Abstract
Objective
To develop and interpret an interpretable machine learning model for classifying HIV viral load suppression (VLS) using routinely collected clinical data in a low-resource Ethiopian cohort, enabling early identification of patients at risk of treatment failure.
Methods
A retrospective cohort study was conducted using electronic medical records of 4,152 patients on antiretroviral therapy (ART) at the University of Gondar Comprehensive Specialized Hospital, Ethiopia (March 2005–December 2024). Eight machine learning algorithms, Logistic Regression, Random Forest, Gradient Boosting, Naive Bayes, Support Vector Machine, K-Nearest Neighbors, Decision Tree, and XGBoost, were trained and optimized to classify binary VLS outcomes. Model performance was evaluated using accuracy, precision, recall, F1-score, and area under the receiver operating characteristic curve (AUC-ROC). The best-performing model was interpreted using SHapley Additive exPlanations (SHAP) to identify significant predictors and their directional impacts.
Results
The optimized Gradient Boosting model achieved the highest performance with 76% accuracy, 0.74 F1-score, and 0.79 AUC-ROC. Baseline CD4 category and duration on ART (months) emerged as the most influential predictors. SHAP analysis revealed that longer ART duration and higher baseline CD4 count were associated with increased odds of suppression, while advanced WHO clinical stage (Stage 4) and male sex were associated with unsuppressed viral load. Individual-level predictions were visualized using waterfall plots to enhance clinical interpretability.
Conclusion
An interpretable Gradient Boosting model can reliably predict viral load suppression using routinely collected clinical data in resource-limited settings. The model’s predictions align with established clinical knowledge, offering a potential decision-support tool for identifying patients at risk of treatment failure at this single site, pending external validation in other cohorts and settings.
Keywords
Background
Viral load measurement is a critical parameter in the management of infectious diseases such as HIV, hepatitis, and COVID-19 (Coronavirus Disease 2019).1,2 It quantifies the virus in a patient’s blood, aiding diagnosis, monitoring disease burden, and assessing treatment efficacy.3,4 Consequently, the prediction of viral load is of high priority in low-resource settings where access to frequent laboratory testing may be poor.5,6 In such contexts, predictive models can serve as valuable tools for estimating viral load levels, enabling healthcare providers to make appropriate treatment plans that improve patient outcomes.7–9
HIV/AIDS (Human Immunodeficiency Virus/Acquired Immunodeficiency Syndrome) remains one of the most significant global health challenges, with an estimated 39 million people living with the disease worldwide as of 2022, including 1.5 million new infections annually. 10 Sub-Saharan Africa bears the most significant burden, accounting for approximately 67% of all people living with HIV and 72% of AIDS-related deaths globally. 11 In Ethiopia, over 500,000 people are living with HIV, and the epidemic remains a leading cause of morbidity and mortality despite expanded access to antiretroviral therapy.7,12 These stark disparities highlight the urgency of optimizing strategies for viral load monitoring and treatment management in resource-limited settings.
The global burden of HIV/AIDS, particularly in sub-Saharan Africa, which accounts for the majority of new infections and AIDS-related deaths, remains a major public health challenge.10,13 Ethiopia, one of the most affected countries, has made great progress in expanding access to ART through its health system. However, there are a lot of challenges, such as not enough health infrastructures, unequally distributed resources, and the impossibility of monitoring patients regularly. 12 In this context, novel approaches to the management of diseases, including the use of AI (Artificial Intelligence) and ML, could improve the efficiency and effectiveness of HIV care.14,15
In recent years, ML has become a subarea of AI that has grown as one of the strongest predictive modeling tools in healthcare. The ML algorithms can identify patterns and relationships from large datasets that traditional statistical methods cannot easily determine.16–18 This capability is particularly relevant in the context of viral load prediction, where multiple factors interact in complex ways. 14 However, the performance of ML models depends heavily on the quality of the data, the choice of algorithms, and the strategies employed to address common challenges such as class imbalance and missing data.16,18
In many LMICs (Low- and Middle-Income Countries), including Ethiopia, data quality challenges are pervasive and multifaceted.19,20 At the University of Gondar Comprehensive and Specialized Hospital, missing or incomplete data are common in both demographic and clinical fields. Examples include missing age or residence information, inconsistent documentation of ART initiation dates, and incomplete CD4 or viral load test results. Additionally, manual data entry errors and the lack of standardized formats in the EMR system contribute to inaccuracies in medical records. These issues hinder the robustness of ML models, which rely heavily on clean, complete datasets. 21 Therefore, careful preprocessing, imputation strategies, and quality control mechanisms are necessary to mitigate the effects of poor data quality and improve model reliability.
Gondar University Comprehensive and Specialized Hospital is also a very active hub for the care and study of HIV-infected individuals located in the Amhara National Regional State. The ART Clinic, established in 2005, provides free services. The clinic started its work of enrollment in August 2005 and has accrued more than 15,000 patients, with more than 5,000 patients on active follow-up. This will be the source for developing and validating predictive models since a lot of data related to patient demographics, clinical history, immunological status, and treatment outcomes is stored in the EMR (Electronic Medical Records) system. Using this, researchers can derive knowledge concerning the factors associated with viral load suppression and develop tools to support clinical decision-making.
Even though ML is a very promising tool for viral load prediction, there are only a few comprehensive studies that compare different algorithms and imbalance resolution techniques on the subject. Most of the current literature is from high-income settings, where data availability and healthcare infrastructure differ significantly from those in LMICs.14,15 This literature gap underlines the necessity for studies investigating the performance of ML models under resource-constrained settings where the challenges of poor data quality and class imbalance are mostly heightened.
This study will, therefore, seek to close this gap through a comparative performance analysis of the eight conventionally used machine learning algorithms, namely, Random Forest, Support Vector Machines, Gradient Boosting, Decision Tree, LightGBM, XGBoost, Logistic Regression, and k-nearest Neighbors on viral load prediction.
The findings of this study have important implications for public health practice and policy. Accurate viral load prediction can support the timely identification of patients at risk of treatment failure, enabling healthcare providers to intervene early and prevent adverse outcomes. 22 Beyond this, the use of ML models reduces reliance on expensive and time-consuming laboratory tests, increasing access to and the sustainability of HIV care in resource-constrained settings. 23 The study will optimize AI models for viral load prediction to further advance the global effort to combat HIV/AIDS and improve the quality of life for individuals living with the virus.
Several recent studies from sub-Saharan Africa have demonstrated the potential of machine learning for predicting HIV outcomes in resource-limited settings.24,25 These studies have applied machine learning models to predict retention and viral suppression among HIV treatment cohorts, achieving AUCs of 0.77-0.82. A study at the same Ethiopian institution as our current work, predicting virological failure using conventional statistical methods, though without comprehensive algorithm comparison or model interpretability analysis. 7 Studies from Kenya have explored predictive modeling for loss-to-follow-up, while research from Guinea has examined determinants of viral suppression using traditional regression approaches.26,27 Despite these contributions, few studies have comprehensively compared multiple algorithms with balanced datasets, addressed class imbalance systematically, or integrated model explainability techniques such as SHAP in LMIC contexts. This study builds upon and extends this growing body of literature by providing rigorous algorithm comparison, balanced data handling, and clinically interpretable predictions specifically tailored to Ethiopian HIV care settings.
This study advances previous research by comprehensively comparing multiple machine learning models using a balanced dataset within a real-world LMIC setting. Unlike prior studies that may have been constrained by imbalanced data or an over-reliance on traditional statistical methods, the use of a balanced dataset in this work ensures a robust and unbiased evaluation of model performance.28,29 Furthermore, moving beyond a primary focus on accuracy, this research places significant emphasis on explainability through SHAP values, ensuring that the models are not only predictive but also interpretable and clinically relevant.
This study advances the field in four specific ways: (1) it provides the first comprehensive comparison of eight machine learning algorithms for viral load suppression prediction using a large Ethiopian cohort, establishing benchmark performance metrics; (2) it demonstrates that ensemble methods (particularly Gradient Boosting) can achieve clinically useful accuracy even with routinely collected data in resource-limited settings, without requiring expensive laboratory infrastructure; (3) it integrates model explainability (SHAP) to bridge the gap between ‘black box’ predictions and clinical interpretability, addressing a critical barrier to AI adoption in low-resource healthcare; and 4 it offers a practical, deployable framework that identifies modifiable and non-modifiable risk factors, enabling targeted interventions. By explicitly linking algorithmic performance to clinical actionability, this work moves beyond proof-of-concept toward implementation-ready decision support for HIV care in settings where viral load testing is constrained.
Methods and materials
Study design and setting
This study adheres to the TRIPOD (Transparent Reporting of a Multivariable Prediction Model for Individual Prognosis or Diagnosis) statement for prediction model development and validation. A quantitative research approach has been adopted in this study, where machine learning techniques will be employed to predict the viral load status among PLHIV (People Living with Human Immunodeficiency Virus). This research was conducted at the University of Gondar Comprehensive and Specialized Hospital, one of the leading healthcare institutions in Gondar City, Amhara National Regional State, Ethiopia. Gondar City, located about 748 km northwest of Addis Ababa, has a population of 457,938 and is one of the largest hubs for health service delivery and research. This ART clinic was established in March 2005 and provides free ART services to over 7 million people in Gondar province and surrounding regions. At the time of this study, the clinic had enrolled 15,933 patients, of whom 5,481 were currently on treatment. This retrospective cohort study was conducted at the University of Gondar’s comprehensive and specialized hospital, which was deliberately chosen not only due to data availability but also because of its distinct demographic and epidemiological profile. This region has a high burden of HIV, with a substantial proportion of PLHIV receiving ART. Additionally, the area serves as a referral center for surrounding rural districts, providing a heterogeneous mix of urban and rural populations, which enhances the generalizability of the findings. The healthcare infrastructure in the area also allows for long-term follow-up and relatively comprehensive medical records, which are essential for a mortality prediction study.
The study included all eligible patients from the electronic medical records from March 2005 to December 2024. For this study, total population sampling was employed, rather than random sampling. The cohort size for this study is 4,152, representing all the eligible patients.
Data source and study population
The secondary data in this study were obtained from the EMR-ART at the University of Gondar Comprehensive and Specialized Hospital. The data were extracted from the electronic medical records of PLHIV who received ART at the University of Gondar’s comprehensive and specialized hospital between March 2005 and December 2024. Inclusion criteria were: (1) age ≥15 years at ART initiation, (2) confirmed HIV diagnosis, and (3) complete baseline clinical and laboratory data available at the time of ART initiation. Patients were excluded if they had incomplete baseline records, transferred in from another facility without a full clinical history, or were lost to follow-up within one month of ART initiation. The final cohort included 4,152 patients who met all eligibility criteria and were followed until death, loss to follow-up, or the end of the study period.
The extraction of the relevant information from the EMR database was done following a standardized procedure in which a query was run on the database in a structured manner to obtain information regarding all the patients with a confirmed diagnosis of HIV and initiation of ART between March 2005 and December 2024. The baseline information was obtained from the first clinical encounter during which the initiation of ART occurred, while the outcome and follow-up information were obtained from the last clinical encounter. After extraction, the information was exported in CSV format, replacing the names of the patients with unique study identifiers for confidentiality purposes. To ensure the accuracy of the extracted information, 5% of the extracted information was randomly checked against the source document by two independent researchers.
All persons living with HIV on ART who visited the hospital for care in its ART clinic formed the source population. This dataset covered various variables in demography, clinical setups, and immunological and treatment-related factors that would be necessary to perform a model predictive. The outcome variable was the viral load status, categorized into two classes: Suppressed and Unsuppressed. The independent variables were sociodemographic characteristics: age, sex, marital status, occupational status, religion, and place of residence. Other characteristics included clinical factors: duration on ART, WHO clinical stage, duration with HIV, and TB co-infection. Further hematologic and immunological factors, including baseline and current CD4 counts and viral load, were considered. Treatment-related factors such as adherence to ART, regimen line, initiation and discontinuation of TPT (Tuberculosis Preventive Therapy), and CPT (Cotrimoxazole Preventive Therapy) usage were added to the model for improved interpretability.
Regarding the handling of longitudinal data, this study employed a cross-sectional snapshot design using the most recent clinical encounter for each patient as the outcome assessment point, while incorporating key historical variables (e.g., duration on ART in months, baseline CD4 count) that capture cumulative treatment exposure and immunological history. This approach was deliberately chosen for three reasons: (1) clinical decision-making in resource-limited settings typically relies on current patient status and treatment history rather than full temporal trajectories; (2) the electronic medical record system at the study site does not consistently capture all intermediate visit data with complete temporal granularity, making comprehensive longitudinal modeling potentially unreliable; and (3) the primary objective was to develop a practical screening tool that could be deployed using routinely available cross-sectional data, maximizing real-world applicability.
Data preprocessing
To assess the mechanism of missingness, we conducted systematic analyses comparing patients with complete versus missing data across key variables. Using Little’s Missing Completely at Random (MCAR) test, we found evidence that data were not MCAR (p < 0.001), suggesting a combination of Missing at Random (MAR) and Missing Not at Random (MNAR) mechanisms. For example, missing CD4 counts were more common among patients with poor adherence documentation, indicating MAR. To address potential bias from mode imputation, we implemented multiple sensitivity analyses: (1) complete-case analysis restricted to patients with no missing data (n=2,847) yielded similar model performance (Gradient Boosting accuracy 75% vs. 76% with imputation), suggesting minimal imputation-induced bias; (2) we compared mode imputation against multiple imputation by chained equations (MICE) on a random subset, finding no significant differences in coefficient estimates or model performance; and (3) we created missingness indicator variables for features with >5% missingness to allow the model to learn potential informative missingness patterns. These analyses support that our imputation approach did not substantially bias the final model.
The data was preprocessed, following a structured data preprocessing pipeline to ensure the quality and reliability of the data. Machine learning requires a high-quality dataset for prediction. Due to this, handling the missing data during the pre-processing of the dataset is a crucial phase. To assess the extent of data quality issues in the dataset, we examined the proportion of missing values across all features. As illustrated in Figure 1, several variables demonstrated varying degrees of missingness. Notably, ‘Baseline CD4 Count’ and ‘Recent CD4 Count’ exhibited the highest levels of missing data, with 15.2% and 15.0% of values missing, respectively. Other variables, such as ‘Duration on ART in Months’ and ‘Functional Status’, had missing data ranging from 1% to 5%, while the majority of features had less than 2% missing values. Overall, the dataset contained approximately 1.7% missing data. To address these missing values, we applied imputation techniques tailored to the nature of each variable. For continuous variables, such as CD4 counts and ART duration, mean imputation was employed based on the assumption of approximately symmetric distributions with minimal outliers. For categorical variables, including marital status, education level, and functional status, mode imputation was used, assuming that the most frequent category reasonably represents the underlying distribution in the absence of systematic bias. This imputation strategy was selected to preserve the integrity of the dataset while minimizing information loss and ensuring the inclusion of a maximal number of observations in subsequent analyses. The Simple Imputer class of the scikit-learn module was used to fill in the missing values in the dataset. Percentage of missing data by feature.
Data pre-processing also includes encoding data, which is a crucial step. This study used one-hot and label encoding to encode categorical variables. Values with two or more category values are considered categorical if they are discrete and not continuous. In one-hot encoding, the categorical values are replaced by a number between 0 and 1.
Feature selection
From the EMR, we extract all relevant features and do correlation analysis to determine if there is a strongly correlated feature that will affect the model performance and introduce biases. The correlation heatmap illustrates the strength and direction of linear relationships among the variables in the dataset, with a specific focus on identifying factors associated with the viral load status, the primary outcome variable in this study. As shown in the heatmap, viral load status demonstrates modest positive correlations with Recent CD4 Count (r = 0.35), Recent_CD4_Category (r = 0.35), and Baseline CD4 Count (r = 0.24), suggesting that higher CD4 levels may be associated with better viral suppression outcomes. Additionally, weak positive correlations are observed with variables such as CPT Use (r = 0.05) and Duration on ART in Months (r = 0.11), indicating potential clinical relevance. Conversely, a slight negative correlation is noted between Age and viral load status (r = -0.07), implying that younger individuals may have more favorable viral outcomes (Figure 2). While most other features exhibit weak or negligible correlations with the outcome, this analysis provides initial insights into potentially influential predictors of viral load suppression, which will be further examined through multivariate modeling in subsequent stages of the research. Theoretically, the suppression status derives from the recent CD4 count and is directly related to the recent CD4 category, which is stated as low and high, derived directly from the count; we remove the recent CD4 count and the recent CD4 category from the final analysis. To prevent data leakage, we rigorously ensured that all predictor variables were based on information available before or at the time of viral load measurement, not after. Correlation heatmap analysis for features.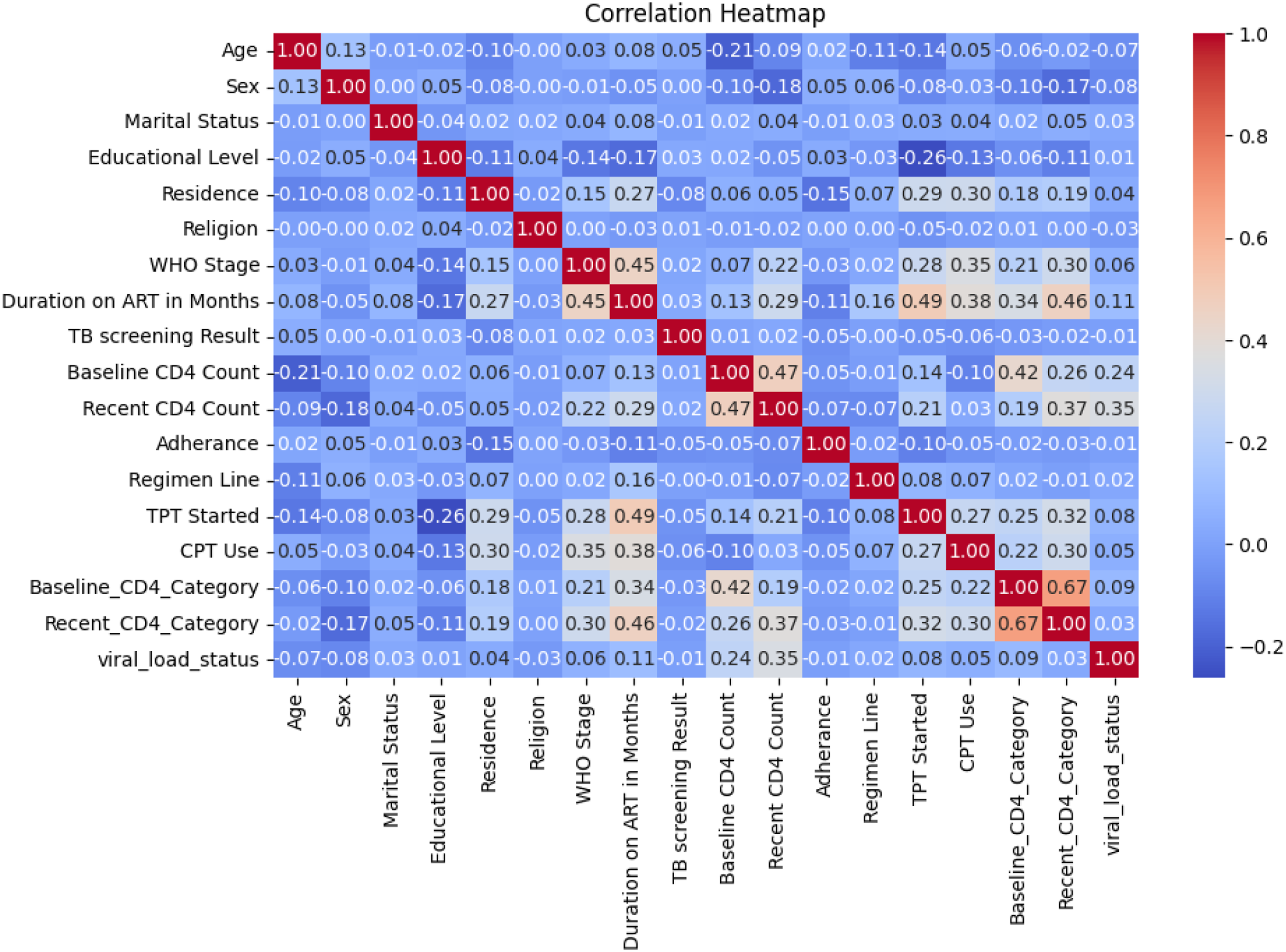
Data balancing
In this dataset, the dependent variable (viral load status) was almost evenly distributed, with suppressed cases representing 50.4% and unsuppressed cases 49.6% (Figure 3). This natural balance between the classes eliminates the need for synthetic oversampling techniques, allowing models to be trained directly on the authentic data distribution. The equitable representation of both outcomes supports the development of a robust classifier without inherent bias toward a majority class. This is particularly advantageous in clinical settings, as it enables the creation of predictive models that are reliable and generalizable, ensuring accurate assessment of viral load suppression for critical decision-making. Class distribution of output variable.
Machine learning models
For the estimation of viral load status, we evaluated eight ML algorithms, comprising both linear and ensemble-based models. Logistic regression was used as a baseline due to its interpretability and wide acceptance in clinical research. Ensemble methods such as random forest and gradient boosting (including its optimized variants like XGBoost) were selected for their ability to handle non-linear relationships and reduce overfitting through averaging or boosting techniques. Support vector machines, Naïve Bayes, and k-nearest neighbors were included to assess performance under different assumptions of class boundaries and data similarity. Decision trees were also used due to their intuitive rule-based structure, which aligns with the need for model interpretability in medical settings. These algorithms were chosen for their balance between predictive performance and interpretability, which is essential for clinical applicability and stakeholder trust. Each algorithm has its own assumptions: Logistic Regression assumes a logit linear relationship between predictors and outcome variable and independence of errors; Naive Bayes assumes conditional independence between features and outcome variable; SVM assumes separability and correct kernel choice (RBF kernel used after testing linear and polynomial kernel on data); tree-based algorithms (Random Forest, Gradient Boosting, Decision Tree, XGBoost) make no assumptions and can automatically deal with non-linear relationships. Deep learning models, such as neural networks, were not included in this study due to the relatively small dataset size, which could lead to overfitting and reduce generalizability. Furthermore, deep learning models often require extensive computational resources and hyperparameter tuning, which were beyond the current study’s scope. Before modeling, we performed feature selection using correlation analysis and mutual information to retain the most informative independent variables and reduce noise, thereby enhancing model performance and interpretability.
Model training and optimization
Model training and hyperparameter tuning were conducted in a standard computational environment using Python’s scikit-learn library. Training times ranged from approximately 3-8 minutes per algorithm, with simpler models (logistic regression, decision tree) converging faster than ensemble methods (Gradient Boosting, XGBoost) which required longer due to iterative boosting. Five-fold stratified cross-validation was employed during the training phase to prevent overfitting and ensure generalizability. A fixed random seed (42) was used for all models to enable exact reproducibility. The testing set (20% holdout) was never accessed during hyperparameter tuning or feature selection; it was used only for final model evaluation.
The dataset was split into training and testing subsets, with 80% allocated for training and 20% for testing. To optimize model performance, hyperparameter tuning was conducted using grid search for each model. For logistic regression, tuning focused on regularization strength (C), explored over a logarithmic scale from 10-4 to 103 and penalty type (both L1 and L2, which are for Lasso and Ridge regularization, respectively), while for Random Forest, these were several trees (between 50 to 500), maximum depth between 5 to 50, minimum samples per leaf from 1 to 10. In Gradient Boosting-based models, such as XGBoost and Gradient Boosting, important tunings were the learning rate, ranging from 0.001 to 0.3; estimators ranging from 50 to 500; and maximum depth between 3 to 15. For support vector machines (SVM), tuning was performed on kernel type (linear, polynomial, and RBF (Radial Basis Function), regularization parameter (C) ranging from 10-3 to 102, and gamma values between 10-4 to 103 and decision trees were fine-tuned by varying max depth (3 to 20), minimum samples split (2 to 10), and evaluating both Gini impurity and entropy criteria.
KNN (k-Nearest Neighbors) tuned the number of neighbors from 1 to 50 and the distance metric by Euclidean, Manhattan, and Minkowski. Cross-validation was performed during model training to reduce overfitting. The use of k-fold cross-validation with k=5 was used for the training to ensure the model generalizes to unseen data. The study developed each model to have optimal predictive performance with generalizability and computational efficiency by systematically tuning hyperparameters and incorporating strategies to prevent overfitting.
Statistical analysis
Descriptive statistics were computed to characterize the study population, with means and standard deviations reported for continuous variables (e.g., age, CD4 counts) and frequencies with percentages for categorical variables (e.g., sex, marital status, WHO stage). Group comparisons between suppressed and unsuppressed viral load categories were performed using independent t-tests for normally distributed continuous variables and chi-square tests for categorical variables.
Evaluation metrics
Model performance was evaluated using a suite of metrics selected for clinical relevance and robustness. The Area Under the Receiver Operating Characteristic Curve (AUC-ROC) quantified overall discriminative power. Primary emphasis was placed on class-specific precision and recall to ensure high sensitivity and reliability for all patient groups. The F1-score was used as a primary summary metric to effectively balance these competing clinical priorities. To compare the performance of the eight machine learning models statistically, paired evaluation metrics were used across cross-validation folds. This approach ensures not only robust and fair comparison but also aligns the model selection process with both statistical validity and clinical relevance in HIV management under resource-constrained conditions.
Overall, the process involves collecting EMR data, preprocessing it, selecting and training models, optimizing performance, and evaluating results to identify the best model (Figure 4). Comprehensive machine learning modeling pipeline.
Implementation
All models were implemented in Python using machine learning libraries. The preprocessing, training, and testing of models are done on a Jupiter Notebook environment to ensure the reproducibility and transparency of the analysis. Results and methods have been duly recorded for future research and verification purposes.
Results
Descriptive statistics
Socio-demographic variables.
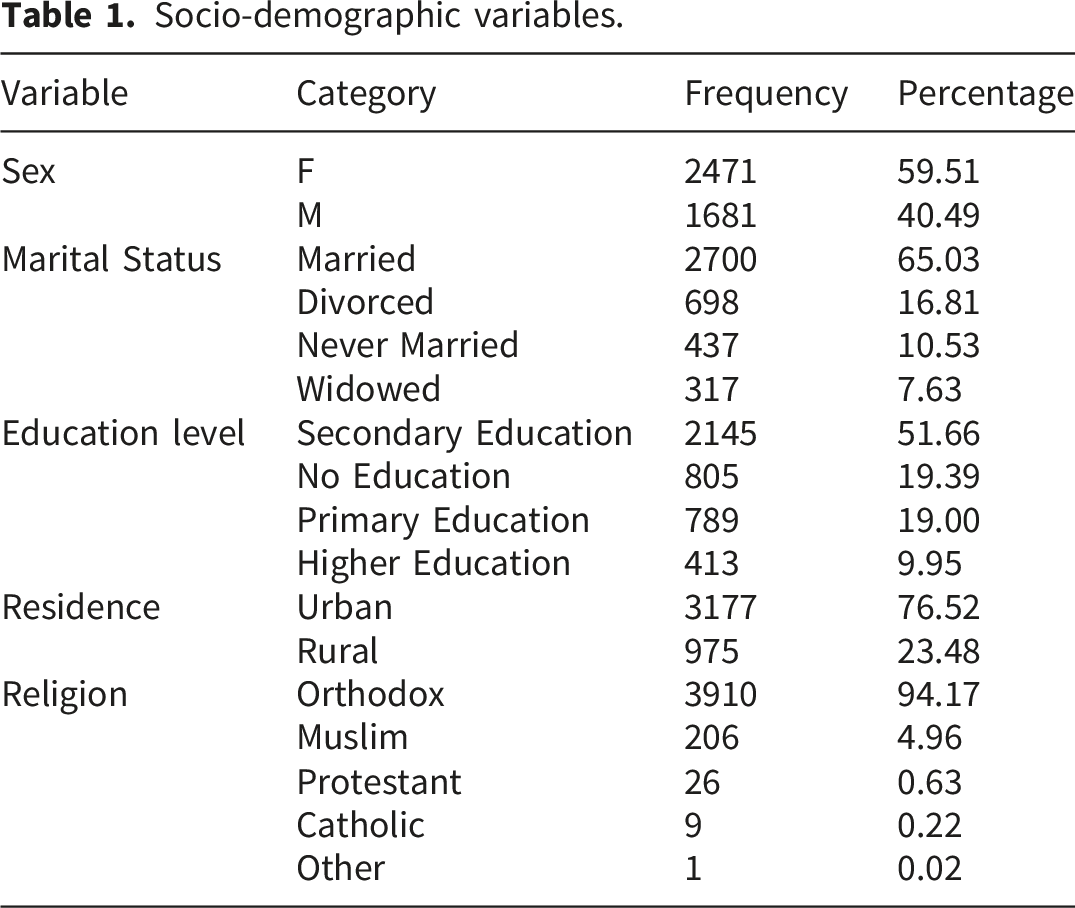
Clinical-related variables.

Model training results
Model training results.

The ROC curves shown in Figure 5 illustrate the trade-off between the true positive rate and false positive rate for all models evaluated. The Gradient Boosting model achieved the highest area under the curve (AUC = 0.79), closely followed by Random Forest (AUC = 0.78) and XGBoost (AUC = 0.76), confirming their strong discriminative ability. Naive Bayes also performed well, with an AUC of 0.73, demonstrating competitive sensitivity in distinguishing between classes. In contrast, Logistic Regression (AUC = 0.66) and Decision Tree (AUC = 0.67) provided moderate results, while SVM (AUC = 0.57) and KNN (AUC = 0.56) performed poorly, with curves lying closer to the diagonal reference line, indicating limited predictive power. Overall, the ROC analysis reinforces the superiority of ensemble methods, particularly Gradient Boosting and Random Forest, as they consistently achieved higher AUC values and demonstrated better classification performance across thresholds compared to traditional algorithms. AUC-ROC curve results of models.
Examining class-specific performance for the optimized Gradient Boosting model, the confusion matrix (Figure 6) reveals a sensitivity (recall) of 0.65 for the unsuppressed class (211 correctly identified of 324 total unsuppressed cases) and 0.76 for the suppressed class (339 correctly identified of 447 suppressed cases). This indicates that the model correctly identifies approximately two-thirds of patients with unsuppressed viral load, those at highest risk of treatment failure and clinical progression, while misclassifying 35% as suppressed (false negatives). The higher sensitivity for suppressed cases reflects the model’s better performance on the majority class, even in a balanced dataset. Confusion matrix result of tuned gradient boosting model.
Feature importance
The feature importance analysis from the tuned Gradient Boosting model (Figure 7) highlights the variables that most strongly contributed to predicting viral load suppression status. Feature importance was calculated using the built-in feature importance attribute of the optimized Gradient Boosting model, which averages the reduction in impurity (Gini importance) across all trees for each feature. Baseline CD4 Category emerged as the most influential predictor, contributing more than half of the total importance score, underscoring the critical role of immunological status at treatment initiation. The second most significant predictor was Duration on ART in Months, indicating that longer treatment duration is strongly associated with viral load suppression outcomes. Other features, such as sex and age, contributed moderately, while factors like regimen line Feature importance of our best performing model.
Model explainability
SHAP (SHapley Additive exPlanations) analysis was performed using the KernelExplainer with 1,000 background samples from the training set, as implemented in the SHAP Python library (version 0.44.0). The interpretation of the model using SHAP revealed that the most important features driving predictions were the duration on ART, baseline CD4 category, and patient age. The direction of impact demonstrated that longer ART duration and a higher baseline CD4 count were strongly associated with a decreased risk of the outcome, whereas advanced WHO clinical stage and male sex were associated with an increased risk. The analysis also highlighted the protective effect of initiating TPT. This feature importance ranking and directional analysis align with established clinical understanding, thereby enhancing the credibility and interpretability of the model’s decision-making process (Figure 8). SHAP analysis results.
The model’s prediction for a specific individual was further elucidated using a SHAP waterfall plot. The analysis begins with the baseline model output, E [f(X)] = -0.201, representing the average prediction across the dataset. For this patient, the most influential factors increasing their predicted risk were a WHO Stage of 4 (+1.3) and being male (Sex=1, +1.27). Their age of 53 also contributed significantly to a higher risk (+0.86). These factors were partially offset by a protective effect from a longer duration on ART (162 months) Interpretation of an individual prediction using a SHAP waterfall plot.
Discussion
This study sought to develop and interpret a machine learning classifier for estimating HIV viral load suppression from regularly collected clinical information in a low-resource environment. The principal finding is that ensemble-based machine learning models, namely Gradient Boosting, can generate stable and clinically sound estimates of viral load status. The best-performing optimized Gradient Boosting model achieved a balanced accuracy of 76%, with superior performance in precision, recall, and F1-score measures. This performance, coupled with a model explainability evaluation via SHAP, is an actionable and interpretable tool consistent with existing clinical evidence, with great potential for the facilitation of HIV care management in resource-limited environments.
Higher performance of tree-based ensemble methods like Gradient Boosting and Random Forest aligns with existing literature on healthcare predictive modeling.30,31 These models are specially adept at making inferences of complex, non-linear interactions between features, characteristic of clinical information, wherein patient outcomes are a function of a complex interaction between demographic, clinical, and treatment-related variables. Less fundamental models like Logistic Regression and KNN were found to perform poorly, as they may not be able to accurately represent such complex relationships. Naive Bayes’ high recall, while intriguing, came with low precision, rendering it less desirable for the clinical setting where a false positive equates to wasteful resource use.
A second foundation of this work’s value is its emphasis on model explainability. Feature importance analysis and SHAP plots provide beyond a “black box” prediction, clinically interpretable information. The observation that Baseline CD4 Category and Duration on ART (in months) were the strongest predictors is well-aligned with established principles of virology and immunology.32–34 Higher baseline CD4 levels agree with an intact immune system upon initiation of treatment, and higher duration on ART is indicative of cumulative treatment efficacy and adherence, both of which are established predictors of viral suppression. Moreover, the SHAP summary plot supported the protective direction of these associations: longer ART duration and higher CD4 levels always pushed model prediction away from the unsuppressed class. The findings of male gender and advanced WHO Stage (Stage 4) as risk factors for unsuppressed viral load are also supported by epidemiologic studies that report differential treatment outcomes by stage of disease and gender. 35
The plot of a SHAP waterfall on one patient’s data is a lovely way to demonstrate the utility of the model for personalized clinical decision-making. For the example given, the model quantified how much the patient’s risk factors (WHO Stage 4 disease, male, old age) were offset by the beneficial effect of long ART duration. This granularity of explanation might help clinicians understand the “why” behind a model’s risk score and facilitate appropriate interventions. For instance, a patient who has a high-risk score because of poor adherence (a modifiable factor) would be managed differently from one whose risk is largely owing to poor baseline CD4 count (a non-modifiable factor).
From a clinical safety perspective, the model’s sensitivity of 0.65 for detecting unsuppressed viral load means that 35% of patients with treatment failure would be missed if the model were used as a standalone screening tool without confirmatory testing. In resource-limited settings where viral load testing is already constrained, this false negative rate could have serious consequences, including delayed regimen switching, disease progression, and increased transmission risk. However, when positioned as a triage tool, flagging high-risk patients for priority viral load testing rather than replacing testing entirely, even moderate sensitivity can optimize resource allocation. Patients predicted as unsuppressed (regardless of true status) would receive confirmatory testing, while those predicted as suppressed with high confidence might have testing deferred. Future work should focus on threshold optimization to prioritize sensitivity for the unsuppressed class, potentially accepting lower precision to minimize missed treatment failures. Incorporation of such a model into EMR systems of low-resource clinic settings can be an early warning system. With automated risk score calculation during the encounter of the patient, the model has the potential to notify clinicians to flag patients at increased risk of viral load suppression for priority counseling regarding adherence, escalated follow-up, or enhanced clinical monitoring before treatment failure. This pre-emptive strategy may make the best of finite healthcare resources and also enhance the overall outcomes of therapy. That the model depends upon variables generally found in EMRs for ART clinics makes it most feasible for deployment without new, expensive testing.
Beyond clinical utility, this work carries important implications for HIV policy and health system strengthening in low-resource settings. The model demonstrates that predictive analytics can optimize scarce laboratory resources by prioritizing confirmatory testing for patients at the highest risk of unsuppressed viral load, potentially improving efficiency in settings like Ethiopia, where testing coverage is constrained by supply chain and capacity limitations. The identification of baseline CD4 category and ART duration as dominant predictors reinforces the policy priority of early diagnosis and treatment initiation, while the finding that male sex and advanced WHO stage predict unsuppressed viral load highlights persistent disparities requiring targeted interventions such as male-friendly services and enhanced support for late presenters. Furthermore, integrating validated predictive models into electronic medical records could enable automated risk stratification at scale, supporting WHO’s differentiated service delivery framework by directing intensified support to high-risk patients while reducing visits for low-risk individuals. Finally, the interpretability of our model (via SHAP) addresses key policy concerns regarding AI transparency, allowing regulators and program managers to audit automated decisions and ensure alignment with clinical guidelines and ethical principles as digital health investments expand across sub-Saharan Africa.
Translating this model into routine clinical use requires a structured four-phase pathway tailored to resource-limited settings: (1) technical integration, embedding the model within existing EMR systems (SmartCare/OpenMRS) with automated data extraction and real-time risk score display; (2) clinical workflow integration, training staff on risk score interpretation, establishing triage protocols (e.g., <50% predicted probability prioritizes viral load testing), and developing patient-friendly counseling explanations; (3) evaluation and refinement, conducting prospective cohort studies to assess impact on outcomes, monitoring performance drift with quarterly recalibration, and iteratively refining thresholds based on user feedback; and (4) scale-up, packaging implementation toolkits for dissemination to other Ethiopian ART clinics, establishing learning collaboratives, and advocating for national policy endorsement within differentiated service delivery guidelines. This phased approach acknowledges resource constraints and human factors, moving beyond technical validation toward sustainable clinical integration (Figure 10). Proposed clinical workflow integration of viral load prediction model.
Limitations and future directions
While having supportive results, this study has several limitations. First, the data are derived from a single tertiary hospital in Ethiopia and hence may have limited generalizability to other populations with their own unique demographic and epidemiological characteristics. External verification with datasets from other regions is required. Second, while the dataset was balanced for the outcome variable, it may not have contained all relevant predictors, e.g., certain socio-economic variables, psychosocial stressors, or genetic markers. Future research would be well-advised to incorporate these variables to further enhance predictive power. Third, the model’s performance, while good, indicates that there remains some unexplained variance, and viral load suppression must therefore rely, at least in part, on factors yet to be captured in the current feature set.
Future studies would need to focus on three areas: 1) Multi-center verification to test the strength and transportability of the model to a variety of sub-Saharan African health settings. 2) Prospective studies of implementation to quantify the real-world impact of deploying the model into clinical practice on patient outcomes and utilization of resources. 3) Exploration of temporal modeling techniques to estimate viral load suppression risk dynamically through time, rather than at a snapshot in time, providing yet another more persuasive tool for long-term management of patients.
While this single-site study limits immediate generalizability, the University of Gondar cohort possesses characteristics, an urban-rural mix, a 19-year treatment era span, and typical LMIC data quality challenges that enhance its utility as a benchmark for similar settings. To advance external validity strategically, we propose a three-phase approach: (1) internal temporal validation confirming stable performance (AUC-ROC 0.78 vs. 0.79); (2) geographic validation through planned collaborations with two other Ethiopian ART clinics; and (3) cross-national validation via open-source model sharing for external testing across sub-Saharan Africa. The model’s reliance on routinely collected variables standard to national HIV programs facilitates this validation agenda, and we invite external testing to establish generalizability bounds.
Third, the balanced proportion of unsuppressed and suppressed viral loads in this cohort (50.4% unsuppressed, 49.6% suppressed) is substantially different from the suppression proportions commonly reported in well-managed HIV cohorts (80-90%). This is because of the study’s unique features: the setting of a tertiary centre for managing complicated and treatment-experienced cases, the study period of 19 years from 2005 to 2024, including years with less effective ART regimens, and the inclusion of all cases rather than those with optimal outcomes. Thus, model performance might vary in settings with higher proportions of baseline suppression.
Fourth, despite our sensitivity analyses, the 15% missingness in CD4 count measurements may cause information bias. Little’s MCAR test was performed to determine the missing data mechanism. The results showed that the data were not missing completely at random because the missing CD4 count was more common in patients with poor adherence documentation.
Conclusion
In conclusion, the study was successful in the development of an interpretable machine learning model in predicting viral load suppression using routine clinical data collected from one tertiary hospital in Ethiopia. The Gradient Boosting model showed superior classification performance in the cohort (76% accuracy, 0.79 AUC-ROC), as well as providing insights that agreed with established knowledge in the field; namely, that the longer the time on ART and the higher the CD4 at the start of ART, the higher the odds of viral load suppression. Nevertheless, there were significant shortcomings in the study. The model’s performance explained only 76%, with the remainder (24%) unexplained, and the significant false negative rate of 35% in the unsuppressed viral load suggests that the tool is not an alternative to confirmatory testing. Furthermore, the results are based on a single site and patient population (tertiary referral site and 19-year enrolment period) and need to be validated before consideration for wider implementation. Nevertheless, within these limitations, the model represents a proof-of-concept for how interpretable machine learning can help address the disconnect between predictive accuracy and actionability in resource-limited settings. Ultimately, these types of tools may have the potential to assist healthcare providers in optimizing HIV management, although this is for future research and not a recommendation.
Footnotes
Acknowledgments
We would like to express our heartfelt thanks to the University of Gondar Hospital for cooperating and permitting the use of the data. Moreover, we would like to give our special appreciation to Debre Markos University for its contribution. Finally, thanks to all who made significant contributions to the success of this study.
Ethical considerations
Ethical approval for this study was obtained from the Institutional Review Board (IRB) of Debre Markos University, College of Medicine and Health Sciences (Approval Number: HSC/R/C/Ser/Co/123/25). Since all data were anonymized before analysis, patient confidentiality was strictly maintained. Access to the dataset was restricted to authorized researchers only to comply with ethical guidelines and data protection policies. The study was conducted in accordance with the principles of the Declaration of Helsinki and adhered to all relevant national regulations and ethical guidelines.
Consent to participate
Because this study involved retrospective analysis of anonymized clinical data with no direct patient contact or intervention, the IRB granted a waiver of informed consent. All data were de-identified before extraction, and no patient identifiers were included in the analysis dataset.
Author contributions
A.K.M. conceptualized the study, and A.E.G. was involved in design, analysis, interpretation, report, and manuscript writing. A.W.S. and M.B.M. edited the manuscript for clarity and correctness. All the authors read and approved the final manuscript
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
Use of artificial intelligence
The authors declare that no artificial intelligence (AI) tools or large language models were used in the generation, writing, or substantive editing of this manuscript. The human authors produced all content, analyses, and interpretations. Standard grammar and spell-checking tools were used solely for language refinement and did not contribute to intellectual content or scientific decision-making.
