Abstract
Introduction
Lung cancer is the most common malignant tumor in terms of morbidity and mortality. 1 Non-small-cell lung cancer (NSCLC), including lung squamous cell carcinoma (LUSC), lung adenocarcinoma (LUAD), and large-cell carcinoma, accounts for 80%–85% of lung cancers. 2 LUAD is the most common and aggressive subtype of lung cancer and has the highest heterogeneity and invasiveness. LUAD is diagnosed mostly at an advanced stage, owing to the lack of early typical clinical symptoms and effective diagnostic methods, 3 and does not meet the conditions for surgical resection, which is a major reason for the poor overall prognosis of LUAD. Therefore, identifying novel prognostic predictors and therapeutic targets for LUAD is crucial.
Interleukin-2 (IL2) is one of the earliest cytokines discovered and was initially considered to stimulate lymphocyte proliferation. Subsequently, cytokines were found to be primarily produced by activated T cells. IL2 can act as either an autocrine growth factor for T cells or as a paracrine growth factor for natural killer (NK) cells. IL2-mediated activation of the IL2 Rβγ complex induces proliferative and activating signals via STAT5 phosphorylation, thereby driving the expansion and tumor clearance of effector T and NK cells. 4 Therefore, IL2 is approved by the Food and Drug Administration (FDA) for the treatment of renal cell carcinoma and metastatic melanoma. 5 However, the role of IL2 in LUAD remains unclear.
Interleukin-2 regulates T-cell metabolism and tumor microenvironment (TME) regulation through a variety of signaling pathways by binding to the IL2 receptor. 6 The TME is a complex system, and immune and stromal cells are the two major types of non-tumor factors. They promote the development of diagnostic and prognostic assessment methods for lung cancer. 7 A high density of mature dendritic cells and CD8+ T lymphocytes is related to improved outcomes in NSCLC. 8 Thus, the absence of immune cells and immune escape play important roles in the occurrence and development of tumors. LUAD has a high TMB and strong immunogenicity. Therefore, it is an ideal indication of immunotherapy. 9 Taube et al. found that programmed cell death-ligand 1 (PD-L1) is expressed in cancer cells and immune-infiltrating cells, and its expression reflects an immune-active microenvironment. 10 Clinically, immune checkpoint inhibitors (ICIs) targeting PD-1 and PD-L1 exhibit potent and durable antitumor activity in patients with LUAD. 11 However, the overall response rate to ICIs is relatively low, and only a subset of patients with LUAD benefits from ICI treatment. 12 Although only a minority of patients present desirable responses to this therapeutic intervention, evaluating the mechanisms that modulate the TME during immunotherapy is urgently required. 13 A recent study showed that co-culture with IL2-treated papillary thyroid cancer (PTC) cells significantly increased proliferation and cytokine production but reduced PD-1 expression in activated CD8+ T cells and reduced PD-L1 expression in PTC cells. Accordingly, IL2 increased the expression of HLA class I molecules and the immunogenicity of PTC cells to enhance T-cell immunity, thereby limiting the progression of PTC. 14 Based on existing research, we hypothesized that IL2 may play an important role in carcinogenesis by regulating immune cell infiltration in LUAD.
In this study, we investigated the expression of IL2 in LUAD and found that IL2 expression was associated with a poor prognosis. Additionally, we observed a significant association between the expression of IL2 and the infiltration levels of B cells, CD4+ T cells, CD8+ T cells, macrophages, neutrophils, and dendritic cells in LUAD. IL2 appeared to affect the prognosis of patients with lung cancer partially through immune cell infiltration. These observations emphasize the role of low IL2 expression in carcinogenesis and indicate that IL2 may play an important role in the regulation of immune cell infiltration in LUAD.
Materials and methods
The University of Alabama at Birmingham Cancer data analysis Portal
UALCAN (http://ualcan.path.uab.edu/) was used to investigate IL2 expression and the association between IL2 expression and various clinicopathological parameters (cancer stage, node metastasis, TP53 mutation status, histological subtypes, age, sex, smoking habits, and patient race) of LUAD and for performing survival analysis for IL2 in bladder cancer (BLCA), breast cancer (BRCA), colon cancer (COAD), kidney chromophobe (KICH), LUAD, LUSC, rectal cancer (READ), and thyroid cancer (THCA).
Human protein atlas
The Human Protein Atlas (https://www.proteinatlas.org/) was used to explore IL2 expression in LUAD and for performing survival analysis for IL2 in LUAD.
PrognoScan database analysis
The correlation between IL2 expression and survival in LUAD was analyzed using the PrognoScan database (http://www.abren.net/PrognoScan/). The associations between IL2 expression and patient prognosis, including overall survival (OS), across a large collection of publicly available cancer microarray datasets were investigated using PrognoScan. To select the datasets to be included in this study, the screening parameters were set as follows: “Cancer Type” as lung cancer and “Subtype” as “adenocarcinoma.” Hazard ratios (HRs) with 95% confidence interval were calculated. The threshold was adjusted to a Cox p-value of < .05.
Gene expression profiling interactive analysis
Gene expression profiling interactive analysis (http://gepia.cancer-pku.cn/index.html) is a web portal for gene expression analysis based on TCGA and GTEx data. In this study, relapse-free survival analysis of IL2 was performed using TCGA-LUAD datasets.
Analysis of IL2-interacting genes and proteins
The GeneMANIA database (http://www.genemania.org) was used to construct an IL2 interaction network. The STRING online database (https://string-db.org/) was used to construct a protein–protein interaction network for IL2.
Gene ontology term and Kyoto encyclopedia of genes and genomes pathway enrichment analyses
The DAVID (https://david.ncifcrf.gov/summary.jsp) online analytical tool was used for KEGG Pathway Enrichment and GO analyses, including Biological Process, Cellular Component, and Molecular Function analyses, of genes interacting with IL2. Enrichment analysis results were obtained using online tools for visualization (http://www.bioinformatics.com.cn).
Tumor immune estimation resource
Tumor immune estimation resource (https://cistrome.shinyapps.io/timer/) is a web server used for the comprehensive analysis of tumor-infiltrating immune cells. TIMER was used to analyze the correlation of IL2 expression in pan-cancer and the correlation of IL2 expression with immune cell infiltration and immune checkpoint expression levels in LUAD. TIMER was also used to investigate the relationship between IL2 expression and different gene marker sets of immune cells through the “Correlation” module. The correlations between IL2 expression and immune infiltration were evaluated using purity-correlated partial Spearman’s correlation and statistical significance.
Kaplan–Meier plotter database analysis
Kaplan–Meier Plotter (http://kmplot.com) was used to analyze the prognostic value of IL2 in LUAD. Patient samples were separated into two groups by median expression (high and low expression) to analyze OS and progression-free survival (PFS) in terms of the HRs, 95% CIs, and log-rank p-values. Correlations between IL2 expression and OS in different immune cell subgroups of patients with LUAD were also estimated using the Kaplan–Meier plotter.
Results
Pan-cancer analysis of IL2 expression
Using the TIMER online database, the mRNA expression of IL2 was explored in human pan-cancers. Reduced expression of IL2 was observed in BLCA, BRCA, COAD, KICH, LUAD, LUSC, READ, and THCA tissues compared to that in the corresponding normal tissues; however, high IL2 expression was observed in kidney clear cell carcinoma (Figure 1(a)). Consistently, we found lower mRNA levels of IL2 in LUAD than in normal tissues in the UALCAN database (Figure 1(b)). The protein expression of IL2 was further investigated in LUAD using the HPA database. We found the ratio of IL2 staining intensity was lower in LUAD and LUSC compared to those in normal lung tissues (Figure 1(c)). These findings indicate that IL2 expression is reduced in LUAD and that IL2 may play an important regulatory role in suppressing lung cancer progression. Expression of IL2 in LUAD. (a) The mRNA of IL2 expression in different types of cancer was investigated with the TIMER database. (b) The mRNA of IL2 expression in LUAD was examined by using the UALCAN database. (c) The protein of IL2 expression in LUAD and LUSC was examined by using the HPA database. *p value < 0.05; **p value < 0.01; ***p value < .001; ****p value < .0001.
Prognostic value of IL2 in LUAD
To explore the prognostic value of IL2 in human cancers with differential IL2 expression, survival analysis of IL2 in BLCA, BRCA, COAD, KICH, LUAD, LUSC, READ, and THCA was conducted using the UALCAN database. The results showed that IL2 expression was not statistically significant in determining the prognosis of patients with BLCA, BRCA, COAD, KICH, LUSC, READ, and THCA (Supplement Figure 1A–G); however, patients with LUAD with higher expression of IL2 showed improved prognosis (Supplement Figure 1H). Moreover, patients with LUAD with reduced IL2 expression exhibited poor overall survival (OS) according to the HPA (Figure 2(a)) and Kaplan–Meier plotter databases (Figure 2(c)). The PrognoScan database also demonstrated that a reduced expression of IL2 was significantly associated with poor OS in the Jocab-0018-UM cohort (Figure 2(b)). However, IL2 expression had no statistical significance in predicting the PFS of patients with LUAD (Supplement Figure 2A, B). A reduced IL2 expression was significantly correlated with poor OS in male, female, and white races; both high and low mutation rates; and high neoantigen load in patients with LUAD (Figure 2(d)). These findings indicate that IL2 expression is significantly associated with the prognosis of patients with LUAD. Survival curve evaluating the prognostic value of IL2 for LUAD. (a) Survival curves using the HPA database is shown for OS. (b) Survival curves using the PrognoScan database is shown for OS. (c) Survival curves using the Kaplan–Meier plotter is shown for OS. (d) A forest plot shows the correlation between IL2 expression and clinicopathological parameters in LUAD patients.
IL2 expression and clinical parameters of patients with LUAD
Interleukin 2 expression among groups of patients with different clinical parameters was investigated using the UALCAN database. Based on the tumor stage, a significant decrease in IL2 expression was observed in patients with LUAD at stages 2, 3, and 4 compared with that in the corresponding normal controls (Figure 3(a)). Based on the node metastasis status, IL2 expression was lower in patients with LUAD classified as N0 or N3 (Figure 3(b)). Down-regulation of IL2 was observed in patients with TP53-mutant LUAD compared with that in normal controls (Figure 3(c)). The expression of IL2 was lower in the NOS, mixed, and mucinous carcinoma histological subtypes than in the normal group (Figure 3(d)). IL2 levels were significantly reduced in patients from different age groups (21–40 and 41–60 years) with LUAD (Figure 3(e)). In terms of sex, IL2 expression was significantly lower in LUAD samples from males than in those from females (Figure 3(f)). This finding may be correlated with smoking. IL2 expression was remarkably lower in the smoking group than in the normal controls (Figure 3(g)). Finally, IL2 expression in patients of different races was investigated. Remarkably reduced expression of IL2 was observed in the Asian group with eight patients (Figure 3(h)). These results confirm that IL2 expression is closely associated with LUAD progression and metastasis. Box plots evaluating IL2 expression among different groups of patients based on clinical parameters using the UALCAN database. Analysis is shown for cancer stage (a), nodes metastasis (b), TP53 mutation status (c), histological subtypes (d), age (e), gender (f), smoking habits (g), and patient’s race (h). *p < .05, **p < .01, ***p < .001, ****p value < .0001.
Identification of IL2-interacting genes and proteins and gene ontology and Kyoto encyclopedia of genes and genomes pathway analyses of IL2-interacting genes
Using the GeneMANIA and STRING online databases, we identified 20 genes and proteins that interact with IL2; these included IL2RA, IL2RG, IL2RB, IL21R, FOXK2, IL17A, PTK2B, FOXP1, XRCC5, XRCC6, CREM, HSPG2, CREB1, CSF2, IL3, IL5, BATF3, GAB2, ILF3, and IL5RA (Figure 4(a) and (b)). GO and KEGG enrichment analyses were performed for IL2-interaction genes. The GO analysis showed that IL2 and the associated genes were involved in biological processes related to the immune response, innate immune response, and other biological processes (Figure 4(a) and (c)). KEGG analysis showed that IL2 and the associated genes were involved in signaling pathways associated with the immune response, including Th17 cell differentiation, Th1 and Th2 cell differentiation, cytokine and receptor interaction, and the T-cell receptor signaling pathway (Figure 4(d)). These results strongly imply that IL2 is involved in the regulation of immune response in LUAD. (a) The gene-gene interaction network of IL2 was constructed using GeneMANIA database. (b) The PPI network of IL2 was generated using STRING. (c), (d) GO and KEGG enrichment analysis for IL2.
Correlation between IL2 expression and the infiltrating immune cells
As GO and KEGG analyses revealed that IL2 was involved in regulation of the immune response in LUAD, we analyzed the correlation between IL2 expression and infiltrating immune cells using the TIMER database. The results showed that IL2 expression was significantly and positively correlated with the infiltration of B cells, CD8+ T cells, CD4+ T cells, macrophages, neutrophils, and dendritic cells in LUAD (Figure 5(a)). Correlation of IL2 expression with immune infiltration level. (a) IL2 is significantly associated with tumor purity and is positively correlated with the infiltration of different immune cells using the TIMER database. (b), (c), (d), (e) Spearman correlation of IL2 with expression of PD-1, PD-L1, CTLA-4, and IDO1in LUAD adjusted by purity using TIMER.
Correlation between IL2 expression and various immune markers in LUAD
Correlation analysis between IL2 and biomarkers of immune cells in LUAD determined by TIMER database.
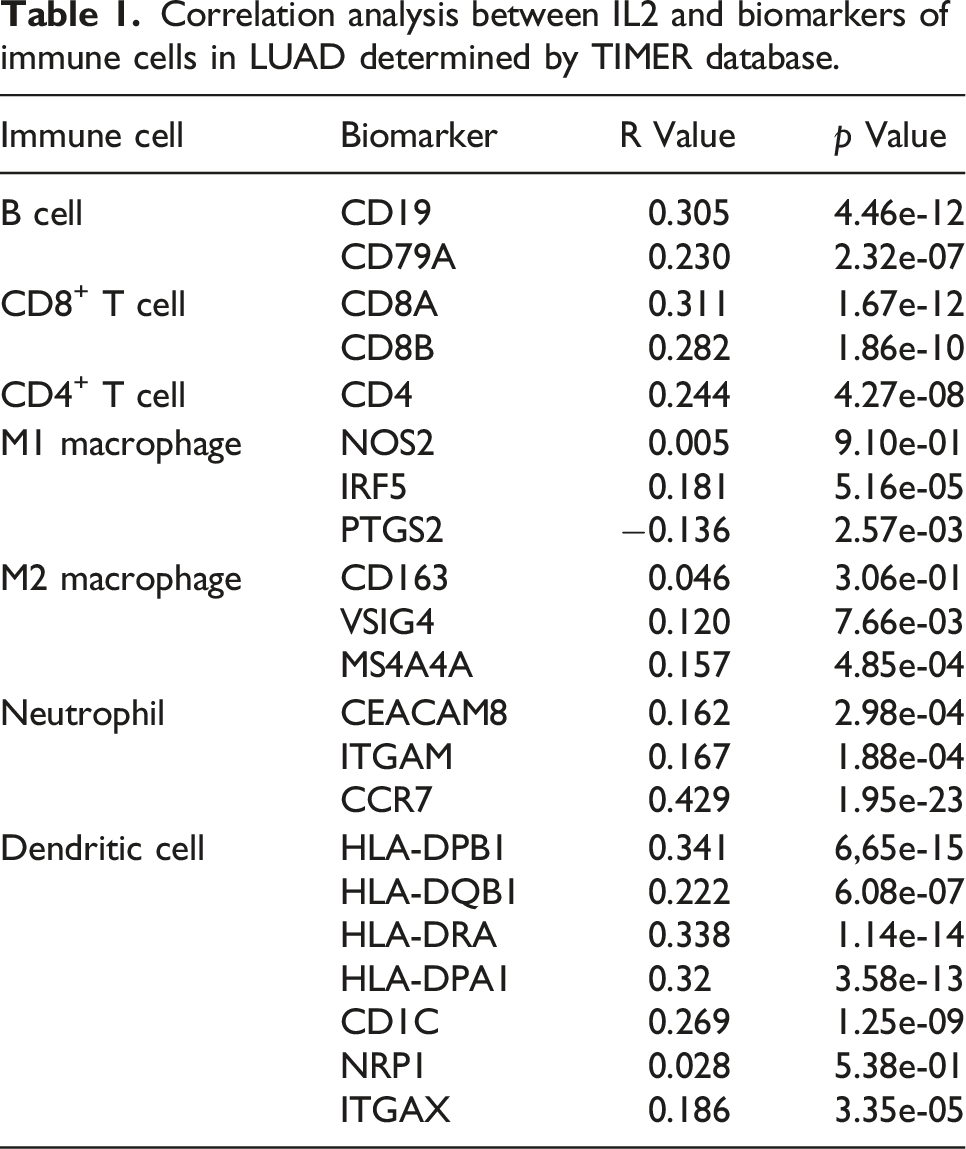
Correlation analysis between IL2 and biomarkers of different types of T cells in LUAD determined by TIMER database.

Prognostic analysis of IL2 expression based on immune cells in patients with LUAD
Finally, we investigated whether IL2 affects the OS of patients with LUAD by influencing the infiltration of different immune cells. We analyzed the expression levels of IL2 in LUSC and associated immune cell subgroups using the Kaplan–Meier plotter database. The results showed that patients with high IL2 expression and enriched infiltration of B cells, CD4+ memory T cells, CD8+ T cells, macrophages, mesenchymal stem cells, and Th2 cells had improved prognoses. However, regardless of whether basophils, natural killer cells, and Th1 cells were enriched or reduced, patients with high IL2 expression had improved prognoses. Patients with high IL2 expression and reduced eosinophil and regulatory T cell counts had improved prognoses (Figure 6(a)–(l)). These results indicate that IL2 may affect the prognosis of patients with LUAD partly owing to immune infiltration. Kaplan–Meier survival curves according to high and low expression of IL2 in immune cell subgroups in LUAD. (a) A forest plot shows the prognostic value of IL2 expression according to different immune cell subgroups in LUAD patients. (b)–(l) Correlations between IL2 expression and OS in different immune cell subgroups in LUAD patients were estimated by Kaplan–Meier plotter.
Discussion
Lung adenocarcinoma is a common type of lung cancer. The prognosis of patients with LUAD is not as good as expected with the current treatment methods. Despite advances in early diagnosis and targeted immune therapies, LUAD is often diagnosed at an advanced stage and is associated with a poor prognosis. 3 Therefore, identifying stable biomarkers to estimate prognosis and guide individualized treatment is important. Growing evidence has shown that the TME plays a key role in tumorigenesis and progression, particularly in those of ICIs that have a significant impact on LUAD. 15 IL2 is a cytokine that stimulates lymphocyte proliferation and plays an important role in the immune microenvironment. 16 In the present study, bioinformatic analysis results obtained from the TIMER, UALCAN, and HPA public databases showed that IL2 expression was lower in patients with LUAD than in the normal control group. A previous study showed that all-trans retinoic acid enhances the cytotoxicity of CIK cells against human lung adenocarcinoma by upregulating MICA and IL2 secretion. 17 These findings indicate that IL2 expression is decreased in LUAD and that IL2 may play an important regulatory role in suppressing lung cancer progression. Furthermore, we found that a low IL2 expression was significantly correlated with tumor stage, node metastasis status, TP53-mutant, histological subtype, age, sex, and smoking in patients with LUAD. Moreover, patients with LUAD with reduced IL2 expression exhibited poor overall survival. These results substantiated that IL2 may be an independent prognostic biomarker in LUAD and that it may suppress the development of targeted precision oncology.
The most commonly mutated genes in LUAD include oncogenes (KRAS and EGFR) and tumor-suppressor genes (TP53, KEAP1, STK11, and NF1).18,19 The JAK–STAT and PI3K–Akt signaling pathways can facilitate the proliferation, migration, and invasion of lung adenocarcinoma.20,21 These findings provide insights into the molecular mechanisms underlying LUAD. To further clarify the role of IL2 in LUAD, we investigated IL2-interacting genes and proteins using the GeneMANIA and STRING online databases. Consistent with the results of previous studies,22,23 KEGG enrichment analysis showed that the IL2-interacting genes were involved in the JAK–STAT and PI3K–Akt signaling pathways. This finding may explain why IL2 is significantly correlated with tumor stage and node metastasis status. Moreover, GO functional enrichment and KEGG enrichment analyses showed that IL2 was mainly involved in the biological processes of innate immunity and immune response and was closely associated with immune response-related pathways in LUAD.
Several studies have shown that IL2 plays an important role in the regulation of immune cells. IL2 is a pleiotropic cytokine with immunostimulatory or immunoinhibitory activity, depending on the target cell, and a well-established regulator of T-cell development and homeostasis. 24 Tregs do not produce IL2 but critically rely on their differentiation from immature single-positive CD4+ T cells in the thymus or from naive CD4+ T cells in the periphery. 25 IL2 regulates the lineage commitment of CD4+ Th cell subsets. Th1 lymphocytes promote cellular immune responses against intracellular microbes and cancer cells. Th1 cell differentiation is triggered by IL2-mediated expression of IL-12Rβ in naive CD4+ T cells. 26 In naive CD8+ T lymphocytes, IL2 mediates the acquisition of the effector cytotoxic phenotype following antigen encounter by promoting the secretion of IFN-γ, TNF-α/β, granzyme B, and perforin. The intensity of IL2/IL2R signaling shifts the fate of CD8+ T cells toward a short-lived effector or long-lived memory phenotype. 27 These results demonstrate that IL2 is closely correlated with immune cell infiltration. Numerous studies have confirmed that tumor immune cell infiltration can influence the efficacy of chemotherapy, radiotherapy, and immunotherapy and the prognosis of cancer patients.28–30 Recent studies have shown that IL2 can exert antitumor effects by influencing immune cell infiltration. 5 Our findings suggest that IL2 is significantly and positively correlated with various immune cells, including B cells, CD8+ T cells, CD4+ T cells, macrophages, neutrophils, and dendritic cells, in LUAD. Moreover, IL2 is markedly positively associated with the biomarkers of these infiltrated immune cells. These findings indicate that tumor immune escape may partially account for the low IL2-mediated oncogenic roles in LUAD. Consistent with this, IL-2 enhanced MHC class I expression in papillary thyroid cancer with Hashimoto’s thyroiditis, thereby overcoming the immune escape. 14
The mechanism of immune escape is complex. IDO1 is a rate-limiting metabolic enzyme that converts tryptophan into downstream catabolites known as kynurenines. Kynurenines can inhibit the activity and proliferation of T cells and other immune cells, such as dendritic cells, leading to successful tumor escape from immune surveillance. 31 IDO1 has been reported to elicit immunosuppressive effects and favor tumor progression in animal models of lung cancer. 32 IDO1 can be regulated by inflammatory molecules through multiple signaling pathways, such as the Janus kinase (JAK)/transducer and activator of transcription (STAT), NF-κB, phosphoinositide 3-kinase (PI3K), IFN regulatory factor 1 (IRF1), and RAS–PKC pathways. 33 Our study that showed IL2-interacting genes were involved in the JAK–STAT and PI3K–Akt signaling pathways. Moreover, IL2 expression was significantly correlated with the expression of IDO1 in LUAD, suggesting that IL2 may be involved in immune escape through the IDO1 signaling pathway.
The efficacy of immunotherapy not only depends on the availability of adequate immune cells to infiltrate the TME but also on the expression of immune checkpoints. 34 Elevated PD-1, PD-L1, and CTLA4 expression of Tregs causing increased IDO1 secretion by DCs has been reported in the literature. IDO1-induced expansion and activation of Tregs, MDSCs, and tolerogenic DCs suppress the activity of antitumor effector T cells, and the induced MDSCs can further inhibit the function of CD8+ T effector cells and NK cells by creating an inflammatory environment and inducing cancer cell migration. 35 Therefore, we speculated whether IL2 can be used as an ICI in the treatment of LUAD. In this study, IL2 expression was positively correlated with PD-1, PD-L1, and CTLA-4 expression. More importantly, IL2 influenced the survival time of patients with LUAD partially through immune cell infiltration, which indicated that IL2 could be a novel immune-related therapeutic target in LUAD. Consistent with this, in a study of PTC, the researchers found that IL-2 can overcome immune escape though decreasing PD-1/PDL1 expression by enhanced MHC class I expression, suggesting that IL-2 may be a valuable ICI for PTC. 14 ICI activates antitumor immune responses and promotes immune-mediated tumor cell clearance mainly by blocking co-inhibitory signaling pathways. However, the clinical benefits of ICI are accompanied with extensive inflammatory toxicity, and associated inflammatory side effects, such as cutaneous inflammatory toxicity, colonitis, and immune pneumonia, have been reported in almost all organ systems. 36 The high levels of ICI cytokines (such as IL-1α, IL-2, IL-3, IL-6, IL-10, IP-10, IL-15, IFN-γ, and IL2-Rα) induced by ICIs are associated with neurotoxicity. 37 In summary, ICI therapy is not a completely safe alternative to chemotherapy.
However, there are some limitations in our study. First, although we investigated the correlation between IL2 and immune escape in LUAD patients, there is a lack of interpretation of the immune analysis according to the different subgroups. Second, the molecular mechanisms and roles of IL2 in tumor growth, metastasis and immune escape need to be explored in further studies. Third, most of the analyses were performed based on mRNA levels of IL2 in the present study. A deeper analysis, based on protein levels, would make the data more convincing. Fourth, the data in this paper come from the public database, and there is no basic experiment for verification. And due to limited funds, clinical data supporting the diagnostic value of IL2 in LUAD are lacking. In future studies, we will focus on the verification of these data.
Conclusion
In summary, we elucidated that IL2 was lowly expressed in LUAD and negatively correlated with an unfavorable prognosis in LUAD. Furthermore, our findings indicate that IL2 may exert its tumor-suppressing role by increasing tumor immune cell infiltration and immune checkpoint expression. Overall, our results indicate that IL2 could serve as a potential novel prognostic biomarker for LUAD. These findings will advance our current understanding of not only the role of IL2 but also its translational use in LUAD prognosis and immunotherapy.
Supplemental Material
Supplemental Material - Down-regulation of interleukin-2 predicts poor prognosis and associated with immune escape in lung adenocarcinoma
Supplemental Material for Down-regulation of interleukin-2 predicts poor prognosis and associated with immune escape in lung adenocarcinoma by Yongwang Hou, Baoli Xiang, Zhicong Yang, Jiangmin Liu, Dandan Xu, Lina Geng, Minghua Zhan, Yuhuan Xu, and Bin Zhang in International Journal of Immunopathology and Pharmacology
Footnotes
Acknowledgments
We thank The First Affiliated Hospital of Hebei North University for their approval of the study.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Hebei Health Commission Scientific Research Foundation Project (20220596).
Supplemental Material
Supplemental material for this article is available online.
Appendix
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
