Abstract
To elucidate the mechanism of the multi-target action of Epimedii Folium on Alzheimer’s disease, this study focuses on the analysis of network pharmacology. Based on a bioinformatics approach, this study obtained the effective components of Epimedium through the Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform, predicted the compound targets through the Pharmapper and Swiss target prediction database and then through Gene Expression Omnibus Datasets and Therapeutic Target Database. We collected and analysed of heral and disease targets, constructed the network. Through the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway, Gene Ontology enrichment, then the key targets and pathways of Epimedii Folium to cope with Alzheimer’s disease have been identified. Twenty-three bioactive components and 477 potential target genes of Epimedii Folium were identified. A total of 1612 target diseases were identified. Through network module analysis, 30 hub target genes were identified. Through enrichment analysis of the KEGG pathway, hub target genes were largely enriched in the PI3K-AKT signaling pathway. Through the analysis of network pharmacology, it was found that Epimedii Folium might play the role of multi-compound and multi-target therapy through the PI3K-AKT signaling pathway. These findings provide helpful directions for future clinical studies.
Introduction
Alzheimer’s disease (AD) is a neurodegenerative disease. Its clinical manifestations include progressive memory loss, cognitive dysfunction, and personality changes. Currently, approximately 50 million people suffer from dementia worldwide, and more than 35% of the population over 80 years of age suffer from this disease. 1 Approximately five million new cases of dementia are reported annually, and the population is expected to increase to 118 million by 2050. 2 The pathological characteristics of AD include neuronal loss, plaque formation in the elderly, neurofibrillary tangles involving amyloid β and tau protein, oxidative stress, and inflammation. 3 AD leads to memory loss, inability in decision making, and ultimately, the inability to take care of themselves. AD impacts patients and their caregivers and relatives, who have to bear the burden of expensive and time-consuming care. Therapies based on multiple actions and targets showed better efficacy than any single-target drug in the treatment of AD.4,5 Therefore, it is essential to study the treatment of AD using multi-target therapy.
Traditional Chinese medicine has been treating AD-related diseases for thousands of years and has accumulated important clinical evidence. Because of its reliable efficacy, it is gradually becoming popular in Western countries. 6 As a crucial part of complementary and alternative medicine systems, traditional Chinese medicine treats numerous diseases through potential multiple drug interactions.7,8
Network pharmacology, which was first proposed in 2007, 9 has become a practical tactic for systematically analyzing the multi-component mechanism of traditional Chinese medicine. Network pharmacology has become one of the most important ways to study the mechanism of traditional Chinese medicine and promote its development. 10 Previous studies have shown that network pharmacology has successfully predicted traditional Chinese medicine’s potential targets and pathways.11-13 Therefore, through the analysis of biological system networks, an effective method of network pharmacology to explore traditional Chinese medication’s potential targets and approach was proposed.
However, studies on the effect of Epimedii Folium on AD are lacking. This study explored the protective effect of Epimedii Folium on AD through network pharmacology and bioinformatics analysis. The workflow of this study is shown in detail in Figure 1. Workflow.
Methods
Collection of Epimedii Folium active compounds
Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform (TCMSP) (http://lsp.nwu.edu.cn/tcmsp.php, version 2.3) provides information on herbal ingredients and their structure and provides parameters related to the absorption, distribution, metabolism, and excretion (ADME) of herbal components, such as oral bioavailability (OB) and drug similarity (DL).14,15 After collecting the TCMSP database, OB and DL were used to screen the bioactive components of Epimedii Folium. OB is the main pharmacokinetic parameter of oral drugs and is used to measure the rate and degree of drug entry into the blood circulation system. As a qualitative principle, DL predicts the possibility of a compound becoming a drug that can be used to help optimize pharmacokinetics and drug properties in drug development. Only compounds with OB greater than or equal to 30% and DL greater than or equal to 0.18 are retained.
Establishment of a database of bioactive compounds and potential targets
All the targets connected to the bioactive compounds of Epimedii Folium were collected from Pharm Mapper (http://lilab-ecust.cn/pharmmapper/, Version 2017) and Swiss Target Prediction (http://www.Swiss.Target.Prediction.ch/, 2019 version) by uploading the structure of bioactive compounds, which were acquired from the PubChem Compound Database (https://www.ncbi.nlm.nih.gov/pccompound) or drawn using Chem3D 16.0. Pharm Mapper and Swiss Target Prediction are web servers for potential drug target prediction by reversing the pharmacophore-matching query compound against an in-house pharmacophore model database. 16
Construction of Alzheimer’s disease–related targets
AD-related genes were screened using Gene Expression Omnibus (GEO) databases (http://www.ncbi.nlm.nih.gov/geo), Therapeutic Target Database (TTD) (http://bidd.nus.edu.sg/group/cjttd/), and Gene Cards (https://www.genecards.org/): (1)GEO databases: we downloaded the gse5281 dataset, which contained brain tissue samples of 87 patients with AD and 74 normal patients, and screened the differentially expressed genes between the AD group and normal control group through limma R-package, the screening criteria were as follows: logFC > 1 and adj.
Common targets mapping
We obtained the targets of Epimedii Folium–related drugs and AD-related diseases through the above process and mapped the two targets. The common targets were those that Epimedii Folium may act on in AD.
Network construction and topology analysis
We constructed the network of active parts and prediction targets of Epimedii Folium first and then constructed the network of “drug components main pathway prediction targets” after the analysis of drug targets with the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway and Gene Ontology (GO) enrichment. After obtaining the common targets, the protein–protein interaction network (PPI) mapping relationship was performed using the STRING database, and the PPI network of the common target was constructed. The key sub-modules of the PPI network were screened using a network topology analysis. According to the target points in the key sub-modules, KEGG enrichment analysis was also carried out, and the “drug composition main pathway key targets” network was also constructed. In this study, network visualization was performed using Cytoscape 3.7.1, and network topology analysis was performed using the Cytoscape MCODE plug-in.
Enrichment analysis of Gene Ontology and Kyoto Encyclopedia of Genes and Genomes
The KEGG pathway and GO enrichment were analyzed using the R-package cluster profiler, and GO included MF, BP, and CC. A
Results
Active components and prediction targets of Epimedii Folium
Information on 23 compounds in Epimedium.

OB, oral bioavailability; D, drug similarity.

Potential targets of Epimedium compounds.
The GO enrichment analysis of the targets.

The enrichment analysis of the targets showed that GO-MF: neurotransmitter receptor activity, protein serine/threonine kinase activity, protein tyrosine kinase activity, etc.; GO-BP: peptide receptor activity, peptidyl-serine modification, peptidyl-serine phosphorylation, etc.; GO-CC: neuronal cell body, an integral component of the presynaptic membrane, an intrinsic component of the presynaptic membrane, etc.; and KEGG: neuroactive ligand-receptor interaction, calcium signaling pathway, EGFR tyrosine kinase inhibitor resistance, cAMP signaling pathway, PI3K-AKT signaling pathway, etc. (Figure 2(b)). In addition, we constructed a network of active ingredients, targets, and KEGG signaling pathways (Figure 2(c)).
Alzheimer’s disease targets
There were 1132 differentially expressed genes (DEGs) between the AD group and the normal group in the GSE5281 dataset,
17
which included 582 upregulated genes and 550 downregulated genes. Heatmaps and volcano maps of the DEGs are shown in Figures 3(a) and (b). In addition, 59 targets were obtained from the TTD database, and the targets were obtained from Gene Cards. We screened the first 500 disease targets according to the score and combined the above disease targets to obtain 1612 disease targets. Screening differential genes using the Gene Expression Omnibus (GEO) Database. 
Common targets
After mapping the disease targets and drug targets, we identified 132 potential AD-related targets of Epimedii Folium constituents (Figure 3(c)).
Protein–protein interaction network construction and topology analysis
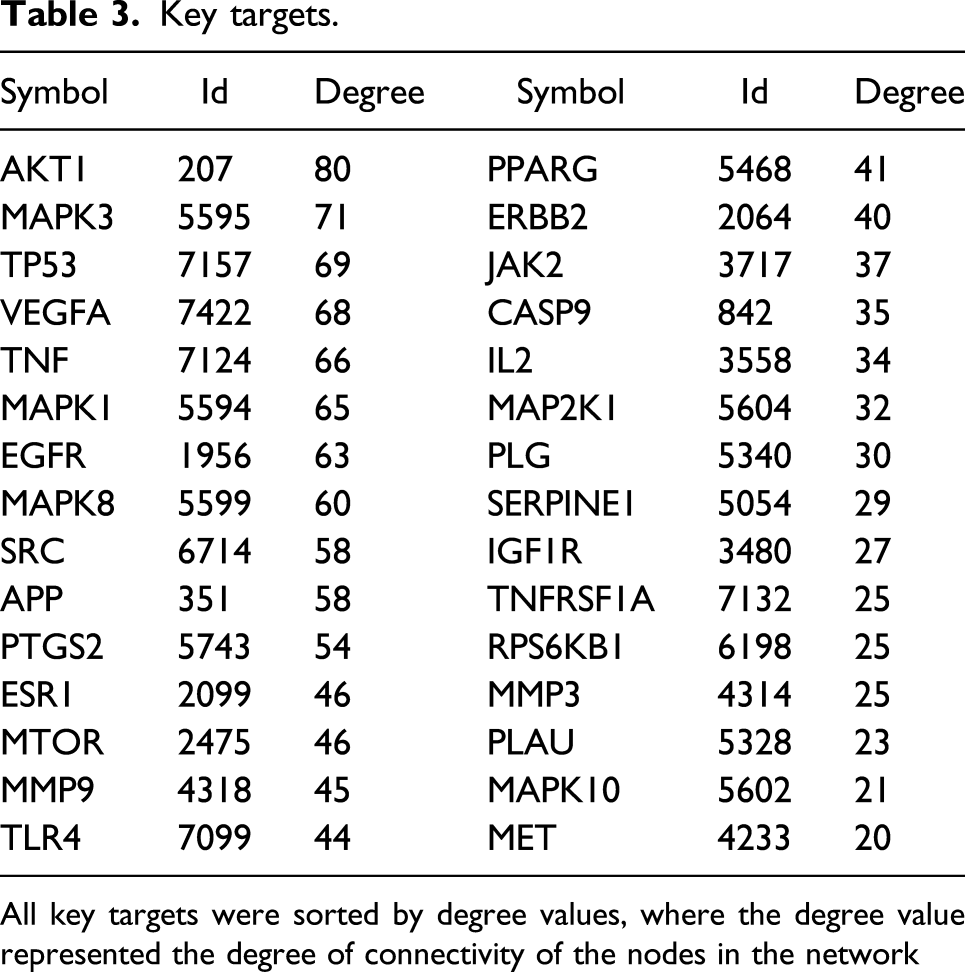
After 132 common targets were imported into the string database, a mapping relationship was obtained. Network visualization was performed using Cytoscape (Figure 4(a)), and the network sub-module was further screened using MCODE. The key network sub-module included 30 nodes, 347 edges, and an MCODE score of 23.9. The key network sub-module contains 30 molecules, which are key molecules, including AKT1, MAPK3, TP53, VEGFA, TNF, MAPK1, and EGFR (Figure 4(b)). The details of the key targets are presented in Table 3. From the most significant terms, the functions of key targets are mainly related to protease activity regulation and oxidative stress response, and the most relevant pathways are the PI3K-AKT signaling pathway, endocrine resistance, and MAPK signaling pathway. Common target and key module networks. Key targets. All key targets were sorted by degree values, where the degree value represented the degree of connectivity of the nodes in the network
GO enrichment analysis of genes in the key network sub-module showed GO-MF: protein serine/threonine/tyrosine kinase activity, phosphatase binding, protein tyrosine kinase activity, etc.; GO-BP: peptidyl-tyrosine phosphorylation, peptidyl-tyrosine modification, response to oxidative stress, etc.; and GO-CC: membrane raft, membrane microdomain, membrane region, etc. (Figure 5(a)). The main enriched KEGG pathways of key molecules included the PI3K-AKT signaling pathway, endocrine resistance, EGFR tyrosine kinase inhibitor resistance, etc. (Figure 5(b)). Moreover, we constructed a network of active ingredients, hub targets, and KEGG signaling pathways (Figure 6). Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway and Gene Ontology (GO) enrichment analysis of key targets. Key targets—pathways—compounds network. The purple node represents the potential target of Epimedium, the green node represents the active compound in Epimedium, and the yellow node represents the enrichment pathways.

All key targets were sorted by degree values, where the degree value represented the degree of connectivity of the nodes in the network.
Critical pathways analysis
The PI3K-AKT signaling pathway was the most significant enrichment of the common targets. The pathway was visualized using the Pathview package, and the nodes involved in the Epimedium target were labeled. There were 16 targets in the pathway: EGFR, TLR4, MAPK1, CASP9, MAP2K1, MET, VEGFA, TP53, ERBB2, RPS6KB1, IGF1R, IL2, AKT1, MAPK3, mTOR, and JAK2. The KEGG pathway map showed that the nodes involved in these targets might be related to the mechanisms of protein synthesis, cell promotion, angiogenesis, DNA repair, and apoptosis (Figure 7). Further analysis revealed that Epimedii Folium interferes with 16 important targets in the PI3K-AKT signaling pathway, and 13 compounds play an active role (Table 4). They are luteolin; quercetin; kaempferol; magnograndiolide; linoleyl acetate; DFV; and chryseriol; 8-Isopentenyl-kaempferol; anhydroicaritin; C-homoerythrinan; 1,6-didehydro-3,15,16-trimethoxy-(3.beta.)-6-hydroxy-11, 12-dimethoxy-2,2-dimethyl-1,8-dioxo-2,3,4,8-tetrahydro-1H-isochromeno [3,4-h]isoquinolin-2-ium; 8-(3-methylbut-2-enyl)-2-phenyl-chromone; and icariin. These 13 compounds can be used as important research objects for future experimental verification. Distribution of key targets in PI3K-AKT signaling pathways. Main components and targets of Epimedii Folium interfering with PI3K-AKT signaling pathways. OB, oral bioavailability; DL, drug similarity.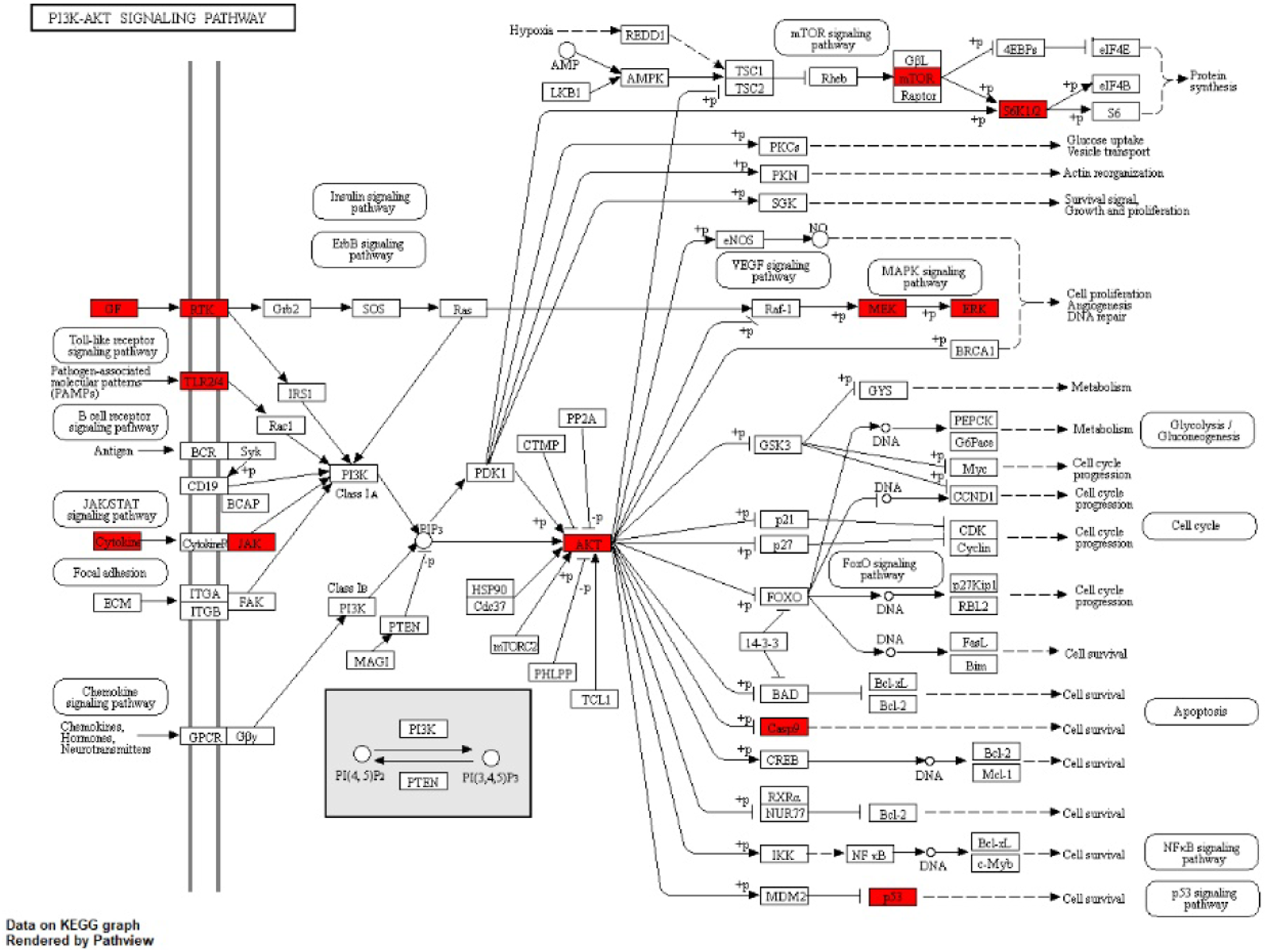

Discussion
As a complicated neurodegenerative disease, AD is characterized by the progressive loss of cognitive function related to neuronal injury and apoptosis, neuritis, lack of neurotransmitters, aβ-plaque deposition, and neurofibrillary tangles. TCM has been empirically used to treat depression and mild cognitive impairment for hundreds of years in China. To clarify the practical effect of Epimedii Folium on AD, we used network pharmacology to study the presumed active components and potential mechanisms.
Twenty-three active compounds and 477 compound-related targets of Epimedii Folium were identified in the public database. A total of 1612 AD-related targets were identified through multiple database searches. Among these targets, 132 were shared between compound-related targets and AD-related targets, suggesting that Epimedii Folium may have an anti-AD effect. Through network construction and central network evaluation, 30 key objectives of the AD target network, potential targets of Epimedii Folium, were selected. They significantly enriched several AD-related pathways, such as the PI3K-AKT signaling pathway, endocrine resistance, EGFR tyrosine kinase inhibitor resistance, HIF-1 signaling pathway, ERBB signaling pathway, TNF signaling pathway, MAPK signaling pathway, prolactin signaling pathway, focal adhesion, and VEGF signaling pathway. This study comprehensively elucidated the presumed active components and multi-target mechanism of Epimedii Folium in treating Alzheimer’s disease, which provides a theoretical basis for the clinical application and further research of Epimedii Folium in the treatment of AD.
Some of the active components of Epimedium obtained in this study have been shown to play a therapeutic role in AD. β-sitosterol has strong anticholinesterase and antioxidant activities, improving memory and learning disorders in APP/PS1 double transgenic AD mice.18,19 Kaempferol is a flavonoid found in foods of plant origin and medicinal plants. It plays a protective role in AD development. By microinjection of kaempferol into the brain’s ventricles, we found that kaempferol may affect memory retention in passive avoidance learning by regulating the cholinergic mechanism. Kaempferol may have a neuroprotective effect on LPS-induced striatum injury. The possible mechanism is related to anti-neuroinflammation, maintenance of blood–brain barrier integrity, and downregulation of the HMGB1/TLR4 pathway.20,21 Icariin is a flavonoid found in Epimedium and has many biological activities. Many studies have shown that icariin and its metabolites may be helpful for AD by reducing the production of extracellular amyloid plaques and intracellular NFTs and inhibiting the activity of phosphodiesterase-5. In addition, there is increasing evidence that icariin also plays a protective role in AD by limiting the potential risk factors of AD, such as inflammation, oxidative stress, and atherosclerosis. Icariin improves cognitive impairment by reducing β-amyloid deposition and inhibiting neuronal apoptosis in SAMP8 mice. Its protective effect on sodium azide neurotoxicity is mediated by the activation of the PI3K-AKT signaling pathway. It affects the axon regeneration, learning, and memory of hippocampal neurons in rats with chronic cerebral hypoperfusion. M1 activation and β-plaque accumulation in microglia in the hippocampus and prefrontal cortex of APP/PS1 mice by upregulating PPARγ.22-26 Exercise pretreatment combined with quercetin injection can improve the memory impairment caused by streptozotocin. Quercetin could improve the survival rate of PC12 cells after a β-injury, promote cell proliferation, resist the toxicity of aβ, and have a neuroprotective effect. Therefore, quercetin is considered a drug for the treatment of AD27,28 Other active chemical components may also have this corresponding intervention effect, which requires further experimental verification.
In this study, the key subnetwork of Epimedii Folium in the treatment of AD was identified, including 30 key targets and 10 KEGG pathways. The PI3K-AKT signaling pathway was the most significant enrichment for the common targets. There were 16 targets: EGFR, TLR4, MAPK1, CASP9, MAPK2K1, MET, VEGFA, TP53, ERBB2, RPS6KB1, IGF1R, IL2, AKT1, MAPK3, mTOR, and JAK2. The KEGG pathway map showed that the nodes involved in these targets might be related to protein synthesis, cell promotion, angiogenesis, DNA repair, apoptosis, and other mechanisms.
The pathology of AD is closely related to oxidative stress, inflammatory responses, and β-deposition. This study revealed that the PI3K-AKT pathway is the most critical pathway for the regulation of AD by Epimedii Folium. Several studies have shown that PI3K-AKT plays a critical role in AD. The PI3K-AKT pathway is crucial for neuronal survival. This can lead to autophagy and decrease the progression of AD. The PI3K-AKT signaling pathway inhibits apoptosis and autophagy, regulates oxidative stress, and plays a neuroprotective role. The PI3K-AKT signaling pathway protects PC12 cells from oxidative stress induced by hydrogen peroxide.29-31 It has been shown that the drug can enhance the understanding and memory of AD mice, inhibit apoptosis, and alleviate the pathological degeneration of the hippocampus by regulating the PI3K-AKT pathway. 32
This study has some limitations. First, compounds screened based on ADME principles may be missing other important compounds. Second, the compounds, targets, and pathways contained in these databases may not be exhaustive. Finally, experimental results need to be further validated using experimental and clinical studies.
Conclusion
In conclusion, Epimedii Folium can regulate PI3K-AKT and other signal transduction pathways through EGFR, TLR4, mapk1, CASP9, MAP2K1, MET, VEGFA, TP53, ERBB2, RPS6KB1, IGF1R, IL2, AKT1, MAPK3, mTOR, and JAK2, which provide a basis for the treatment of AD by Epimedii Folium. The target and pathway of action of Epimedii Folium in the treatment of AD are important for its application. For future anti-AD drugs, new clues are provided by this research.
Footnotes
Author Contributions
Data collection and analysis: ZY and GL; Figures and table designs: YY, YS; Writing—review and editing: ZY and QH; all authors have read and agreed to the final article.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by the General Program of Xuzhou Science and Technology Fund, Xuzhou, China (grant number KC19041).
