Abstract
Introduction
Hip fractures are among the most common major orthopaedic injuries globally, with one in three women and one in twelve men projected to sustain a hip fracture in their lifetime. Identifying genetic factors that contribute to hip fracture risk could improve risk stratification and inform prevention strategies. This study aims to identify genetic variants associated with hip fracture susceptibility through a genome-wide association study (GWAS).
Materials and Methods
A GWAS was undertaken using the UK Biobank to identify risk loci for hip fractures.
Results
At the time of analysis, 2165 neck of femur fractures were identified among the 502 507 participants. Thirteen SNPs in five putative haplotypes were identified as significantly associated with hip fracture using the stringent GWAS threshold of 5E-8. Two of these loci appear to affect HOXC8, either by influencing the 3’ UTR (rs4142680[T]) or via the miRNA hsa-miR-196a (rs11614913[T]). These two SNPs were also found to be expression quantitative trait loci for homeobox-C cluster genes (HOXC6, HOXC9, and HOXC-AS1).
Conclusions
Polymorphisms affecting homeobox-C cluster genes influence hip fracture risk in the general population. Future research should focus on validating these genetic associations and exploring optimal therapeutic interventions that could mitigate fracture risk in subpopulations carrying these polymorphisms.
Keywords
Introduction
Hip fractures are among the most common major orthopaedic injuries globally, with one in three women and one in twelve men projected to sustain a hip fracture in their lifetime. 1 The number of hip fractures is expected to reach approximately 4.5 million by the year 2050. 2 Although primarily a condition affecting the elderly, hip fractures are life-changing events associated with negative outcomes such as disability, cognitive impairment and increased dependence. The secondary care costs associated with these injuries in the UK alone are estimated at £1.1 billion annually. 3 Readmission significantly contributes to healthcare costs, with 5%-12% of hip fracture patients discharged to post-acute care facilities being readmitted within six weeks. 4 Furthermore, it should be noted that around 50% of patients who lived independently before sustaining a hip fracture never regain their independence.5,6
The significant mortality associated with a neck of femur fracture is not fully captured by headline costs to health systems. Older adults have a 5- to 8-fold increased risk for all-cause mortality during the first 3 months after hip fracture. 7 Despite advances in perioperative geriatric care, one third of adults who have a hip fracture die within 12 months, with an increased risk of death persisting for at least 10 years. 8 Common causes of death include deconditioning, pneumonia, ischaemic heart disease and frailty resulting from the event. 9 Therefore, it is imperative to identify and address risk factors for this significant condition for individuals and society as a whole.
Genome-wide association studies (GWAS) are genetic studies that focus on identifying genetic variations associated with particular traits or diseases across the entire genome. GWAS analyse genetic variations across the entire genome to identify associations between specific genetic variants and a specific outcome. These variants, typically single nucleotide polymorphisms (SNPs), are tested for their frequency and distribution among individuals with the outcome and without. 10 In the past decade, GWAS have investigated a variety of variables, with such work significantly enhancing our understanding of cancer types and anthropometric traits. 11 GWAS also hold potential for advancing our biological understanding of hip fractures. A previous study examining hip geometry identified associations between several genes and hip structural measures, explaining 12% to 22% of heritability at different sites. 12 However, GWAS in this field are still rare, highlighting the need for further research to uncover additional genetic determinants of fracture risk and bone morphology.
Here we independently interrogate one of the largest cohort studies in existence, the UK Biobank to identify and reaffirm SNPs associated with hip fracture risk. The UK Biobank is a large-scale biomedical database and offers a resource for investigating the intricate interplay between genetics, lifestyle factors, and health outcomes. This comprehensive resource presents an exciting opportunity to investigate if relationships exist between genetic polymorphisms and hip fractures.
Methods
UK Biobank
The UK Biobank is a large-scale biomedical database and research resource that collects and shares health data to support scientific research aimed at improving the prevention, diagnosis, and treatment of a wide range of diseases. Established in 2006, it is one of the most comprehensive and ambitious studies of its kind in the world.
The UK Biobank consists of approximately 500 000 UK residents who consented to share their clinical, lifestyle, anthropometric, and genetic data for research purposes. Participants were between the ages of 40 and 69 at the time of enrolment into the UK Biobank cohort study. Initially on enrolment participants underwent face-to-face interviews and sample collection. At the start they underwent a wide range of physical measures, provided information on their lifestyle and medical history, donated blood, urine and saliva samples for future analysis and agreed to have their health followed up through linkage to their health-related records. Over the subsequent 18 years, patient information was sought from inpatient hospital records, primary care data, and centralised clinical registries, including the Cancer and Death registers. Hip fracture occurrence was part of these data (mean age of occurrence is included in the results). The UK Biobank secured informed consent from all individuals, and all study protocols received approval from the National Research Ethics Service Committee. This research will be performed as per an approved UK Biobank application (number 79840) and in compliance with the Declaration of Helsinki. 13
To obtain genetic data, blood samples were taken at the time of participant enrolment, and DNA was subsequently extracted. 14 This DNA underwent genotyping using either the Affymetrix UK BiLEVE Axiom array or the Affymetrix UK Biobank Axiom array. The imputation process utilised a combined reference panel consisting of approximately 90 million biallelic variants derived from the 1000 Genomes Phase 3 and the UK10K haplotype panels.15,16 The imputation was carried out using the software IMPUTE2 resulting in data for 488 295 genotyped participants.14,17
Quality Control
Participants were excluded from the study according to the following criteria: discrepancies between self-reported and genetically determined sex (data-field: 22001 and 31), a genotype missingness rate exceeding 0.05 (data-field: 22005), genetic relatedness (kinship coefficient greater than 0.0442), presence of sex chromosome aneuploidy (data-field: 22019) and anomalies in heterozygosity or missing rates (described in data-field: 22027, with heterozygosity over 0.1903). Only individuals of European descent were included, determined by self-reported ethnicity (data-field: 21000) and the exclusion of non-white backgrounds. This approach has been used successfully elsewhere.11,18
SNP exclusions were based on several criteria: Hardy-Weinberg equilibrium with a P-value smaller than 1E-10, a minor allele frequency (MAF) above 0.001, a missingness level exceeding 0.05, and an imputation score below 0.8. After exclusions were carried out in Python (v3.10), 13 380 287 SNPs remained. To link the SNPs to possible genes, we utilised eQTL datasets curated by the eQTL Catalog 19 and GTEX consortium, 20 focusing on data derived from bone/muscle/adipose/cartilage tissues and associations with P value less that 1E-05.
Genomic Association Analyses
Exploratory data analyses were undertaken in both R (v4.3) and Python (v3.10). SNP based genetic association analysis of the dichotomous variable of hip fracture was carried out across the entire genome using PLINK. Age, sex and genetic principal components (PCs) were used as covariates (1-10 PCs). 21 Genetic principal components were calculated by the curators of UK Biobank and provided as part of the UK Biobank dataset. 14 Hip fractures were identified from hospital records using the following International Classification of Diseases (ICD) codes: 8200, 8202, 8208, S72.0, S72.00, S72.01.
Results
Through mining the UK biobank, 2165 neck of femur fractures were identified in the 502 507 participants. The mean age of hip fracture was 70 years in women and 67 years in men. To narrow down to the most likely biologically relevant SNPs, SNPs were filtered to those who had a MAF >0.001. 13 SNPs were identified as significantly associating with hip fracture using the stringent GWAS threshold of 5E-8 (Figure 1). SNPs Associated with Neck of Femur Fracture in the UK Biobank. Manhattan Plot Displaying Chromosome Loci on the Horizontal Axis and the –log10(p) Values for Independence on the Vertical Axis, where Higher Vertical Points Suggest Stronger Associations. The Dotted Line Marks the Widely Accepted GWAS Threshold of 5E-8.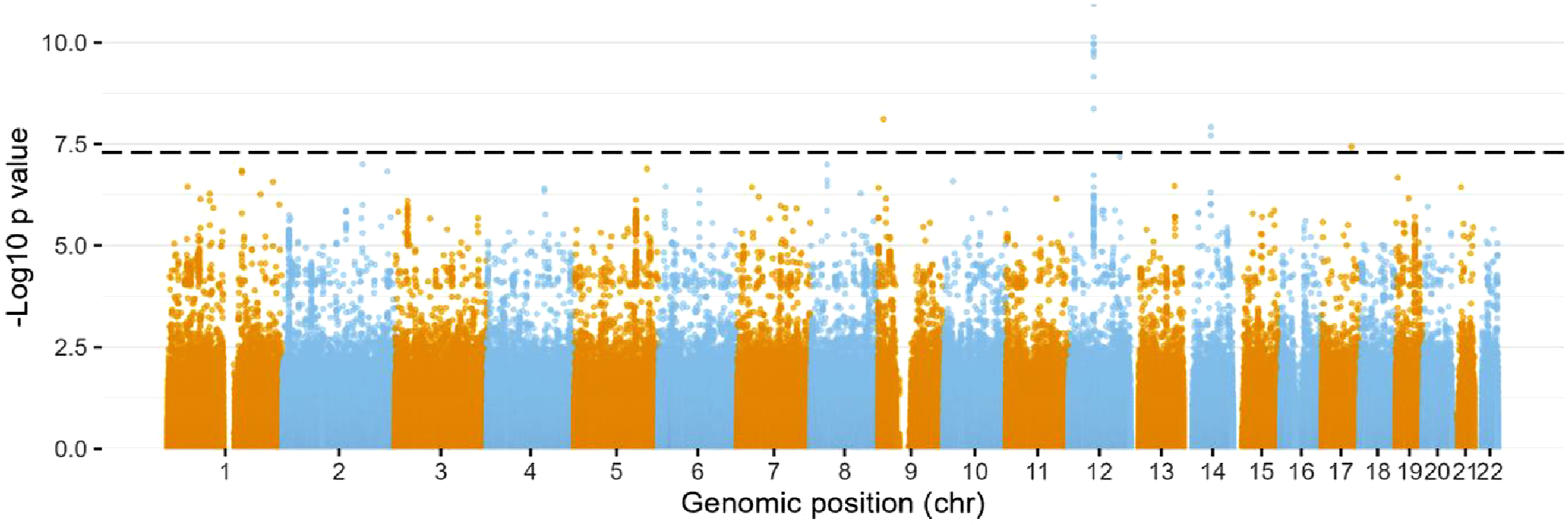
SNPs Associated with Neck of Femur Fracture in the UK BIOBANK (MAF > 0.001) Part (a) Displays SNPs and Recognised Haplotypes as Described in the 1000 Genomes Project, Whereas Part (b) Shows Several SNPs in a Putative Haplotype that is also Associated with Neck of Femur Fracture.

Previous work has shown rs11614913[T] induces a wobble bond in the primary miR-196a-2 hairpin transcript, changing the pairing from U to G instead of the usual C to G, leaving the seed sequences of both the 5p and 3p strands of duplex unaffected. 22 Thus, the variant might affect miR-196a-5p target gene suppression by affecting the thermodynamic stability or processing of the mature miR-196a-5p duplex.23,28,29
In trying to understand further if rs11614913[T] may influence miR-196a-5p, we turned to other fields. Work in the human embryonic kidney cell line (HEK293) showed the rs11614913[T] suppressed genes significantly influenced gene expression, being less effective in suppressing miR-196a-5p target genes. 22 Further, analysis of breast tumours showed samples homozygous for CC of rs11614913 trended towards having a higher fold increase of mir-196a (3.21 ± 1.00 [CC], 1.19 ± 0.24 [TT], P = 0.075). 30 A study of non-small cell lung cancer tumours showed that homozygous mutation of rs11614913 [CC] was associated with a statistically significant increase in mature hsa-mir-196a expression but not with changes in levels of the precursor, suggesting enhanced processing of the pre-miRNA to its mature form. Furthermore, binding assays revealed that the rs11614913 SNP can affect binding of mature mir-196a-3p to its target mRNA. Survival was significantly decreased in individuals who were homozygous. 31 Thus reduction in miR-196a-5p levels is likely driven by rs11614913[T], which has been shown to modulate the expression of homeobox C8 (HOXC8) across different tissues.22,23
A SNP in a different haplotype, rs4142680[T] is found at the same genomic region as HOXC8, which has been described as affecting the 3’ UTR. This SNP has been described in previous work as associated with hip fracture and decreased femoral neck bone mineral density in a study which included 11 516 hip fracture cases and 723 838 controls from the the Trøndelag Health Study (HUNT, Norway), Umeå Fracture and Osteoporosis Study (UFO, Sweden), UK Biobank, Estonian Biobank, and FinnGen biobanks (Finland). 32 This result underlines how genetic polymorphisms affecting HOXC8 seem to influence hip fracture risk in the general population. Both rs11614913[T] and rs4142680[T] were found to be expression quantitative trait loci (eSNPs) for other homeobox-C cluster genes, including homeobox C6 (skin), homeobox C9 (adrenal gland, lung, tibial artery, tibial nerve), and HOXC Cluster Antisense RNA 1 (HOXC-AS1). However, current literature on their involvement in biological processes related to bone development is sparse.
Other loci associated with hip fractures were found on chromosome 9 with lead SNP rs575603778[A] which is an Intron Variant of LOC102724027 (9p21.1). Little is known about this RNA gene and it has previously not been associated with traits in the GWAS catolog. Two SNPs from the other associating locus, rs538670058[T] and rs561165601[C] have no reported gene consequence as does rs143842828[C], found in a separate haplotype. 33
Discussion
Our study highlights SNPs affecting the homeobox-C cluster genes influence hip fracture risk in the general population (HOXC6, HOXC8, HOXC9, and HOXC-AS1). The HOX-C cluster has been shown to influence skeletal development, 32 with HOXC8 appearing to be particularly important.
HOXC8 has been shown to be important in murine skeletal development, with correct temporal expression of HOXC8 being key for determining the correct identity of the vertebral column in early embryos. 34 Aberrant expression of HOXC8 is also associated with homeotic transformation of thoracic vertebrae.35,36 The tissue-specific overexpression of HOXC8 has been shown to inhibit chondrocyte maturation and stimulate chondrocyte proliferation. 36 HOXC8 has been described as a major mechanism of osteoblast differentiation in bone morphogenetic protein (BMP) induced skeletal development, thus anything impeding its function even slightly may influence osteoblastic activity, bone development and fracture risk.37,38
Whilst this study specifically focuses on hip fractures, we acknowledge that SNPs associated with fracture risk may also contribute to fractures in other skeletal regions. Future studies should explore whether individuals carrying these SNPs exhibit an increased overall fracture risk beyond the femoral neck. Additionally, the UK Biobank does not provide data on femoral neck shaft angle, femoral physiological curvature radius, or bone density for all participants. Future research should consider incorporating these structural parameters, as well as gait analysis, to better understand the biomechanical impact of homeobox-C cluster gene variants on fracture susceptibility.
The effect of SNPs on HOXC8 has only been shown to influence fracture risk in European populations to date, which is one of the major limitations of many GWAS cohorts.39-42 This population is narrower, limited to those residing in the UK during recruitment. Further the participants do have a lower mean age of fracture than is typical for the UK (∼77 years), perhaps highlighting a recruitment bias towards more active individuals. Another limitation is that some samples were removed following the quality control covered in methods, therefore perhaps losing valuable cases. Further, it should be noted that this GWAS represents a snapshot in time. This consideration applies to all genome-wide studies where ongoing data collection may influence future findings.
Currently, the detailed characterisation of fractures in individuals with SNPs influencing the HOX-C cluster remains unclear. It is possible that aberrant expression of genes within this cluster may predispose individuals to specific fracture patterns in the hip and other bones. Additionally, whether screening for HOX-C-affecting SNPs in European populations could offer clinical benefit and whether traditional fracture prevention strategies remain effective in SNP carriers require further investigation. While this work provides a foundation for exploring personalised hip fracture prevention, further validation studies and clinical research are necessary to fully assess its implications.
ORCID iD
Louis John Koizia https://orcid.org/0000-0003-4074-6926
Statements and Declarations
Ethical Approval
Approval was received from the UK Biobank (application number 79840). The UK Biobank secured approval from the National Research Ethics Service Committee.
Footnotes
Author Contributions
LJK: Devised the idea for the study, applied for the data, analysed the data, and wrote the article, MDG: Advised on data analysis and aided with writing the manuscript, PZ: Advised on data analysis and aided with writing the manuscript, BHLH: Advised on data analysis, supervised the work, and aided with writing the manuscript, MBF: Supervised the work and aided with writing the manuscript.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
The datasets generated and/or analysed during the current study are not publicly available because the data for this study was made available under an application to the UK Biobank consortium. We are not at liberty to make their data public. Access is controlled and researchers can apply for this data directly from the UK Biobank consortium. Data are available through an application to the UK Biobank (![]() ) and also are available from the corresponding author with blessing from the UK Biobank consortium on request.
) and also are available from the corresponding author with blessing from the UK Biobank consortium on request.
