Abstract
Introduction: Increasing evidence indicates that lncRNA TUG1 represents an oncogenic factor in cancer. But, the mechanisms by which lncRNA TUG1 contributes to lung adenocarcinoma (LAC) remain undocumented. Methods: The relationship between lncRNA TUG1/miR-138-5p expression and clinical outcomes in patients with LAC was indicated by qPCR, FISH, and TCGA cohort. Gain- or loss-of-function experiments and in vivo tumorigenesis were used to assess the role of lncRNA TUG1 in LAC. The interplay between TUG1 and miR-138-5p was validated by luciferase gene report and RIP assays. qPCR and Western blot analyses were used to investigate the effects of TUG1 on miR-138-5p/HIF1A axis in LAC cells. Results: We found that upregulation of TUG1 or downregulation of miR-138-5p was associated with lymph node or distant metastasis and indicated a poor survival in LAC. Reduced expression of TUG1 restrained the growth of LAC cells, while restored expression of TUG1 had the opposite effects. TUG1 was identified to negatively regulate miR-138-5p expression, and miR-138-5p reversed TUG1-induced cell proliferation by targeting HIF1A. Elevated expression of HIF1A predicted a poor survival in LAC. Conclusion: Our findings demonstrate that lncRNA TUG1 promotes the growth of LAC by regulating miR-138-5p-HIF1A axis.
Introduction
Lung cancer configures as one of the malignant tumors and is the most commonly diagnosed cancer in China. 1 Lung cancer consists of small cell lung cancer (SCLC) and non-small cell lung cancer (NSCLC), and NSCLC histologically contains lung adenocarcinoma (LAC), squamous cell carcinoma, and large cell carcinoma. 2 The latest decade has witnessed the progress in the treatment of LAC, but the ineffective treatment and poor outcomes in some cases are always attributable to the tumor distant metastases. 3 It has been reported that the abnormal expression of lncRNA is related to the development of LAC.4,5 Thus, discovery of lncRNA-related biomarkers may be helpful to the early diagnosis of LAC.
Increasing evidence indicates that taurine-upregulated gene 1 (TUG1) has been shown to participate in the pathogenesis of multiple malignancies. Upregulation of TUG1 is reported in a variety of cancers including prostate cancer (PCa), 6 colorectal cancer (CRC), 7 hepatocellular carcinoma, 8 esophageal squamous cell carcinoma (ESCC), 9 laryngocarcinoma, 10 melanoma, 11 bladder cancer, 12 and osteosarcoma, 13 and indicates a poor prognosis in patients with PCa, ESCC, and melanoma.6,9,11 Ectopic expression of TUG1 facilitates tumor cell growth, metastasis, cytoskeleton rearrangement, and induces cell apoptosis,6,8,10,11,13 and its downregulation favors the radio-sensitivity6,9 and chemotherapy.12,14,15
It has been shown that lncRNA TUG1 can not only activate the multiple signaling pathways such as TGF-β/TWIST1/EMT, 7 JAK2/STAT3, 16 and YAP 17 to accelerate tumor metastasis but also sponge miRNAs to promote cancer progression.8,10–13 For example, TUG1 acts as a tumor-promoting factor by regulating miR-524-5p in oral squamous cell carcinoma, 18 miR-299-3p in renal cell carcinoma, 19 and miR-26a-5p in CRC. 20 In addition, miR-1299 and TUG1 can form the feedback loop to lead to the growth of ovarian cancer. 21
TUG1 is also reported to be downregulated in LAC, 22 but its underlying mechanisms about how TUG1 contributes to LAC remain elusive. We herein found that upregulation of TUG1 or downregulation of miR-138-5p was associated with lymph node metastasis and indicated a worse prognosis in LAC. LncRNA TUG1 contributed to the tumorigenesis of LAC by regulating miR-138-5p-HIF1A axis.
Methods
Tissue samples
The clinical data for LAC as well as the levels of TUG1, miRNAs (miR-17-5p, miR-18a-5p, miR-20a-5p, miR-93-5p, miR-106a-5p, miR-199a-5p, miR-199b-5p, miR-138-5p, miR-106b-5p, miR-18b-5p, and miR-20b-5p), and HIF1A were downloaded from TCGA dataset. 8 paired LAC tissue samples were stored and frozen at −80°C. The tissue microarray (TMA, HLugS030PG02) containing 24 paired LAC tissues was from Shanghai Outdo Biotech. The expression of TUG1 in LAC tissues was measured by qPCR analysis in 8 paired LAC tissues, FISH analysis in 24 paired LAC tissues, and TCGA cohort in 58 paired and 515 unpaired LAC tissues. Our study was approved by the Ethics Committee of Kunming Medical University (YS-2019-005).
Cell transfection
Lentiviruses-mediated short hairpin RNA (shRNA) targeting TUG1 (sh-TUG1, GGCTATCAGAATAACCACTCACCTA) and sh-NC (GGCGACTATAAACCAAC TCCATCTA) was constructed by annealing single-strand hairpin cDNA. When H1975, A549, and SPC-A1 cells reached 60–80% confluence, they were transfected with sh-TUG1 (TUG1) or sh-NC.
Quantitative real-time PCR (qRT-PCR)
To measure the expression of TUG1 in LAC tissues, qRT-PCR was executed. Total RNA was extracted using TRIzol (Invitrogen, Karlsruhe, Germany). Reverse transcription was conducted by PrimeScript RT Reagent Kit and cDNA amplification by using SYBR Premix Ex Taq (TaKaRa, Dalian, China). GAPDH or U6 gene was used as an endogenous control. The primers used are listed in Supplementary Table S1.
Western blot analysis
Western blot analysis was conducted as previously reported. 22 The primary antibody against HIF1A (Cat #PA1-184, Invitrogen, Carlsbad, CA, USA) was diluted (1:1000) according to the instructions and incubated overnight at 4°C.
MTT, colony formation, and transwell assays
MTT, colony formation, and Transwell assays were executed as previously reported. 22
Dual-luciferase reporter assay
H1975 and A549 cells were seeded into 24-well plates. After 24 h incubation, luciferase report vectors embodying WT or Mut TUG1/HIF1A 3’UTR were co-transfected with miR-138-5p mimic into H1975 and A549 cells. After the transfection for 48 h, the luciferase activities of WT or Mut TUG1/HIF1A 3’UTR were examined.
RNA immunoprecipitation (RIP) assay
RIP assay was executed as previously described. 22
In vivo tumorigenesis
BALB/c (nu/nu) nude mice (6 weeks) were from Shanghai Laboratory Animals Center (Shanghai, China). A subcutaneous xenograft tumor model was executed by injecting sh-TUG1 or sh-NC stably transfected 1 × 107 A549 cells, and the mice were divided into sh-TUG1 (n = 5) and sh-NC (n = 5) groups. After 3 weeks, when the mice were sacrificed, the tumor tissue samples were collected and the tumor size and weight were detected. The tumor volume was acquired depending on the formula: length × width2/2. This animal study was approved by the Animal Ethics Committee of Kunming Medical University.
Immunochemistry analysis
Immunochemistry (IHC) analysis Ki-67 index in mouse tumor tissues was executed as previously described. 22
Statistical analysis
Statistical tests were performed by SPSS 20.0 (IBM, SPSS, Chicago, IL, USA) and GraphPad version 5.0 Prism (GraphPad Software, La Jolla, CA, USA). The continuous variables were described as the mean ± standard deviations. The significant differences were evaluated via Student’s t test, chi-square test, or one-way ANOVA for comparisons of two groups. Survival and recurrence curves were assessed by the Kaplan–Meier and log-rank testing. p value < 0.05 was considered statistically significant.
Results
Upregulation of TUG1 was lined to a unfavorable survival in LAC
We measured the expression levels of TUG1 in 8 paired LAC tissue samples by qPCR and found that TUG1 harbored a dramatically increased expression in LAC tissues (p = 0.0031, Figure 1(A)). The consistent results were affirmed respectively in 24 paired LAC by FISH (p = 0.0006, Figures 1(B) and (C)) and TCGA cohort (Figure 1(D)). Elevated TUG1 expression was linked to poor survival in LAC patients. 
The correlation of TUG1 expression with clinicopathological characteristics in LAC patients
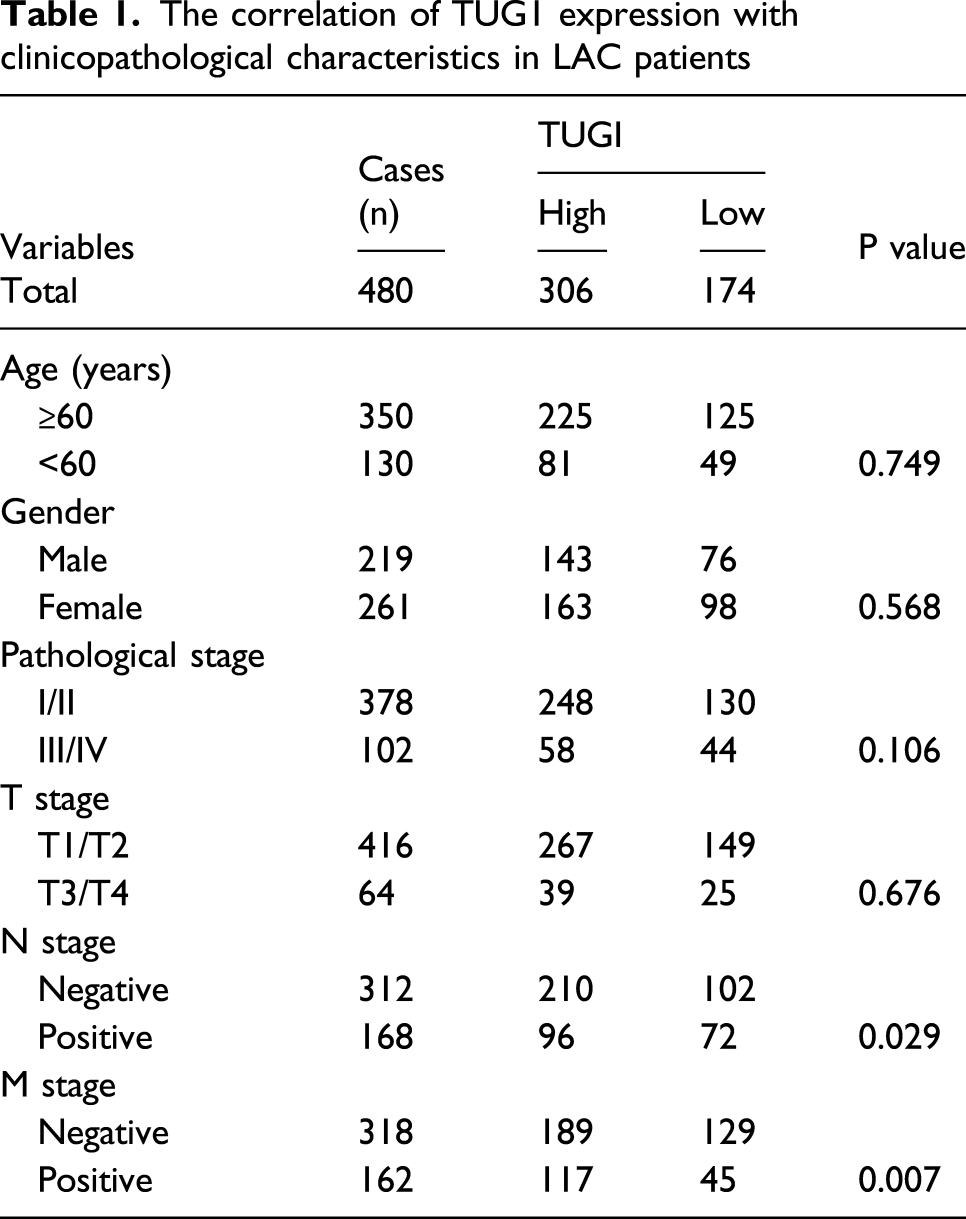
TUG1 accelerated in vitro cell growth and invasion
The levels of TUG1 were tested in various LAC cell lines and TUG1 harbored a higher expression level in H1975 and A549 cell lines but a lowered expression in SPC-A1 cell line when compared with the normal BEAS-2B (Figure 2(A)). Then, the transfection efficiencies of sh-TUG1 lentiviruses in H1975 and A549 cell lines were determined by qPCR analysis (Figure 2(B)). It was shown that silencing of TUG1 reduced cell proliferative viability (Figure 2(C)), colony formation (Figure 2(D)), and cell invasive capability (Figure 2(E)). Restored expression of TUG1 in SPC-A1 (Figure 2(F)) displayed the opposite effects (Figure 2(G) and Supplementary Figure S2). TUG1 accelerated the proliferation and invasion of LAC cells. 
Downregulated miR-138-5p was linked to the poor survival in LAC
We identified 11 miRNAs that have the potential to bind with TUG1 by StarBasev2.0, of which miR-138-5p was found to exhibit an obviously downregulated expression in 39 paired (p < 0.001) and 447 non-matched LAC tissues (p < 0.001) when compared with the other miRNAs in TCGA cohort (Figure 3(A)). Further qPCR analysis also showed such a result (p = 0.0072, Figure 3(B)) and miR-138-5p harbored a negative correlation with TUG1 expression in LAC tissues (Figure 3(C)). According to miR-138-5p expression levels and the prognostic information (n = 400), we acquired a cutoff value (1.274) of miR-138-5p in LAC, and categorized the cases into high-miR-138-5p and low-miR-138-5p expression groups (Figure 3(D), Supplementary Figure S3(A)). As indicated in Supplementary Table S2, low-miR-138-5p expression was linked to lymph node (p = 0.047) in LAC. Kaplan–Meier indicated that the cases with low-miR-138-5p expression harbored a worse survival (p = 0.0215, Figure 3(E)), but had no difference in tumor recurrence (p = 0.8611, Supplementary Figure S3(B)). Multivariate factor analyses unveiled that miR-138-5p was not an independent prognostic factor in LAC (Supplementary Table S3). Low expression of miR-138-5p was associated with poor prognosis in LAC patients. 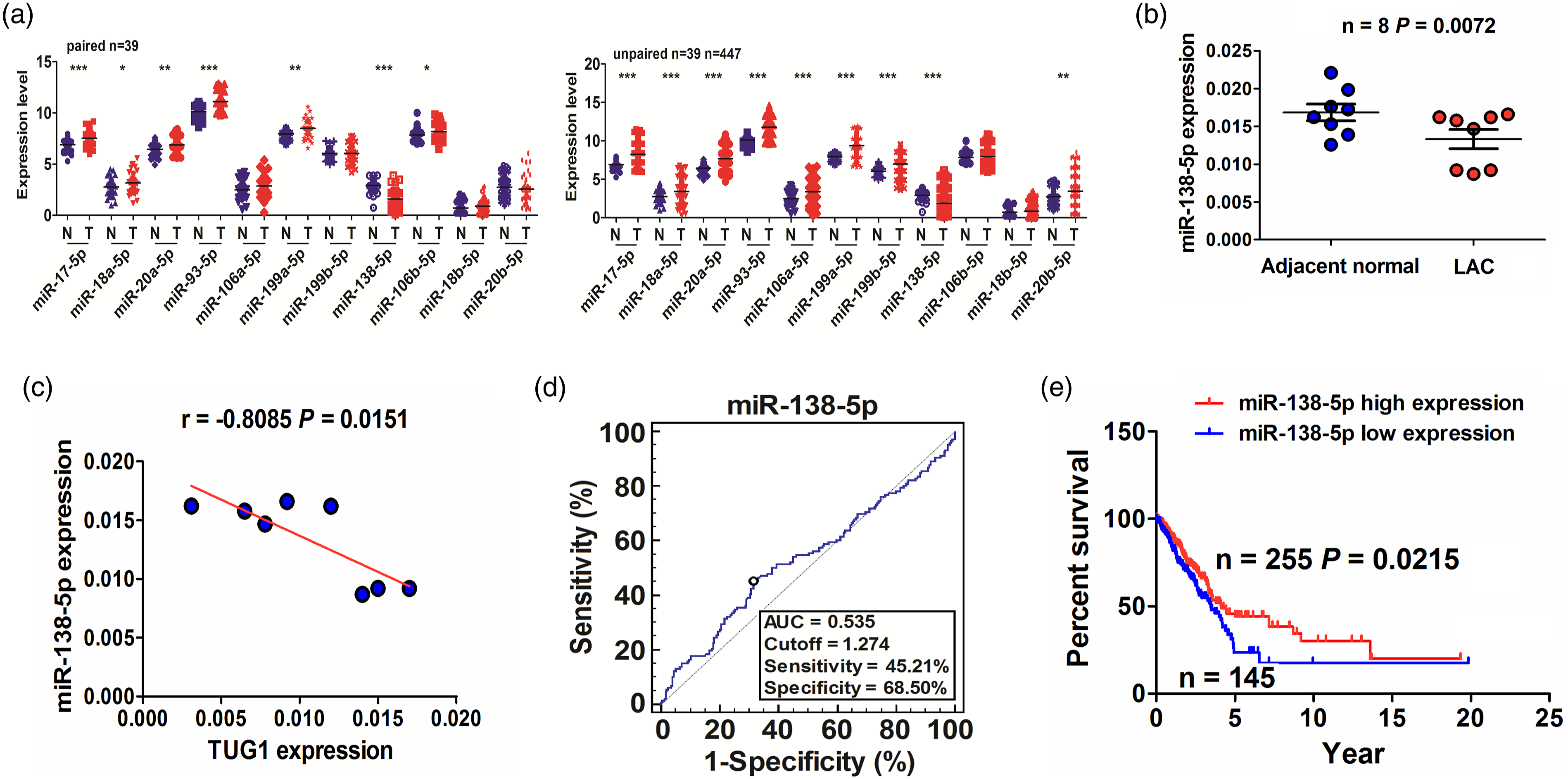
TUG1 could negatively regulate miR-138-5p in LAC cells
The binding between miR-138-5p and WT/Mut TUG1 3’UTR is shown in Figure 4(A). H1975 and A549 cell lines were treated with miR-138-5p mimic and WT (Mut) TUG1 3’UTR for 48 h and miR-138-5p could remarkably reduce the luciferase activity of WT TUG1 3’UTR rather than the Mut TUG1 3’UTR when compared with the control group (Figures 4(B) and (C)). Silencing of TUG1 increased the expression levels of miR-138-5p by qPCR analysis (Figure 4(D)), while miR-138-5p inhibitor exhibited no effect on TUG1 expression in H1975 and A549 cell lines (Supplementary Figure S4). RIP analysis indicated that Ago2 antibody precipitated the Ago2 protein from the cell lysates, and TUG1 and miR-138-5p harbored higher enrichment levels in Ago2 pull-down RNA from H1975 and A549 cell lines when compared with the control IgG pull-down RNA (Figures 4(E) and (F)). In addition, we treated H1975 and A549 cell lines with sh-TUG1 lentiviruses and miR-138-5p inhibitor for 72 h by MTT analysis, and found that miR-138-5p inhibitor prompted cell proliferative viability and attenuated TUG1-induced proliferation promoting effect (Figure 4(G)). TUG1 negatively regulated miR-138-5p in LAC cells. 
Upregulated HIF1A was linked to a poor survival in LAC
HIF1A is identified to bind with miR-138-5p by StarBasev2.0, and HIF1A expression was increased and harbored a negative correlation with miR-138-5p expression in LAC tissues by qPCR analysis (p < 0.05, Figure 5(A)). This result was further validated in 50 paired (p < 0.0001) and 456 non-matched LAC tissues (p < 0.0001) in TCGA cohort (Figures 5(B) and (C)). Kaplan–Meier indicated that the cases with high-HIF1A expression harbored a worse survival (p = 0.0347), but had no difference in tumor recurrence (p = 0.1246, Figure 5(D)). High expression of HIF1A was linked to the poor survival in LAC patients. 
miR-138-5p attenuated TUG1-induced HIF1A expression in LAC cells
The binding between miR-138-5p and WT (Mut) HIF1A 3’UTR is shown in Figure 6(A). We treated H1975 and A549 cell lines with miR-138-5p mimic and WT (Mut) HIF1A 3’UTR for 48 h and found that miR-138-5p significantly reduced the luciferase activity of WT HIF1A 3’UTR rather than the Mut HIF1A 3’UTR when compared with the control group (Figures 6(B) and (C)). In addition, we treated H1975 and A549 cell lines with sh-TUG1 lentiviruses and miR-138-5p inhibitor for 48h by qPCR and Western blot analyses, and found that miR-138-5p inhibitor increased HIF1A expression and attenuated TUG1 silencing-induced reduction in HIF1A expression (Figures 6(D) and (E)). miR-138-5p reversed TUG1-induced HIF1A expression in LAC cells. 
Silencing of TUG1 repressed the in vivo tumorigenesis of LAC
The effects of TUG1 on the in vivo tumorigenesis of LAC were further observed. As indicated in Figure 7(A), we constructed a subcutaneous xenograft tumor model by sh-TUG1 or sh-NC stably transfected A549 cells, and found that the tumor volume in sh-TUG1 group was smaller than that in control group, and the tumor growth was markedly restricted in sh-TUG1 group. After 3 weeks, when the tumor tissues were harvested, we found that the tumor weight, rather than the body weight, was much slighter in sh-TUG1 group than that in sh-NC group (Figures 7(B) and (C)). In addition, IHC analysis showed that the Ki-67 index was dramatically decreased in sh-TUG1 group (Figure 7(D)). We also found that downregulation of TUG1 could upregulate miR-138-5p and reduce HIF1A expression in tumor tissues relative to the control group (Figure 7(E)). TUG1 knockdown suppressed in vivo tumorigenesis of LAC. 
Discussion
Increasing number of studies indicate that lncRNA TUG1 expression is elevated in various cancers and indicates a poor survival in ESCC 9 and melanoma. 11 However, previous studies showed that TUG1 expression level is reduced in LAC 23 and low expression of TUG1 is associated with TNM stage and tumor size. 24 Probably, the downregulation of TUG1 in LAC was reported in a small sample size, 23 and its expression level was measured in a large sample size in TMA and TCGA cohorts which indicated that TUG1 expression was increased in LAC tissues and associated with lymph node and distant metastases and indicated a poor survival in LAC.
Functionally, a great many studies support that TUG1 promotes the tumorigenesis, metastasis and chemotherapy resistance,6–15 and activates the cancer-associated signaling pathways.16,17 On the other hand, it was indicated that TUG1 represses the tumorigenesis in breast cancer 25 and glioma. 26 Herein, it was found that TUG1 knockdown inhibited the growth of LAC cells in vitro and in vivo, but its overexpression displayed the opposite effects, implying that TUG1 might function as an oncogene in LAC.
TUG1 can sponge miRNAs to participate in tumor progression. 27 In the present study, we found that TUG1 could be bound with Ago2/miR-138-5p complex and inhibit miR-138-5p expression, indicating that TUG1 might act as a miR-138-5p sponge to affect LAC cell growth. In addition to TUG1, lncRNA MCM3AP-AS1 and HOTAIR also sponge miR-138-5p to promote cell growth and cisplatin resistance in pancreatic and ovarian cancers.28,29 It has been shown that downregulation of miR-138-5p displays a higher Gleason score and poor prognosis in PCa, 30 and miR-138-5p can inhibit the tumor growth and invasion by targeting FOXC1 in PCa, 30 RHBDD1 in breast cancer, 31 and ZEB2 in LAC. 32 In consistence with the report, 32 miR-138-5p expression was found decreased in LAC tissues and its reduced expression was related to a poor survival in LAC. We found that miR-138-5p attenuated TUG1-caused cell proliferation, but the effects of miR-138-5p on cell growth and invasion in vitro and in vivo need to be further validated in the further work.
Hypoxia-inducible factor 1 (HIF-1) as a transcription factor acts a crucial role in tumor progression. Upregulation of HIF1A indicates a poor prognosis in LAC, 33 promotes proliferation and angiogenesis of lung cancer,34,35 and inhibition of HIF1A restrains the metastasis of LAC. 36 We herein found that HIF1A exhibited a negative correlation with miR-138-5p expression and indicated a poor survival in LAC, and HIF1A was validated as a direct target of miR-138-5p in LAC cells. We also found that miR-138-5p attenuated TUG1-caused HIF1A expression. Previous studies showed that TUG1 can favor cancer progression by regulating multiple signaling pathways including JAK2/STAT3, 16 YAP, 17 MMP14/p38 MAPK/Hsp27 axis, 20 Wnt/β-catenin, 37 and miR-138-5p-SIRT1 axis. 38 We also found that downregulation of TUG1 upregulated miR-138-5p and reduced HIF1A expression in tumor tissues. Our findings also suggested that TUG1 contributed to the tumorigenesis of LAC by regulating miR-138-5p-HIF1A axis.
Our present study had the limitations. In addition to the TCGA cohort, the association of lncRNA TUG1 and miR-138-5p with clinicopathological characteristics and prognosis in patients with LAC should be confirmed in a large sample size by PCR and FISH. The direct interaction between lncRNA TUG1 and miR-138-5p needs to be further clarified. The calculation and justification of the sample size for animal models had not been indicated in our study and more nude mice should be used to verify the role of lncRNA TUG1 in LAC tumorigenesis.
Conclusion
Our findings unveiled that upregulation of TUG1/HIF1A or downregulation of miR-138-5p was linked to lymph node or distant metastasis and indicated a poor survival in LAC. TUG1 could accelerate the tumorigenesis of LAC by regulating miR-138-5p-HIF1A axis.
Supplemental Material
sj-pdf-1-iji-10.1177_20587384211048265 – Supplemental Material for LncRNA TUG1 contributes to the tumorigenesis of lung adenocarcinoma by regulating miR-138-5p-HIF1A axis
Supplemental Material, sj-pdf-1-iji-10.1177_20587384211048265 for LncRNA TUG1 contributes to the tumorigenesis of lung adenocarcinoma by regulating miR-138-5p-HIF1A axis by Ke Li, Huatao Niu, Ying Wang, Ruilei Li, Yuan Zhao, Chao Liu, Honghua Cao, Haitao Chen, Ran Xie and Li Zhuang in International Journal of Immunopathology and Pharmacology
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by 2017 Joint Applied Basic Research Projects from the Yunnan Provincial Science and Technology Department and Kunming Medical University [2017FE467(-070), 2017FE468(-213)], The 2015 Doctoral Scientific Research Fund Project of Yunnan Cancer Hospital (2015Y181), and 2017 Medical Oncology Academic leader training program from the Health and Family Planning Commission of Yunnan province (D-2017001).
Ethics approval
Ethical approval for this study was obtained from Ethics Committee of Kunming Medical University (YS-2019-005).
Animal welfare
The present study followed international, national, and/or institutional guidelines for humane animal treatment and complied with relevant legislation.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
