Abstract
It is indicated that the dysregulation of long noncoding RNAs (lncRNAs) is implicated in cancer progression. However, the clinical significance of lncRNA small nucleolar RNA host gene 1 (SNHG1) in gastric cancer remains elusive. The expression levels of SNHGs and the association of SNHG1/10/11 with the clinical characteristics in patients with gastric cancer were analyzed by The Cancer Genome Atlas RNA-seq data. A Cox proportional hazard regression model was used to evaluate the association of SNHG1/10/11 expression with the clinical outcomes in patients with gastric cancer. It was demonstrated that SNHG1/10/11 expression levels were dramatically elevated in gastric cancer tissue samples as compared with the adjacent normal tissues. Increased expression of SNHG1 had no correlation with the clinicopathological parameters, but acted as an independent prognostic factor of poor survival (hazard ration (HR) = 0.590, 95% confidence interval (CI) = 0.399–0.872,
Introduction
Due to the widespread popularization of gastroscopy and eradication of
Long noncoding RNAs (lncRNAs) have been shown to participate in the progression of multiple malignancies, of which small nucleolar RNA host gene 1 (SNHG1) as a novel lncRNA is upregulated in a variety of cancers including non–small cell lung cancer (NSCLC), 4 colorectal cancer (CRC), 5 liver cancer, 6 breast cancer, 7 laryngeal cancer, 8 and pancreatic cancer (PC). 9 Elevated expression of SNHG1 is linked to lymph node metastasis, 5 pathological stage, 6 and poor survival,5,8,9 in cancer patient. Moreover, restoration of SNHG1 expression facilitates the tumor growth, migration, and invasion by sponging miR-145-5p in NSCLC, 4 miR-154-5p in CRC, 5 miR-382 in breast cancer, 7 and miR-195/Cyclin D1 axis in PC. 9 Besides, SNHG1 can bind to DNA methyltransferase 1 (DNMT1) to repress p53 activity 6 and activate the notch receptor 1 (Notch1) signaling 8 in liver and laryngeal cancers. These findings may provide a potential therapeutic marker for cancer patients.
LncRNA SNHG1 can also result in the growth and migration of gastric cancer by regulating DNMT1 10 and miR-140. 11 However, the association of SNHG1 expression with the prognosis of patients with gastric cancer remains elusive. Herein, we found that SNHG1 expression was dramatically increased in gastric cancer, and its elevated expression was an independent prognostic factor of poor survival and tumor recurrence in patients with gastric cancer. Knockdown of SNHG1 in vitro inhibited the proliferation and invasion of gastric cancer cells, indicating that SNHG1 might be a potential therapeutic target for gastric cancer.
Materials and methods
TCGA analysis
The clinical information including the overall survival time, survival status, recurrence time, as well as SNHG1/3/5/6/10/11/12 expression levels in gastric cancer is collected from The Cancer Genome Atlas (TCGA) database (https://genome-cancer.ucsc.edu). The clinicopathological parameters of gastric cancer patients are indicated in Supplementary Table S1. Ethical approval for this study was obtained from the Ethics Committee of Shanghai Sixth People’s Hospital (YS-2017-003).
Quantitative reverse transcription polymerase chain reaction analysis
The gastric cancer cell lines (AGS, HGC-27, SGC-7901, MKN-28) and GES-1 were stored in our laboratory. Total RNA was isolated by Trizol reagent (Invitrogen, Carlsbad, CA, USA). The complementary DNA (cDNA) was amplified by using a PrimeScript™ Reverse Transcription Kit (TakaRa, Japan) in an ABI 7500 System (Applied Biosystems; Thermo Fisher Scientific, Waltham, MA, USA). The primer sequences of SNHG1 and GAPDH were synthesized by Shanghai Sangon Biotech (Shanghai, China) and listed in Supplementary Table S2. GAPDH was used as the internal control. All quantitative polymerase chain reaction (qPCR) reactions were performed in duplicate.
SiRNA transfection
Plasmids-mediated siRNA targeting SNHG1 vector (si-SNHG1) were purchased from GenePharma (Shanghai, China) and the negative control (si-NC) was used as the control vector. SGC-7901 cell line was planted in six-well plates 24 h prior to si-SNHG1 transfection with 50%–60% confluence, and then were transfected with Lipofectamine 2000 (Invitrogen) according to the manufacture instructions.
MTT and Transwell assays
MTT and Transwell assays were performed as previously reported. 3
Statistical analysis
Statistical analyses were conducted by SPSS 20.0 (IBM; SPSS, Chicago, IL, USA) and GraphPad Prism 7.0. Student’s t test or chi-square analysis was used to evaluate the statistical significance for the comparisons of two groups. Survival or recurrence curves were drawn by the Kaplan–Meier and log-rank test. Univariate analysis and multivariate models were conducted by a Cox proportional hazards regression model. The value of
Results
The expression of SNHGs in gastric cancer tissues
We assessed the expression levels of lncRNA SNHGs (SNHG1/3/5/6/10/11/12) in gastric cancer tissue samples from TCGA dataset and found that SNHG1/10/11 displayed significantly higher expression levels than SNHG3/5/6//12 in pair-matched gastric cancer tissues (n = 32, Figure 1(a)) as compared with the adjacent normal tissues. The similar results were further validated in 415 cases of gastric cancer tissues (Figure 1(b)).

The expression of SNHGs in gastric cancer: (a) TCGA analysis of the expression levels of SNHGs in pair-matched gastric cancer (n = 32). (b) TCGA validation of the expression levels of SNHGs in 415 cases of gastric cancer.
Upregulation of SNHG1 was associated with poor prognosis in patients with gastric cancer
As shown in Figure 2(a), in the light of the survival time, survival status, and SNHG1 expression levels, we obtained the cutoff value (9.237) of SNHG1 in gastric cancer using a cutoff finder (http://molpath.charite.de/cutoff/load.jsp) and divided the patients into SNHG1 high-expression (n = 169) and low-expression groups (n = 136, Figure 2(b)). The receiver operating characteristic curve (ROC) demonstrated that the sensitivity and specificity of SNHG1 in gastric cancer were 61.2% and 55.8%, indicating SNHG1 as a potential diagnostic marker for gastric cancer.

The association of SNHG1 with the prognosis of gastric cancer patients. (a) The cutoff value, sensitivity, and specificity of SNHG1 were acquired in gastric cancer samples. (b) The gastric cancer patients were divided into SNHG1 high-expression group and SNHG1 low-expression group. (c, d) Kaplan–Meier analysis of the correlation of SNHG1 high or low expression with the overall survival and tumor recurrence in patients with gastric cancer.
As indicated in Supplementary Table S3, we estimated the association of SNHG1 with the clinicopathological parameters and found that SNHG1 high expression possessed no association with clinicopathological factors in patients with gastric cancer. According to the gastric cancer data from TCGA, Kaplan–Meier analysis unveiled that the patients with SNHG1 high expression possessed a poorer survival (
Univariate analysis indicated that age, pathological stage, lymph node metastasis, and SNHG1 high expression were related to the overall survival (Table 1), while gender and SNHG1 high expression had an association with tumor recurrence (Table 2) in patients with gastric cancer. Further multivariate Cox regression analysis revealed that SNHG1 high expression acted as an independent prognostic factor of poor survival and tumor recurrence in patients with gastric cancer (Tables 1 and 2).
Cox regression analysis of SNHG1 expression as a survival predictor in GC.

SNHG: small nucleolar RNA host gene 1; NA: not analyzed; RR: risk ratio; CI: confidence interval; GC: gastric cancer.
Cox regression analysis of SNHG1 expression as a recurrence predictor in GC.

SNHG: small nucleolar RNA host gene 1; NA: not analyzed; RR: risk ratio; CI: confidence interval; GC: gastric cancer.
In addition, we also analyzed the association of SNHG10/11 with the prognosis in patients with gastric cancer and found that the increased expression of SNHG10/11 was incompatible with the favorable survival in gastric cancer (Supplementary Figure S2). Therefore, SNHG1 was selected for further investigations.
Knockdown of SNHG1 inhibited the proliferation and invasion of gastric cancer cells
We detected the expression levels of SNHG1 in gastric cancer cell lines and found that SNHG1 had a relatively higher expression level in SGC-7901 cell line as compared with the other cell lines (Figure 3(a)). The silencing efficiency (56.3%) of si-SNHG1 in SGC-7901 cell line was validated by quantitative reverse transcription polymerase chain reaction (qRT-PCR) analysis (Figure 3(b)). MTT and Transwell assays demonstrated that knockdown of SNHG1 repressed the cell viability (Figure 3(c)) and invasion (Figure 3(d)) of gastric cancer cells as compared with the control group.
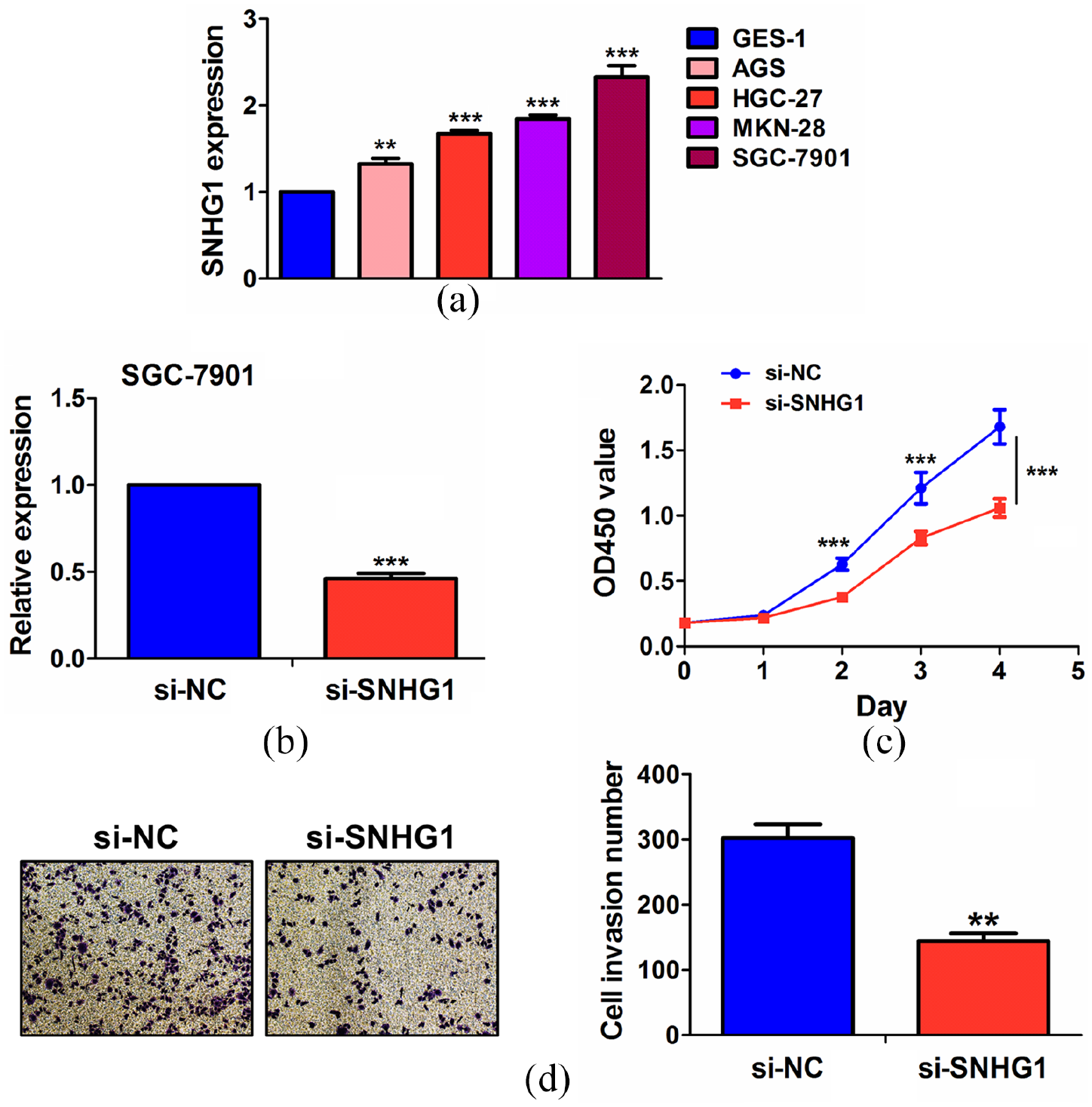
The effects of SNHG1 silencing on the proliferation and invasion of gastric cancer cells: (a) qRT-PCR analysis of the expression levels of SNHG1 in different gastric cancer cell lines. (b) qRT-PCR analysis of the silencing efficiency of si-SNHG1 in gastric cancer SGC-7901 cell line. (c) MTT and (d) Transwell assessment of the effects of SNHG1 silencing on cell viability and invasion.
Discussion
A sea of studies have shown that the dysregulation of lncRNA SNHG1 is implicated in the cell growth and invasion and is related to the poor survival in a variety of cancers.5–9 Herein, we found that SNHG1 expression levels were remarkably increased in gastric cancer tissues as compared with the adjacent normal, but had no association with the clinicopathological parameters. Nevertheless, multifactor analysis revealed that high expression of SNHG1 was an independent prognostic factor of poor survival and tumor recurrence in patients with gastric cancer,
Accumulating evidence indicated that SNHG1 acts as an oncogenic factor in multiple tumors.4,5,7,9 SNHG1 can stimulate the pathogenesis and progression in CRC by downregulating miR-137, 12 and in glioma via miR-154-5p/miR-376b-3p 13 and lead to cisplatin resistance in NSCLC by modulating miR-140-5p/Wnt/β-catenin signaling. 14 In the present study, we also found that knockdown of SNHG1 repressed the proliferation and invasion of gastric cancer, and our results exerted a consistence with those reported by other studies.10,11 These findings suggested that SNHG1 might be a oncogenic factor in gastric cancer.
In addition, some studies showed that lncRNA SNHG10 expression is elevated in liver cancer and favors the tumorigenesis and metastasis by regulating SCARNA13.15,16 Likewise, circulating lncRNA SNHG11 has been considered as a new candidate for early detection and poor prognosis in CRC 17 and prostate cancer. 18 In the present study, we also found that SNHG10/11 possessed higher expression levels in gastric cancer as compared with the adjacent normal tissues, but displayed no relationship with the prognosis in patient with gastric cancer. In addition, more cohorts should be validated for the association of SNHG10/11 with the outcomes of the patients with gastric cancer. In addition, the comprehensive role in vitro and in vivo and regulatory mechanisms of SNHG1 in gastric cancer need to be further explored.
Conclusion
Altogether, our findings demonstrated that increased expression of SNHG1 acted as an independent prognostic factor of poor survival and tumor recurrence in patients with gastric cancer, and knockdown of SNHG1 in vitro inhibited the proliferation and invasion of gastric cancer cells, Our results might provide a potential therapeutic target for gastric cancer.
Supplemental Material
Author-declaration-template-HUMAN-RESEARCH-AND-IN-VITROSTUDIES – Supplemental material for Upregulation of long noncoding RNA SNHG1 indicates a poor prognosis in patients with gastric cancer
Supplemental material, Author-declaration-template-HUMAN-RESEARCH-AND-IN-VITROSTUDIES for Upregulation of long noncoding RNA SNHG1 indicates a poor prognosis in patients with gastric cancer by Hua-Li Zhu and Jing Zou in European Journal of Inflammation
Supplemental Material
Supplementary_data – Supplemental material for Upregulation of long noncoding RNA SNHG1 indicates a poor prognosis in patients with gastric cancer
Supplemental material, Supplementary_data for Upregulation of long noncoding RNA SNHG1 indicates a poor prognosis in patients with gastric cancer by Hua-Li Zhu and Jing Zou in European Journal of Inflammation
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
Ethical approval for this study was obtained from the Ethics Committee of Shanghai Sixth People’s Hospital (YS-2017-003).
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by General Project of Zhejiang Medical and Health Science and Technology Plan (2019KY216).
Informed consent
Informed consent was not sought for the present study because the data were downloaded from the TCGA dataset.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
