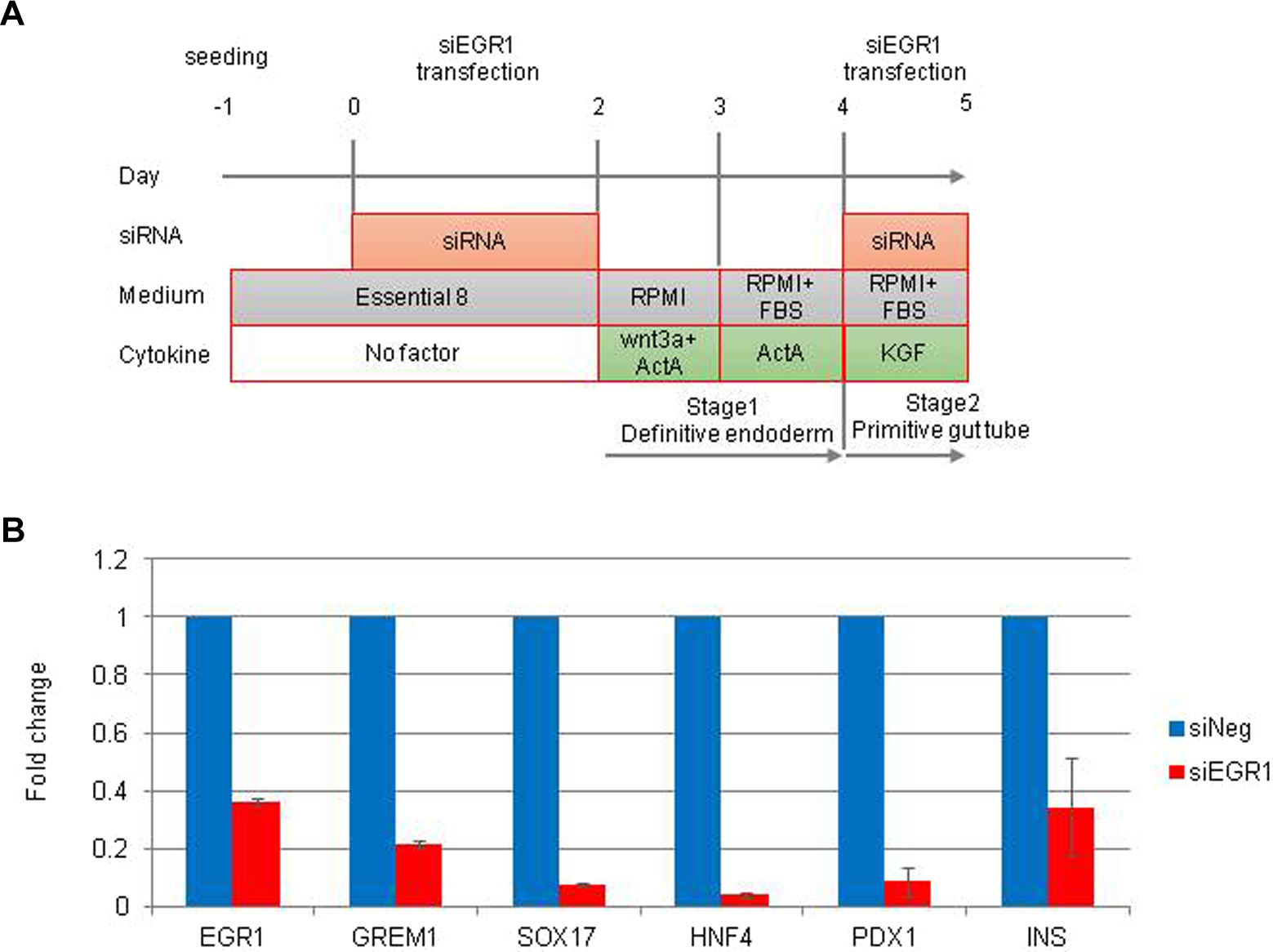
Transfection of small interfering RNA (siRNA) of early growth response protein 1 (Egr1) (siEgr1) in human-induced pluripotent stem cells (iPSCs) derived from fibroblast (FiPSCs) in the early and late phases and treatment with growth factors. (A) Time schedules for the induction of FiPSCs by siEgr1 and growth factors. (B) A quantitative reverse transcription polymerase chain reaction analysis of Egr1 and endoderm- and pancreas-specific genes in FiPSCs treated with siEgr1. The data are expressed as the genes to glycelaldehyde-3-phosphate dehydrogenase ratio, with that of FiPSCs treated with siNeg arbitrarily set at 1.
Upregulation of Egr1 Inhibits Endoderm Differentiation
To investigate whether or not the overexpression of Egr1 reduces the differentiation of iPSCs into insulin-producing cells, mEgr1 was transfected into PiPSCs, which were derived from pancreatic cells. The expression of Egr1 was lower than that in FiPSCs. We used mRNA of EGFP as a control for mEgr1. PiPSCs were incubated with mEgr1 for 24 h and then treated with Wnt3A and activin A for another 24 h. Subsequently, the cells were transfected with mEgr1 again (Fig. 5A). After the second mEgr1 transfection, the expression of Egr1 was increased, and the endoderm- (except GREM1 and HNF4) and pancreatic-specific genes were expressed to a lesser degree in mEgr1-transfected PiPSCs than in EGFP-transfected PiPSCs (Fig. 5B). These data suggest that the overexpression of Egr1 suppressed the differentiation of PiPSCs into pancreatic endoderm and insulin-producing cells.
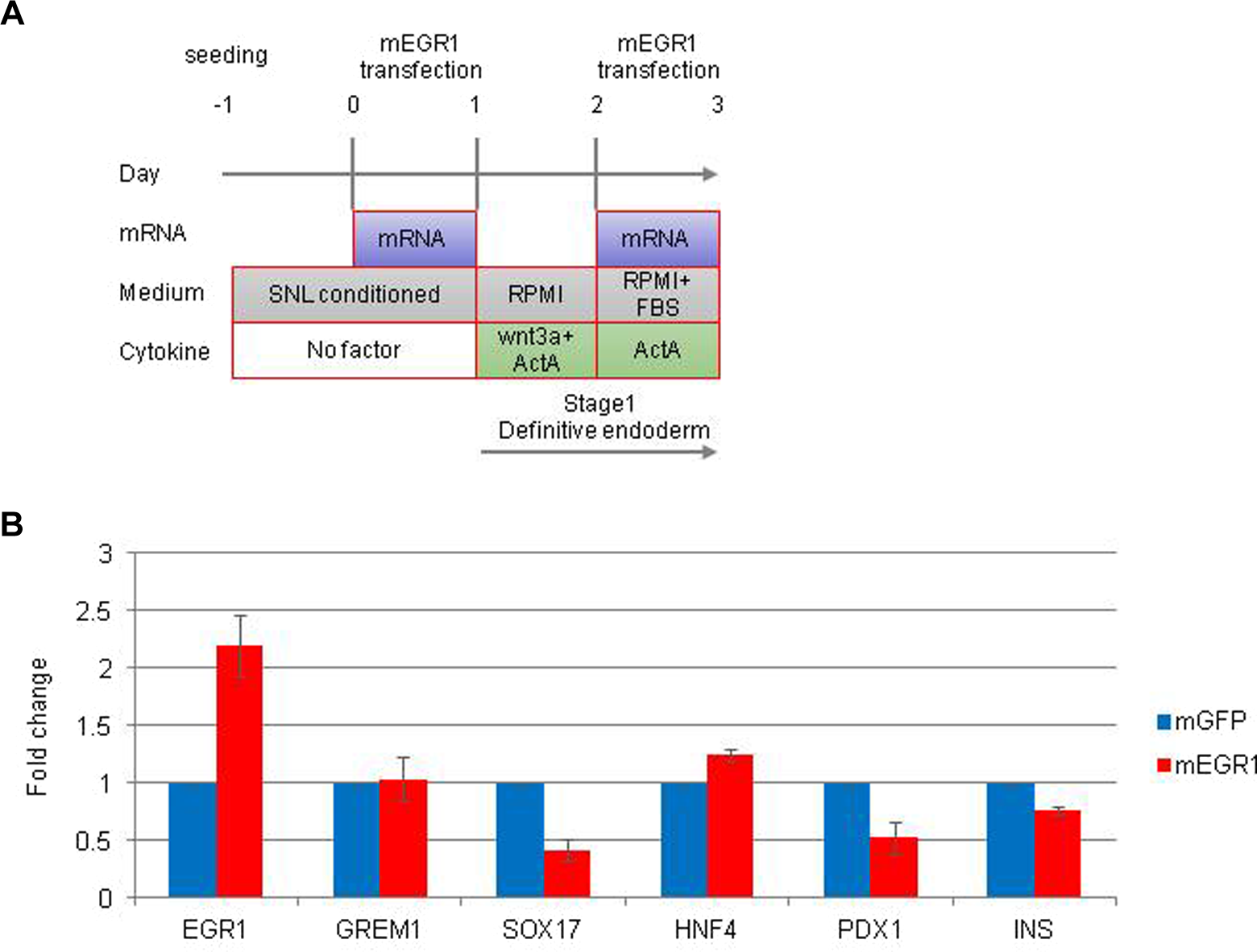
Transfection of messenger RNA of early growth response protein 1 (mEgr1) in human-induced pluripotent stem cells (iPSCs) derived from pancreatic cells (PiPSCs) and treatment with growth factors. (A) Time schedules for the induction of PiPSCs by mEgr1 and growth factors. (B) A quantitative reverse transcription polymerase chain reaction analysis of Egr1 and endoderm- and pancreas-specific genes in PiPSCs treated with mEgr1. The data are expressed as the genes to glycelaldehyde-3-phosphate dehydrogenase ratio, with that of PiPSCs treated with green fluorescent protein (GFP) mRNA arbitrarily set at 1.
Discussion
In this study, we showed that the suppression of Egr1 in the early phase induced the differentiation of iPSCs into pancreatic endoderm and insulin-producing cells. It has been reported that Wnt signaling inhibits Egr1 15 and that Egr1 acts as an upstream regulator of β-catenin signaling 16 . Since Wnt/β-catenin signaling is important for endodermal differentiation, the suppression of Egr1 in the early phase may also be important for differentiation. In contrast, the suppression of Egr1 in the late phase suppressed the differentiation of iPSCs into pancreatic endoderm and insulin-producing cells. Eto et al. reported that Egr1 activates the insulin promoter in part by increasing the expression of Pdx1 and enhancing the Pdx1 binding to insulin promoter sequences 12 . The same group reported that Egr1 regulates the expression of Pdx1 in pancreatic β cells by both direct and indirect activation of the Pdx1 promoter, suggesting that the Egr1 expression may act as a sensor in pancreatic β cells to translate extracellular signals into changes in the Pdx1 expression and pancreatic β cell function 13 . These reports support our data on the suppression of Egr1 in the late phase.
ESCs and iPSCs are capable of differentiation into cells from the 3 embryonic germ layers that constitute the body, making them particularly useful for regenerative medicine research at many institutions 17 –19 . It was recently shown that, following the reprogramming of mouse/human iPSCs, epigenetic memory inherited from parental cells affects the differentiation capacity of iPSC lines 20 –22 . It has also been shown that this epigenetic memory predisposes iPSCs derived from pancreatic β cells to differentiate more readily into insulin-producing cells 14 . These findings demonstrate that the iPSC phenotype is influenced by the cells of origin. We previously reported that PiPSCs differentiated into insulin-producing cells more efficiently than FiPSCs and that the expression of Egr1 in FiPSCs was higher than in PiPSCs 11 . These data suggest that the inhibition of Egr1 may be an important factor for the differentiation of PSCs into insulin-producing cells. We therefore used FiPSCs and PiPSCs in the present study.
In conclusion, the inhibition of Egr1 in the early phase induced the differentiation of PSCs into insulin-producing cells, and the inhibition of Egr1 in the late phase suppressed the differentiation of FiPSCs into pancreatic endoderm and insulin-producing cells. The role of Egr1 may be important for the differentiation of PSCs into insulin-producing cells.
Footnotes
Ethical Approval
This study was approved by our institutional review board.
Statement of Human and Animal Rights
This article does not contain any studies with human or animal subjects.
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article. This work was supported in part by the Japan Society for the Promotion of Science and the Ministry of Health, Labour and Welfare.
